Abstract
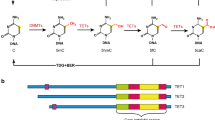
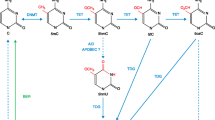
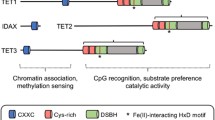
The ten–eleven translocation (TET) family of dioxygenases consists of three members, TET1, TET2, and TET3. All three TET enzymes have Fe+2 and α-ketoglutarate (α-KG)-dependent dioxygenase activities, catalyzing the 1st step of DNA demethylation by converting 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC), and further oxidize 5hmC to 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC). Gene knockout studies demonstrated that all three TET proteins are involved in the regulation of fetal organ generation during embryonic development and normal tissue generation postnatally. TET proteins play such roles by regulating the expression of key differentiation and fate-determining genes via (1) enzymatic activity-dependent DNA methylation of the promoters and enhancers of target genes; and (2) enzymatic activity-independent regulation of histone modification. Interacting partner proteins and post-translational regulatory mechanisms regulate the activities of TET proteins. Mutations and dysregulation of TET proteins are involved in the pathogenesis of human diseases, specifically cancers. Here, we summarize the research on the interaction partners and post-translational modifications of TET proteins. We also discuss the molecular mechanisms by which these partner proteins and modifications regulate TET functioning and target gene expression. Such information will help in the design of medications useful for targeted therapy of TET-mutant-related diseases.





Similar content being viewed by others
Availability of data and materials
This is not applicable for this review.
Code availability
This is not applicable for this review.
Abbreviations
- TET:
-
Ten–eleven translocation
- α-KG:
-
α-Ketoglutarate
- 5mC:
-
5-Methylcytosine
- 5hmC:
-
5-Hydroxymethylcytosine
- 5fC:
-
5-Formylcytosine
- 5caC:
-
5-Carboxylcytosine
- TFs:
-
Transcription factors
- ARCH:
-
Age-related clonal hematopoiesis
- MDS:
-
Myelodysplastic syndromes
- AML:
-
Acute myeloid leukemia
- ALL:
-
Acute lymphoblastic leukemia
- DLBCLs:
-
Diffuse large B-cell lymphomas
- PTCL:
-
Peripheral T-cell lymphoma
- IFN-γ:
-
Interferon-γ
- TNF-α:
-
Tumor necrosis factor-α
- TIS:
-
Transcriptional initiation sites
- DSBH:
-
Double-stranded beta-helix domain
- TDG:
-
Thymine DNA glycosylase
- BER:
-
Base excision repair
- ES:
-
Embryonic stem cells
- IDHs:
-
Isocitrate dehydrogenases
- AP site:
-
Apyrimidinic site
- AID:
-
Activation-induced cytidine deaminase
- 5hmU:
-
5-Hydroxymethyluracil
- GC:
-
Germinal center
- CSR:
-
Class-switch recombination
- DSBs:
-
Double-strand breaks
- PGCs:
-
Primordial germ cells
- CGIs:
-
CpG islands
- OGT:
-
O-linked GlcNAc transferase
- SID:
-
Sin3A interacts with the Sin3-interaction domain
- CoA:
-
Coactivators
- PTM:
-
Post-translational modifications
- AMPK:
-
AMP-activated protein kinase
- C/EBPα:
-
CCAAT/enhancer-binding protein alpha
- KLF4:
-
Kruppel-like factor-4
- TFCP2l1:
-
Transcription factor CP2 like-1
- YBX1:
-
Y box-binding protein-1
- FOXK2:
-
Forkhead box protein K-2
- KZF1:
-
DNA-binding protein Ikaros 1
- NFIL3:
-
Nuclear factor interleukin-3-regulated protein
- ATRX:
-
Alpha-thalassemia/mental retardation syndrome X-linked transcriptional regulator
- CUX1:
-
Homeobox protein cut-like-1
- YY2:
-
Yin and yang-2 transcription factor
- IκBζ:
-
Inhibitory-kappa-B-zeta
References
McKinney-Freeman S et al (2012) The transcriptional landscape of hematopoietic stem cell ontogeny. Cell Stem Cell 11:701–714. https://doi.org/10.1016/j.stem.2012.07.018
Long HK, Prescott SL, Wysocka J (2016) Ever-changing landscapes: transcriptional enhancers in development and evolution. Cell 167:1170–1187. https://doi.org/10.1016/j.cell.2016.09.018
Kulis M et al (2015) Whole-genome fingerprint of the DNA methylome during human B cell differentiation. Nat Genet 47:746–756. https://doi.org/10.1038/ng.3291
Barwick BG, Scharer CD, Bally APR, Boss JM (2016) Plasma cell differentiation is coupled to division-dependent DNA hypomethylation and gene regulation. Nat Immunol 17:1216–1225. https://doi.org/10.1038/ni.3519
Smith ZD, Meissner A (2013) DNA methylation: roles in mammalian development. Nat Rev Genet 14:204–220. https://doi.org/10.1038/nrg3354
Apostolou E, Hochedlinger K (2013) Chromatin dynamics during cellular reprogramming. Nature 502:462–471. https://doi.org/10.1038/nature12749
Yin Y et al (2017) Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science. https://doi.org/10.1126/science.aaj2239
Schubeler D (2015) Function and information content of DNA methylation. Nature 517:321–326. https://doi.org/10.1038/nature14192
Hu S et al (2013) DNA methylation presents distinct binding sites for human transcription factors. Elife 2:e00726. https://doi.org/10.7554/eLife.00726
Klose RJ, Bird AP (2006) Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci 31:89–97. https://doi.org/10.1016/j.tibs.2005.12.008
Greenberg MVC, Bourc’his D (2019) The diverse roles of DNA methylation in mammalian development and disease. Nat Rev Mol Cell Biol 20:590–607. https://doi.org/10.1038/s41580-019-0159-6
Yamazaki J et al (2015) TET2 mutations affect non-CpG island DNA methylation at enhancers and transcription factor-binding sites in chronic myelomonocytic leukemia. Cancer Res 75:2833–2843. https://doi.org/10.1158/0008-5472.CAN-14-0739
Lea AJ et al (2018) Genome-wide quantification of the effects of DNA methylation on human gene regulation. Elife. https://doi.org/10.7554/eLife.37513
Rasmussen KD et al (2019) TET2 binding to enhancers facilitates transcription factor recruitment in hematopoietic cells. Genome Res 29:564–575. https://doi.org/10.1101/gr.239277.118
Chen Z, Zhang Y (2020) Role of mammalian DNA methyltransferases in development. Annu Rev Biochem 89:135–158. https://doi.org/10.1146/annurev-biochem-103019-102815
Wu X, Zhang Y (2017) TET-mediated active DNA demethylation: mechanism, function and beyond. Nat Rev Genet 18:517–534. https://doi.org/10.1038/nrg.2017.33
Broome R et al (2021) TET2 is a component of the estrogen receptor complex and controls 5mC to 5hmC conversion at estrogen receptor cis-regulatory regions. Cell Rep 34:108776. https://doi.org/10.1016/j.celrep.2021.108776
Cruz-Rodriguez N, Combita AL, Zabaleta J (2018) Epigenetics in hematological malignancies. Methods Mol Biol 1856:87–101. https://doi.org/10.1007/978-1-4939-8751-1_5
Hu D, Shilatifard A (2016) Epigenetics of hematopoiesis and hematological malignancies. Genes Dev 30:2021–2041. https://doi.org/10.1101/gad.284109.116
Jiang Y, Dominguez PM, Melnick AM (2016) The many layers of epigenetic dysfunction in B-cell lymphomas. Curr Opin Hematol 23:377–384. https://doi.org/10.1097/MOH.0000000000000249
Abdel-Wahab O et al (2009) Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood 114:144–147. https://doi.org/10.1182/blood-2009-03-210039
Delhommeau F et al (2009) Mutation in TET2 in myeloid cancers. N Engl J Med 360:2289–2301. https://doi.org/10.1056/NEJMoa0810069
Smith AE et al (2010) Next-generation sequencing of the TET2 gene in 355 MDS and CMML patients reveals low-abundance mutant clones with early origins, but indicates no definite prognostic value. Blood 116:3923–3932. https://doi.org/10.1182/blood-2010-03-274704
Kosmider O et al (2009) TET2 gene mutation is a frequent and adverse event in chronic myelomonocytic leukemia. Haematologica 94:1676–1681. https://doi.org/10.3324/haematol.2009.011205
Swierczek SI et al (2011) Extent of hematopoietic involvement by TET2 mutations in JAK2V(6)(1)(7)F polycythemia vera. Haematologica 96:775–778. https://doi.org/10.3324/haematol.2010.029678
Tefferi A et al (2009) TET2 mutations and their clinical correlates in polycythemia vera, essential thrombocythemia and myelofibrosis. Leukemia 23:905–911. https://doi.org/10.1038/leu.2009.47
Tefferi A et al (2009) Detection of mutant TET2 in myeloid malignancies other than myeloproliferative neoplasms: CMML, MDS, MDS/MPN and AML. Leukemia 23:1343–1345. https://doi.org/10.1038/leu.2009.59
Jankowska AM et al (2009) Loss of heterozygosity 4q24 and TET2 mutations associated with myelodysplastic/myeloproliferative neoplasms. Blood 113:6403–6410. https://doi.org/10.1182/blood-2009-02-205690
Langemeijer SM et al (2009) Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat Genet 41:838–842. https://doi.org/10.1038/ng.391
Asmar F et al (2013) Genome-wide profiling identifies a DNA methylation signature that associates with TET2 mutations in diffuse large B-cell lymphoma. Haematologica 98:1912–1920. https://doi.org/10.3324/haematol.2013.088740
Reddy A et al (2017) Genetic and functional drivers of diffuse large B cell lymphoma. Cell 171:481–494. https://doi.org/10.1016/j.cell.2017.09.027 (e415)
Dominguez PM et al (2018) TET2 deficiency causes germinal center hyperplasia, impairs plasma cell differentiation, and promotes B-cell lymphomagenesis. Cancer Discov 8:1632–1653. https://doi.org/10.1158/2159-8290.CD-18-0657
Li Z et al (2011) Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 118:4509–4518. https://doi.org/10.1182/blood-2010-12-325241
Lemonnier F et al (2012) Recurrent TET2 mutations in peripheral T-cell lymphomas correlate with TFH-like features and adverse clinical parameters. Blood 120:1466–1469. https://doi.org/10.1182/blood-2012-02-408542
Sakata-Yanagimoto M et al (2014) Somatic RHOA mutation in angioimmunoblastic T cell lymphoma. Nat Genet 46:171–175. https://doi.org/10.1038/ng.2872
Odejide O et al (2014) A targeted mutational landscape of angioimmunoblastic T-cell lymphoma. Blood 123:1293–1296. https://doi.org/10.1182/blood-2013-10-531509
Palomero T et al (2014) Recurrent mutations in epigenetic regulators, RHOA and FYN kinase in peripheral T cell lymphomas. Nat Genet 46:166–170. https://doi.org/10.1038/ng.2873
Schmitz R et al (2018) Genetics and pathogenesis of diffuse large B-cell lymphoma. N Engl J Med 378:1396–1407. https://doi.org/10.1056/NEJMoa1801445
Soucie E et al (2012) In aggressive forms of mastocytosis, TET2 loss cooperates with c-KITD816V to transform mast cells. Blood 120:4846–4849. https://doi.org/10.1182/blood-2011-12-397588
Tefferi A et al (2009) Frequent TET2 mutations in systemic mastocytosis: clinical, KITD816V and FIP1L1-PDGFRA correlates. Leukemia 23:900–904. https://doi.org/10.1038/leu.2009.37
Coltro G et al (2020) Clinical, molecular, and prognostic correlates of number, type, and functional localization of TET2 mutations in chronic myelomonocytic leukemia (CMML)-a study of 1084 patients. Leukemia 34:1407–1421. https://doi.org/10.1038/s41375-019-0690-7
Yao WQ et al (2020) Angioimmunoblastic T-cell lymphoma contains multiple clonal T-cell populations derived from a common TET2 mutant progenitor cell. J Pathol 250:346–357. https://doi.org/10.1002/path.5376
Shih AH, Abdel-Wahab O, Patel JP, Levine RL (2012) The role of mutations in epigenetic regulators in myeloid malignancies. Nat Rev Cancer 12:599–612. https://doi.org/10.1038/nrc3343
Xie M et al (2014) Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med 20:1472–1478. https://doi.org/10.1038/nm.3733
Jaiswal S et al (2014) Age-related clonal hematopoiesis associated with adverse outcomes. N Engl J Med 371:2488–2498. https://doi.org/10.1056/NEJMoa1408617
Genovese G et al (2014) Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med 371:2477–2487. https://doi.org/10.1056/NEJMoa1409405
Zhang CRC et al (2019) Inflammatory cytokines promote clonal hematopoiesis with specific mutations in ulcerative colitis patients. Exp Hematol 80:36–41. https://doi.org/10.1016/j.exphem.2019.11.008 (e33)
Hormaechea-Agulla D et al (2021) Chronic infection drives Dnmt3a-loss-of-function clonal hematopoiesis via IFNgamma signaling. Cell Stem Cell. https://doi.org/10.1016/j.stem.2021.03.002
Buscarlet M et al (2017) DNMT3A and TET2 dominate clonal hematopoiesis and demonstrate benign phenotypes and different genetic predispositions. Blood 130:753–762. https://doi.org/10.1182/blood-2017-04-777029
Jaiswal S, Ebert BL (2019) Clonal hematopoiesis in human aging and disease. Science. https://doi.org/10.1126/science.aan4673
Steensma DP et al (2015) Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126:9–16. https://doi.org/10.1182/blood-2015-03-631747
Sun D et al (2014) Epigenomic profiling of young and aged HSCs reveals concerted changes during aging that reinforce self-renewal. Cell Stem Cell 14:673–688. https://doi.org/10.1016/j.stem.2014.03.002
Zhang W et al (2016) Isoform switch of TET1 regulates DNA demethylation and mouse development. Mol Cell 64:1062–1073. https://doi.org/10.1016/j.molcel.2016.10.030
Good CR et al (2017) A novel isoform of TET1 that lacks a CXXC domain is overexpressed in cancer. Nucleic Acids Res 45:8269–8281. https://doi.org/10.1093/nar/gkx435
Iyer LM, Abhiman S, Aravind L (2011) Natural history of eukaryotic DNA methylation systems. Prog Mol Biol Transl Sci 101:25–104. https://doi.org/10.1016/B978-0-12-387685-0.00002-0
Sohni A et al (2015) Dynamic switching of active promoter and enhancer domains regulates Tet1 and Tet2 expression during cell state transitions between pluripotency and differentiation. Mol Cell Biol 35:1026–1042. https://doi.org/10.1128/MCB.01172-14
Liu N et al (2013) Intrinsic and extrinsic connections of Tet3 dioxygenase with CXXC zinc finger modules. PLoS One 8:e62755. https://doi.org/10.1371/journal.pone.0062755
Ito S et al (2011) Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333:1300–1303. https://doi.org/10.1126/science.1210597
He YF et al (2011) Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333:1303–1307. https://doi.org/10.1126/science.1210944
Ito S et al (2010) Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466:1129–1133. https://doi.org/10.1038/nature09303
Xu Y et al (2011) Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol Cell 42:451–464. https://doi.org/10.1016/j.molcel.2011.04.005
Hahn MA et al (2013) Dynamics of 5-hydroxymethylcytosine and chromatin marks in Mammalian neurogenesis. Cell Rep 3:291–300. https://doi.org/10.1016/j.celrep.2013.01.011
Jin SG et al (2016) Tet3 reads 5-carboxylcytosine through Its CXXC domain and is a potential guardian against neurodegeneration. Cell Rep 14:493–505. https://doi.org/10.1016/j.celrep.2015.12.044
Shekhawat J et al (2021) Ten-eleven translocase: key regulator of the methylation landscape in cancer. J Cancer Res Clin Oncol 147:1869–1879. https://doi.org/10.1007/s00432-021-03641-3
Kunimoto H, Nakajima H (2021) TET2: a cornerstone in normal and malignant hematopoiesis. Cancer Sci 112:31–40. https://doi.org/10.1111/cas.14688
Bray JK, Dawlaty MM, Verma A, Maitra A (2021) Roles and regulations of TET enzymes in solid tumors. Trends Cancer 7:635–646. https://doi.org/10.1016/j.trecan.2020.12.011
Dziaman T et al (2018) Characteristic profiles of DNA epigenetic modifications in colon cancer and its predisposing conditions-benign adenomas and inflammatory bowel disease. Clin Epigenetics 10:72. https://doi.org/10.1186/s13148-018-0505-0
Pei YF et al (2016) TET1 inhibits gastric cancer growth and metastasis by PTEN demethylation and re-expression. Oncotarget 7:31322–31335. https://doi.org/10.18632/oncotarget.8900
Wu J et al (2019) TET1-mediated DNA hydroxymethylation activates inhibitors of the Wnt/beta-catenin signaling pathway to suppress EMT in pancreatic tumor cells. J Exp Clin Cancer Res 38:348. https://doi.org/10.1186/s13046-019-1334-5
Eyres M et al (2021) TET2 drives 5hmc marking of GATA6 and epigenetically defines pancreatic ductal adenocarcinoma transcriptional subtypes. Gastroenterology 161:653–668. https://doi.org/10.1053/j.gastro.2021.04.044 (e616)
Spans L et al (2016) Genomic and epigenomic analysis of high-risk prostate cancer reveals changes in hydroxymethylation and TET1. Oncotarget 7:24326–24338. https://doi.org/10.18632/oncotarget.8220
Chen Q et al (2017) MicroRNA-29a induces loss of 5-hydroxymethylcytosine and promotes metastasis of hepatocellular carcinoma through a TET-SOCS1-MMP9 signaling axis. Cell Death Dis 8:e2906. https://doi.org/10.1038/cddis.2017.142
Lian CG et al (2012) Loss of 5-hydroxymethylcytosine is an epigenetic hallmark of melanoma. Cell 150:1135–1146. https://doi.org/10.1016/j.cell.2012.07.033
Liu J et al (2019) Global DNA 5-hydroxymethylcytosine and 5-formylcytosine contents are decreased in the early stage of hepatocellular carcinoma. Hepatology 69:196–208. https://doi.org/10.1002/hep.30146
Zahid OK et al (2021) Solid-state nanopore analysis of human genomic DNA shows unaltered global 5-hydroxymethylcytosine content associated with early-stage breast cancer. Nanomed Nanotechnol Biol Med 35:102407. https://doi.org/10.1016/j.nano.2021.102407
Haffner MC et al (2011) Global 5-hydroxymethylcytosine content is significantly reduced in tissue stem/progenitor cell compartments and in human cancers. Oncotarget 2:627–637. https://doi.org/10.18632/oncotarget.316
Kraus TF et al (2015) Loss of 5-hydroxymethylcytosine and intratumoral heterogeneity as an epigenomic hallmark of glioblastoma. Tumour Biol 36:8439–8446. https://doi.org/10.1007/s13277-015-3606-9
Thomson JP et al (2016) Loss of Tet1-associated 5-hydroxymethylcytosine is concomitant with aberrant promoter hypermethylation in liver cancer. Cancer Res 76:3097–3108. https://doi.org/10.1158/0008-5472.CAN-15-1910
Yang H et al (2013) Tumor development is associated with decrease of TET gene expression and 5-methylcytosine hydroxylation. Oncogene 32:663–669. https://doi.org/10.1038/onc.2012.67
Tsai KW et al (2015) Reduction of global 5-hydroxymethylcytosine is a poor prognostic factor in breast cancer patients, especially for an ER/PR-negative subtype. Breast Cancer Res Treat 153:219–234. https://doi.org/10.1007/s10549-015-3525-x
Deng M et al (2017) TET-mediated sequestration of miR-26 drives EZH2 expression and gastric carcinogenesis. Cancer Res 77:6069–6082. https://doi.org/10.1158/0008-5472.CAN-16-2964
Chen LY et al (2019) TET1 reprograms the epithelial ovarian cancer epigenome and reveals casein kinase 2α as a therapeutic target. J Pathol 248:363–376. https://doi.org/10.1002/path.5266
Filipczak PT et al (2019) p53-suppressed oncogene TET1 prevents cellular aging in lung cancer. Cancer Res 79:1758–1768. https://doi.org/10.1158/0008-5472.CAN-18-1234
Muller T et al (2012) Nuclear exclusion of TET1 is associated with loss of 5-hydroxymethylcytosine in IDH1 wild-type gliomas. Am J Pathol 181:675–683. https://doi.org/10.1016/j.ajpath.2012.04.017
Good CR et al (2018) TET1-mediated hypomethylation activates oncogenic signaling in triple-negative breast cancer. Cancer Res 78:4126–4137. https://doi.org/10.1158/0008-5472.CAN-17-2082
Lio CJ, Yuita H, Rao A (2019) Dysregulation of the TET family of epigenetic regulators in lymphoid and myeloid malignancies. Blood 134:1487–1497. https://doi.org/10.1182/blood.2019791475
Cimmino L et al (2015) TET1 is a tumor suppressor of hematopoietic malignancy. Nat Immunol 16:653–662. https://doi.org/10.1038/ni.3148
Morin RD et al (2011) Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476:298–303. https://doi.org/10.1038/nature10351
Pasqualucci L et al (2011) Analysis of the coding genome of diffuse large B-cell lymphoma. Nat Genet 43:830–837. https://doi.org/10.1038/ng.892
Okosun J et al (2014) Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat Genet 46:176–181. https://doi.org/10.1038/ng.2856
Li H et al (2014) Mutations in linker histone genes HIST1H1 B, C, D, and E; OCT2 (POU2F2); IRF8; and ARID1A underlying the pathogenesis of follicular lymphoma. Blood 123:1487–1498. https://doi.org/10.1182/blood-2013-05-500264
De Keersmaecker K et al (2013) Exome sequencing identifies mutation in CNOT3 and ribosomal genes RPL5 and RPL10 in T-cell acute lymphoblastic leukemia. Nat Genet 45:186–190. https://doi.org/10.1038/ng.2508
Cancer Genome Atlas Research N et al (2013) Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. New Eng J Med 368:2059–2074. https://doi.org/10.1056/NEJMoa1301689
Quesada V et al (2012) Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat Genet 44:47–52. https://doi.org/10.1038/ng.1032
Nickerson ML et al (2017) TET2 binds the androgen receptor and loss is associated with prostate cancer. Oncogene 36:2172–2183. https://doi.org/10.1038/onc.2016.376
Neri F et al (2015) TET1 is controlled by pluripotency-associated factors in ESCs and downmodulated by PRC2 in differentiated cells and tissues. Nucleic Acids Res 43:6814–6826. https://doi.org/10.1093/nar/gkv392
Tsai YP et al (2014) TET1 regulates hypoxia-induced epithelial-mesenchymal transition by acting as a co-activator. Genome Biol 15:513. https://doi.org/10.1186/s13059-014-0513-0
Yang YA et al (2016) FOXA1 potentiates lineage-specific enhancer activation through modulating TET1 expression and function. Nucleic Acids Res 44:8153–8164. https://doi.org/10.1093/nar/gkw498
Yosefzon Y et al (2017) An epigenetic switch repressing Tet1 in gonadotropes activates the reproductive axis. Proc Natl Acad Sci USA 114:10131–10136. https://doi.org/10.1073/pnas.1704393114
Li L et al (2016) Epigenetic inactivation of the CpG demethylase TET1 as a DNA methylation feedback loop in human cancers. Sci Rep 6:26591. https://doi.org/10.1038/srep26591
Agirre X et al (2015) Whole-epigenome analysis in multiple myeloma reveals DNA hypermethylation of B cell-specific enhancers. Genome Res 25:478–487. https://doi.org/10.1101/gr.180240.114
Sun M et al (2013) HMGA2/TET1/HOXA9 signaling pathway regulates breast cancer growth and metastasis. Proc Natl Acad Sci USA 110:9920–9925. https://doi.org/10.1073/pnas.1305172110
Forloni M et al (2016) Oncogenic EGFR represses the TET1 DNA demethylase to induce silencing of tumor suppressors in cancer cells. Cell Rep 16:457–471. https://doi.org/10.1016/j.celrep.2016.05.087
Collignon E et al (2018) Immunity drives TET1 regulation in cancer through NF-kappaB. Sci Adv 4:eaap7309. https://doi.org/10.1126/sciadv.aap7309
Noreen F et al (2019) DNA methylation instability by BRAF-mediated TET silencing and lifestyle-exposure divides colon cancer pathways. Clin Epigenetics 11:196. https://doi.org/10.1186/s13148-019-0791-1
Hsu CH et al (2012) TET1 suppresses cancer invasion by activating the tissue inhibitors of metalloproteinases. Cell Rep 2:568–579. https://doi.org/10.1016/j.celrep.2012.08.030
Teslow EA et al (2019) Obesity-induced MBD2_v2 expression promotes tumor-initiating triple-negative breast cancer stem cells. Mol Oncol 13:894–908. https://doi.org/10.1002/1878-0261.12444
Wang H et al (2017) MiR-29b/TET1/ZEB2 signaling axis regulates metastatic properties and epithelial-mesenchymal transition in breast cancer cells. Oncotarget 8:102119–102133. https://doi.org/10.18632/oncotarget.22183
Chen N et al (2018) A novel FLI1 exonic circular RNA promotes metastasis in breast cancer by coordinately regulating TET1 and DNMT1. Genome Biol 19:218. https://doi.org/10.1186/s13059-018-1594-y
Wu Y et al (2013) Oct4 and the small molecule inhibitor, SC1, regulates Tet2 expression in mouse embryonic stem cells. Mol Biol Rep 40:2897–2906. https://doi.org/10.1007/s11033-012-2305-5
Koh KP et al (2011) Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8:200–213. https://doi.org/10.1016/j.stem.2011.01.008
Fischer AP, Miles SL (2017) Silencing HIF-1alpha induces TET2 expression and augments ascorbic acid induced 5-hydroxymethylation of DNA in human metastatic melanoma cells. Biochem Biophys Res Commun 490:176–181. https://doi.org/10.1016/j.bbrc.2017.06.017
Zhang LY, Li PL, Wang TZ, Zhang XC (2015) Prognostic values of 5-hmC, 5-mC and TET2 in epithelial ovarian cancer. Arch Gynecol Obstet 292:891–897. https://doi.org/10.1007/s00404-015-3704-3
Tucker DW et al (2018) Epigenetic reprogramming strategies to reverse global loss of 5-hydroxymethylcytosine, a prognostic factor for poor survival in high-grade serous ovarian cancer. Clin Cancer Res 24:1389–1401. https://doi.org/10.1158/1078-0432.CCR-17-1958
Rawluszko-Wieczorek AA et al (2015) Clinical significance of DNA methylation mRNA levels of TET family members in colorectal cancer. J Cancer Res Clin Oncol 141:1379–1392. https://doi.org/10.1007/s00432-014-1901-2
Carella A et al (2020) Epigenetic downregulation of TET3 reduces genome-wide 5hmC levels and promotes glioblastoma tumorigenesis. Int J Cancer 146:373–387. https://doi.org/10.1002/ijc.32520
Mo HY, An CH, Choi EJ, Yoo NJ, Lee SH (2020) Somatic mutation and loss of expression of a candidate tumor suppressor gene TET3 in gastric and colorectal cancers. Pathol Res Pract 216:152759. https://doi.org/10.1016/j.prp.2019.152759
Ye Z et al (2016) TET3 inhibits TGF-beta1-induced epithelial-mesenchymal transition by demethylating miR-30d precursor gene in ovarian cancer cells. J Exp Clin Cancer Res 35:72. https://doi.org/10.1186/s13046-016-0350-y
Cao T, Pan W, Sun X, Shen H (2019) Increased expression of TET3 predicts unfavorable prognosis in patients with ovarian cancer-a bioinformatics integrative analysis. J Ovarian Res 12:101. https://doi.org/10.1186/s13048-019-0575-4
Song SJ et al (2013) MicroRNA-antagonism regulates breast cancer stemness and metastasis via TET-family-dependent chromatin remodeling. Cell 154:311–324. https://doi.org/10.1016/j.cell.2013.06.026
Cheng J et al (2013) An extensive network of TET2-targeting microRNAs regulates malignant hematopoiesis. Cell Rep 5:471–481. https://doi.org/10.1016/j.celrep.2013.08.050
Fu X et al (2013) MicroRNA-26a targets ten eleven translocation enzymes and is regulated during pancreatic cell differentiation. Proc Natl Acad Sci USA 110:17892–17897. https://doi.org/10.1073/pnas.1317397110
Loriot A et al (2014) A novel cancer-germline transcript carrying pro-metastatic miR-105 and TET-targeting miR-767 induced by DNA hypomethylation in tumors. Epigenetics 9:1163–1171. https://doi.org/10.4161/epi.29628
Chuang KH et al (2015) MicroRNA-494 is a master epigenetic regulator of multiple invasion-suppressor microRNAs by targeting ten eleven translocation 1 in invasive human hepatocellular carcinoma tumors. Hepatology 62:466–480. https://doi.org/10.1002/hep.27816
Zhang W et al (2015) MiR-520b suppresses proliferation of hepatoma cells through targeting ten-eleven translocation 1 (TET1) mRNA. Biochem Biophys Res Commun 460:793–798. https://doi.org/10.1016/j.bbrc.2015.03.108
Song SJ et al (2013) The oncogenic microRNA miR-22 targets the TET2 tumor suppressor to promote hematopoietic stem cell self-renewal and transformation. Cell Stem Cell 13:87–101. https://doi.org/10.1016/j.stem.2013.06.003
Takeshima H et al (2020) TET repression and increased DNMT activity synergistically induce aberrant DNA methylation. J Clin Invest 130:5370–5379. https://doi.org/10.1172/JCI124070
Scherm MG et al (2019) miRNA142-3p targets Tet2 and impairs Treg differentiation and stability in models of type 1 diabetes. Nat Commun 10:5697. https://doi.org/10.1038/s41467-019-13587-3
Jiang S, Yan W, Wang SE, Baltimore D (2019) Dual mechanisms of posttranscriptional regulation of Tet2 by Let-7 microRNA in macrophages. Proc Natl Acad Sci USA 116:12416–12421. https://doi.org/10.1073/pnas.1811040116
Maiti A, Drohat AC (2011) Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J Biol Chem 286:35334–35338. https://doi.org/10.1074/jbc.C111.284620
Cortellino S et al (2011) Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. Cell 146:67–79. https://doi.org/10.1016/j.cell.2011.06.020
Bhutani N et al (2010) Reprogramming towards pluripotency requires AID-dependent DNA demethylation. Nature 463:1042–1047. https://doi.org/10.1038/nature08752
Popp C et al (2010) Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463:1101–1105. https://doi.org/10.1038/nature08829
Nawy T (2013) Dynamics of DNA demethylation. Nat Methods 10:466. https://doi.org/10.1038/nmeth.2506
Muramatsu M et al (2000) Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell 102:553–563. https://doi.org/10.1016/s0092-8674(00)00078-7
Xu Z, Zan H, Pone EJ, Mai T, Casali P (2012) Immunoglobulin class-switch DNA recombination: induction, targeting and beyond. Nat Rev Immunol 12:517–531. https://doi.org/10.1038/nri3216
Barreto G et al (2007) Gadd45a promotes epigenetic gene activation by repair-mediated DNA demethylation. Nature 445:671–675. https://doi.org/10.1038/nature05515
Schmitz KM et al (2009) TAF12 recruits Gadd45a and the nucleotide excision repair complex to the promoter of rRNA genes leading to active DNA demethylation. Mol Cell 33:344–353. https://doi.org/10.1016/j.molcel.2009.01.015
Ma DK et al (2009) Neuronal activity-induced Gadd45b promotes epigenetic DNA demethylation and adult neurogenesis. Science 323:1074–1077. https://doi.org/10.1126/science.1166859
Hajkova P et al (2010) Genome-wide reprogramming in the mouse germ line entails the base excision repair pathway. Science 329:78–82. https://doi.org/10.1126/science.1187945
Dominguez PM et al (2015) DNA methylation dynamics of germinal center B cells are mediated by AID. Cell Rep 12:2086–2098. https://doi.org/10.1016/j.celrep.2015.08.036
Lio CJ et al (2019) TET enzymes augment activation-induced deaminase (AID) expression via 5-hydroxymethylcytosine modifications at the Aicda superenhancer. Sci Immunol. https://doi.org/10.1126/sciimmunol.aau7523
Delker RK, Fugmann SD, Papavasiliou FN (2009) A coming-of-age story: activation-induced cytidine deaminase turns 10. Nat Immunol 10:1147–1153. https://doi.org/10.1038/ni.1799
Kunimoto H et al (2017) Aid is a key regulator of myeloid/erythroid differentiation and DNA methylation in hematopoietic stem/progenitor cells. Blood 129:1779–1790. https://doi.org/10.1182/blood-2016-06-721977
Bachman M et al (2014) 5-Hydroxymethylcytosine is a predominantly stable DNA modification. Nat Chem 6:1049–1055. https://doi.org/10.1038/nchem.2064
Bachman M et al (2015) 5-Formylcytosine can be a stable DNA modification in mammals. Nat Chem Biol 11:555–557. https://doi.org/10.1038/nchembio.1848
Wu H, Wu X, Shen L, Zhang Y (2014) Single-base resolution analysis of active DNA demethylation using methylase-assisted bisulfite sequencing. Nat Biotechnol 32:1231–1240. https://doi.org/10.1038/nbt.3073
Spruijt CG et al (2013) Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 152:1146–1159. https://doi.org/10.1016/j.cell.2013.02.004
Mellen M, Ayata P, Dewell S, Kriaucionis S, Heintz N (2012) MeCP2 binds to 5hmC enriched within active genes and accessible chromatin in the nervous system. Cell 151:1417–1430. https://doi.org/10.1016/j.cell.2012.11.022
Takai H et al (2014) 5-Hydroxymethylcytosine plays a critical role in glioblastomagenesis by recruiting the CHTOP-methylosome complex. Cell Rep 9:48–60. https://doi.org/10.1016/j.celrep.2014.08.071
Yildirim O et al (2011) Mbd3/NURD complex regulates expression of 5-hydroxymethylcytosine marked genes in embryonic stem cells. Cell 147:1498–1510. https://doi.org/10.1016/j.cell.2011.11.054
Chen R et al (2017) The 5-hydroxymethylcytosine (5hmC) reader UHRF2 is required for normal levels of 5hmC in mouse adult brain and spatial learning and memory. J Biol Chem 292:4533–4543. https://doi.org/10.1074/jbc.M116.754580
Tsagaratou A et al (2014) Dissecting the dynamic changes of 5-hydroxymethylcytosine in T-cell development and differentiation. Proc Natl Acad Sci U S A 111:E3306-3315. https://doi.org/10.1073/pnas.1412327111
Pastor WA et al (2011) Genome-wide mapping of 5-hydroxymethylcytosine in embryonic stem cells. Nature 473:394–397. https://doi.org/10.1038/nature10102
Stroud H, Feng S, Morey Kinney S, Pradhan S, Jacobsen SE (2011) 5-Hydroxymethylcytosine is associated with enhancers and gene bodies in human embryonic stem cells. Genome Biol 12:R54. https://doi.org/10.1186/gb-2011-12-6-r54
Williams K et al (2011) TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473:343–348. https://doi.org/10.1038/nature10066
Madzo J et al (2014) Hydroxymethylation at gene regulatory regions directs stem/early progenitor cell commitment during erythropoiesis. Cell Rep 6:231–244. https://doi.org/10.1016/j.celrep.2013.11.044
Guilhamon P et al (2013) Meta-analysis of IDH-mutant cancers identifies EBF1 as an interaction partner for TET2. Nat Commun 4:2166. https://doi.org/10.1038/ncomms3166
Mellen M, Ayata P, Heintz N (2017) 5-hydroxymethylcytosine accumulation in postmitotic neurons results in functional demethylation of expressed genes. Proc Natl Acad Sci USA 114:E7812–E7821. https://doi.org/10.1073/pnas.1708044114
He B et al (2021) Tissue-specific 5-hydroxymethylcytosine landscape of the human genome. Nat Commun 12:4249. https://doi.org/10.1038/s41467-021-24425-w
Dawlaty MM et al (2014) Loss of Tet enzymes compromises proper differentiation of embryonic stem cells. Dev Cell 29:102–111. https://doi.org/10.1016/j.devcel.2014.03.003
Li C et al (2015) Overlapping requirements for Tet2 and Tet3 in normal development and hematopoietic stem cell emergence. Cell Rep 12:1133–1143. https://doi.org/10.1016/j.celrep.2015.07.025
Dawlaty MM et al (2013) Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev Cell 24:310–323. https://doi.org/10.1016/j.devcel.2012.12.015
Hu L et al (2015) Structural insight into substrate preference for TET-mediated oxidation. Nature 527:118–122. https://doi.org/10.1038/nature15713
Huang Y et al (2014) Distinct roles of the methylcytosine oxidases Tet1 and Tet2 in mouse embryonic stem cells. Proc Natl Acad Sci USA 111:1361–1366. https://doi.org/10.1073/pnas.1322921111
Tan L, Shi YG (2012) Tet family proteins and 5-hydroxymethylcytosine in development and disease. Development 139:1895–1902. https://doi.org/10.1242/dev.070771
Song CX et al (2011) Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine. Nat Biotechnol 29:68–72. https://doi.org/10.1038/nbt.1732
Putiri EL et al (2014) Distinct and overlapping control of 5-methylcytosine and 5-hydroxymethylcytosine by the TET proteins in human cancer cells. Genome Biol 15:R81. https://doi.org/10.1186/gb-2014-15-6-r81
Yamaguchi S, Shen L, Liu Y, Sendler D, Zhang Y (2013) Role of Tet1 in erasure of genomic imprinting. Nature 504:460–464. https://doi.org/10.1038/nature12805
Wu H, Zhang Y (2014) Reversing DNA methylation: mechanisms, genomics, and biological functions. Cell 156:45–68. https://doi.org/10.1016/j.cell.2013.12.019
Gu TP et al (2011) The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477:606–610. https://doi.org/10.1038/nature10443
Pastor WA, Aravind L, Rao A (2013) TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nat Rev Mol Cell Biol 14:341–356. https://doi.org/10.1038/nrm3589
Dawlaty MM et al (2011) Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9:166–175. https://doi.org/10.1016/j.stem.2011.07.010
Wu H et al (2011) Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473:389–393. https://doi.org/10.1038/nature09934
Hon GC et al (2014) 5mC oxidation by Tet2 modulates enhancer activity and timing of transcriptome reprogramming during differentiation. Mol Cell 56:286–297. https://doi.org/10.1016/j.molcel.2014.08.026
Fidalgo M et al (2016) Zfp281 coordinates opposing functions of Tet1 and Tet2 in pluripotent states. Cell Stem Cell 19:355–369. https://doi.org/10.1016/j.stem.2016.05.025
Piccolo FM et al (2013) Different roles for Tet1 and Tet2 proteins in reprogramming-mediated erasure of imprints induced by EGC fusion. Mol Cell 49:1023–1033. https://doi.org/10.1016/j.molcel.2013.01.032
Jin C et al (2014) TET1 is a maintenance DNA demethylase that prevents methylation spreading in differentiated cells. Nucleic Acids Res 42:6956–6971. https://doi.org/10.1093/nar/gku372
Iyer LM, Tahiliani M, Rao A, Aravind L (2009) Prediction of novel families of enzymes involved in oxidative and other complex modifications of bases in nucleic acids. Cell Cycle 8:1698–1710. https://doi.org/10.4161/cc.8.11.8580
Frauer C et al (2011) Different binding properties and function of CXXC zinc finger domains in Dnmt1 and Tet1. PLoS One 6:e16627. https://doi.org/10.1371/journal.pone.0016627
Zhang H et al (2010) TET1 is a DNA-binding protein that modulates DNA methylation and gene transcription via hydroxylation of 5-methylcytosine. Cell Res 20:1390–1393. https://doi.org/10.1038/cr.2010.156
Greer CB et al (2021) Tet1 isoforms differentially regulate gene expression, synaptic transmission, and memory in the mammalian brain. J Neurosci 41:578–593. https://doi.org/10.1523/JNEUROSCI.1821-20.2020
Chandru A, Bate N, Vuister GW, Cowley SM (2018) Sin3A recruits Tet1 to the PAH1 domain via a highly conserved Sin3-Interaction Domain. Sci Rep 8:14689. https://doi.org/10.1038/s41598-018-32942-w
Zhu F et al (2018) Sin3a-Tet1 interaction activates gene transcription and is required for embryonic stem cell pluripotency. Nucleic Acids Res 46:6026–6040. https://doi.org/10.1093/nar/gky347
Vella P et al (2013) Tet proteins connect the O-linked N-acetylglucosamine transferase Ogt to chromatin in embryonic stem cells. Mol Cell 49:645–656. https://doi.org/10.1016/j.molcel.2012.12.019
Deplus R et al (2013) TET2 and TET3 regulate GlcNAcylation and H3K4 methylation through OGT and SET1/COMPASS. EMBO J 32:645–655. https://doi.org/10.1038/emboj.2012.357
Chen Q, Chen Y, Bian C, Fujiki R, Yu X (2013) TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493:561–564. https://doi.org/10.1038/nature11742
Hrit J et al (2018) OGT binds a conserved C-terminal domain of TET1 to regulate TET1 activity and function in development. Elife. https://doi.org/10.7554/eLife.34870
Teif VB et al (2014) Nucleosome repositioning links DNA (de)methylation and differential CTCF binding during stem cell development. Genome Res 24:1285–1295. https://doi.org/10.1101/gr.164418.113
Dubois-Chevalier J et al (2014) A dynamic CTCF chromatin binding landscape promotes DNA hydroxymethylation and transcriptional induction of adipocyte differentiation. Nucleic Acids Res 42:10943–10959. https://doi.org/10.1093/nar/gku780
Wiehle L et al (2019) DNA (de)methylation in embryonic stem cells controls CTCF-dependent chromatin boundaries. Genome Res 29:750–761. https://doi.org/10.1101/gr.239707.118
Perera A et al (2015) TET3 is recruited by REST for context-specific hydroxymethylation and induction of gene expression. Cell Rep 11:283–294. https://doi.org/10.1016/j.celrep.2015.03.020
Ko M et al (2013) Modulation of TET2 expression and 5-methylcytosine oxidation by the CXXC domain protein IDAX. Nature 497:122–126. https://doi.org/10.1038/nature12052
Ayaz G et al (2020) CXXC5 as an unmethylated CpG dinucleotide binding protein contributes to estrogen-mediated cellular proliferation. Sci Rep 10:5971. https://doi.org/10.1038/s41598-020-62912-0
Ravichandran M et al (2019) Rinf regulates pluripotency network genes and TET enzymes in embryonic stem cells. Cell Rep 28:1993–2003. https://doi.org/10.1016/j.celrep.2019.07.080 (e1995)
Abbas S, Erpelinck-Verschueren CA, Goudswaard CS, Lowenberg B, Valk PJ (2010) Mutant Wilms’ tumor 1 (WT1) mRNA with premature termination codons in acute myeloid leukemia (AML) is sensitive to nonsense-mediated RNA decay (NMD). Leukemia 24:660–663. https://doi.org/10.1038/leu.2009.265
Wang Y et al (2015) WT1 recruits TET2 to regulate its target gene expression and suppress leukemia cell proliferation. Mol Cell 57:662–673. https://doi.org/10.1016/j.molcel.2014.12.023
Rampal R et al (2014) DNA hydroxymethylation profiling reveals that WT1 mutations result in loss of TET2 function in acute myeloid leukemia. Cell Rep 9:1841–1855. https://doi.org/10.1016/j.celrep.2014.11.004
Pan F, Weeks O, Yang FC, Xu M (2015) The TET2 interactors and their links to hematological malignancies. IUBMB Life 67:438–445. https://doi.org/10.1002/iub.1389
Lazarenkov A, Sardina JL (2022) Dissecting TET2 regulatory networks in blood differentiation and cancer. Cancers (Basel). https://doi.org/10.3390/cancers14030830
Muller U, Bauer C, Siegl M, Rottach A, Leonhardt H (2014) TET-mediated oxidation of methylcytosine causes TDG or NEIL glycosylase dependent gene reactivation. Nucleic Acids Res 42:8592–8604. https://doi.org/10.1093/nar/gku552
Cartron PF et al (2013) Identification of TET1 partners that control Its DNA-demethylating function. Genes Cancer 4:235–241. https://doi.org/10.1177/1947601913489020
Costa Y et al (2013) NANOG-dependent function of TET1 and TET2 in establishment of pluripotency. Nature 495:370–374. https://doi.org/10.1038/nature11925
Zheng L et al (2016) Modification of Tet1 and histone methylation dynamics in dairy goat male germline stem cells. Cell Prolif 49:163–172. https://doi.org/10.1111/cpr.12245
Zheng L et al (2016) The modification of Tet1 in male germline stem cells and interact with PCNA, HDAC1 to promote their self-renewal and proliferation. Sci Rep 6:37414. https://doi.org/10.1038/srep37414
Zhang X, Yang C, Peng X, Chen X, Feng Y (2017) Acute WT1-positive promyelocytic leukemia with hypogranular variant morphology, bcr-3 isoform of PML-RARalpha and Flt3-ITD mutation: a rare case report. Sao Paulo medical journal = Revista paulista de medicina 135:179–184. https://doi.org/10.1590/1516-3180.2016.020104102016
Zhang Q et al (2015) Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature 525:389–393. https://doi.org/10.1038/nature15252
Chen LL et al (2018) SNIP1 recruits TET2 to regulate c-MYC target genes and cellular DNA damage response. Cell Rep 25:1485–1500. https://doi.org/10.1016/j.celrep.2018.10.028 (e1484)
Sardina JL et al (2018) Transcription factors drive Tet2-mediated enhancer demethylation to reprogram cell fate. Cell Stem Cell 23:905–906. https://doi.org/10.1016/j.stem.2018.11.001
Zhang YW et al (2017) Acetylation enhances TET2 function in protecting against abnormal DNA methylation during oxidative stress. Mol Cell 65:323–335. https://doi.org/10.1016/j.molcel.2016.12.013
Song C et al (2018) PML recruits TET2 to regulate DNA modification and cell proliferation in response to chemotherapeutic agent. Cancer Res 78:2475–2489. https://doi.org/10.1158/0008-5472.CAN-17-3091
Nakagawa T et al (2015) CRL4(VprBP) E3 ligase promotes monoubiquitylation and chromatin binding of TET dioxygenases. Mol Cell 57:247–260. https://doi.org/10.1016/j.molcel.2014.12.002
Qiao Y et al (2015) AF9 promotes hESC neural differentiation through recruiting TET2 to neurodevelopmental gene loci for methylcytosine hydroxylation. Cell Discovery 1:15017. https://doi.org/10.1038/celldisc.2015.17
Lio CW et al (2016) Tet2 and Tet3 cooperate with B-lineage transcription factors to regulate DNA modification and chromatin accessibility. Elife. https://doi.org/10.7554/eLife.18290
de la Rica L et al (2013) PU.1 target genes undergo Tet2-coupled demethylation and DNMT3b-mediated methylation in monocyte-to-osteoclast differentiation. Genome Biol 14:R99. https://doi.org/10.1186/gb-2013-14-9-r99
Suzuki T et al (2017) RUNX1 regulates site specificity of DNA demethylation by recruitment of DNA demethylation machineries in hematopoietic cells. Blood Adv 1:1699–1711. https://doi.org/10.1182/bloodadvances.2017005710
Zeng Y et al (2016) Lin28A binds active promoters and recruits Tet1 to regulate gene expression. Mol Cell 61:153–160. https://doi.org/10.1016/j.molcel.2015.11.020
Sun Z et al (2019) EGR1 recruits TET1 to shape the brain methylome during development and upon neuronal activity. Nat Commun 10:3892. https://doi.org/10.1038/s41467-019-11905-3
Guan W et al (2017) Methylcytosine dioxygenase TET3 interacts with thyroid hormone nuclear receptors and stabilizes their association to chromatin. Proc Natl Acad Sci USA 114:8229–8234. https://doi.org/10.1073/pnas.1702192114
Fujiki K et al (2013) PPARgamma-induced PARylation promotes local DNA demethylation by production of 5-hydroxymethylcytosine. Nat Commun 4:2262. https://doi.org/10.1038/ncomms3262
Baubec T, Ivanek R, Lienert F, Schubeler D (2013) Methylation-dependent and -independent genomic targeting principles of the MBD protein family. Cell 153:480–492. https://doi.org/10.1016/j.cell.2013.03.011
Peng L et al (2016) MBD3L2 promotes Tet2 enzymatic activity for mediating 5-methylcytosine oxidation. J Cell Sci 129:1059–1071. https://doi.org/10.1242/jcs.179044
Okashita N et al (2014) PRDM14 promotes active DNA demethylation through the ten-eleven translocation (TET)-mediated base excision repair pathway in embryonic stem cells. Development 141:269–280. https://doi.org/10.1242/dev.099622
Soufi A et al (2015) Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161:555–568. https://doi.org/10.1016/j.cell.2015.03.017
Boller S et al (2016) Pioneering activity of the C-terminal domain of EBF1 shapes the chromatin landscape for B Cell programming. Immunity 44:527–541. https://doi.org/10.1016/j.immuni.2016.02.021
Liu Y et al (2014) Structural basis for Klf4 recognition of methylated DNA. Nucleic Acids Res 42:4859–4867. https://doi.org/10.1093/nar/gku134
Hashimoto H et al (2014) Wilms tumor protein recognizes 5-carboxylcytosine within a specific DNA sequence. Genes Dev 28:2304–2313. https://doi.org/10.1101/gad.250746.114
Vanzan L et al (2021) High throughput screening identifies SOX2 as a super pioneer factor that inhibits DNA methylation maintenance at its binding sites. Nat Commun 12:3337. https://doi.org/10.1038/s41467-021-23630-x
Quenneville S et al (2011) In embryonic stem cells, ZFP57/KAP1 recognize a methylated hexanucleotide to affect chromatin and DNA methylation of imprinting control regions. Mol Cell 44:361–372. https://doi.org/10.1016/j.molcel.2011.08.032
Wang D et al (2017) MAX is an epigenetic sensor of 5-carboxylcytosine and is altered in multiple myeloma. Nucleic Acids Res 45:2396–2407. https://doi.org/10.1093/nar/gkw1184
Domcke S et al (2015) Competition between DNA methylation and transcription factors determines binding of NRF1. Nature 528:575–579. https://doi.org/10.1038/nature16462
Xiong J et al (2016) Cooperative action between SALL4A and TET proteins in stepwise oxidation of 5-methylcytosine. Mol Cell 64:913–925. https://doi.org/10.1016/j.molcel.2016.10.013
Looney TJ et al (2014) Systematic mapping of occluded genes by cell fusion reveals prevalence and stability of cis-mediated silencing in somatic cells. Genome Res 24:267–280. https://doi.org/10.1101/gr.143891.112
Polo JM et al (2012) A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151:1617–1632. https://doi.org/10.1016/j.cell.2012.11.039
Okashita N et al (2016) PRDM14 drives OCT3/4 recruitment via active demethylation in the transition from primed to naive pluripotency. Stem Cell Reports 7:1072–1086. https://doi.org/10.1016/j.stemcr.2016.10.007
Iwafuchi-Doi M, Zaret KS (2016) Cell fate control by pioneer transcription factors. Development 143:1833–1837. https://doi.org/10.1242/dev.133900
Mayran A, Drouin J (2018) Pioneer transcription factors shape the epigenetic landscape. J Biol Chem 293:13795–13804. https://doi.org/10.1074/jbc.R117.001232
Larson ED, Marsh AJ, Harrison MM (2021) Pioneering the developmental frontier. Mol Cell 81:1640–1650. https://doi.org/10.1016/j.molcel.2021.02.020
Balsalobre A, Drouin J (2022) Pioneer factors as master regulators of the epigenome and cell fate. Nat Rev Mol Cell Biol. https://doi.org/10.1038/s41580-022-00464-z
Iwafuchi-Doi M (2019) The mechanistic basis for chromatin regulation by pioneer transcription factors. Wiley Interdisciplin Rev 11:e1427. https://doi.org/10.1002/wsbm.1427
Chronis C et al (2017) Cooperative binding of transcription factors orchestrates reprogramming. Cell 168:442–459. https://doi.org/10.1016/j.cell.2016.12.016 (e420)
Mayran A et al (2019) Pioneer and nonpioneer factor cooperation drives lineage specific chromatin opening. Nat Commun 10:3807. https://doi.org/10.1038/s41467-019-11791-9
Sonmezer C et al (2021) Molecular co-occupancy identifies transcription factor binding cooperativity in vivo. Mol Cell 81:255–267. https://doi.org/10.1016/j.molcel.2020.11.015 (e256)
Rishi V et al (2010) CpG methylation of half-CRE sequences creates C/EBPα binding sites that activate some tissue-specific genes. Proc Natl Acad Sci USA 107:20311–20316. https://doi.org/10.1073/pnas.1008688107
van Oevelen C et al (2015) C/EBPα activates pre-existing and de novo macrophage enhancers during induced pre-B cell transdifferentiation and myelopoiesis. Stem Cell Reports 5:232–247. https://doi.org/10.1016/j.stemcr.2015.06.007
Xue S et al (2016) TET3 inhibits type I IFN production independent of DNA demethylation. Cell Rep 16:1096–1105. https://doi.org/10.1016/j.celrep.2016.06.068
Zhong J et al (2017) TET1 modulates H4K16 acetylation by controlling auto-acetylation of hMOF to affect gene regulation and DNA repair function. Nucleic Acids Res 45:672–684. https://doi.org/10.1093/nar/gkw919
Bauer C et al (2015) Phosphorylation of TET proteins is regulated via O-GlcNAcylation by the O-Linked N-acetylglucosamine transferase (OGT). J Biol Chem 290:4801–4812. https://doi.org/10.1074/jbc.m114.605881
Li X et al (2021) SIRT1 deacetylates TET2 and promotes its ubiquitination degradation to achieve neuroprotection against parkinson’s disease. Front Neurol 12:652882. https://doi.org/10.3389/fneur.2021.652882
Wu D et al (2018) Glucose-regulated phosphorylation of TET2 by AMPK reveals a pathway linking diabetes to cancer. Nature 559:637–641. https://doi.org/10.1038/s41586-018-0350-5
Rao VK et al (2020) Phosphorylation of Tet3 by cdk5 is critical for robust activation of BRN2 during neuronal differentiation. Nucleic Acids Res 48:1225–1238. https://doi.org/10.1093/nar/gkz1144
Coulter JB et al (2017) TET1 deficiency attenuates the DNA damage response and promotes resistance to DNA damaging agents. Epigenetics 12:854–864. https://doi.org/10.1080/15592294.2017.1359452
Zhang T et al (2019) Phosphorylation of TET2 by AMPK is indispensable in myogenic differentiation. Epigenetics Chromatin 12:32. https://doi.org/10.1186/s13072-019-0281-x
Fiedler EC, Shaw RJ (2018) AMPK regulates the epigenome through phosphorylation of TET2. Cell Metab 28:534–536. https://doi.org/10.1016/j.cmet.2018.09.015
Chen H et al (2019) TET2 stabilization by 14-3-3 binding to the phosphorylated Serine 99 is deregulated by mutations in cancer. Cell Res 29:248–250. https://doi.org/10.1038/s41422-018-0132-5
Kundu A et al (2020) 14–3-3 proteins protect AMPK-phosphorylated ten-eleven translocation-2 (TET2) from PP2A-mediated dephosphorylation. J Biol Chem 295:1754–1766. https://doi.org/10.1074/jbc.RA119.011089
Giovannucci E et al (2010) Diabetes and cancer: a consensus report. Cancer J Clin 60:207–221. https://doi.org/10.3322/caac.20078
Brower V (2012) Illuminating the diabetes-cancer link. J Natl Cancer Inst 104:1048–1050. https://doi.org/10.1093/jnci/djs322
Wang T et al (2021) Diabetes risk reduction diet and survival after breast cancer diagnosis. Cancer Res 81:4155–4162. https://doi.org/10.1158/0008-5472.CAN-21-0256
Jeong JJ et al (2019) Cytokine-regulated phosphorylation and activation of TET2 by JAK2 in hematopoiesis. Cancer Discov 9:778–795. https://doi.org/10.1158/2159-8290.CD-18-1138
Jin Z et al (2020) FGFR3 big up tri, open7-9 promotes tumor progression via the phosphorylation and destabilization of ten-eleven translocation-2 in human hepatocellular carcinoma. Cell Death Dis 11:903. https://doi.org/10.1038/s41419-020-03089-2
Mancini M et al (2012) Cytoplasmatic compartmentalization by Bcr-Abl promotes TET2 loss-of-function in chronic myeloid leukemia. J Cell Biochem 113:2765–2774. https://doi.org/10.1002/jcb.24154
Jiang D, Wei S, Chen F, Zhang Y, Li J (2017) TET3-mediated DNA oxidation promotes ATR-dependent DNA damage response. EMBO Rep 18:781–796. https://doi.org/10.15252/embr.201643179
Ito R et al (2014) TET3-OGT interaction increases the stability and the presence of OGT in chromatin. Genes Cells 19:52–65. https://doi.org/10.1111/gtc.12107
Zhang Q et al (2014) Differential regulation of the ten-eleven translocation (TET) family of dioxygenases by O-linked β-N-acetylglucosamine transferase (OGT). J Biol Chem 289:5986–5996. https://doi.org/10.1074/jbc.M113.524140
Shi FT et al (2013) Ten-eleven translocation 1 (Tet1) is regulated by O-linked N-acetylglucosamine transferase (Ogt) for target gene repression in mouse embryonic stem cells. J Biol Chem 288:20776–20784. https://doi.org/10.1074/jbc.M113.460386
Lv L et al (2018) Vpr targets TET2 for degradation by CRL4(VprBP) E3 ligase to sustain IL-6 expression and enhance HIV-1 replication. Mol Cell 70:961–970. https://doi.org/10.1016/j.molcel.2018.05.007 (e965)
Sun J et al (2018) SIRT1 activation disrupts maintenance of myelodysplastic syndrome stem and progenitor cells by restoring TET2 function. Cell Stem Cell 23:355–369. https://doi.org/10.1016/j.stem.2018.07.018 (e359)
Wang Y, Zhang Y (2014) Regulation of TET protein stability by calpains. Cell Rep 6:278–284. https://doi.org/10.1016/j.celrep.2013.12.031
Bamezai S et al (2021) TET1 promotes growth of T-cell acute lymphoblastic leukemia and can be antagonized via PARP inhibition. Leukemia 35:389–403. https://doi.org/10.1038/s41375-020-0864-3
Ciccarone F et al (2014) Poly(ADP-ribosyl)ation is involved in the epigenetic control of TET1 gene transcription. Oncotarget 5:10356–10367. https://doi.org/10.18632/oncotarget.1905
Ciccarone F, Valentini E, Zampieri M, Caiafa P (2015) 5mC-hydroxylase activity is influenced by the PARylation of TET1 enzyme. Oncotarget 6:24333–24347. https://doi.org/10.18632/oncotarget.4476
Thienpont B et al (2016) Tumour hypoxia causes DNA hypermethylation by reducing TET activity. Nature 537:63–68. https://doi.org/10.1038/nature19081
Cimmino L et al (2017) Restoration of TET2 function blocks aberrant self-renewal and leukemia progression. Cell 170:1079–1095. https://doi.org/10.1016/j.cell.2017.07.032 (e1020)
Chen J et al (2013) Vitamin C modulates TET1 function during somatic cell reprogramming. Nat Genet 45:1504–1509. https://doi.org/10.1038/ng.2807
Yin R et al (2013) Ascorbic acid enhances Tet-mediated 5-methylcytosine oxidation and promotes DNA demethylation in mammals. J Am Chem Soc 135:10396–10403. https://doi.org/10.1021/ja4028346
Tahiliani M et al (2009) Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324:930–935. https://doi.org/10.1126/science.1170116
Hu L et al (2013) Crystal structure of TET2-DNA complex: insight into TET-mediated 5mC oxidation. Cell 155:1545–1555. https://doi.org/10.1016/j.cell.2013.11.020
Sciacovelli M et al (2016) Fumarate is an epigenetic modifier that elicits epithelial-to-mesenchymal transition. Nature 537:544–547. https://doi.org/10.1038/nature19353
Bledea R et al (2019) Functional and topographic effects on DNA methylation in IDH1/2 mutant cancers. Sci Rep 9:16830. https://doi.org/10.1038/s41598-019-53262-7
Morin A et al (2020) TET-mediated hypermethylation primes SDH-deficient cells for HIF2α-driven mesenchymal transition. Cell Rep 30:4551–4566. https://doi.org/10.1016/j.celrep.2020.03.022 (e4557)
Chen LL et al (2022) Itaconate inhibits TET DNA dioxygenases to dampen inflammatory responses. Nat Cell Biol 24:353–363. https://doi.org/10.1038/s41556-022-00853-8
Jakubek M et al (2019) Hydrazones as novel epigenetic modulators: correlation between TET 1 protein inhibition activity and their iron(II) binding ability. Bioorg Chem 88:102809. https://doi.org/10.1016/j.bioorg.2019.02.034
Vissers MCM, Das AB (2018) Potential mechanisms of action for vitamin C in cancer: reviewing the evidence. Front Physiol 9:809. https://doi.org/10.3389/fphys.2018.00809
Brabson JP, Leesang T, Mohammad S, Cimmino L (2021) Epigenetic regulation of genomic stability by vitamin C. Front Genet 12:675780. https://doi.org/10.3389/fgene.2021.675780
Lee Chong T, Ahearn EL, Cimmino L (2019) Reprogramming the epigenome with vitamin C. Front Cell Dev Biol 7:128. https://doi.org/10.3389/fcell.2019.00128
Agathocleous M et al (2017) Ascorbate regulates haematopoietic stem cell function and leukaemogenesis. Nature 549:476–481. https://doi.org/10.1038/nature23876
Chrysanthou S et al (2022) The DNA dioxygenase Tet1 regulates H3K27 modification and embryonic stem cell biology independent of its catalytic activity. Nucleic Acids Res 50:3169–3189. https://doi.org/10.1093/nar/gkac089
Ito K et al (2019) Non-catalytic roles of Tet2 are essential to regulate hematopoietic stem and progenitor cell homeostasis. Cell Rep 28:2480–2490. https://doi.org/10.1016/j.celrep.2019.07.094 (e2484)
Ko M et al (2010) Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468:839–843. https://doi.org/10.1038/nature09586
Rasmussen KD et al (2015) Loss of TET2 in hematopoietic cells leads to DNA hypermethylation of active enhancers and induction of leukemogenesis. Genes Dev 29:910–922. https://doi.org/10.1101/gad.260174.115
Thurman RE et al (2012) The accessible chromatin landscape of the human genome. Nature 489:75–82. https://doi.org/10.1038/nature11232
Neri F et al (2015) TET1 is a tumour suppressor that inhibits colon cancer growth by derepressing inhibitors of the WNT pathway. Oncogene 34:4168–4176. https://doi.org/10.1038/onc.2014.356
Prasad P, Mittal SA, Chongtham J, Mohanty S, Srivastava T (2017) Hypoxia-mediated epigenetic regulation of stemness in brain tumor cells. Stem Cells 35:1468–1478. https://doi.org/10.1002/stem.2621
Herrmann A et al (2020) Integrin alpha6 signaling induces STAT3-TET3-mediated hydroxymethylation of genes critical for maintenance of glioma stem cells. Oncogene 39:2156–2169. https://doi.org/10.1038/s41388-019-1134-6
Wu MZ et al (2015) Hypoxia drives breast tumor malignancy through a TET-TNFα-p38-MAPK signaling axis. Cancer Res 75:3912–3924. https://doi.org/10.1158/0008-5472.CAN-14-3208
Puig I et al (2018) TET2 controls chemoresistant slow-cycling cancer cell survival and tumor recurrence. J Clin Invest 128:3887–3905. https://doi.org/10.1172/JCI96393
Su PH et al (2019) TET1 promotes 5hmC-dependent stemness, and inhibits a 5hmC-independent epithelial-mesenchymal transition, in cervical precancerous lesions. Cancer Lett 450:53–62. https://doi.org/10.1016/j.canlet.2019.01.033
Hart GW, Housley MP, Slawson C (2007) Cycling of O-linked β-N-acetylglucosamine on nucleocytoplasmic proteins. Nature 446:1017–1022. https://doi.org/10.1038/nature05815
Fujiki R et al (2011) GlcNAcylation of histone H2B facilitates its monoubiquitination. Nature 480:557–560. https://doi.org/10.1038/nature10656
Maury JJ et al (2015) RING1B O-GlcNAcylation regulates gene targeting of polycomb repressive complex 1 in human embryonic stem cells. Stem Cell Res 15:182–189. https://doi.org/10.1016/j.scr.2015.06.007
Chu CS et al (2014) O-GlcNAcylation regulates EZH2 protein stability and function. Proc Natl Acad Sci USA 111:1355–1360. https://doi.org/10.1073/pnas.1323226111
Capotosti F et al (2011) O-GlcNAc transferase catalyzes site-specific proteolysis of HCF-1. Cell 144:376–388. https://doi.org/10.1016/j.cell.2010.12.030
Li HJ et al (2021) Roles of ten-eleven translocation family proteins and their O-linked β-N-acetylglucosaminylated forms in cancer development. Oncol Lett 21:1. https://doi.org/10.3892/ol.2020.12262
Iwafuchi-Doi M, Zaret KS (2014) Pioneer transcription factors in cell reprogramming. Genes Dev 28:2679–2692. https://doi.org/10.1101/gad.253443.114
Funding
This work was supported by NIH grants R01 HL133560 and R01 CA223194 through Loyola University Chicago, as well as Loyola program development funds to Jiwang Zhang.
Author information
Authors and Affiliations
Contributions
Kanak Joshi and Shanhui Liu drafted the first version of this review. All of the authors contributed to the writing of this manuscript. Peter Breslin did the final editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no competing financial or professional interests.
Ethics approval
This is not applicable for this review.
Consent to participate
This is not applicable for this review.
Consent for publication
This is not applicable for this review.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Joshi, K., Liu, S., Breslin S.J., P. et al. Mechanisms that regulate the activities of TET proteins. Cell. Mol. Life Sci. 79, 363 (2022). https://doi.org/10.1007/s00018-022-04396-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04396-x