Abstract
Beta-site amyloid precursor protein cleaving enzyme 1 (BACE1), also known as β-secretase, is an aspartic protease. The sorting of this enzyme into Rab11-positive recycling endosomes regulates the BACE1-mediated cleavage of its substrates, however, the mechanisms underlying this targeting remain poorly understood. The neural cell adhesion molecule 2 (NCAM2) is a substrate of BACE1. We show that BACE1 cleaves NCAM2 in cultured hippocampal neurons and NCAM2-transfected CHO cells. The C-terminal fragment of NCAM2 that comprises the intracellular domain and a small portion of NCAM2’s extracellular domain, associates with BACE1. This association is not affected in cells with inhibited endocytosis, indicating that the interaction of NCAM2 and BACE1 precedes the targeting of BACE1 from the cell surface to endosomes. In neurons and CHO cells, this fragment and BACE1 co-localize in Rab11-positive endosomes. Overexpression of full-length NCAM2 or a recombinant NCAM2 fragment containing the transmembrane and intracellular domains but lacking the extracellular domain leads to an increase in BACE1 levels in these organelles. In NCAM2-deficient neurons, the levels of BACE1 are increased at the cell surface and reduced in intracellular organelles. These effects are correlated with increased levels of the soluble extracellular domain of BACE1 in the brains of NCAM2-deficient mice, suggesting increased shedding of BACE1 from the cell surface. Of note, shedding of the extracellular domain of Sez6, a protein cleaved exclusively by BACE1, is reduced in NCAM2-deficient animals. These results indicate that the BACE1-generated fragment of NCAM2 regulates BACE1 activity by promoting the targeting of BACE1 to Rab11-positive endosomes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The neural cell adhesion molecule 2 (NCAM2) is a cell surface glycoprotein of the immunoglobulin superfamily of cell adhesion molecules [1]. Two NCAM2 isoforms, generated via alternative splicing, have identical extracellular domains comprising five immunoglobulin-like (Ig) and two fibronectin type III (Fn) repeats but differ in their membrane anchorage. The shorter isoform is glycosylphosphatidylinositol anchored, whereas the longer isoform is a transmembrane protein with an intracellular domain of 119 amino acids [2,3,4]. During development, NCAM2 promotes neurite outgrowth as well as branching and guidance of dendrites and axons of differentiating neurons [5,6,7,8]. NCAM2 also contributes to maturation of developing synapses [9]. In the mature brain, NCAM2 accumulates in excitatory synapses, where it forms trans-synaptic homophilic adhesive bonds involved in synapse maintenance [10, 11]. Deletions of the NCAM2 gene are found in individuals with neurodevelopmental disorders, intellectual disability, and autism and other deficits [12,13,14]. NCAM2 is encoded by a gene located on chromosome 21 in humans, which is triplicated and thereby overexpressed in Down’s syndrome [15]. The overall levels of NCAM2 are increased in sporadic Alzheimer’s disease (AD), whereas its synaptic levels are reduced [11]. Single nucleotide polymorphisms (SNPs) in the NCAM2 gene are associated with an increased risk of late-onset AD [16]. SNPs in the NCAM2 gene are also associated with increased cerebrospinal fluid levels of Aβ, an insoluble toxic peptide accumulating in AD brains [17], suggesting that NCAM2 is involved in regulating the expression and activity of proteolytic enzymes involved in amyloid β (Aβ) production.
One such proteolytic enzyme is the beta-site amyloid precursor protein cleaving enzyme 1 (BACE1). This aspartic acid protease mediates a rate-limiting step in Aβ production. BACE1 activity is highest in the acidic environment of endosomes. To reach endosomes, BACE1 is transported via the Golgi apparatus and Golgi-derived vesicles to the cell surface [18, 19] from where BACE1 is endocytosed via a clathrin-dependent pathway to reach early and Rab11-positive recycling endosomes [18,19,20]. BACE1 then shuttles between the plasma membrane and the endosomal system [18]. BACE1 can also be targeted from the endosomal system to lysosomes for degradation [21] or retrogradely transported to the trans-Golgi network [22]. Trafficking of newly synthesized BACE1 is polarized in neurons, where it is initially targeted to the cell surface of dendrites, wherefrom it is endocytosed into early and recycling endosomes [23,24,25]. From dendrites, BACE1 redistributes to axons and synaptic terminals via the cell soma, a process in which Rab11-positive slowly recycling endosomes appear to play a major role [24, 26]. In contrast, BACE1-containing early endosomes are stationary in dendrites [27]. Rab11-dependent transport of BACE1 is required for its activity towards some of its substrates, including amyloid precursor protein (APP) [25, 28]. The removal of BACE1 from synaptic terminals occurs via dynein-snapin-mediated retrograde axonal transport [26]. While the role of endosomes in the axonal trafficking of BACE1 and its activity towards APP is well established, mechanisms regulating BACE1 targeting to the recycling endosomes are less understood. The activity of BACE1 towards its substrates can also be regulated via the shedding of the ectodomain of BACE1 from the plasma membrane [29, 30], since soluble BACE1 is unable to cleave APP [31]. Mechanisms regulating BACE1shedding by proteolysis remain also mostly unknown.
BACE1 cleaves NCAM2 in the second Fn domain at amino acids 662–663, and this cleavage results in the release of a part of the extracellular domain of NCAM2 from the cell surface and the formation of a C-terminal fragment comprising the whole intracellular and transmembrane domain in conjunction with the residual 35 amino acids of the extracellular domain of NCAM2 [32]. In the present study, we show that the BACE1-generated C-terminal fragment of NCAM2 associates with BACE1 and regulates the targeting of BACE1 to Rab11-positive endosomes. NCAM2 deficiency leads to the accumulation of BACE1 at the dendritic cell surface and enhances the shedding of BACE1. Our results suggest that the cleavage of NCAM2 by BACE1 represents a yet unknown mechanism by which BACE1 can be sorted to recycling endosomes.
Materials and methods
Antibodies and inhibitors
Goat polyclonal antibodies against the C-terminal intracellular amino acids 822–836 of human NCAM2 (14 of 15 amino acids are conserved in mouse NCAM2) from GeneTex (cat# GTX89311) were used in the proximity ligation (PL) assay (PL, 1:100), immunocytochemistry (IC, 1:100), and Western blot (WB, 1:1000). The mouse monoclonal antibody recognizing an extracellular epitope located within the first Fn repeat of mouse NCAM2 (Supplementary Fig. S1) from Santa Cruz Biotechnology (cat# sc-136328) was used in IC (1:100) and WB (1:1000). The specificity of these antibodies had been confirmed [9, 11] and herein by using NCAM2−/− tissue (Fig. 9B). The rabbit monoclonal antibody against the intracellular domain of BACE1 (D10E5, cat# 5606) from Cell Signaling Technology [33] was used for PL (1:100). Rabbit polyclonal antibodies against the N-terminus of BACE1 (amino-acids 38–70 of human BACE1, cat# AP14566PU-N) from OriGene were used for IC (1:100). The mouse monoclonal antibody against synaptophysin (cat# sc-17750, IC at 1:100) was from Santa Cruz Biotechnology. The mouse monoclonal antibody against the FLAG tag (cat# F1804, IC at 1:1000), rabbit polyclonal antibodies against the human influenza hemagglutinin (HA) tag (cat # H6908, IC at 1:100), mouse monoclonal antibody against glyceraldehyde 3-phosphate dehydrogenase (GAPDH, cat # G8795, WB at 1:1000), and mouse monoclonal antibody against actin (cat# A1978, WB at 1:3000) were from Sigma-Aldrich. The mouse monoclonal antibody against Rab11 (cat# 610656, IC at 1:50, WB at 1:1000), mouse monoclonal antibody against Rab4 (cat# 610888, WB at 1:1000), and mouse monoclonal antibody against Rab5 (cat# 610725, WB at 1:1000) were from BD Biosciences. β-Amyloid (1–42 specific) (D3E10) rabbit monoclonal antibodies (cat# 12843, Dot Blot at 1:2000) were from Cell Signaling Technology. Rabbit polyclonal antibodies against Sez6 were as described [34]. Fluorochrome- and HRP-conjugated secondary antibodies were from Jackson ImmunoResearch. BACE1 inhibitor (β-Secretase Inhibitor IV, cat# sc-222304), which inhibits BACE1 (IC50 = 15 nM) and BACE2 (IC50 = 0.23 µM), was from Santa Cruz Biotechnology. Ethylenediaminetetraacetic acid (EDTA)-free complete inhibitors were from Roche.
DNA constructs and small interfering RNAs (siRNAs)
Bace1 (Myc-Flag-tagged, cat# MR208042) and Bace2 (Myc-Flag-tagged, cat# MR220631) were from OriGene. HA-tagged full-length transmembrane isoform of human NCAM2 was as described [11] and used as a template to produce HA-tagged NCAM2ΔED comprising transmembrane and intracellular domains of NCAM2 (aa 694–837). pcDNA3 was from Life Technologies. BACE1 siRNA (cat# sc-37225), BACE2 siRNA (cat# sc-29777) and control siRNA (cat# sc-37007) were from Santa Cruz Biotechnology. The efficiency of knock-down was verified in transfected CHO cells and cultured hippocampal neurons (Supplementary Fig. S2). BACE1 fused to the C-terminal amino acid residues 155–238 of the Venus fluorescent protein (BACE1-VC) was a kind gift of Subhojit Roy [35]. Rab11-DsRed (Addgene, plasmid # 12679 [36]), DsRed-Golgi [37], Lamp1-EYFP [37], and GFP-Sec61B (Addgene, plasmid # 121159 [38]) were as described. mTagBFP-Nucleus-7 (Addgene, plasmid # 55265) was a kind gift of Michael Davidson (unpublished). Cherry was a kind gift from Roger Tsien [39]. DNA constructs coding for Ig1, Ig2, Ig3, Ig4, Ig5, Fn1, Fn2 and intracellular domain (ID) of NCAM2 were synthesized and subcloned into the pET100 vector using GeneArt services (Thermo Fisher Scientific). HA-NCAM2-VN and HA-NCAM2ΔED-VN were generated by inserting the first 155 amino acids of Venus (VN) at the C-terminus of HA-NCAM2 and HA-NCAM2ΔED, respectively, with an intervening 10 amino acid linker (PRARDPPVAT). Briefly, DNA encoding VN bearing an I152L point mutation was PCR amplified from pBiFC-VN155(I152L) (Addgene, plasmid # 27097 [40]) and inserted at the extreme C-terminal end of NCAM2 using the NEBuilder HiFi DNA Assembly kit (New England Biolabs) according to the manufacturer’s instructions. Both constructs were cloned in frame and fidelity confirmed by sequencing.
Animals
NCAM2-deficient mice (STOCK Ncam2tm1Mom/MomJ; Stock No: 006706) were from the Jackson Laboratory. For biochemical analyses of adult 2-month-old brain tissue and dot blot analysis of Aβ1-42 levels in 5–6-month-old brains, NCAM2+/+, NCAM2+/− and NCAM2−/− littermates from heterozygous breeding pairs were used. 0- to 3-day-old NCAM2 + / + and NCAM2−/− mice for cell culture and biochemical analysis were obtained using homozygous breeding pairs of respective genotypes. We also used C57BL/6 mice in experiments, which were not aimed at comparing mice of different genotypes. Experiments were approved by the Animal Care and Ethics Committee of the University of New South Wales (permit 18/99A (for work with C57BL/6 mice) and 17/102A & 20/93B (for work with transgenic mice)).
Neuronal cell culture and transfection
Mouse hippocampal neurons were prepared from 0- to 3-day-old NCAM2 + / + and NCAM2−/− mice or C57BL/6 mice and cultured as described [9, 41]. Neurons were maintained in Neurobasal A supplemented with 2% B-27, glutamine, and 2 ng/ml FGF-2 on glass coverslips coated with poly-d-lysine (100 μg/ml) (all reagents were from Thermo Fisher Scientific). Neurons were transfected before plating by electroporation using a Neon transfection system (Thermo Fisher Scientific). Alternatively, neurons were transfected using the calcium phosphate method essentially as described [42]. Briefly, coverslips with neurons maintained for 7 to 10 days in culture were transferred 30 min prior to transfection to a 24-well-plate containing 500 µl/well culture medium. One µg of plasmid DNA and 3.1 µl of 2 M CaCl2 were mixed with water to a final volume of 25 µl per coverslip. DNA-Ca2+-phosphate precipitate was prepared by adding DNA/CaCl2 solution to 25 µl of 2 × HEPES-buffered saline (HBS, 280 mM NaCl, 1.5 mM Na2HPO4, 50 mM HEPES, pH 7.10) (1/8th at a time and mixing briefly between each addition). The solution was incubated for 10 min at 37 °C and the resulting suspension was applied dropwise to the coverslips. Neurons were incubated with the precipitate for 3 h, washed with acidified Tyrode’s solution (pH 6.7–6.8) to remove the precipitate, and transferred back to the wells containing the original conditioned culture medium.
CHO cell culture and transfection
CHO cells were maintained in Dulbecco’s Modified Eagle’s Medium/Nutrient Mixture F-12 HAM (Sigma) supplemented with 5% newborn calf serum (Sigma) in an incubator at 37 °C and 5% CO2. For immunofluorescence analysis, CHO cells were plated on glass coverslips in a 24-well-plate in a culture medium and transfected using Lipofectamine 3000 transfection reagent (Thermo Fisher Scientific) according to the manufacturer’s instructions. Alternatively, cells were transfected using the calcium phosphate method as described in Neuronal culture and transfection. Neuronal cultures were supplied with fresh culture medium prior to transfection, and CHO cells were treated with culture medium without serum. After transfection, neurons were washed with Tyrode’s solution, and CHO cells were treated with 15% glycerol dissolved in phosphate-buffered saline (PBS) for 1.5 min, washed with PBS and supplied with fresh culture medium containing 5% serum. Cells were incubated for 24 h with BACE1 inhibitor IV (30 nM), dynasore (80 µM [39]) or vehicle (0.1% DMSO in culture medium) at 24 h after transfection.
Immunofluorescence labeling of BACE1 at the cell surface of cultured cells
Live CHO cells and neurons were incubated with rabbit polyclonal antibodies against the N-terminus of BACE1 diluted in culture medium for 20 min on ice, fixed with 4% formaldehyde in PBS for 15 min, washed with PBS, blocked with 1% donkey serum diluted in PBS and incubated with fluorochrome conjugated anti-rabbit secondary antibodies for 45 min at room temperature. Cells were then washed with PBS and postfixed in 4% formaldehyde in PBS for 5 min at room temperature. Fixed cells were immunolabeled as described in “Immunofluorescence labeling of fixed cultured cells”.
Immunofluorescence labeling of cultured cells
The labeling was done essentially as described [43]. In brief, cultured neurons and CHO cells were fixed with 4% formaldehyde in PBS for 15 min at room temperature and washed with PBS. Cells were permeabilized with 0.25% Triton X-100 in PBS for 5 min and blocked with 1% donkey serum in PBS for 20 min at room temperature. Cells were then incubated with primary antibodies at 4 °C overnight, washed with PBS and incubated with fluorochrome-conjugated secondary antibodies for 45 min at room temperature. All antibodies were diluted in PBS containing 1% donkey serum. Thereafter, cells were embedded in ProLong Gold Antifade Mountant (Thermo Fisher Scientific).
Analysis of BACE1 endocytosis in cultured neurons and CHO cells
Cells were incubated with rabbit polyclonal antibodies against the N-terminus of BACE1 diluted in culture medium for 30 min in the incubator, washed with PBS, fixed with 4% formaldehyde in PBS for 15 min at room temperature, washed again with PBS, blocked with 1% donkey serum diluted in PBS for 20 min at room temperature and incubated with fluorochrome-conjugated anti-rabbit secondary antibodies for 1 h at room temperature to detect cell surface BACE1. To detect endocytosed BACE1–BACE1 antibody complexes, cells were then washed with PBS, postfixed with 2% formaldehyde in PBS for 5 min at room temperature, washed with PBS, permeabilized with 0.25% Triton X-100 for 5 min, blocked with 1% donkey serum diluted in PBS for 20 min at room temperature and incubated with anti-rabbit secondary antibodies conjugated to a fluorochrome, which differed from the one used to detect cell surface BACE1, for 45 min at room temperature. All secondary antibodies were diluted in PBS containing 1% donkey serum. Labeling for other antigens was done as described in “Immunofluorescence labeling of fixed cells”.
Proximity ligation assay (PL assay)
The assay was performed essentially as described [44]. Cultured neurons were fixed in 4% formaldehyde in PBS, washed with PBS, permeabilized with 0.25% Triton X-100 in PBS for 5 min, and blocked with 1% bovine serum albumin (BSA) in PBS for 20 min. Antibodies against the intracellular domain of NCAM2 and BACE1 were applied to the cells in 0.1% BSA in PBS overnight at 4 °C. Further steps were performed using secondary antibodies conjugated with oligonucleotides (PL assay probes, Olink Bioscience, Uppsala, Sweden) and Duolink II fluorescence kit (Olink Bioscience) in accordance with the manufacturer’s instructions.
Confocal microscopy and image analysis
Fluorescence images were acquired at room temperature using the confocal laser scanning microscope Nikon C1si, NIS Elements software and CFI Plan Apochromat VC 60XH objective (numerical aperture 1.4) all from Nikon Corporation (Tokyo, Japan). Image analysis was done with ImageJ.
Analysis of the density of intracellular BACE1 clusters in cultured neurons
Surface and total BACE1 clusters/puncta were outlined using the threshold function, with a constant threshold set to 4 times the average mean intensity along dendrites in each channel. To count intracellular puncta, a mask of all surface labeling above the threshold was subtracted from the images for determination of the total BACE1 labeling. Thereafter, the remaining dendritic puncta, which were not detectable at the cell surface, were counted using the analyze particles tool in ImageJ. The numbers were normalized to dendritic length to obtain the density of intracellular BACE1 puncta.
Analysis of BACE1 targeting to organelles in CHO cells
Organelles were outlined using the threshold function of ImageJ. The enrichment of BACE1 in these organelles was calculated by dividing the mean intensity of BACE1 labeling within the outlined organelles by the mean intensity of BACE1 labeling in the entire cell. The percentage of BACE1 labeling in organelles was calculated by dividing the integrated density of BACE1 labeling within the outlined organelles by the integrated density of BACE1 labeling within the entire cell.
Analysis of BACE1 targeting to Rab11-positive endosomes in cultured hippocampal neurons
Rab11-positive endosomes were outlined using the threshold function in ImageJ. The enrichment of BACE1 was calculated by dividing the mean intensity of BACE1 labeling within the outlined endosomes by the mean intensity of BACE1 labeling in the entire dendrite. The percentage of BACE1 labeling in Rab11-positive endosomes was calculated by dividing the integrated density of BACE1 labeling in endosomes by the integrated density of BACE1 labeling in the entire dendrite.
Preparation of brain homogenates, synaptosomes and soluble protein fractions
Brain tissue homogenates (10%, w/v) were prepared in HOMO-A buffer which was prepared from HOMO buffer (1 mM MgCl2, 1 mM CaCl2, 1 mM NaHCO3, 5 mM Tris, pH 7.4) and contained 0.32 M sucrose, EDTA-free complete inhibitors and 1 mM phenylmethylsulfonyl fluoride (PMSF). Homogenates were used for synaptosome isolation as described [39, 45]. All steps were performed at 4 °C. Briefly, homogenates were centrifuged at 1400g for 10 min. The supernatant and pellet were resuspended in HOMO-A buffer and centrifuged for 10 min at 700g. The resulting supernatants were pooled and centrifuged at 17,500g for 15 min. The supernatant was centrifuged at 200,000g for 1 h and used as the soluble fraction. The 17,500g pellet was resuspended in HOMO-A buffer and applied to the top of a step gradient with interfaces of 0.65 M, 0.85 M, 1.0 M, 1.2 M sucrose in HOMO buffer. The 700 g pellets were combined, adjusted to 1.0 M sucrose in HOMO buffer and layered onto 1.2 M sucrose in HOMO buffer. HOMO-A buffer was then applied to the top of the gradient. The crude synaptosomal fractions were collected at the 1.0 M/1.2 M interface after centrifugation for 2 h at 100,000 g and pooled. The crude synaptosomal fraction was again adjusted to 1.0 M sucrose and applied onto the 1.2 M sucrose cushion. HOMO-A buffer was then applied to the top of the gradient. After centrifugation for 2 h at 100,000g, synaptosomes were collected at the 1.0 M/1.2 M interface, resuspended in HOMO-A buffer, pelleted by centrifugation for 30 min at 100,000g and resuspended in HOMO-A buffer.
Isolation of the transport vesicle-enriched fraction
Isolation of transport vesicles was performed essentially as described [46] (Fig. 2E). In brief, brain homogenates were prepared in 20 mM HEPES, 40 mM KCl, 5 mM EGTA, 5 mM EDTA, 1 mM PMSF and EDTA-free complete inhibitors. The homogenate was centrifuged at 1000g for 20 min at 4 °C. Pellet P1 was enriched in nuclei and plasma membranes. Supernatant S1 was then centrifuged at 10,200g for 20 min at 4 °C to produce crude synaptosomes (pellet P2). Supernatant S2 containing soluble proteins and transport vesicles was centrifuged at 100,000g for 1 h at 4 °C. The pellet P100 contained transport vesicles and supernatant S100 contained soluble proteins.
Analysis of NCAM2 levels in neuronal culture medium and cell lysates
Hippocampal neurons were maintained in 6-well-plates for 14 days and then treated with β-Secretase Inhibitor IV (30 nM) or vehicle (0.1% dimethyl sulfoxide (DMSO)) for 24 h. Culture media were collected. Neurons were washed with ice-cold PBS and treated with lysis buffer (1% sodium deoxycholate, 1% Triton X-100, 50 mM Tris–Cl, 150 mM NaCl, 1 mM EDTA, EDTA-free complete inhibitors and 1 mM PMSF). Lysates were collected, incubated on ice for 30 min and centrifuged at 16,000g for 15 min at 4 °C. Supernatants were then collected for Western blot analysis. Culture media were centrifuged at 200,000g for 1 h at 4 °C, and the supernatants therefrom were collected. Proteins were precipitated on ice by adding 2% sodium deoxycholate to a final concentration of 0.03% and 20% trichloroacetic acid to a final concentration of 12% for 30 min. Precipitates were collected by centrifuging at 10,000g for 15 min at 4 °C, washed twice with ice-cold acetone, reconstituted in 50 µl lysis buffer and 50 µl 12 M urea, and used for Western blot analysis.
Western blot analysis
Proteins and protein markers (Bio-Rad) were separated in 8% Bis–Tris bolt mini gel (Thermo Fisher Scientific) and electroblotted onto PVDF membranes (0.2 µm; Amersham). Membranes were washed with PBS, blocked with 5% skim milk in PBS, and incubated with appropriate primary antibodies overnight at 4 °C, followed by incubation with corresponding HRP-conjugated secondary antibodies for 1.5 h at room temperature. The antibodies were diluted in PBS containing 0.05% Tween. Protein bands were visualized by applying Luminata Forte Western HRP substrate (Merck Millipore) to the membrane. The chemiluminescence images were taken using Micro-Chemi 4.2 (DNR Bio-Imaging Systems) and analyzed with ImageJ.
Dot blot analysis
Mouse brain homogenates or BSA as negative control were diluted in Tris-buffered saline (TBS), pH 7.4 and 10 µg of total protein was applied to a nitrocellulose membrane (0.2 µm, Bio-Rad). After drying for 15 min at room temperature, the membrane was stained with Ponceau to visualize total protein and then blocked using 5% skim milk in TBS. The membrane was then incubated overnight with rabbit monoclonal antibodies specific for Aβ42 (D3E10), followed by incubation with HRP-conjugated anti-rabbit secondary antibodies for 2 h at room temperature. Dots were visualized using Pierce ECL, imaged using Micro-Chemi 4.2 (DNR Bio-Imaging Systems) and analyzed with ImageJ.
Statistical analysis
Unless indicated otherwise, all experiments were independently performed at least three times. Statistical analyses were done in GraphPad Prism 7. In experiments with immunofluorescence analysis, differences between two groups were analyzed using an unpaired t test, while differences between three or more groups were analyzed using one-way ANOVA and Dunnett’s multiple comparisons test. In each biochemical experiment, samples were organized into sets containing one sample of each kind, and each set was separately analyzed. Littermates were always included in the same set. Fold change relative to the level in the control sample was then calculated for each set, and the data from all sets were combined. One-sample t test was then used to estimate the statistical significance of the fold change relative to the control sample level.
Results
BACE1 is involved in the proteolytic processing of NCAM2 in hippocampal neurons
A recent report demonstrated that NCAM2 is cleaved by BACE1 at a membrane-proximal site within the second Fn repeat of the extracellular domain (ED) of NCAM2 in transfected HEK cells and in the olfactory bulb of mice (Fig. 1A, [32]). This study did not find changes in the levels of full-length NCAM2 or its cleavage products in the hippocampus of BACE1 knock-out mice. However, since BACE1 and NCAM2 are expressed in the hippocampus, the effect of BACE1 deficiency on NCAM2 processing could have been obscured by yet unidentified compensatory proteolytic mechanisms. To determine whether BACE1 plays a direct role in the proteolytic processing of NCAM2 in hippocampal neurons, cultured hippocampal neurons were treated for 24 h with β-secretase inhibitor IV. Western blot analysis with an antibody against the first Fn repeat of NCAM2-ED (Supplementary Fig. S1) which is located N-terminally to the BACE1-cleavage site (Fig. 1A, [32]) showed that BACE1 inhibition causes an increase in NCAM2 levels in lysates of neurons (Fig. 1B). In the supernatant collected from neuronal cultures, this antibody detected soluble NCAM2, which represents NCAM2-ED, as indicated by molecular weight determination (Fig. 1B). BACE1 inhibition reduced levels of soluble NCAM2 in the culture medium (Fig. 1B) indicating that NCAM2-ED was proteolytically removed from the cell surface by BACE1.
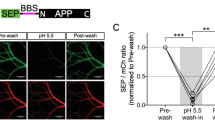
BACE1 is involved in proteolytic processing of NCAM2 in hippocampal neurons. A Schematic diagram showing locations of the domains recognized by antibodies against NCAM2-ED, NCAM2-ID, and HA tag (only in overexpressed NCAM2) used in this study. Orange arrow denotes the BACE1 cleavage site. B Lysates and medium from cultured hippocampal neurons treated with vehicle (0.1% DMSO) or BACE inhibitor analyzed by Western blot with anti-NCAM2-ED antibodies. Levels of NCAM2 are increased in lysates and reduced in the culture medium of neurons treated with the inhibitor. Graph shows NCAM2 levels in lysates (n = 14) and the ratios of medium/lysate levels (n = 8) from the inhibitor-treated neurons relative to levels in vehicle-treated neurons set to 100%. Means ± SEM are indicated. *p, one sample t test compared to the vehicle level. C NCAM2-ED and NCAM2-ID labeling in hippocampal neurons co-transfected with GFP and BACE1 siRNA, BACE2 siRNA or control siRNA. Bar = 20 μm. Graphs show mean + SEM NCAM2-ED and NCAM2-ID labeling intensities along dendrites of transfected neurons and their ratios (n > 46 neurons per group) normalized to the mean of control siRNA-transfected neurons set to 100%. *p, one-way ANOVA with Dunnett’s multiple comparisons test. D NCAM2-ED and NCAM2-ID labeling of CHO cells co-transfected with GFP and NCAM2 and treated with the BACE inhibitor or vehicle (0.1% DMSO). Bar = 20 μm. Graphs show mean + SEM NCAM2-ED and NCAM2-ID labeling intensities and their ratios (n > 173 cells per group) normalized to the mean of GFP-transfected cells set to 100%. *p, unpaired t-test. E NCAM2-ID and HA tag labeling in CHO cells co-transfected with HA-NCAM2 and GFP or BACE1-FLAG. Note the strongly reduced HA tag labeling in BACE1 co-transfected cells. Bar, 20 μm. Graph shows mean + SEM ratios of the HA tag and NCAM2-ID labeling intensities in CHO cells co-transfected with GFP, BACE1 or BACE2 (n > 188 cells per group) normalized to the means of GFP-transfected cells set to 100%. *p, one-way ANOVA and Dunnett’s multiple comparisons test
To investigate the effect of BACE1 deficiency on NCAM2 levels, the levels of NCAM2-ED immunoreactivity were analyzed along dendrites of cultured hippocampal neurons co-transfected with GFP together with BACE1 or control siRNAs. Unexpectedly, a ⁓50% reduction in BACE1 levels (Supplementary Fig. S2) did not affect the NCAM2-ED levels along dendrites (Fig. 1C). The intensities of labeling with antibodies against the very C-terminus of the intracellular domain (ID) of NCAM2 (Fig. 1A) were, however, reduced along dendrites (Fig. 1C) and the ratios of NCAM2-ED/NCAM2-ID levels were increased in cultures treated with BACE1 siRNA vs control siRNA (Fig. 1C). A similar effect was observed in neurons transfected with BACE2 siRNA (Fig. 1C), with BACE2 being also expressed in hippocampal neurons although at a lower level than BACE1 [47, 48].
To further investigate whether changes in the NCAM2-ED/NCAM2-ID ratio reflect BACE1 activity levels, we used CHO cells, which had been used for the analysis of the cleavage of APP, a well-studied BACE1 substrate [20, 49,50,51]. Inhibition of BACE1 with β-secretase inhibitor IV did not affect the levels of NCAM2-ED in NCAM2-transfected CHO cells but led to a reduction in the levels of NCAM2-ID and an increase in the ratios of NCAM2-ED/NCAM2-ID labeling intensities (Fig. 1D), as seen in cultured neurons. In contrast, co-expression of BACE1 with human NCAM2 containing an HA tag at the extracellular N-terminus (HA-NCAM2, Fig. 1A) reduced the NCAM2-ED/NCAM2-ID ratio in transfected cells (Supplementary Fig. S3). Similarly, the HA tag/NCAM2-ID ratio was reduced in BACE1-co-trasfected CHO cells (Fig. 1E). This reduction was reflected by a nearly complete loss of the HA tag and NCAM2-ED immunoreactivity in BACE1-co-transfected cells, whereas NCAM2-ID was retained (Fig. 1E, Supplementary Fig. S3). Results indicate that the NCAM2-ED/NCAM2-ID ratio inversely correlates with BACE1 activity.
Altogether, we conclude that BACE1 is involved in the proteolytic processing of NCAM2 in hippocampal neurons.
A fragment of NCAM2 accumulates in BACE1-containing endosomes
In cultured hippocampal neurons analyzed by confocal microscopy, accumulations of NCAM2-ID and BACE1 immunoreactivities co-localized in vesicle-like structures in dendritic shafts, occasionally in dendritic spines (Fig. 2A) and in somata (Fig. 2B). Accumulations of NCAM2-ID immunoreactivity co-localizing with BACE1 were often not detectable with NCAM2-ED antibodies (Fig. 2B). Out of 313 clusters of BACE1 identified manually in dendrites, 128 clusters colocalized with a cluster of NCAM2-ID immunoreactivity and only 64 clusters also colocalized with a cluster of NCAM2-ED immunoreactivity (31 neurons from 2 cultures were analyzed).
NCAM2-ID and BACE1 co-localize in endosomes. A Distribution of NCAM2-ID and BACE1 immunoreactivity in a cultured hippocampal neuron transfected with GFP to visualize its morphology. Note overlapping accumulations of NCAM2-ID and BACE1 in the shaft (arrows) and spine (arrowhead) in the dendrite. Bars, 10 µm (low magnification), 5 µm (high magnification). B A confocal slice through a soma showing NCAM2-ED-negative intracellular accumulations of NCAM2-ID immunoreactivity co-localizing with BACE1 (arrows). Bar, 5 µm. C The grayscale image shows NCAM2-ID/BACE1-ID proximity ligation (PL) products (black aggregates) in a cultured hippocampal neuron. NCAM2-ID/BACE1-ID PL products are visible in the soma (arrowheads) and in the dendritic shaft (arrows). Co-labeling for synaptophysin shows that the dendritic PL-labeled proteins are detectable in the vicinity of synaptophysin. Bar, 10 µm. D Co-localization of internalized BACE1 and NCAM2-ID accumulations (arrows) in a CHO cell co-transfected with BACE1 and NCAM2. Accumulations of internalized BACE1 are seen as red clusters of BACE1 detected after surface membrane permeabilization (surf. + int. BACE1). These clusters do not overlap with clusters of BACE1 detected before permeabilizing membranes (surf. BACE1). Bars, 10 μm (low magnification), 5 µm (high magnification). E Scheme of the subcellular fractionation protocol used to produce fractions analyzed in F. F Western blot analysis of the brain homogenate (BH) and fractions obtained as shown in E. Note that BACE1 and the ~ 32 kDa NCAM2 fragment detected with NCAM2-ID antibodies, which was previously described as the cleavage product of BACE1 [32], are present in fraction P100 enriched in transport vesicles. This fragment is not detectable in fraction S100 containing soluble proteins. Graphs show the enrichment of this fragment relative to full-length NCAM2 levels (NCAM2 fragment/FL) and relative to BACE1 levels (NCAM2 fragment/BACE1) in corresponding fractions (mean ± SEM, n = 3) normalized to the enrichment in BH set to 1. *p, ANOVA with Tukey’s multiple comparisons test
Proximity ligation (PL) using antibodies against NCAM2-ID and BACE1-ID produced a positive reaction (Fig. 2C). The reaction was not observed when the BACE1 antibodies were omitted (Supplementary Fig. S4). The NCAM2-ID/BACE1-ID ligation products were present along dendrites, occasionally near synaptophysin-positive synaptic boutons, and in somata (Fig. 2C) showing that NCAM2-ID and BACE1-ID are in close proximity (40 nm or less) and may even form a complex containing the two molecules.
To confirm that BACE1 co-localizes with NCAM2-ID in endosomes, CHO cells co-transfected with BACE1 and NCAM2 were incubated live with antibodies against BACE1-ED for 30 min at 37 °C. BACE1-ED antibodies bound to BACE1 remaining at the cell surface were then visualized with fluorochrome-conjugated secondary antibodies applied to live cells on ice. BACE1-ED antibodies internalized together with BACE1 into endosomes were then visualized after fixing and detergent-permeabilizing the cells using secondary antibodies conjugated to another fluorochrome. Co-labeling of cells for NCAM2-ID showed that accumulations of NCAM2-ID immunoreactivity indeed co-localized with clusters of internalized BACE1 (Fig. 2D).
To determine whether NCAM2 co-localizes with BACE1 in transport vesicles of the brain, the transport vesicle enriched fraction (P100), containing vesicles involved in BACE1 transport in neurons [27], was isolated from brain tissue by differential centrifugation [27, 46], by sequentially depleting the mouse brain tissue of nuclei and plasma membranes (fraction P1) [52], synaptosomes (fraction P2) and soluble proteins [46] (Fig. 2E). Western blot analysis of these fractions with NCAM2-ID antibody showed that full-length NCAM2 was enriched in fractions P1 and P2 (Fig. 2F) in accordance with NCAM2 localization at the plasma membrane (Fig. 2B) and synapses [11]. Low levels of full-length NCAM2 were also detected in fraction P100 enriched in transport vesicles (Fig. 2F). The NCAM2-ID antibody also detected a fragment of NCAM2 previously described as a BACE1-generated cleavage product and detectable at ~ 32 kDa [32], which was present at similar levels in fractions P1, P2 and P100 (Fig. 2F). This fragment was not detectable in the soluble protein fraction S100 (Fig. 2F), indicating that it is a membrane-bound NCAM2 fragment. The enrichment of the 32 kDa fragment relative to full-length NCAM2, determined as the ratio of their levels in respective fractions was highly increased in transport vesicles (Fig. 2F). Furthermore, the enrichment of this fragment relative to BACE1, determined as the ratio of the levels of these two proteins was also highly increased in transport vesicles (Fig. 2F).
Altogether, our data indicate that the NCAM2 fragment most likely representing the previously described BACE1-generated cleavage product accumulates in vesicles involved in BACE1 transport.
NCAM2-ED is not required for the NCAM2-ID/BACE1 complex formation and its targeting to endosomes
We next analyzed whether NCAM2-ED is required for targeting of NCAM2-ID to BACE1. In CHO cells co-transfected with the NCAM2 fragment comprising only the transmembrane and intracellular domains (NCAM2ΔED), NCAM2-ID and BACE1 immunolabeling highly co-localized at the plasma membrane and in vesicular-like structures (Fig. 3A).
NCAM2-ED is not required for targeting of NCAM2-ID to BACE1. A NCAM2-ID and BACE1 distribution in a CHO cell co-transfected with NCAM2ΔED and BACE1. Graph shows the distribution of labeling intensities along the dashed line. Bar, 10 µm. B Schematic diagram showing the method of BiFC. Reconstitution of the fluorescent protein Venus from VN and VC fragments fused to the C-termini of NCAM2ΔED and BACE1 results in fluorescence. C BiFC signals in CHO cells co-transfected with BACE1-VC and NCAM2ΔED-VN and cherry fluorescent protein. Bar, 20 μm. Note BiFC signals at the plasma membrane and cytoplasm. D BiFC staining in a cultured hippocampal neuron co-transfected with BACE1-VC, NCAM2ΔED-VN, and cherry fluorescent protein, and co-labelled for synaptophysin. Arrowheads show BiFC in the soma (low magnification image; bar, 10 µm), dendrites (i) and axon (ii) (high magnification images; bar, 5 µm). Axonal BiFC signals co-localize with synaptophysin accumulations. E Co-localization of internalized BACE1 and NCAM2-ID accumulations (arrows) in a CHO cell co-transfected with BACE1 and NCAM2ΔED. Accumulations of internalized BACE1 are seen as red clusters of BACE1 detected after the permeabilization of membranes (surf. + int. BACE1), which do not overlap with clusters of BACE1 detected before permeabilizing membranes (surf. BACE1). Bars, 10 μm (low magnification), 5 µm (high magnification)
Bimolecular fluorescence complementation (BiFC) assay, allowing to visualize proteins located from each other within ⁓ 4.2 nm, i.e. the size of the Venus fluorescent protein [35], was then used to test whether BACE1 and NCAM2ΔED form a complex. In this assay, non-fluorescent N-terminal (VN) and C-terminal (VC) fragments of Venus were fused to the cytosolic C-termini of NCAM2ΔED and BACE1, respectively, and the re-formation of the fluorescent full-length Venus protein from VN and VC tags at sites of proximity between NCAM2 and BACE1 was then analyzed. Co-transfection of CHO cells with NCAM2ΔED-VN and BACE1-VC led to the reconstitution of Venus and generation of BiFC signals at the plasma membrane and in the cytoplasm (Fig. 2C). Formation of NCAM2ΔED-VN/BACE1-VC BiFC was inhibited in a dominant-negative manner in cells co-transfected with BACE1 without the VC tag (Supplementary Fig. S5), indicating that BiFC depended on the interaction between NCAM2ΔED and BACE1 and that this interaction was not caused by the non-specific association of the VN and VC tags.
Co-transfection of cultured hippocampal neurons with NCAM2ΔED-VN and BACE1-VC also resulted in BiFC signals detected along dendrites, in somata and synaptophysin-positive structures in axons (Fig. 3D) indicating that NCAM2ΔED associates with BACE1 and that the NCAM2-ED is not required for this association.
In CHO cells, NCAM2ΔED also co-localized with clusters of internalized BACE1 detected by incubating live cells with antibodies against BACE1-ED for 30 min at 37 °C (Fig. 3E).
The combined observations allow us to conclude that the transmembrane and intracellular domains of NCAM2 are sufficient for the interaction with BACE1, which occurs independently of NCAM2-ED.
The association of NCAM2 and BACE1 precedes endocytosis
BACE1 is associated with its well-characterized substrate APP in endosomes and their approximation is blocked by the endocytosis inhibitor dynasore [27]. In contrast, the NCAM2ΔED/BACE1 BiFC complexes were found at the plasma membrane (Fig. 3C) suggesting that their formation precedes endocytosis. Indeed, dynasore did not reduce the co-localization of NCAM2ΔED and BACE1 in transfected CHO cells (Fig. 4A) and did not affect the BiFC signal levels produced in CHO cells co-transfected with NCAM2ΔED-VN/BACE1-VC (Fig. 4B). Similarly, co-transfection of CHO cells with the VN-tagged full-length NCAM2 (NCAM2-VN) and BACE1-VC also led to the reconstitution of Venus and generation of BiFC signals, which were not affected by dynasore (Fig. 4C). Our results suggest that the association of NCAM2 with BACE1 precedes endocytosis and might occur at the cell surface.
Inhibition of endocytosis does not reduce the interaction of NCAM2 and BACE1. A BACE1 and NCAM2-ID in CHO cells co-transfected with BACE1 and NCAM2ΔED and treated with dynasore or vehicle (0.1% DMSO). Graph shows mean + SEM and Pearson’s coefficient of BACE1 and NCAM2-ID co-localization (n > 20 cells). B, C BiFC in CHO cells co-transfected with BACE1-VC and NCAM2ΔED-VN (B) or NCAM2-VN (C). Cells were either treated with dynasore or vehicle (0.1% DMSO), and co-labelled for NCAM2-ID and BACE1. Graph shows mean + SEM BiFC fluorescence intensities. n = 50 (DMSO) and 22 (dynasore) cells in B, n = 13 (DMSO) and 13 (dynasore) cells in C. Bars = 20 μm
NCAM2ΔED co-accumulates with BACE1 in Rab11-positive recycling endosomes
BACE1 endocytosed from the dendritic cell surface is incorporated into early or recycling endosomes [20, 25, 27]. The BACE1-containing early endosomes are mostly stationary in dendrites [27], whereas the recycling Rab11-positive endosomes are involved in the transport of BACE1 from the dendrites and its trafficking to axons and axonal terminals [24]. Presence of NCAM2ΔED-VN/BACE1-VC BiFC in axons (Fig. 3D) suggested that NCAM2ΔED co-accumulates with BACE1 in the recycling Rab11-positive endosomes. Indeed, in cultured hippocampal neurons co-transfected with NCAM2ΔED and Rab11-DsRed, NCAM2-ID immunoreactivity accumulated in Rab11-positive endosomes in both dendrites and axons identified morphologically (Fig. 5A). BiFC complexes formed by NCAM2ΔED-VN and BACE1-VC along dendrites, axons, and in somata were also found in Rab11-positive endosomes as detected with antibodies against endogenous Rab11 (Fig. 5B). In CHO cells co-transfected with BACE1, Rab11-DsRed and NCAM2 or NCAM2ΔED, NCAM2-ID labeling also highly co-localized with BACE1 in Rab11-DsRed-positive recycling endosomes (Fig. 5C). In summary, our data indicates that NCAM2ΔED co-accumulates with BACE1 in Rab11-positive recycling endosomes.
NCAM2ΔED associates with BACE1 in Rab11-positive recycling endosomes. A NCAM2-ID labeling in a cultured hippocampal neuron co-transfected with Rab11-DsRed and NCAM2ΔED. High magnification images show examples of NCAM2-ID accumulations in Rab11-DsRed positive endosomes (arrowheads) in dendrites (i) and axon (ii). Bar = 10 µm (low magnification), 5 µm (high magnification). B BiFC in cultured hippocampal neurons co-transfected with cherry, NCAM2ΔED-VN and BACE1-VC and labelled for Rab11. Arrowheads show examples of BiFC co-localized with Rab11 in the soma (low magnification image) and along dendrites (high magnification image). Bar, 10 µm (low magnification), 5 µm (high magnification). C BiFC in CHO cells co-transfected with Rab11-DsRed, BACE1-VC and NCAM2-VN or NCAM2ΔED-VN. Bar, 10 µm
NCAM2 promotes targeting of BACE1 to Rab11-positive recycling endosomes
The observation of the close association of NCAM2ΔED with BACE1 prompted us to analyze whether NCAM2 regulates the targeting of BACE1 to recycling endosomes. First, we compared the levels of BACE1 in Rab11-positive endosomes in CHO cells co-transfected with BACE1 and either the control pcDNA3 vector or NCAM2. In NCAM2-transfected cells, accumulations of NCAM2-ID immunoreactivity co-localized with BACE1 in Rab11-positive endosomes (Fig. 6A). The enrichment of BACE1 in individual Rab11-positive recycling endosomes relative to the total BACE1 levels in the cell and the proportion of BACE1 in these organelles were increased in NCAM2-transfected vs control vector-transfected cells (Fig. 6A). A similar increase was found in CHO cells co-transfected with NCAM2ΔED instead of NCAM2 (Fig. 6A), indicating that the targeting of BACE1 to Rab11-positive endosomes is mediated by amino acid sequences present in NCAM2ΔED independently of the extracellular domain of NCAM2. From the endosomal compartment, BACE1 can be transported to the trans-Golgi [22] or to lysosomes for degradation [21]. The levels of BACE1 were similar in Golgi organelles (Supplementary Fig. S6) and only slightly increased in lysosomes (Supplementary Fig. S7) in NCAM2- or NCAM2ΔED-transfected vs control vector-transfected CHO cells. The levels of BACE1 in the endoplasmic reticulum were not changed in NCAM2- or NCAM2ΔED-transfected CHO cells (Supplementary Fig. S8). Overexpression of NCAM2ΔED in cultured hippocampal neurons also enriched BACE1 in individual Rab11-positive recycling endosomes and increased the proportion of BACE1 in these organelles (Fig. 6B).
NCAM2ΔED promotes targeting of BACE1 to Rab11-positive endosomes. A NCAM2-ID and BACE1 labeling in CHO cells co-transfected with BACE1, Rab11-DsRed and full-length NCAM2, NCAM2ΔED or empty pcDNA3 vector. Graphs show mean + SEM of BACE1 levels in endosomes relative to the total BACE1 levels and percentages of BACE1 in endosomes (n > 36). *p, one-way ANOVA and Dunnett’s multiple comparisons test, compared to pcDNA3. Bars, 10 µm (low magnification), 5 µm (high magnification). B BACE1 levels in cultured hippocampal neurons co-transfected with Rab11-DsRed and NCAM2ΔED or empty pcDNA3 vector. Note higher co-localization of BACE1 with Rab11-positive endosomes in NCAM2ΔED overexpressing neurons (arrows). Graphs show mean + SEM of levels of BACE1 in Rab11-positive endosomes relative to total BACE1 levels and percentages of BACE1 in Rab11-positive endosomes (n = 78). *p, Mann–Whitney test. Bar, 10 µm (low magnification), 5 µm (high magnification)
Altogether, our data indicate that NCAM2 targets BACE1 to the recycling endosomes, does not increase its targeting to other organelles and does not change its export from the endoplasmic reticulum.
Rab11 levels are downregulated in NCAM2-deficient mice
Rab11-positive slowly recycling endosomes are involved in regulating the turnover of proteins in the post-synaptic density [53, 54], where NCAM2 is present [11]. Western blot analysis demonstrated that Rab11 is enriched in synaptosomes from NCAM2 + / + brains (Fig. 7A). The levels of Rab11 were, however, strongly reduced in NCAM2−/− brain homogenates and synaptosomes and were also reduced in NCAM2+/− mice (Fig. 7A). In contrast, the levels of Rab4 and Rab5, which localize to the rapid recycling and early endosomes, respectively, were not affected (Fig. 7B,C). Altogether, these data indicate that NCAM2 specifically regulates Rab11-positive recycling endosomes.
Rab11 expression is downregulated in brains of NCAM2−/− mice. A–C Levels of Rab11 (A), Rab4 (B), and Rab5 (C) in brain homogenates (h) and synaptosomes (s) from NCAM2 + / + , NCAM2+/− and NCAM2−/− mice as indicated by Western blot analysis. Graphs show mean ± SEM changes (in fold) in homogenates and synaptosomes of NCAM2+/− and NCAM2−/− mice relative to NCAM2 + / + levels (n = 8). *p, one sample t test, compared to + / +
NCAM2 deficiency causes accumulation of BACE1 at the dendritic cell surface
BACE1 is targeted to the endosomal compartment from the cell surface and can be recycled back to the cell surface from endosomes [18]. To determine how changes in endosomal targeting affect the cell surface localization of BACE1, BACE1 at the neuronal cell surface was visualized by labeling live NCAM2 + / + and NCAM2−/− hippocampal neurons with antibodies against BACE1-ED applied on ice. We found that cell surface BACE1 was predominantly detected in morphologically identified dendrites (Fig. 8A). The ratio of cell surface BACE1 levels to total BACE1 levels detected by co-labeling neurons with BACE1 antibodies after detergent permeabilization was higher in dendrites of NCAM2−/− than NCAM2 + / + neurons (Fig. 8A), indicating that NCAM2 deficiency causes accumulation of BACE1 at the dendritic cell surface. An increase of BACE1 levels at the cell surface was accompanied by a reduction in numbers of intracellular clusters of BACE1 in NCAM2−/− neurons (Fig. 8B), further suggesting a redistribution of BACE1 to the dendritic cell surface.
NCAM2 reduces cell surface levels of BACE1. A, B Cell surface and total BACE1 visualized in cultured NCAM2 + / + and NCAM2−/− hippocampal neurons (A). Higher magnification of dendrites is shown in B. Note higher cell surface BACE1 levels in NCAM2−/− neurons (A, B), and higher numbers of intracellular BACE1 accumulations (arrows) in dendrites of NCAM2 + / + neurons (B). Graphs show mean + SEM ratios of cell surface and total BACE1 labeling intensities along dendrites (A) and densities of intracellular BACE1 accumulations (B). *p, unpaired t test (n = 120). Bar, 20 µm (A), 5 µm (B). C Cell surface and total BACE1 visualized in CHO cells co-transfected with BACE1 and pcDNA3 (control), full-length NCAM2 or NCAM2ΔED and co-labelled for NCAM2-ID. Note reduced cell surface BACE1 levels in NCAM2 and NCAM2ΔED co-transfected cells. Graph shows mean ± SEM ratios of cell surface and total BACE1 levels. *p, one-way ANOVA and Dunnett’s multiple comparisons test, compared to pcDNA3 (n = 37 (pcDNA3), 34 (NCAM2), 46 (NCAM2ΔED)). D Labeling of the cell surface bound pool of BACE1 antibody (surface BACE1) and combined cell surface bound and internalized pool of BACE1 antibody (surface + internalized BACE1) in cultured NCAM2 + / + and NCAM2−/− hippocampal neurons incubated with antibodies against the extracellular domain of BACE1 for 30 min at 37 °C. Graph shows mean + SEM ratios of the cell surface and combined pools (n = 180 (+ / +), 193 (−/−))
In CHO cells transfected with BACE1-FLAG, co-transfection with NCAM2 also led to a reduction in the levels of BACE1 at the cell surface relative to the total levels of BACE1 (Fig. 8C). A similar effect was found in CHO cells co-transfected with NCAM2ΔED (Fig. 8C), indicating that this NCAM2 fragment is sufficient to reduce BACE1 levels at the cell surface.
To analyze whether the accumulation of BACE1 at the cell surface of NCAM2−/− neurons is caused by changes in endocytosis rates, live hippocampal neurons of NCAM2 + / + and NCAM2−/− mice were incubated with antibodies against the extracellular domain of BACE1 for 30 min at 37 °C to allow antibody internalization via endocytosis. Neurons were then gently fixed and cell surface and total pools of BACE1 antibody were visualized with secondary antibodies of different colors applied to neurons before and after permeabilization. Analysis of the confocal images of neurons showed that, while the cell surface levels of BACE1/antibody complexes were higher in NCAM2−/− neurons, the total pool of the BACE1/antibody complexes comprising the cell surface and endocytosed pools was also increased, and the ratio of the cell surface and total levels of BACE1/antibody complexes was similar in NCAM2 + / + and NCAM2−/− neurons (Fig. 8D) indicating similar endocytosis rates.
Our results thus indicate that reduced targeting of BACE1 to Rab11-positive endosomes in NCAM2−/− neurons leads to the accumulation of BACE1 at the cell surface most likely caused by increased recycling of BACE1 from the endosomal compartment to the cell surface.
BACE1 shedding is increased, and its activity is reduced in NCAM2−/− brains
While the cell surface levels of BACE1 were increased in dendrites of NCAM2−/− neurons (Fig. 8A), the total pool of BACE1 detected by immunolabeling of detergent-permeabilized cultured hippocampal neurons with antibodies against BACE1 was reduced in dendrites of NCAM2−/− neurons (Fig. 9A).
Shedding of BACE1 from the cell surface is increased and BACE1 activity is reduced in NCAM2−/− mice. A Total BACE1 visualized in cultured NCAM2 + / + and NCAM2−/− hippocampal neurons. Graph shows mean + SEM labeling intensities of BACE1 in dendrites of neurons (n = 60). *p, Mann–Whitney test. Bar, 10 µm. B Total BACE1 levels in brain homogenates (hom.) and soluble BACE1 levels in soluble protein fractions (soluble) from 2-day-old and 2-month-old NCAM2 + / + , NCAM2+/− and NCAM2−/− mice as detected by Western blot analysis with BACE1-ED antibodies. Actin served as a loading control. NCAM2-ID labeling shows higher levels of NCAM2 in the adult brain. Glycosylated (*) and non-glycosylated (**) BACE1 [56] are enriched in brain homogenates from 2-day-old and 2-month-old mice, respectively. Bands corresponding to soluble BACE1-ED in the soluble protein fraction are ~ 6 kDa smaller than full-length BACE1. A BACE1-immunoreactive band at ~ 110 kDa most likely represents dimers of BACE1-ED [86]. Lower molecular weight degradation products of BACE1 are also detected in the soluble protein fraction. Graph shows mean + SEM change (in fold) of the ratio of soluble vs total BACE1 levels relative to 2-month-old NCAM2 + / + brains set to 100% (n = 6). *p, one sample t test. C Total levels of Sez6 levels in brain homogenates and soluble Sez6 levels in soluble protein fractions from 2-month-old NCAM2 + / + , NCAM2+/− and NCAM2−/− mice as detected by Western blot analysis. GAPDH served as a loading control. Graph shows mean ± SEM change (in fold) in the ratio of soluble and total Sez6 levels relative to + / + brains set to 100% (n = 8 (−/−), 4 (+/−)). *p, one sample t test, compared to + / + . D Dot blot analysis of Aβ42 levels in brain homogenates from 5–6 months-old NCAM2 + / + , NCAM2+/− and NCAM2−/− mice. Samples from each animal were analyzed in triplicates. Graph shows mean ± SEM change (in fold) in Aβ42 levels relative to + / + brains set to 100% (n = 6 mice per group). *p, one sample t-test, compared to + / +
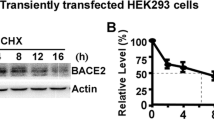
Since BACE1-ED can be shed from the cell surface via proteolytic cleavage of BACE1 near the plasma membrane [29, 30], we asked whether the accumulation of BACE1 at the cell surface of NCAM2−/− dendrites causes increased shedding of BACE1-ED. Western blot analysis with antibodies against BACE1-ED indicated that the ratio of BACE1 cleavage products in the soluble protein fraction versus BACE1 levels in total brain homogenates was increased in 2-month-old NCAM2−/− and NCAM2+/− mice (Fig. 9B), indicating an increase in BACE1 shedding. The levels of the transmembrane NCAM2 isoform increase with age (6.3 ± 1.5-fold higher (n = 3) in the brains of 2-month-old versus 2-day-old NCAM2 + / + mice (Fig. 9B)). The levels of BACE1 shedding were similar in brain homogenates of 2-day-old NCAM2 + / + and NCAM2−/− mice containing predominantly the highly glycosylated form of BACE1 as detected by its mass ~ 65 kDa [55, 56], further indicating that shedding of BACE1 is inversely correlated with NCAM2 levels.
BACE1 is most active in the acidic environment of endosomes and trans-Golgi network, rather than at the cell surface [57]. Furthermore, soluble BACE1 is unable to proteolytically process its substrates [31]. To determine whether an increase in BACE1 cell surface localization and shedding affects the activity of the enzyme, shedding of Sez6, which is shed exclusively via the BACE1-mediated cleavage [58, 59] was analyzed and found to be strongly reduced in the brains of 2-month-old NCAM2−/− mice (Fig. 9C). Since BACE1 is a rate-limiting enzyme in the pathway leading to Aβ generation, the levels of Aβ42 were determined and found to be also strongly reduced in brain homogenates from NCAM2−/− vs NCAM2 + / + mice, and an intermediate effect was found in NCAM2+/− mice (Fig. 9D). We interpret these findings to show that the loss of NCAM2 leads to reduced activity of BACE1.
Our observations thus indicate that NCAM2 deficiency causes a reduction in BACE1 activity, most likely due to its increased shedding from the cell surface.
Discussion
Substrate cleavage by BACE1 is increasingly appreciated to play important roles in modifying or activating the function of substrates [60,61,62,63]. Whether the cleavage products generated by BACE1 regulate BACE1 is however unknown. In our work, we confirm a previous study showing that NCAM2 is a BACE1 substrate [32]. Our data showing that the inhibition of BACE1 in hippocampal neurons results in the reduced release of soluble NCAM2 with the molecular weight corresponding to the size of NCAM2 ectodomain indicates that BACE1 cleaves NCAM2 in these neurons at the membrane-proximal extracellular site as reported previously [32]. Furthermore, we demonstrate that the BACE1-generated NCAM2 fragment and the recombinant fragment of NCAM2 comprising only the transmembrane and intracellular domains associate with and regulate the trafficking and activity of BACE1. These results suggest a scenario in which BACE1 interacts with NCAM2 and cleaves it, with the transmembrane product of this cleavage remaining associated with BACE1 and regulating its activity.
Several lines of evidence indicate that BACE1-mediated NCAM2 cleavage precedes the cleavage of other BACE1 substrates. Delivery of BACE1 to endosomes is required for the interaction with and processing of APP by BACE1 [27]. In contrast, we demonstrate that inhibition of endocytosis does not reduce the interaction of NCAM2 with BACE1. The cleavage of Sez6 by BACE1 most likely also occurs in endosomes since the levels of Sez6 are increased in the brains of Niemann Pick C1 intracellular cholesterol transporter (NPC1) knockout mice, where BACE1 and Sez6 accumulate in recycling endosomes [64]. The cleavage of Sez6 is reduced in brains of NCAM2 deficient mice, indicating that it depends on NCAM2.
BACE1 is found in many different organelle preparations, including the endoplasmic reticulum, Golgi apparatus, cell surface plasma membrane, lysosomes and several types of endosomes [18, 56, 65, 66]. BACE1 activity is higher in acidic environments [67], including the Golgi apparatus [31]. We cannot exclude that BACE1 interacts with NCAM2 and cleaves it early in the biosynthetic pathway since other CAMs are known to interact with cargo proteins and regulate their transport within the biosynthetic pathway [68]. However, this proposition is unlikely since high levels of NCAM2 in the plasma membrane of neurons indicate that the full-length NCAM2 protein reaches the cell surface and therefore does not meet with and is not cleaved by BACE1 on its way to the cell surface. It is however possible that NCAM2 is cleaved by BACE1 at the cell surface, and the suboptimal non-acidic extracellular environment precludes its excessive cleavage by BACE1.
In dendrites, a large pool of BACE1 is detected in the early and recycling endosomes [24, 25, 27]. In non-neuronal cells, cargo from early endosomes can recycle directly to the cell surface via a Rab4-mediated route or can be targeted to the cell surface via a Rab11-mediated recycling route via the perinuclear compartment [69]. Substantial evidence indicates that in neurons Rab11-positive endosomes are involved in the targeting of BACE1 from dendrites to axons through the soma resulting in a concomitant reduction in the dendritic pool of BACE1 available for recycling [24, 25, 70]. Overexpression of the NCAM2 fragment consisting of the transmembrane and intracellular domains of NCAM2 in CHO cells or cultured neurons leads to an increase in BACE1 levels in Rab11-positive endosomes, indicating that the BACE1-generated NCAM2 cleavage product can target BACE1 to Rab11-positive recycling endosomes. An inhibition of the Rab11-dependent transport of BACE1 is also suggested by increased levels of dendritic cell surface BACE1 in NCAM2−/− neurons. This increase cannot be explained by altered endocytosis, since endocytosis rates are similar in NCAM2−/− and NCAM2 + / + neurons, but rather indicates a larger pool of BACE1 to be available for dendritic recycling. In agreement, a similar increase in the cell surface BACE1 levels was found when the sorting of BACE1 from early to recycling endosomes was blocked by the S498A mutation leading to BACE1 accumulation in early endosomes [66].
The proteins destined for different types of endosomes are sorted at the plasma membrane before endocytosis [71, 72]. Our results suggest that NCAM2 associates with BACE1 at the plasma membrane and mediates molecular sorting that leads to the transport of BACE1 to Rab11-positive endosomes. It may also be possible that NCAM2 promotes the transport of BACE1 from early endosomes to Rab11-positive recycling endosomes. Since Rab11-positive recycling endosomes also deliver cargo to the dendritic surface [53, 54], NCAM2 could promote the accumulation of BACE1 in Rab11-positive endosomes by inhibiting the fusion of these endosomes with the dendritic cell surface, thus retaining BACE1 in these endosomes. Finally, NCAM2 can target BACE1 from the plasma membrane directly to Rab11-positive recycling endosomes, thereby bypassing the passage through early endosomes. Recycling endosomes fuse with the dendritic plasma membrane in two modes, either via “full fusion”, which releases all cargo to the plasma membrane, or “display fusion”, in which only a small pore connects the endosome with the plasma membrane, while soluble proteins are released via the pore, with the transmembrane molecules being retained in the endosome [73, 74]. Experiments in non-neuronal cells indicate that transmembrane proteins can be targeted to the recycling endosomes not only via early endosomes but also independently of the early endosomes [75] possibly by recruitment from the plasma membrane to the recycling endosomes after fusion of the endosome with the plasma membrane. Interestingly, full fusion of recycling endosomes with the cell surface is promoted by signaling through L-type voltage-gated calcium channels [73], which are activated by NCAM2 [6, 9]. NCAM2 may therefore act to bias BACE1 transport via a route which bypasses early endosomes and preferentially targets BACE1 to recycling endosomes from the plasma membrane. Future work should distinguish these possibilities.
We demonstrate that NCAM2 deficiency causes increased cell surface accumulation of BACE1, which correlates with higher shedding of BACE1 in adult NCAM2 deficient mice. The shedding of the BACE1 ectodomain is mediated by the metalloproteinase ADAM10, which functions at the neuronal cell surface [30, 76]. We, therefore, propose that recycling of BACE1 to the dendritic cell surface leads to an increase in the cell surface pool of BACE1 in NCAM2−/− neurons, thereby likely facilitating the ADAM10-dependent shedding of BACE1. Whether changes in the BACE1 shedding and inactivation are coupled to changes in NCAM2 expression levels in the broad spectrum of neuronal functions will be interesting to investigate.
BACE1 activity is increased in the brains of individuals with Alzheimer’s disease [77, 78] and Down syndrome [79]. The levels of NCAM2 are also increased in brains of individuals with AD [11]. NCAM2 is encoded by a gene on chromosome 21 and is also overexpressed in individuals with Down’s syndrome [15]. The results of our present study indicate that an increase in NCAM2 levels can lead to enhanced BACE1 activity. Enhanced processing of BACE1 substrates via increased targeting of BACE1 to Rab11-positive endosomes can also explain a functional link between NCAM2 and Alzheimer’s disease as suggested by studies showing that single nucleotide polymorphisms in the NCAM2 gene are associated with increased risk of late-onset AD [16] and increased levels of amyloid protein β in the cerebrospinal fluid [17].
NCAM2 deficiency causes neurodevelopmental disorders in humans via mechanisms that remain poorly understood. The current results suggest that, at least partially, abnormalities in brain development and function caused by NCAM2 deficiency can be linked to the reduced function of BACE1, which has been suggested to be associated with changes in synaptic plasticity [80], memory deficits [48, 81], spontaneous seizures [82], defective myelination [83] and guidance of axons [84, 85]. We propose that NCAM2-mediated pathways of BACE1 expression are interesting to investigate as therapeutic targets for treating conditions associated with NCAM2 deficiency or overexpression in humans.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- Aβ:
-
Amyloid β
- AD:
-
Alzheimer’s disease
- ANOVA:
-
Analysis of variance
- APP:
-
Amyloid precursor protein
- BACE1:
-
Beta-site amyloid precursor protein cleaving enzyme 1
- BACE1-VC:
-
VC-tagged BACE1
- BiFC:
-
Bimolecular fluorescence complementation
- BSA:
-
Bovine serum albumin
- CHO cells:
-
Chinese hamster ovary cells
- DMSO:
-
Dimethyl sulfoxide
- DNA:
-
Deoxyribonucleic acid
- ED:
-
Extracellular domain
- EDTA:
-
Ethylenediaminetetraacetic acid
- EGTA:
-
Ethylene glycol-bis(β-aminoethyl ether)-N,N,N′,N′-tetraacetic acid)
- EYFP:
-
Enhanced yellow fluorescent protein
- FGF-2:
-
Fibroblast growth factor-2
- Fn:
-
Fibronectin type III homologous repeat
- GAPDH:
-
Glyceraldehyde 3-phosphate dehydrogenase
- GFP:
-
Green fluorescent protein
- HA tag:
-
Human influenza hemagglutinin tag
- HBS:
-
HEPES buffered saline
- HEPES:
-
4-(2-Hydroxyethyl)-1-piperazineethanesulfonic acid
- HRP:
-
Horseradish peroxidase
- IC:
-
Immunocytochemistry
- ID:
-
Intracellular domain
- Ig:
-
Immunoglobulin-like
- LAMP1:
-
Lysosomal-associated membrane protein 1
- min:
-
Minute
- NCAM2:
-
Neural cell adhesion molecule 2
- NCAM2ΔED:
-
NCAM2 fragment comprising transmembrane and intracellular domains of NCAM2
- NCAM2-VN:
-
VN-tagged NCAM2
- NCAM2ΔED-VN:
-
VN-tagged NCAM2ΔED
- PBS:
-
Phosphate-buffered saline
- PL:
-
Proximity ligation
- PMSF:
-
Phenylmethylsulfonyl fluoride
- PVDF:
-
Polyvinylidene difluoride
- siRNAs:
-
Small interfering RNAs
- SNPs:
-
Single nucleotide polymorphisms
- TBS:
-
Tris-buffered saline
- VC:
-
C-terminal amino acid residues 155–238 of the fluorescent protein Venus
- VN:
-
N-terminal 155 amino acid residues of the fluorescent protein Venus
- WB:
-
Western blot
References
Sytnyk V, Leshchyns’ka I, Schachner M (2017) Neural cell adhesion molecules of the immunoglobulin superfamily regulate synapse formation, maintenance, and function. Trends Neurosci 40:295–308
Paoloni-Giacobino A, Chen H, Antonarakis SE (1997) Cloning of a novel human neural cell adhesion molecule gene (NCAM2) that maps to chromosome region 21q21 and is potentially involved in Down syndrome. Genomics 43:43–51
Yoshihara Y, Kawasaki M, Tamada A et al (1997) OCAM: A new member of the neural cell adhesion molecule family related to zone-to-zone projection of olfactory and vomeronasal axons. J Neurosci 17:5830–5842
Alenius M, Bohm S (1997) Identification of a novel neural cell adhesion molecule-related gene with a potential role in selective axonal projection. J Biol Chem 272:26083–26086
Parcerisas A, Pujadas L, Ortega-Gasco A et al (2020) NCAM2 regulates dendritic and axonal differentiation through the cytoskeletal proteins MAP2 and 14-3-3. Cereb Cortex 30:3781–3799
Sheng L, Leshchyns’ka I, Sytnyk V (2015) Neural cell adhesion molecule 2 promotes the formation of filopodia and neurite branching by inducing submembrane increases in Ca2+ levels. J Neurosci 35:1739–1752
Walz A, Mombaerts P, Greer CA et al (2006) Disrupted compartmental organization of axons and dendrites within olfactory glomeruli of mice deficient in the olfactory cell adhesion molecule, OCAM. Mol Cel Neurosci 32:1–14
Alenius M, Bohm S (2003) Differential function of RNCAM isoforms in precise target selection of olfactory sensory neurons. Development 130:917–927
Sheng L, Leshchyns’ka I, Sytnyk V (2019) Neural cell adhesion molecule 2 (NCAM2)-induced c-Src-dependent propagation of submembrane Ca2+ spikes along dendrites inhibits synapse maturation. Cereb Cortex 29:1439–1459
Kulahin N, Kristensen O, Rasmussen KK et al (2011) Structural model and trans-interaction of the entire ectodomain of the olfactory cell adhesion molecule. Structure 19:203–211
Leshchyns’ka I, Liew HT, Shepherd C et al (2015) Abeta-dependent reduction of NCAM2-mediated synaptic adhesion contributes to synapse loss in Alzheimer’s disease. Nat Commun 6:8836
Petit F, Plessis G, Decamp M et al (2015) 21q21 deletion involving NCAM2: report of 3 cases with neurodevelopmental disorders. Eur J Med Genet 58:44–46
Haldeman-Englert CR, Chapman KA, Kruger H et al (2010) A de novo 8.8-Mb deletion of 21q21.1-q21.3 in an autistic male with a complex rearrangement involving chromosomes 6, 10, and 21. Am J Med Genet A 152A:196–202
Scholz C, Steinemann D, Malzer M et al (2016) NCAM2 deletion in a boy with macrocephaly and autism: cause, association or predisposition? Eur J Med Genet 59:493–498
Lockstone HE, Harris LW, Swatton JE et al (2007) Gene expression profiling in the adult down syndrome brain. Genomics 90:647–660
Kimura R, Kamino K, Yamamoto M et al (2007) The DYRK1A gene, encoded in chromosome 21 Down syndrome critical region, bridges between beta-amyloid production and tau phosphorylation in Alzheimer disease. Hum Mol Genet 16:15–23
Han MR, Schellenberg GD, Wang LS (2010) Genome-wide association reveals genetic effects on human Abeta42 and tau protein levels in cerebrospinal fluids: a case control study. BMC Neurol 10:90
Huse JT, Pijak DS, Leslie GJ et al (2000) Maturation and endosomal targeting of beta-site amyloid precursor protein-cleaving enzyme. The Alzheimer’s disease beta-secretase. J Biol Chem 275:33729–33737
Tan JZA, Fourriere L, Wang J et al (2020) Distinct anterograde trafficking pathways of BACE1 and amyloid precursor protein from the TGN and the regulation of amyloid-beta production. Mol Biol Cell 31:27–44
Chia PZ, Toh WH, Sharples R et al (2013) Intracellular itinerary of internalised beta-secretase, BACE1, and its potential impact on beta-amyloid peptide biogenesis. Traffic 14:997–1013
Koh YH, Von Arnim CA, Hyman BT et al (2005) BACE is degraded via the lysosomal pathway. J Biol Chem 280:32499–32504
Wahle T, Prager K, Raffler N et al (2005) GGA proteins regulate retrograde transport of BACE1 from endosomes to the trans-Golgi network. Mol Cell Neurosci 29:453–461
Wu J, Petralia RS, Kurushima H et al (2011) Arc/Arg3.1 regulates an endosomal pathway essential for activity-dependent beta-amyloid generation. Cell 147:615–628
Buggia-Prevot V, Fernandez CG, Riordan S et al (2014) Axonal BACE1 dynamics and targeting in hippocampal neurons: a role for Rab11 GTPase. Mol Neurodegener 9:1
Buggia-Prevot V, Fernandez CG, Udayar V et al (2013) A function for EHD family proteins in unidirectional retrograde dendritic transport of BACE1 and Alzheimer’s disease Abeta production. Cell Rep 5:1552–1563
Ye X, Feng T, Tammineni P et al (2017) Regulation of synaptic amyloid-beta generation through BACE1 retrograde transport in a mouse model of Alzheimer’s Disease. J Neurosci 37:2639–2655
Das U, Scott DA, Ganguly A et al (2013) Activity-induced convergence of APP and BACE-1 in acidic microdomains via an endocytosis-dependent pathway. Neuron 79:447–460
Udayar V, Buggia-Prévot V, Guerreiro RL et al (2013) A paired RNAi and RabGAP overexpression screen identifies Rab11 as a regulator of β-amyloid production. Cell Rep 5:1536–1551
Benjannet S, Elagoz A, Wickham L et al (2001) Post-translational processing of beta-secretase (beta-amyloid-converting enzyme) and its ectodomain shedding. The pro- and transmembrane/cytosolic domains affect its cellular activity and amyloid-beta production. J Biol Chem 276:10879–10887
Hussain I, Hawkins J, Shikotra A et al (2003) Characterization of the ectodomain shedding of the beta-site amyloid precursor protein-cleaving enzyme 1 (BACE1). J Biol Chem 278:36264–36268
Yan R, Han P, Miao H et al (2001) The transmembrane domain of the Alzheimer’s β-secretase (BACE1) determines its late Golgi localization and access to β-amyloid precursor protein (APP) substrate. J Biol Chem 276:36788–36796
Kim W, Watanabe H, Lomoio S et al (2021) Spatiotemporal processing of neural cell adhesion molecules 1 and 2 by BACE1 in vivo. J Biol Chem 296:100372
Lombardo S, Chiacchiaretta M, Tarr A et al (2019) BACE1 partial deletion induces synaptic plasticity deficit in adult mice. Sci Rep 9:19877
Gunnersen JM, Kim MH, Fuller SJ et al (2007) Sez-6 proteins affect dendritic arborization patterns and excitability of cortical pyramidal neurons. Neuron 56:621–639
Das U, Wang L, Ganguly A et al (2016) Visualizing APP and BACE-1 approximation in neurons yields insight into the amyloidogenic pathway. Nat Neurosci 19:55–64
Choudhury A, Dominguez M, Puri V et al (2002) Rab proteins mediate Golgi transport of caveola-internalized glycosphingolipids and correct lipid trafficking in Niemann-Pick C cells. J Clin Investigation 109:1541–1550
Du X, Kumar J, Ferguson C et al (2011) A role for oxysterol-binding protein-related protein 5 in endosomal cholesterol trafficking. J Cell Biol 192:121–135
Ma W, Mayr C (2018) A membraneless organelle associated with the endoplasmic reticulum enables 3’UTR-mediated protein-protein interactions. Cell 175(1492–1506):e1419
Shetty A, Sytnyk V, Leshchyns’ka I et al (2013) The neural cell adhesion molecule promotes maturation of the presynaptic endocytotic machinery by switching synaptic vesicle recycling from adaptor protein 3 (AP-3)- to AP-2-dependent mechanisms. J Neurosci 33:16828–16845
Kodama Y, Hu CD (2010) An improved bimolecular fluorescence complementation assay with a high signal-to-noise ratio. Biotechniques 49:793–805
Wobst H, Schmitz B, Schachner M et al (2015) Kinesin-1 promotes post-Golgi trafficking of NCAM140 and NCAM180 to the cell surface. J Cell Sci 128:2816–2829
Jiang M, Chen G (2006) High Ca2+-phosphate transfection efficiency in low-density neuronal cultures. Nat Protoc 1:695–700
Bliim N, Leshchyns’ka I, Keable R et al (2019) Early transcriptome changes in response to chemical long-term potentiation induced via activation of synaptic NMDA receptors in mouse hippocampal neurons. Genomics 111:1676–1686
Tian N, Leshchyns’ka I, Welch JH et al (2012) Lipid raft-dependent endocytosis of close homolog of adhesion molecule L1 (CHL1) promotes neuritogenesis. J Biol Chem 287:44447–44463
Andreyeva A, Leshchyns’ka I, Knepper M et al (2010) CHL1 is a selective organizer of the presynaptic machinery chaperoning the SNARE complex. PLoS ONE 5:e12018
Scott DA, Das U, Tang Y et al (2011) Mechanistic logic underlying the axonal transport of cytosolic proteins. Neuron 70:441–454
Voytyuk I, Mueller SA, Herber J et al (2018) BACE2 distribution in major brain cell types and identification of novel substrates. Life Sci Alliance 1:e201800026
Laird FM, Cai H, Savonenko AV et al (2005) BACE1, a major determinant of selective vulnerability of the brain to amyloid-beta amyloidogenesis, is essential for cognitive, emotional, and synaptic functions. J Neurosci 25:11693–11709
Nunan J, Williamson NA, Hill AF et al (2003) Proteasome-mediated degradation of the C-terminus of the Alzheimer’s disease beta-amyloid protein precursor: effect of C-terminal truncation on production of beta-amyloid protein. J Neurosci Res 74:378–385
Sharples RA, Vella LJ, Nisbet RM et al (2008) Inhibition of gamma-secretase causes increased secretion of amyloid precursor protein C-terminal fragments in association with exosomes. FASEB J 22:1469–1478
Walsh DM, Klyubin I, Fadeeva JV et al (2002) Naturally secreted oligomers of amyloid beta protein potently inhibit hippocampal long-term potentiation in vivo. Nature 416:535–539
Liu X, Fagotto F (2011) A method to separate nuclear, cytosolic, and membrane-associated signaling molecules in cultured cells. Sci Signal 4:2
Esteves Da Silva M, Adrian M, Schatzle P et al (2015) Positioning of AMPA receptor-containing endosomes regulates synapse architecture. Cell Rep 13:933–943
Wang Z, Edwards JG, Riley N et al (2008) Myosin Vb mobilizes recycling endosomes and AMPA receptors for postsynaptic plasticity. Cell 135:535–548
Kizuka Y, Kitazume S, Fujinawa R et al (2015) An aberrant sugar modification of BACE1 blocks its lysosomal targeting in Alzheimer’s disease. EMBO Mol Med 7:175–189
Capell A, Steiner H, Willem M et al (2000) Maturation and pro-peptide cleavage of beta-secretase. J Biol Chem 275:30849–30854
Cole SL, Vassar R (2007) The basic biology of BACE1: a key therapeutic target for Alzheimer’s disease. Curr Genom 8:509–530
Pigoni M, Wanngren J, Kuhn PH et al (2016) Seizure protein 6 and its homolog seizure 6-like protein are physiological substrates of BACE1 in neurons. Mol Neurodegener 11:67
Kuhn PH, Koroniak K, Hogl S et al (2012) Secretome protein enrichment identifies physiological BACE1 protease substrates in neurons. EMBO J 31:3157–3168
Hampel H, Vassar R, De Strooper B et al (2021) The β-secretase BACE1 in Alzheimer’s disease. Biol Psychiatry 89:745–756
Zhu K, Xiang X, Filser S et al (2018) Beta-site amyloid precursor protein cleaving enzyme 1 inhibition impairs synaptic plasticity via seizure protein 6. Biol Psychiatry 83:428–437
Barão S, Gärtner A, Leyva-Díaz E et al (2015) Antagonistic effects of BACE1 and APH1B-γ-secretase control axonal guidance by regulating growth cone collapse. Cell Rep 12:1367–1376
Willem M, Garratt AN, Novak B et al (2006) Control of peripheral nerve myelination by the ß-secretase BACE1. Science 314:664–666
Causevic M, Dominko K, Malnar M et al (2018) BACE1-cleavage of Sez6 and Sez6L is elevated in Niemann-Pick type C disease mouse brains. PLoS ONE 13:e0200344
Ye X, Cai QJCR (2014) Snapin-mediated BACE1 retrograde transport is essential for its degradation in lysosomes and regulation of APP processing in neurons. Cell Rep 6:24–31
Toh WH, Chia PZC, Hossain MI et al (2018) GGA1 regulates signal-dependent sorting of BACE1 to recycling endosomes, which moderates Aβ production. Mol Biol Cell 29:191–208
Vassar R, Kovacs DM, Yan R et al (2009) The β-secretase enzyme BACE in health and Alzheimer’s disease: regulation, cell biology, function, and therapeutic potential. J Neurosci 29:12787–12794
Keable R, Leshchyns KI, Sytnyk VJTN (2020) Trafficking and activity of glutamate and GABA receptors: regulation by cell adhesion molecules. Neuroscientist 26:415–437
Li X, Difiglia MJPIN (2012) The recycling endosome and its role in neurological disorders. Prog Neurobiol 97:127–141
Buggia-Prevot V, Thinakaran G (2015) Significance of transcytosis in Alzheimer’s disease: BACE1 takes the scenic route to axons. BioEssays 37:888–898
Leonard D, Hayakawa A, Lawe D et al (2008) Sorting of EGF and transferrin at the plasma membrane and by cargo-specific signaling to EEA1-enriched endosomes. J Cell Sci 121:3445–3458
Lakadamyali M, Rust MJ, Zhuang X (2006) Ligands for clathrin-mediated endocytosis are differentially sorted into distinct populations of early endosomes. Cell 124:997–1009
Hiester BG, Bourke AM, Sinnen BL et al (2017) L-type voltage-gated Ca2+ channels regulate synaptic activity-triggered recycling endosome fusion in neuronal dendrites. Cell Rep 21:2134–2146
Jullie D, Choquet D, Perrais D (2014) Recycling endosomes undergo rapid closure of a fusion pore on exocytosis in neuronal dendrites. J Neurosci 34:11106–11118
Puri C, Renna M, Bento CF et al (2013) Diverse autophagosome membrane sources coalesce in recycling endosomes. Cell 154:1285–1299
Murayama KS, Kametani F, Araki W (2005) Extracellular release of BACE1 holoproteins from human neuronal cells. Biochem Biophys Res Commun 338:800–807
Fukumoto H, Cheung BS, Hyman BT et al (2002) β-Secretase protein and activity are increased in the neocortex in Alzheimer disease. Arch Neurol 59:1381–1389
Yang L-B, Lindholm K, Yan R et al (2003) Elevated β-secretase expression and enzymatic activity detected in sporadic Alzheimer disease. Nat Med 9:3–4
Sun X, Tong Y, Qing H et al (2006) Increased BACE1 maturation contributes to the pathogenesis of Alzheimer’s disease in Down syndrome. FASEB J 20:1361–1368
Wang H, Song L, Laird F et al (2008) BACE1 knock-outs display deficits in activity-dependent potentiation of synaptic transmission at mossy fiber to CA3 synapses in the hippocampus. J Neurosci 28:8677–8681
Das B, Singh N, Yao AY et al (2021) BACE1 controls synaptic function through modulating release of synaptic vesicles. Mol Psychiatry 26:6394–6410
Hu X, Zhou X, He W et al (2010) BACE1 deficiency causes altered neuronal activity and neurodegeneration. J Neurosci 30:8819–8829
Hu X, Hicks CW, He W et al (2006) Bace1 modulates myelination in the central and peripheral nervous system. Nat Neurosci 9:1520–1525
Rajapaksha TW, Eimer WA, Bozza TC et al (2011) The Alzheimer’s β-secretase enzyme BACE1 is required for accurate axon guidance of olfactory sensory neurons and normal glomerulus formation in the olfactory bulb. Mol Neurodegener 6:1–9
Hitt B, Riordan SM, Kukreja L et al (2012) beta-Site amyloid precursor protein (APP)-cleaving enzyme 1 (BACE1)-deficient mice exhibit a close homolog of L1 (CHL1) loss-of-function phenotype involving axon guidance defects. J Biol Chem 287:38408–38425
Schmechel A, Strauss M, Schlicksupp A et al (2004) Human BACE forms dimers and colocalizes with APP. J Biol Chem 279:39710–39717
Acknowledgements
We are grateful to Subhojit Roy (University of California, San Diego) for the BACE1-VC construct and comments on the manuscript. The study was supported by the National Health and Medical Research Council (VS, APP1129869). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
VS and IL designed the project. RK, SH, GP, IK, FS, IL and VS designed and performed experiments. All authors were involved in data analysis. VS wrote the manuscript, which was then edited by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Keable, R., Hu, S., Pfundstein, G. et al. The BACE1-generated C-terminal fragment of the neural cell adhesion molecule 2 (NCAM2) promotes BACE1 targeting to Rab11-positive endosomes. Cell. Mol. Life Sci. 79, 555 (2022). https://doi.org/10.1007/s00018-022-04575-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04575-w