Abstract
Ten-eleven translocation (TET) family proteins (TETs), specifically, TET1, TET2 and TET3, can modify DNA by oxidizing 5-methylcytosine (5mC) iteratively to yield 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC), and 5-carboxycytosine (5caC), and then two of these intermediates (5fC and 5caC) can be excised and return to unmethylated cytosines by thymine-DNA glycosylase (TDG)-mediated base excision repair. Because DNA methylation and demethylation play an important role in numerous biological processes, including zygote formation, embryogenesis, spatial learning and immune homeostasis, the regulation of TETs functions is complicated, and dysregulation of their functions is implicated in many diseases such as myeloid malignancies. In addition, recent studies have demonstrated that TET2 is able to catalyze the hydroxymethylation of RNA to perform post-transcriptional regulation. Notably, catalytic-independent functions of TETs in certain biological contexts have been identified, further highlighting their multifunctional roles. Interestingly, by reactivating the expression of selected target genes, accumulated evidences support the potential therapeutic use of TETs-based DNA methylation editing tools in disorders associated with epigenetic silencing. In this review, we summarize recent key findings in TETs functions, activity regulators at various levels, technological advances in the detection of 5hmC, the main TETs oxidative product, and TETs emerging applications in epigenetic editing. Furthermore, we discuss existing challenges and future directions in this field.
Similar content being viewed by others
Introduction
DNA methylation is one of the most common DNA modifications in mammals, and typically occurs at the CpG dinucleotide site where a methyl group is added to the fifth position of cytosine to generate 5-methylcytosine.1,2,3,4 This process is mediated by DNA methyltransferase (DNMTs). Among them, DNMT3a, DNMT3b, and DNMT3c establish de novo methylation by targeting unmethylated CpG sites, while DNMT1 predominantly serves as a maintenance methyltransferase during cell divisions.5,6 Although DNA methylation is generally stable, it can be removed by active demethylation associated with TET dioxygenases (DNA replication-independent) and passive demethylation (DNA replication-dependent). TET dioxygenases, specifically, TET1, TET2, and TET3 oxidize 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC), and 5-carboxycytosine (5caC) in an Fe (II)/α-ketoglutarate-dependent manner.7,8,9 Notably, 5fC and 5caC can be excised by thymine-DNA glycosylase (TDG), and the modified site returns to the unmethylated status through base excision repair (BER).10,11,12,13,14 Therefore, these enzymes regulate active turnover of DNA methylation. Besides, UHRF1 recognizes 5mC:C dyads and recruits DNMT1 to hemi-methylated CpG sites to maintain DNA methylation.15,16 Disruption of this DNA methylation machinery dilutes 5mC during DNA replication. In addition, 5hmC reduces the affinity of UHRF1 towards 5hmC:C dyads and alters the specificity of DNMT1.17,18,19 Additionally, 5fC:C and 5caC:C dyads are capable of reducing the activity of DNMT1 in vitro.20 These observations suggest that all three oxidation products of TETs (5hmC, 5fC, and 5caC) are poor DNMT1 substrates and are involved in passive DNA demethylation.
TET1 was the first identified member of TET family, acting as a fusion partner of MLL gene in acute myeloid leukemia patients bearing the t(10;11)(q22;q23) translocation, and TET2 and TET3 were subsequently identified based on their significant sequence homology to TET1.21,22 The biological function of TET family was unclear until two landmark discoveries by Kriaucionis et al. 23 and Tahiliani et al. 7. They found TET1 could covert 5mC to 5hmC, which was an Fe (II)/α-ketoglutarate-dependent enzyme by homology searching for JBP1, known as enzymes to oxidize methyl-thymine.7,24,25 Further findings revealed that TET2 and TET3 also could catalyze similar reactions.8 In addition to converting 5mC to 5hmC, TETs were capable of oxidating 5hmC to 5fC and further to 5caC.9
The mechanism underlying TET-mediated demethylation of DNA was not clear until 2011, when two important papers identified that the oxidization products of 5mC, 5fC, and 5caC, could be excised by TDG,10,11 suggesting that TET-mediated oxidization was implicated in active DNA demethylation26,27,28 This was supported by the following study that biochemical reconstitution of TET-TDG-BER system could lead to DNA demethylation29(Fig. 1a).
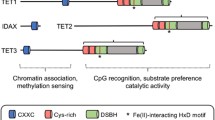
Function and structure of TET proteins. a The dynamic cycle of DNA methylation and demethylation. DNA methyltransferases (DNMTs) catalyzed the formation of 5-methylcytosine (5mC), which can be removed by TET-mediated oxidation, coupled with thymine-DNA glycosylase (TDG)-involved excision and base excision repair (BER). b Domain structure of TET proteins. All TET proteins possessed one core catalytic domain in C-terminal. A CXXC domain, located in the N-terminal of TET1 and TET3, but not in TET2, conferred DNA binding ability directly
Because DNA demethylation and the 5hmC mark involve in various biological reactions, TETs play a very important role in both physiological and pathological processes, which have been elucidated by many studies.13,30,31,32,33,34,35,36 For example, TET2 loss resulted in hypermutagenicity in haematopoietic progenitor cells, unveiling a key role of TET2 in safeguarding cells against genomic mutagenicity.37 Dysfunctions of TET2 in cancer are associated with TET2 mutation and abnormal expression of TET2 regulators.32,38,39,40,41,42 Of note, since 2009, many studies have demonstrated that TET2 mutations were frequently identified in multiple hematologic diseases.43,44,45,46,47,48,49 By genomic sequencing, one study revealed that TET2 mutations were present in 14% (2 of 14) of patients with myelodysplastic syndrome (MDS), 37% (11 of 30) with myelodysplastic/myeloproliferative neoplasms (MDS/MPN), and 43% (6 of 14) with secondary acute myeloid leukemia (sAML) evolved from MDS/MPN. Among the patients harboring TET2 mutations, MDS/MPN accounted for 58% (11 of 19), sAML evolved from MDS/MPN represented 32% (6 of 19), and MDS comprised 10% (2 of 19).44 However, TET2 mutations were infrequent in patients with solid tumors,50 despite somatic mutations in TET1(8 of 74), TET2(5 of 74), and TET3(4 of 74) were identified in colon cancer.51 Other molecular mechanisms underlying the dysregulation of TETs functions in both blood and solid cancers were diverse and complex such as metabolic alterations.52,53 These are discussed in the part of TETs function regulators.
In the following sections, we discuss the structures, functions, and regulators of TETs and summarize the representative methods for 5hmC detection and epigenetic editing.
TET family structure
The primary structure of TETs contains a carboxy-terminal catalytic domain, which is made up of a cysteine-rich domain (CRD), and two double-stranded β-helix (DSBH) regions separated by a large low-complexity insert.7,25 The DSBH domain possesses key residues, responsible for binding to its cofactors (α-ketoglutarate and Fe (II)), which are necessary to its catalytic function.54 Two zinc fingers combine the DSBH and CRD together to form the compact catalytic core.54 Although TET proteins are capable of oxidizing 5mC to 5hmC, 5fC and 5caC, structure analysis revealed that TET2 preferred 5mC substrate, rather than 5hmC and 5fC.55
TET1 and TET3 contain a CXXC domain, located in the amino-terminal region, which is implicated in binding to CpG dinucleotides,56,57 whereas TET2 loses its CXXC domain likely due to a chromosomal inversion (Fig. 1b). Consequently, this allows the ancestral TET2 CXXC domain to be a separate gene called IDAX (also named CXXC4). In the case of TET1 and TET3 with their respective CXXC domains, they can bind with DNA directly. In vitro binding assays revealed that TET1 slightly preferred substrates of unmethylated over that of methylated.58 Further studies showed that, similar to other proteins harboring the CXXC zinc finger domain, TET1 preferentially bound to CpG-enriched promoters of genes, which was certified by chromatin immunoprecipitation of TET1 coupled with DNA sequencing in mouse embryonic stem cells (mESCs).59,60 Similarly, TET3 CXXC-bound regions exhibited a significant enrichment of CpG and more than half of them were enriched in gene promoters.57 This study further revealed that the CXXC domain of TET3 was essential to its biological function through biochemical and structural analysis.57 In contrast, 5hmC regulated by TET2 is mainly located in gene bodies and exons rather than gene promoters.61 Of note, TET1/3 can be also recruited by their binding proteins for context-specific DNA regions.62,63 For example, the pluripotency factor NANOG interacted with TET1 and ChIP-seq analysis identified TET1-NANOG co-binding sites associated with NANOG target genes, suggesting that NANOG guided TET1 to specific sites of chromatin and some DNA-binding proteins were also important to TET1 functions.62
TETs functions and binding partners
The primary functions of TETs are able to oxidate 5mC, and the products are subsequently involved in DNA demethylation.64,65,66 Besides, TET genes expression in different tissues are analyzed using the proteinatlas database67(https://www.proteinatlas.org/) in Fig. 2, which may suggest unique and various functions of TETs in tissues. Evidences also support 5hmC as an epigenetic mark, not only a demethylation intermediate.68 In addition, the non-catalytic activities of TETs are discovered.69 In this section, we discuss the classical and non-classical functions of TETs (Fig. 3). As TETs binding partners appear to be their main regulators, we also summarize their information here (Tables 1–3).
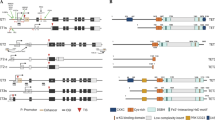
Expression levels of TET1(a), TET2(b), and TET3(c) in different tissues. The data were obtained from the proteinatlas.67 Modified from images available for TET1 (https://www.proteinatlas.org/ENSG00000138336-TET1/tissue), for TET2 (https://www.proteinatlas.org/ENSG00000168769-TET2/tissue), and for TET3(https://www.proteinatlas.org/ENSG00000187605-TET3/tissue). nTPM: consensus normalized expression
TETs working models. a A particular transcription factor (TF) recruited TETs to specific DNA areas of promoters to increase the downstream gene expression in a dioxygenase activity-dependent manner.100,120 b TETs binding with other epigenetic regulatory enzymes, such as HDAC, together regulated the particular gene expression independent of TETs enzyme activity.183,187 c, d TET1/2 oxidated mRNA (c)191,194 and TET2 oxidated tRNA (d)196 to exhibit regulatory functions in RNA levels
Classical functions of TETs
TET1
Studies have shown the important role of TET1 in physiological functions, including development.70 The deficiency of TET1 lowered female germ-cell numbers by controlling meiosis through mediating related-gene DNA demethylation,71 while TET1 loss was dispensable for mice embryonic and postnatal development.72 However, acute deletion of TET1 caused a significant decrease of 5hmC levels and impaired embryonic stem cell identity,73 possibly because a long-term chronic reduction in TET1 led to homeostatic compensation.73 TET1 is also essential for intestinal stem cell functions in vivo74 and implicated in dynamic changes of DNA methylation during the maturation of fetal intestinal epithelial organoids in vitro.75 Epigenetic programming by catalytic-dependent TET1 is implicated in liver regeneration76 and remyelination in mouse brain.77,78 TET1 is involved in regulating iron homeostasis by demethylating the promoter of RNF217 and this ubiquitin ligase is responsible for the degradation of iron exporter ferroportin.79 TET1-deficient mice exhibit impaired spatial learning and memory.80 Findings also support the roles of TET1 in promoting pluripotent stem cell induction.81,82,83,84 In addition, TET1 is required in the reprogramming of fibroblasts to dopaminergic neurons.85
Abnormal expression of TET1 is associated with many diseases.86,87,88,89,90,91,92 Loss of TET1 led to B cell malignancy in aged mice, partly due to aberrant DNA-hypermethylation,93 although TET1 acted as an oncogene in MLL-rearranged leukemia.94 Additionally, insufficient TET1 was implicated in pulmonary arterial hypertension.95 Interestingly, overexpression of TET1 promoted cholangiocarcinoma progress via proliferative and anti-apoptotic signaling pathways,96 while insufficient TET1 accelerated intestinal tumorigenesis.97 Besides, high expression of TET1 appeared to be involved in polycystic ovary syndrome with hypomethylation signatures.98
Protein interactions enable rapid regulation and represent an important regulation in TETs functions, which allow precise modifications in specific DNA loci timely.99 For example, TET1 interacted with STAT1, contributed to the demethylation of IRF1 promoter and transcriptional upregulation of PD-L1, to drive tumor immune evasion.100 In addition, interestingly, FXR1, an m6A reader, guided TET1 to specific genomic loci near m6A RNA to result in DNA demethylation, revealing a novel regulation between RNA modification and DNA demethylation.101 Similar models have been supported by many findings, which are summarized in Table 1.
TET2
Unlike TET1 and TET3, TET2 mutations with high frequency are identified in hematologic malignancies.102,103,104,105 Thus, the relationship between TET2 mutations and overall survival has been investigated. Evidence showed that the patients with TET2 mutations had worse overall survival compared with the patients with wild-type TET2 in 93 patients with AML.38 However, other studies showed no survival association in 111 patients with de novo AML106 and in a cohort of 247 patients with secondary AML.107 Therefore, the significance of TET2 mutations in AML prognosis remains to be elucidated. The effects of TET2 mutations on its functions, such as enzymatic activity and the ability of binding other proteins, and potential confounding variables should be considered. Further studies suggest that TET2 works as a tumor suppressor.108,109,110,111,112 Interestingly, findings also reveal tumor-promoting roles of TET2.113,114 For example, TET2 maintained the immunosuppressive-related gene expression in tumor-associated macrophages.115
As TET2 does not contain the CXXC domain, this raises a question of how TET2 is bound with the chromatin? One reasonable hypothesis is that, IDAX, originating from the ancestral TET2 CXXC domain, mediates its chromatin recruitment. Indeed, biochemical studies demonstrated that IDAX could bind to TET2 directly, suggesting that IDAX was able to recruit TET2 to DNA.116
In addition to IDAX, some other TET2-binding proteins have been identified by biochemical studies (Table 2). For instance, TET2 interacted with NANOG and played an important role in the establishment of pluripotency in a NANOG-dependent manner.117 EBF1, a transcription factor, had also been identified as a TET2-binding protein by co-immunoprecipitation of TET2 and EBF1. Importantly, sequence analyzing revealed that these two proteins were enriched in a proportional way, implying that TET2, without a CXXC DNA-binding motif, exploited a DNA-binding protein, such as a transcription factor, to regulate sequence-specific DNA demethylation.118 This idea was reinforced by the interaction of TET2 with WT1.119,120 Further studies revealed that mutations of TET2 and WT1 were mutual exclusively in AML, and WT1 guided TET2 to a specific DNA sequence, leading to the demethylation and activation of WT1-target genes.120 Many following studies supported this model, in which a DNA-binding factor recruited TET2 to a specific DNA sequence and regulated the expression of this gene in certain contexts.121,122,123,124,125,126,127 For instance, we found that the transcription factor HNF4α could recruit TET2 to FBP1 promoters, resulting in the increase of FBP1 expression, to suppress the tumor growth.127 These models relied on the oxygenase activity of TET2.
TET3
As a member of the TET family, the main role of TET3 is implicated in demethylation in many biological processes such as zygote formation,128,129,130,131 embryogenesis,132 axon regeneration,133 and synaptic transmission.134 For example, TET3-mediated DNA demethylation is necessary for liver tissue maturation via proper hepatocyte gene expression.135 In addition, TET3 deficiency induced by mutations is associated with abnormal growth and intellectual disability,136 indicating the fundamental role of TET3 in development. In adult mice, TET3 ablation is associated with anxiety-like behaviors, although the molecular mechanisms remain to be explored.137
Interestingly, hepatic TET3 was recruited to the promoters of the fetal version of HNF4α by FOXA2, contributing to high expression of HNF4α transcription by promoter demethylation, and this process impaired glucose homeostasis due to HNF4α-mediated gluconeogenesis activation. Thus, these findings linked TET3 to type-2 diabetes.138 In addition, insufficient demethylation of several insulin secretion genes, owing to the maternal inheritance of oocyte TET3 insufficiency, contributed to glucose intolerance.139 These findings demonstrated the distinct roles of TET3 in certain contexts. Similar to TET2, binding partners are involved in TET3 function regulation (Table 3). For example, PGC7 interacted with TET3 and suppressed TET3 enzymatic activity to protect DNA methylation at imprinting loci during early embryogenesis,140 although PGC7 bound to H3K9me2 to block the TET3-mediated conversion of 5mC to 5hmC.141
TET1/2/3
Furthermore, in some biological contexts, TETs cooperate with each other to orchestrate specific functions. For instance, TET1 and TET2 are involved in pluripotent reprogramming and imprint erasure induced by cell fusion,142 erasure of 5mC in mouse primordial germ cells,143 pre-mRNA alternative splicing,144 maintaining stem cell identity,145 reprogramming to recover youthful DNA methylation patterns in aged mice146 and epigenetic reprogramming in offspring caused by maternal exercise.147 Binding proteins are required for desired functions in some cases. For example, upon TGF-β and IL-2 signaling, TET1 and TET2, recruited by SMAD3 and STAT5, bound to and subsequently demethylated FOXP3 promoter to maintain immune homeostasis.148 Similarly, to main bone homeostasis, both TET1 and TET2 were required for demethylating promoters of P2RX7.149 Additionally, TET1 and TET3 are associated with cerebellar circuit formation150 and CD4 expression in peripheral T cells.151
TET2 and TET3 are required for Treg cell stability and immune homeostasis,152 and improve Treg cell efficacy by increasing the stability of FOXP3.153 TET2 and TET3 acted as recruiters of HDACs to suppress CD86 and prevent autoimmunity.154 Findings also reveal the roles of TET2 and TET3 in embryonic heart development155 and in regulating proper development and maturation of invariant natural killer T cells.156 Knockdown of TET2 led to hyper-proliferation of erythroid progenitors, whereas knockdown of TET3 impaired terminal erythroid differentiation. These findings revealed distinct roles of TET2 and TET3 in the regulation of human erythropoiesis.157 Furthermore, the deletion of TET2 and TET3 led to aggressive myeloid cancer in mice.158 Mice with TET2 and TET3 double knockout in mature B cells developed B cell lymphoma, which can be delayed upon DNMT1 deletion,159 suggesting the importance of proper methylome.
TET1, TET2, and TET3 are required for somatic cell reprogramming of fibroblasts to pluripotency,160 telomere homeostasis,161 and early body plan formation.162 Human embryonic stem cells (hESCs) with triple-knockout of TET1, TET2, and TET3 exhibited prominent bivalent promoter hypermethylation, suggesting the role of TETs in maintaining hypomethylation at bivalent promoters to ensure proper lineage-specific transcription during differentiation.163 In mESCs, TETs tended to increase demethylation rates at enhancer elements.164 Distinct roles of TETs in regulating 5hmC formation, DNA demethylation, and gene expression are also explored in cancer cells.165
The overlapping roles of TETs have been explored due to their similar enzymatic activity. Mice with loss of either TET172 or TET2166 are viable, while most TET1/2 double knockout mice die perinatally,167 suggesting that deletion of the individual TET gene can be compensated by other TETs. Interestingly, TET3 knockout leads to neonatal lethality,128 indicating the unique role of TET3 that could not be compensated by the other TETs. Thus, the overlapping roles of TETs in certain contexts have not yet been fully established. In addition, to understand the TETs functions in vivo, mouse models with gene constitutive or conditional knockout have been generated, some of which are summarized in Table 4.
5hmC
TETs-mediated 5hmC formation appears to be an epigenetic mark, although the physiological significance has not been fully elucidated.168,169,170 The 5hmC acquisition occurred in mouse, rabbit, and bovine zygotes,171 indicating that the mark was conserved in these mammalian species. MBD3, required for pluripotency in ESCs,172 preferred to binding 5hmC-containing probes rather than 5mC-containing probes and regulated the expression of genes with 5hmC modifications in ESCs.173 In addition, the acquisition of 5hmC by TET1 in enhancers was associated with enhancer activation,174 implying that 5hmC represented a signal mark rather than an intermediate. The idea was supported by the role of 5hmC in germline reprogramming175 and in drug addiction.176 Interestingly, particular 5hmC acquisition by cocaine lasted at least one month in mouse nucleus accumbens.176 Besides, TET1-mediated 5hmC deposition was also implicated in osteoarthritis.177
Interestingly, 5hmC formation is not required for the loss of paternal 5mC in early mouse zygotes,178 further supporting the fascinating and mysterious role of 5hmC, not just the demethylation intermediate. 5hmC modifications have been reported to affect protein binding,179 and consistently, 5hmC might recruit a chromatin-modifying complex to suppress transcription.180 5hmC formation caused by TET3, prevented spurious transcription, which was critical for maintaining transcriptional fidelity in the lung.181
Moonlighting functions of TETs
Overexpression of either TET1 or catalytic-death TET1 impaired long-term memory in mice, suggesting the catalytic-independent function of TET1.182 Furthermore, TET1 acted as an epigenetic suppressor of thermogenesis in beige adipocytes largely independent of its catalytic activity. Specifically, TET1 interacted with HDAC1 to suppress key thermogenic gene transcription by reducing histone acetylation.183 Consistently, catalytic-independent functions of TET1 in silencing developmental genes by regulating H3K27 modifications,184 supported that TET1 acted as an interaction hub for recruiting different chromatin-modifying complexes in a non-catalytic manner.185 Besides, the non-catalytic function of TET3 in transcriptional repression of SNRPN by binding to SNRPN promoter, was critical for the maintenance of adult neural stem cell state.186
Apart from the ability of DNA oxygenase, studies also unveiled that TET2 could reduce inflammation by repressing IL-6, which is independent of its role in converting DNA 5mC to 5hmC. Specifically, TET2, binding with IκBζ, recruited HDAC2 to promote histone deacetylation, which led to the repression of IL-6 at the transcription level. These findings provided a TET2 enzymatic-independent function in repressing specific gene transcription.187
To explore the enzymatic versus nonenzymatic roles of TET2 in hematopoiesis, Ito et al. performed a comparative analysis of TET2 catalytic mutant mice and TET2 knockout mice. This study found that mice with non-catalytic TET2 mainly developed myeloid malignancies, while mice with complete loss of TET2 developed both myeloid and lymphoid disorders, supporting the unique non-catalytic role of TET2 in the hematopoietic stem and progenitor cell homeostasis.188
Interestingly, besides its well-known function in regulating the modification of DNA, TET2 possessed the activity of oxidating 5mC RNA (m5C) into 5-hydroxymethylcytidine (hm5C). Fu et al. found that the catalytic domain of TET2 could induce the formation of hm5C in HEK293T cells. Considering hm5C accounting for approximately 0.02% of total m5C RNA in tumor samples, this implied the involvement of TET2 in RNA biology.189 Consistently, a study in Drosophila showed that TET protein was involved in the formation of hm5C.190 This study also mapped the distribution of hm5C and revealed hm5C located in coding sequences of many gene transcripts. Importantly, hm5C favors mRNA translation.190 However, the biofunction of hm5C in mammalian RNA is largely unknown until Shen et al. discovered that TET2 was involved in RNA stability.191 These findings uncovered that TET2, depending on its enzymatic activity of mRNA oxidation, promoted pathogen infection-associated myelopoiesis. Specifically, TET2 mediated oxidation of SOCS3 m5C, which led to ADAR1 binding and destabilizing SOCS3 mRNA and consequently repressed SOCS3 expression.191 Meanwhile, by the TET2 interactome in mouse ESCs, Guallar et al. identified that paraspeckle component 1(PSPC1), an RNA-binding protein, could bind to TET2 and this complex recruited HDAC1/2 for repression of MERVL transcription independent of TET2 catalytic activity. More importantly, this study further found that TET2, recruited by PSPC1, catalyzed hm5C modification of MERVL RNAs, facilitating the degradation of MERVL transcripts, and thus provided a new paradigm for TET2-mediated post-transcriptional silencing of the specific gene. Notably, PSPC1 and its RNA-binding domains are essential for TET2 function in regulating MERVL by both transcriptional and post-transcriptional mechanisms.192 Interestingly, using a proteomics approach, Huang et al. discovered PSPC1 also bound to TET1 for bivalent gene regulation in formative pluripotency independent of the catalytic activity of TET1.193 Additionally, TET2 has been shown to function in ESC differentiation by reducing the pluripotency-related mRNA stability, caused by TET2-mediated hm5C.194 Notably, this study confirmed that TET2 contained an RNA-binding domain, which had been identified by a proteomic approach in a previous study.195
In addition to its oxidation of mRNA, recently, He et al. found that TET2 could convert m5C into hm5C in tRNA, subsequently affecting tRNA fragment levels.196 Meanwhile, m5C oxidation in tRNA mediated by TET2 facilitated translation.197 These findings linked TET2-mediated tRNA modification to tRNA processing and mRNA translation,196,197 unveiling novel roles of TET2 in gene regulation at multiple levels. Additionally, findings revealed that TET1/2 could oxidize T to 5hmU in mESCs.198
In this part, the interaction with various binding proteins stably and transiently mainly affects TETs location, including recruiting TETs to specific sites, allowing TETs to recruit other proteins, and stabilizing TETs association with DNA. Besides, TETs are capable of oxidizing both DNA and RNA. Understanding the characteristics of TETs might provide key insights into epigenetic editing, such as DNA demethylation and mRNA modification. Here, we summarize major discoveries in the history of TETs over time (Fig. 4).
TET function regulators
Numerous studies have identified the factors in regulating TETs function, including transcription factors, microRNAs, post-translational modifications, and small molecules in different levels, as summarized in Fig. 5.
Primary factors in positive(red) and negative(blue) regulation of TETs activity and the outmost layer described the involved mechanism. FOXA1,201 Vitamin C,238 CRL4(VprBP),224 P300,222 and AMPK223 enhanced TETs activity, while Vpr,227 IDAX,116 Calpain 1,228 2HG232 and miR-29b207 decreased TETs activity
Transcriptional level
Pluripotency-associated transcriptional factors, such as MYC and NANOG, regulated TET1 expression in hESCs.199 TET1/2 was regulated by Oct4 and SOX2.200 Interestingly, FOXA1 not only transcriptionally regulated TET1, but also interacted with TET1 to mediate DNA demethylation of its targeted enhancers.201 Besides, STAT3/5 transcriptionally activated TET1 expression in AML202 and P53 positively regulates TET1/2 transcription in mESCs.203 SIN3A increased TET1 and TET2 mRNA expression in human pulmonary arterial smooth muscle cells.95 Transcriptional suppression was also identified in the regulation of TETs. NF-κB-mediated repression of TET1 transcription was uncovered in basal-like breast cancer.204 TET3, transcriptionally repressed by the nuclear receptor TLX, acted as a tumor suppressor in glioblastoma.205
microRNAs
Using the TCGA database, the miR-29 family were predicted to regulate DNA demethylation by potentially targeting TET1.206 Indeed, miR-29b directly targeted and repressed TET1 to promote the mesendoderm lineage formation207 and the miR-19b/TET1 axis could be utilized in attenuating osteoarthritis progression.208 Downregulation of TET1 also has been reported by miR-494 in hepatocellular carcinoma tumors209 and by miR-191 in intrahepatic cholangiocarcinoma,210 respectively.
Multiple studies have also demonstrated that microRNAs are involved in downregulating TET2 expression. Biochemistry studies discovered that miRNA-22 could directly bind to TET2 mRNA and negatively regulate TET2 expression, which contributed to myelodysplastic syndrome and hematological malignancies.211 miRNA-29a could also downregulate TET2 expression.212 A thorough analysis of TET2-targeting miRNA by a high-throughout 3’UTR screen, identified extensive miRNAs such as miRNA-29b and miRNA-101, inhibiting TET2 expression and these miRNAs regulated malignant hematopoiesis.213 Further studies identified that TET2 was under control of miRNA-Let7,214 miRNA-210215 miRNA-144-3p,216 miRNA-142-3p,217 miRNA-26a218 and miRNA-10b-5p.219 Besides, TET3 as a miR-150 target was associated with the generation of non-classical monocytes.220
Post-translational modifications
Post-translational modification is also a key process in regulating TETs functions. Bauer et al. found that phosphorylation and O-GlcNAcylation existed in TET2 protein modification,221 indicating that complex modification modulated TET2 functions in different conditions. Indeed, P300-mediated acetylation of conserved lysine residues enhanced TET2 stability, and increased its ability to target chromatin, which reduced aberrant DNA methylation, and thereby protected against abnormal DNA methylation induced by DNA damage.222 Additionally, AMP-activated kinase catalyzed the phosphorylation of TET2 at serine 99, which increased the stability of TET2. While the phosphorylation of TET2 was inhibited under hyperglycaemic conditions such as diabetes, consequently decreasing TET2 levels.223 Monoubiquitylation of TET2 at lysine 1299 mediated by VprBP facilitated TET2 association to chromatin, whereas mutation of TET2 at 1299 blocked its interaction with VprBP and decreased its association with DNA.224 Interestingly, the K1299-linked monoubiquitylation of TET2 could be removed by USP15, decreasing TET2 association to DNA.225 Additionally, phosphorylation of TET3 by CDK5 caused lower binding affinity to histone variant H2A.Z. and contributed to higher level of 5hmC at BRN2 promoter to activate BRN2 expression during neuronal differentiation.226
Protein degradation
Surprisingly, besides the CRL4 E3 ligase mediated TET2 monoubiquitylation promoted TET2 association to chromatin, HIV-1 derived Vpr hijacked CRL4, and this E3 ligase preferred to catalyze polyubiquitylation of TET2, accordingly promoting TET2 degradation to sustain IL-6 expression and facilitate viral replication.227 Unexpectedly, IDAX, the TET2-binding protein, promoted TET2 degradation in a caspase activation-dependent manner.116 With different proteolytic pathway inhibitors, calpains were identified to be involved in TET2 protein regulation. Specifically, calpain 1 was implicated in the degradation of TET2 in ESCs, leading to skewing lineage expression.228
Small molecules
As α-KG is required to maintain the oxygenase activity of TETs, it is plausible that 2HG, generated by the reduction of α-KG catalyzed by IDH enzyme mutants,229 might disrupt TETs function.230 Indeed, biochemistry studies demonstrated that mutant IDH decreased TET2-mediated 5hmC levels.231 Consistently, structure analysis revealed that 2HG occupied the site of α-KG in protein conformational space, suggesting that 2HG served as a competitive inhibitor of α-KG-dependent enzyme activity, including TET2.232 In addition to 2HG, succinate and fumarate were also identified to act as α-KG antagonists, which inhibited TET2 dioxygenase activity.233 Recently, Chen et al. found that itaconate was also a TET2 dioxygenase inhibitor through the competition with α-KG to interact with TET2, resulting in dampening inflammatory responses.234 Besides, the nuclear glutamate dehydrogenase interacted with TET3 to supply TET3 with αKG and increased its demethylation activity in neurons.235
Previous studies have revealed that vitamin C could upregulate the activity of some α-KG-dependent dioxygenases, suggesting that vitamin C might be involved in the modulation of TETs activity. Indeed, vitamin C could enhance TET2 activity and subsequently increase 5hmC levels in ESCs.236,237 Yin et al. found that vitamin C, but not other reducing chemicals such as NADPH and vitamin E, was a unique activator of TET dioxygenases.238 It is possible because vitamin C was capable of binding to the catalytic domain of TET proteins, facilitating protein folding, and accelerating oxidation reactions.238 The idea, that vitamin C acting as a TET agonist, was reinforced by a series of further studies.239,240,241,242,243,244,245 Notably, TET2 deficiency presented in aberrant self-renewal and leukemia progression, which can be blocked by treatment with vitamin C, suggesting that vitamin C treatment might be beneficial to patients with leukemia.246 Specifically, vitamin C restored TETs function and drove the expression of related genes.246
Aside from metabolites, Thienpont et al. found that the activity of TET2 was reduced under hypoxic conditions, leading to DNA-hypermethylation.247 Oxygen levels determined the activity of TET1 in ESCs.248 Redox-active quinones promoted the production of 5hmC by TETs.249
Artificial inhibitors and activators of TETs have also been explored. By screening strategy, a small molecule compound, C35, was identified as a TETs inhibitor. Notably, this compound specifically blocked TETs catalytic activities without abolishing TETs complexes.250 Bobcat339, one of synthesized cytosine derivatives, inhibits TET1 and TET2 activity.251 A small molecule, TETi76, inhibits TETs specifically.252 Interestingly, Nickel (II) exhibits inhibition to TETs enzymatic activities by replacing the cofactor Fe (II) of TETs.253 Additionally, SRT1720, a SIRT1 agonist, by deacetylating TET2, significantly increases TET2 activity.254
Together, similar to other genes, TETs can be regulated at multiple levels, including post-transcriptional and post-translational regulation. Furthermore, it can be modulated by small molecules involved in its enzymatic reaction. This ensured the fine-tuning of TETs enzymatic activity in response to external cues.
Targeted therapy and clinical trials
Given the various roles of TETs in biological processes, it comes as no surprise that it has been proposed as an important therapeutic target for diseases such as cancer.113,114,252,255,256 For example, vitamin C, by improving TETs activity, allows leukemia cells to be more sensitive to PARP inhibitors.246 Interestingly, cells with TET2 mutations, possibly heavily relying on compensatory roles of TET1/3, showed more vulnerable to TETs inhibitors compared with normal ones. These findings provide a new therapeutic strategy for selective targeting of cells bearing TET2 mutations.252 5-azacytidine, a DNA demethylating agent, shows higher cytotoxicity in TET2-silenced cells, probably due to the hypermethylation pattern caused by the loss of TET2.256 In addition, C35, a selective TETs inhibitor, promotes somatic cell reprogramming.250 As a robust TET2 activator, clinical trials are investigating the effects of Vitamin C on hematologic malignancy patients with TET2 mutations (NCT03397173; NCT03433781). Of note, the antitumor effects of vitamin C has been studied for a long time; however, its efficacy against cancers have not been established by clinical trials, possibly because of the complex mechanisms of action of vitamin C.257,258,259,260,261,262 As a new target, the role of TET2 enzymatic activity enhanced by vitamin C in patients with hematological malignancies remains unclear. Besides, clinical trials evaluating the contribution of vitamin C-mediated upregulating TET2 enzymatic activity in solid tumors are urgently required. Notably, high concentrations of vitamin C administration with or without anticancer drugs have not shown serious adverse effects in clinical trials, suggesting that vitamin C is a drug with low toxicity.263,264,265,266,267 Therefore, vitamin C might be a promising anticancer treatment option for cancer patients with dysfunctions of TET2 in the future.
Detection of 5hmC
5hmC plays distinct epigenetic roles in mESCs.268,269 In addition, aberrant levels of 5hmC are associated with various cancers.270,271,272,273,274,275,276,277,278 Furthermore, 5hmC signatures in circulating cell-free DNA can be used as biomarkers for cancer diagnosis.279,280,281,282,283 Together, mapping the distribution of 5hmC in a genome is important not only to elucidate its biology, such as functions in development, but also to use it for clinical potential.284,285,286,287,288 In this section, representative approaches for detecting 5hmC with or without bisulfite treatment are discussed (Fig. 6).
hMeDIP
To investigate the global distribution of 5hmC, anti-5hmC antibodies were utilized to capture 5hmC DNA from genomic DNA followed by sequencing, and this approach was named as hMeDIP.289,290,291 This method is cost-effective and widely used. However, the biggest limitation of this method is the quality of anti-5hmC antibodies. To solve the problem caused by using antibodies of different production batches, Robertson et al. developed a novel 5hmC detection method, based on the selective glycosylation of 5hmC treated with β-glucosyltransferase. This β-glucosyl-5-hydroxymethylcytosine-containing DNA could be efficiently and specifically captured by J-binding protein 1. After enriching 5hmC, further analysis could be performed, such as qPCR and sequencing.292,293 Likewise, 5hmC was converted to cytosine-5-methylenesulfonate (CMS) upon sodium bisulfite treatment, and then the CMS-specific antiserum was used to capture CMS-containing DNA fragments for further analysis.294,295
hMe-Seal
Bisulfite treatment could lead to significant degradation of DNA, and therefore bisulfite-free methods were developed for limited DNA samples. For example, β-glucosyltransferase could convert 5hmC to β-glucosyl-5-hydroxymethylcytosine (5gmC) in the presence of UDP-Glu. The 5hmC can be labeled with an azide group using the modified UDP-Glu with the azide. This allowed biotin moiety containing an alkynyl group to link to 5hmC using click chemistry, followed by affinity enrichment and sequencing.296
TAB-Seq
In 2012, Yu et al. developed a TET-assisted bisulfite sequencing approach, named as TAB-Seq, which enabled the detection of genomic 5hmC sites at single-base resolution. Specifically, 5mC could be oxidized to 5caC with TET proteins and the 5caC could subsequently be deaminated to form U by bisulfite treatment, while the glucosylated-5hmC was protected from TET oxidation and bisulfite deamination and therefore was identified as C. This method allowed discriminating 5hmC from 5mC, in contrast with traditional bisulfite sequencing.297
oxBS-Seq
Meanwhile, Booth et al. also developed a method of quantitatively mapping 5hmC distribution at single-base resolution, known as oxidative bisulfite sequencing (oxBS-Seq). This approach utilized potassium perruthenate to selectively oxidate 5hmC to 5fC that was subsequently converted to U by bisulfite treatment, while 5mC was not oxidized by potassium perruthenate and still detected as C. This method enabled the determination of the amount of specific 5hmC sites by subtracting the readout of traditional bisulfite sequencing.298
hmC-CATCH
Similar to oxBS-Seq, potassium ruthenate was used to convert 5hmC to 5fC, which was further selectively modified with an azido, and this adduct was identified as T during PCR. Therefore, the C-to-T transition was regarded as the readout of 5hmC. Additionally, the azido group rendered it easily for enrichment and sequencing.299
CAPS
Similar to TAB-seq, TETs were employed to convert both 5mC and 5hmC to 5caC, and pyridine borane was subsequently used to convert 5caC to dihydrouracil, that was read as T during PCR. This modified C-to-T transition allowed whole-genome detection of 5mC and 5hmC at single base-level resolution. In contrast, glucosylated-5hmC was inert to TET oxidation and borane reduction, and thus 5mC sites could be analyzed specifically.300 Accordingly, the amount of 5hmC sites could also be determined by comparing the readouts with or without β-glucosyltransferase treatment at the first step. Alternatively, TET proteins could be replaced by potassium perruthenate to selectively oxidate 5hmC to 5fC, allowing specifical sequencing of 5hmC.301
Jump-seq
A new strategy, called Jump-seq, was developed by Hu et al. for detecting 5hmC without sequencing the whole genome at nearly a single-base resolution. This method took advantage of selectively labeling 5hmC with a glucose moiety carrying an azide group, followed by linking a hairpin DNA with an alkyne group. 5hmC positions could be deduced by the connection between genomic DNA sequence and the hairpin sequence after primer extension.302
ACE-seq
APOBEC3A-based 5hmC sequencing method, named ACE-seq, has been developed without bisulfite treatment at single-base resolution. 5hmC was modified with glucose by β-glucosyltransferase and the glucose-modified 5hmC was inert to APOBEC3A, a DNA deaminase, whereas C and 5mC could be converted to U, yielding 5hmC identified particularly.303
CAM-Seq
With a similar strategy, 5hmC was initially converted to 5fC by KRuO4. Then using azi-BP, a compound reported by the same group, 5fC was selectively labeled, rendering it matching with A and identified as T by PCR. Using this method 5hmC loci in genomic DNA could be analyzed at single-base resolution.304
As the findings of the important role of TET families in DNA modification, selective chemical labeling of the hydroxyl group of 5hmC is fast-growing to map the genome-wide distribution of 5hmC. Here, we summarize some characteristics of each method in Table 5. Of note, recently, nanopore sequencing technologies have shown a diverse range of applications, including 5hmC detection.305,306 In addition, hm5C could be detected by mass spectrometry.307,308 Like hMeDIP, hm5C-containing RNA could be captured by the anti-hm5C antibody followed by sequencing and this method was named as hMeRIP-seq.190,194
Demethylation editing tools
Dynamic regulation of DNA methylation and demethylation plays a critical role in many biological processes, including epigenetic memory, genomic imprinting, and development.309,310,311 Dysregulation of this process leads to many diseases such as autoimmune disorders and cancers.312,313,314 In addition, hypermethylation patterns are usually associated with gene silencing. Therefore, developing epigenetic editing tools allow us not only to modify the target locus to evaluate the consequences of epigenetic marks, but also to silence or activate the gene in specific contexts. The general idea of epigenetic editing is that an epigenetic writer or eraser is fused to a sequence-specific DNA-binding domain to rewrite the epigenetic marks in targeted loci or histone315,316(Fig. 7). In this part, we summarize TETs-based epigenetic editing tools (Table 6).
Representative working models of targeted demethylation. a With the gDNA, a DNA modification domain, such as TET1, fused to dCas9, led to the erasure of specific DNA methylation.333 b GCN4 repeats fused to dCas9, recruited many copies of an anti-GCN4 antibody (scFV)-fused TET1, to amplify demethylation efficiently.322 c Multiple effectors were used to increase the efficacy of demethylation. The modified gRNA with PUF binding sites, recruited protein fusions of PUF, TET1, and NEIL2 to particular DNA methylation sites. Among these, PUF were used for binding to the modified gRNA, and TET1 oxidated 5mC and NEIL2 worked as a DNA glycosylase to promote DNA demethylation.324 d Without tethering an effector such as TET1, only gRNA-dCas9 led to specific DNA demethylation, by sterically blocking DNA methyltransferase331
TET1-TALE-fused-based tools were developed for epigenetic editing,317,318 and successfully increased β cell replication, demonstrating a promising approach in therapeutic applications.318 Customized TALE repeat arrays worked as a platform for guiding TET1 to the DNA sequence of interest, therefore leading to the demethylation of targeted loci, and subsequently increasing the related gene expression.317
Additionally, engineered endonuclease-dead Cas9 (dCas9) could also be used as a linker, and recruited indirectly or fused directly to the designed effector domains, such as TET1, to modify the specific target in conjunction with gRNA.319,320,321,322,323 Furthermore, co-delivery of demethylation pathway-related proteins such as GADD45A and NEIL2, with dCas9-TET1, enhanced demethylation editing efficacy.324 Besides, the CRISPR/dCas9-based gene transcription activation system coupled with TET1, activated silenced genes through demethylating.325,326
In addition to dCas9, other DNA-binding domains worked as a target loci modification guider. For example, a synthetic fusion protein, carrying enzymatic domains of TET1 and reverse tetracycline transactivator, exhibited demethylation of Tet promoter, upon doxycycline treatment.327 Similarly, TET2 was fused to a DNA-binding domain to promote the demethylation of targeted loci, and thereby a TET2-based editing approach was developed.328 The engineered protein contained two core domains: TET2 for inducing DNA demethylation and zinc fingers for binding the ICAM-1 promoters.328
Other effectors could also be employed such as TET3 and ROS1. TET3 catalytic domains, fused to dCas9, could produce 5hmC formation.329 Plants DNA demethylases such as ROS1 could replace TET1 to induce demethylation.330 Interestingly, simple CRISPR/dCas9 and gDNA without tethering any other enzymes appeared to demethylate target loci efficiently largely due to steric blockage of DNA methyltransferase.331
Methylation editing tools have shown great potential in clinical research and treatment. Model mice with Silver-Russell syndrome has been successfully generated by TET1-dCas9 based system.332 TET1-based DNA methylation editing could restore the expression of FMR1 by demethylating its promoter, supporting the potential application of epigenome editing in fragile X syndrome treatment.333 Similarly, TET1-dCas9 mediated demethylation of the MECP2 promoter, rescued Rett syndrome neurons.334 Thus, precise and efficient epigenetic editing tools would provide new insights into the functions of the specific DNA modification locus temporal-spatially.
Summary
Here we review the remarkable findings in understanding the function of TETs in modifications of DNA and RNA, and summarize recent advances in the detection of 5hmC and DNA demethylation editing tools. Despite the formation of oxidation products (5hmC, 5fC, and 5caC) and the mechanism of active DNA demethylation have been characterized, some questions have yet to be answered. First, the significance of 5hmC needs to be delineated. Second, in addition to oxidating DNA, recent studies have also demonstrated that TET2 is capable of oxidating RNA. It is still not well-defined what factors determine TET2 in choosing oxidating DNA or RNA. Third, regardless of containing DNA-binding domain, all TETs appear to be recruited to specific DNA sequences by their binding partners. It is worth to further explore how to modulate the binding of TETs to its target DNA sequences in various biological processes. Fourth, loss of function mutations of TET2 are frequently identified in blood malignancies, whereas mutations of TET2 are uncommon in solid tumors. However, significant downregulation of TET2 activity is observed in many solid tumors. The underlying mechanisms are still not clear and require to be explored for the diagnosis and therapy of cancers. We believe that addressing the questions above will help us further understand the roles of TETs in the occurrence and development of many diseases.
References
Parry, A., Rulands, S. & Reik, W. Active turnover of DNA methylation during cell fate decisions. Nat. Rev. Genet. 22, 59–66 (2021).
Moore, L. D., Le, T. & Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 38, 23–38 (2013).
Ballestar, E., Sawalha, A. H. & Lu, Q. Clinical value of DNA methylation markers in autoimmune rheumatic diseases. Nat. Rev. Rheumatol. 16, 514–524 (2020).
Greenberg, M. V. C. & Bourc'his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 20, 590–607 (2019).
Li, E. & Zhang, Y. DNA methylation in mammals. Cold Spring Harb. Perspect. Biol. 6, a019133 (2014).
Chen, Z. & Zhang, Y. Role of mammalian DNA methyltransferases in development. Annu. Rev. Biochem. 89, 135–158 (2020).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Maiti, A. & Drohat, A. C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J. Biol. Chem. 286, 35334–35338 (2011).
Kohli, R. M. & Zhang, Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502, 472–479 (2013).
Rasmussen, K. D. & Helin, K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 30, 733–750 (2016).
Lu, X., Zhao, B. S. & He, C. TET family proteins: oxidation activity, interacting molecules, and functions in diseases. Chem. Rev. 115, 2225–2239 (2015).
Bostick, M. et al. UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317, 1760–1764 (2007).
Sharif, J. et al. The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmt1 to methylated DNA. Nature 450, 908–912 (2007).
Hashimoto, H. et al. Recognition and potential mechanisms for replication and erasure of cytosine hydroxymethylation. Nucleic Acids Res. 40, 4841–4849 (2012).
Otani, J. et al. Cell cycle-dependent turnover of 5-hydroxymethyl cytosine in mouse embryonic stem cells. PLoS ONE 8, e82961 (2013).
Valinluck, V. & Sowers, L. C. Endogenous cytosine damage products alter the site selectivity of human DNA maintenance methyltransferase DNMT1. Cancer Res. 67, 946–950 (2007).
Ji, D., Lin, K., Song, J. & Wang, Y. Effects of Tet-induced oxidation products of 5-methylcytosine on Dnmt1- and DNMT3a-mediated cytosine methylation. Mol. Biosyst. 10, 1749–1752 (2014).
Ono, R. et al. LCX, leukemia-associated protein with a CXXC domain, is fused to MLL in acute myeloid leukemia with trilineage dysplasia having t(10;11)(q22;q23). Cancer Res. 62, 4075–4080 (2002).
Lorsbach, R. B. et al. TET1, a member of a novel protein family, is fused to MLL in acute myeloid leukemia containing the t(10;11)(q22;q23). Leukemia 17, 637–641 (2003).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009).
Yu, Z. et al. The protein that binds to DNA base J in trypanosomatids has features of a thymidine hydroxylase. Nucleic Acids Res. 35, 2107–2115 (2007).
Iyer, L. M., Tahiliani, M., Rao, A. & Aravind, L. Prediction of novel families of enzymes involved in oxidative and other complex modifications of bases in nucleic acids. Cell Cycle 8, 1698–1710 (2009).
Guo, J. U., Su, Y., Zhong, C., Ming, G. L. & Song, H. Hydroxylation of 5-methylcytosine by TET1 promotes active DNA demethylation in the adult brain. Cell 145, 423–434 (2011).
Muller, U., Bauer, C., Siegl, M., Rottach, A. & Leonhardt, H. TET-mediated oxidation of methylcytosine causes TDG or NEIL glycosylase dependent gene reactivation. Nucleic Acids Res. 42, 8592–8604 (2014).
Cortellino, S. et al. Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. Cell 146, 67–79 (2011).
Weber, A. R. et al. Biochemical reconstitution of TET1-TDG-BER-dependent active DNA demethylation reveals a highly coordinated mechanism. Nat. Commun. 7, 10806 (2016).
Zadeh, F. J. et al. The role of molecular mechanism of Ten-Eleven Translocation2 (TET2) family proteins in pathogenesis of cardiovascular diseases (CVDs). Mol. Biol. Rep. 47, 5503–5509 (2020).
Ross, S. E. & Bogdanovic, O. TET enzymes, DNA demethylation and pluripotency. Biochem. Soc. Trans. 47, 875–885 (2019).
Lio, C. J., Yuita, H. & Rao, A. Dysregulation of the TET family of epigenetic regulators in lymphoid and myeloid malignancies. Blood 134, 1487–1497 (2019).
Solary, E., Bernard, O. A., Tefferi, A., Fuks, F. & Vainchenker, W. The Ten-Eleven Translocation-2 (TET2) gene in hematopoiesis and hematopoietic diseases. Leukemia 28, 485–496 (2014).
Cong, B., Zhang, Q. & Cao, X. The function and regulation of TET2 in innate immunity and inflammation. Protein Cell 12, 165–173 (2021).
Jiang, S. Tet2 at the interface between cancer and immunity. Commun. Biol. 3, 667 (2020).
Garcia-Outeiral, V., de la Parte, C., Fidalgo, M. & Guallar, D. The complexity of TET2 functions in pluripotency and development. Front. Cell Dev. Biol. 8, 630754 (2020).
Pan, F. et al. Tet2 loss leads to hypermutagenicity in haematopoietic stem/progenitor cells. Nat. Commun. 8, 15102 (2017).
Abdel-Wahab, O. et al. Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood 114, 144–147 (2009).
Bejar, R. et al. Validation of a prognostic model and the impact of mutations in patients with lower-risk myelodysplastic syndromes. J. Clin. Oncol. 30, 3376–3382 (2012).
Lemonnier, F. et al. Recurrent TET2 mutations in peripheral T-cell lymphomas correlate with TFH-like features and adverse clinical parameters. Blood 120, 1466–1469 (2012).
Huang, Y. & Rao, A. Connections between TET proteins and aberrant DNA modification in cancer. Trends Genet. 30, 464–474 (2014).
Lazarenkov, A. & Sardina, J. L. Dissecting TET2 regulatory networks in blood differentiation and cancer. Cancers 14, 830 (2022).
Delhommeau, F. et al. Mutation in TET2 in myeloid cancers. N. Engl. J. Med. 360, 2289–2301 (2009).
Jankowska, A. M. et al. Loss of heterozygosity 4q24 and TET2 mutations associated with myelodysplastic/myeloproliferative neoplasms. Blood 113, 6403–6410 (2009).
Langemeijer, S. M. et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat. Genet. 41, 838–842 (2009).
Bowman, R. L. & Levine, R. L. TET2 in normal and malignant hematopoiesis. Cold Spring Harb. Perspect. Med. 7, a026518 (2017).
Shih, A. H., Abdel-Wahab, O., Patel, J. P. & Levine, R. L. The role of mutations in epigenetic regulators in myeloid malignancies. Nat. Rev. Cancer 12, 599–612 (2012).
Hellstrom-Lindberg, E. Significance of JAK2 and TET2 mutations in myelodysplastic syndromes. Blood Rev. 24, 83–90 (2010).
Sato, H., Wheat, J. C., Steidl, U. & Ito, K. DNMT3A and TET2 in the pre-leukemic phase of hematopoietic disorders. Front. Oncol. 6, 187 (2016).
Bray, J. K., Dawlaty, M. M., Verma, A. & Maitra, A. Roles and regulations of TET enzymes in solid tumors. Trends Cancer 7, 635–646 (2021).
Seshagiri, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012).
Wu, M. J. et al. Mutant IDH inhibits IFNgamma-TET2 signaling to promote immunoevasion and tumor maintenance in cholangiocarcinoma. Cancer Discov. 12, 812–835 (2022).
Momparler, R. L., Cote, S. & Momparler, L. F. Epigenetic modulation of self-renewal capacity of leukemic stem cells and implications for chemotherapy. Epigenomes 4, 3 (2020).
Hu, L. et al. Crystal structure of TET2-DNA complex: insight into TET-mediated 5mC oxidation. Cell 155, 1545–1555 (2013).
Hu, L. et al. Structural insight into substrate preference for TET-mediated oxidation. Nature 527, 118–122 (2015).
Xu, Y. et al. Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol. Cell 42, 451–464 (2011).
Xu, Y. et al. Tet3 CXXC domain and dioxygenase activity cooperatively regulate key genes for Xenopus eye and neural development. Cell 151, 1200–1213 (2012).
Zhang, H. et al. TET1 is a DNA-binding protein that modulates DNA methylation and gene transcription via hydroxylation of 5-methylcytosine. Cell Res. 20, 1390–1393 (2010).
Wu, H. et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473, 389–393 (2011).
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011).
Huang, Y. et al. Distinct roles of the methylcytosine oxidases Tet1 and Tet2 in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 111, 1361–1366 (2014).
Pantier, R., Mullin, N., Hall-Ponsele, E. & Chambers, I. TET1 interacts directly with NANOG via independent domains containing hydrophobic and aromatic residues. J. Mol. Biol. 432, 6075–6091 (2020).
Perera, A. et al. TET3 is recruited by REST for context-specific hydroxymethylation and induction of gene expression. Cell Rep. 11, 283–294 (2015).
Branco, M. R., Ficz, G. & Reik, W. Uncovering the role of 5-hydroxymethylcytosine in the epigenome. Nat. Rev. Genet. 13, 7–13 (2011).
Wu, H. & Zhang, Y. Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev. 25, 2436–2452 (2011).
Lio, C. J. et al. TET methylcytosine oxidases: new insights from a decade of research. J. Biosci. 45, 21 (2020).
Sjostedt, E. et al. An atlas of the protein-coding genes in the human, pig, and mouse brain. Science 367, eaay5947 (2020).
Thaler, R. et al. Vitamin C epigenetically controls osteogenesis and bone mineralization. Nat. Commun. 13, 5883 (2022).
Khoueiry, R. et al. Lineage-specific functions of TET1 in the postimplantation mouse embryo. Nat. Genet. 49, 1061–1072 (2017).
Yamaguchi, S., Shen, L., Liu, Y., Sendler, D. & Zhang, Y. Role of Tet1 in erasure of genomic imprinting. Nature 504, 460–464 (2013).
Yamaguchi, S. et al. Tet1 controls meiosis by regulating meiotic gene expression. Nature 492, 443–447 (2012).
Dawlaty, M. M. et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9, 166–175 (2011).
Freudenberg, J. M. et al. Acute depletion of Tet1-dependent 5-hydroxymethylcytosine levels impairs LIF/Stat3 signaling and results in loss of embryonic stem cell identity. Nucleic Acids Res. 40, 3364–3377 (2012).
Kim, R., Sheaffer, K. L., Choi, I., Won, K. J. & Kaestner, K. H. Epigenetic regulation of intestinal stem cells by Tet1-mediated DNA hydroxymethylation. Genes Dev. 30, 2433–2442 (2016).
Kraiczy, J. et al. DNA methylation defines regional identity of human intestinal epithelial organoids and undergoes dynamic changes during development. Gut 68, 49–61 (2019).
Aloia, L. et al. Epigenetic remodelling licences adult cholangiocytes for organoid formation and liver regeneration. Nat. Cell Biol. 21, 1321–1333 (2019).
Zhang, M. et al. Ten-eleven translocation 1 mediated-DNA hydroxymethylation is required for myelination and remyelination in the mouse brain. Nat. Commun. 12, 5091 (2021).
Moyon, S. et al. TET1-mediated DNA hydroxymethylation regulates adult remyelination in mice. Nat. Commun. 12, 3359 (2021).
Jiang, L. et al. RNF217 regulates iron homeostasis through its E3 ubiquitin ligase activity by modulating ferroportin degradation. Blood 138, 689–705 (2021).
Zhang, R. R. et al. Tet1 regulates adult hippocampal neurogenesis and cognition. Cell Stem Cell 13, 237–245 (2013).
Gao, Y. et al. Replacement of Oct4 by Tet1 during iPSC induction reveals an important role of DNA methylation and hydroxymethylation in reprogramming. Cell Stem Cell 12, 453–469 (2013).
Wang, T. et al. Subtelomeric hotspots of aberrant 5-hydroxymethylcytosine-mediated epigenetic modifications during reprogramming to pluripotency. Nat. Cell Biol. 15, 700–711 (2013).
Kim, K. P. et al. Permissive epigenomes endow reprogramming competence to transcriptional regulators. Nat. Chem. Biol. 17, 47–56 (2021).
Tran, K. A., Dillingham, C. M. & Sridharan, R. Coordinated removal of repressive epigenetic modifications during induced reversal of cell identity. EMBO J. 38, e101681 (2019).
Jiang, H. et al. Cell cycle and p53 gate the direct conversion of human fibroblasts to dopaminergic neurons. Nat. Commun. 6, 10100 (2015).
Bacos, K. et al. Type 2 diabetes candidate genes, including PAX5, cause impaired insulin secretion in human pancreatic islets. J. Clin. Invest. 133, e163612 (2023).
Liu, L., Wang, H., Xu, G. L. & Liu, L. Tet1 deficiency leads to premature ovarian failure. Front. Cell Dev. Biol. 9, 644135 (2021).
Shirai, K. et al. TET1 upregulation drives cancer cell growth through aberrant enhancer hydroxymethylation of HMGA2 in hepatocellular carcinoma. Cancer Sci. 112, 2855–2869 (2021).
Chen, W. et al. Fetal growth restriction impairs hippocampal neurogenesis and cognition via Tet1 in offspring. Cell Rep. 37, 109912 (2021).
Bamezai, S. et al. TET1 promotes growth of T-cell acute lymphoblastic leukemia and can be antagonized via PARP inhibition. Leukemia 35, 389–403 (2021).
Good, C. R. et al. TET1-mediated hypomethylation activates oncogenic signaling in triple-negative breast cancer. Cancer Res. 78, 4126–4137 (2018).
Filipczak, P. T. et al. p53-Suppressed oncogene TET1 prevents cellular aging in lung cancer. Cancer Res. 79, 1758–1768 (2019).
Cimmino, L. et al. TET1 is a tumor suppressor of hematopoietic malignancy. Nat. Immunol. 16, 653–662 (2015).
Huang, H. et al. TET1 plays an essential oncogenic role in MLL-rearranged leukemia. Proc. Natl Acad. Sci. USA 110, 11994–11999 (2013).
Bisserier, M. et al. Regulation of the methylation and expression levels of the BMPR2 gene by SIN3a as a novel therapeutic mechanism in pulmonary arterial hypertension. Circulation 144, 52–73 (2021).
Bai, X. et al. Ten-eleven translocation 1 promotes malignant progression of cholangiocarcinoma with wild-type isocitrate dehydrogenase 1. Hepatology 73, 1747–1763 (2021).
Tricarico, R. et al. TET1 and TDG suppress inflammatory response in intestinal tumorigenesis: implications for colorectal tumors with the CpG island methylator phenotype. Gastroenterology 164, 921–936.e1 (2023).
Mimouni, N. E. H. et al. Polycystic ovary syndrome is transmitted via a transgenerational epigenetic process. Cell Metab. 33, 513–530 e518 (2021).
Zhang, P. et al. Methyl-CpG binding domain protein 1 regulates localization and activity of Tet1 in a CXXC3 domain-dependent manner. Nucleic Acids Res. 45, 7118–7136 (2017).
Lv, H. et al. NAD(+) metabolism maintains inducible PD-L1 expression to drive tumor immune evasion. Cell Metab. 33, 110–127 e115 (2021).
Deng, S. et al. RNA m(6)A regulates transcription via DNA demethylation and chromatin accessibility. Nat. Genet. 54, 1427–1437 (2022).
Metzeler, K. H. et al. TET2 mutations improve the new European LeukemiaNet risk classification of acute myeloid leukemia: a Cancer and Leukemia Group B study. J. Clin. Oncol. 29, 1373–1381 (2011).
Ko, M. et al. TET proteins and 5-methylcytosine oxidation in hematological cancers. Immunol. Rev. 263, 6–21 (2015).
Cimmino, L., Abdel-Wahab, O., Levine, R. L. & Aifantis, I. TET family proteins and their role in stem cell differentiation and transformation. Cell Stem Cell 9, 193–204 (2011).
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010).
Nibourel, O. et al. Incidence and prognostic value of TET2 alterations in de novo acute myeloid leukemia achieving complete remission. Blood 116, 1132–1135 (2010).
Kosmider, O. et al. TET2 mutations in secondary acute myeloid leukemias: a French retrospective study. Haematologica 96, 1059–1063 (2011).
Li, Z. et al. Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 118, 4509–4518 (2011).
Li, S. et al. TET2 promotes anti-tumor immunity by governing G-MDSCs and CD8(+) T-cell numbers. EMBO Rep. 21, e49425 (2020).
Dominguez, P. M. et al. TET2 deficiency causes germinal center hyperplasia, impairs plasma cell differentiation, and promotes B-cell lymphomagenesis. Cancer Discov. 8, 1632–1653 (2018).
Lyu, R. et al. Tumour suppressor TET2 safeguards enhancers from aberrant DNA methylation and epigenetic reprogramming in ERalpha-positive breast cancer cells. Epigenetics 17, 1180–1194 (2022).
Bonvin, E. et al. TET2-dependent hydroxymethylome plasticity reduces melanoma initiation and progression. Cancer Res. 79, 482–494 (2019).
Zhang, J. et al. p53-dependent autophagic degradation of TET2 modulates cancer therapeutic resistance. Oncogene 38, 1905–1919 (2019).
Puig, I. et al. TET2 controls chemoresistant slow-cycling cancer cell survival and tumor recurrence. J. Clin. Invest. 128, 3887–3905 (2018).
Pan, W. et al. The DNA methylcytosine dioxygenase Tet2 sustains immunosuppressive function of tumor-infiltrating myeloid cells to promote melanoma progression. Immunity 47, 284–297 e285 (2017).
Ko, M. et al. Modulation of TET2 expression and 5-methylcytosine oxidation by the CXXC domain protein IDAX. Nature 497, 122–126 (2013).
Costa, Y. et al. NANOG-dependent function of TET1 and TET2 in establishment of pluripotency. Nature 495, 370–374 (2013).
Guilhamon, P. et al. Meta-analysis of IDH-mutant cancers identifies EBF1 as an interaction partner for TET2. Nat. Commun. 4, 2166 (2013).
Rampal, R. et al. DNA hydroxymethylation profiling reveals that WT1 mutations result in loss of TET2 function in acute myeloid leukemia. Cell Rep. 9, 1841–1855 (2014).
Wang, Y. et al. WT1 recruits TET2 to regulate its target gene expression and suppress leukemia cell proliferation. Mol. Cell 57, 662–673 (2015).
de la Rica, L. et al. PU.1 target genes undergo Tet2-coupled demethylation and DNMT3b-mediated methylation in monocyte-to-osteoclast differentiation. Genome Biol. 14, R99 (2013).
Ichiyama, K. et al. The methylcytosine dioxygenase Tet2 promotes DNA demethylation and activation of cytokine gene expression in T cells. Immunity 42, 613–626 (2015).
Song, C. et al. PML recruits TET2 to regulate DNA modification and cell proliferation in response to chemotherapeutic agent. Cancer Res. 78, 2475–2489 (2018).
Sardina, J. L. et al. Transcription factors drive Tet2-mediated enhancer demethylation to reprogram cell fate. Cell Stem Cell 23, 727–741 e729 (2018).
Xu, Y. P. et al. Tumor suppressor TET2 promotes cancer immunity and immunotherapy efficacy. J. Clin. Invest. 129, 4316–4331 (2019).
Chen, L. L. et al. SNIP1 recruits TET2 to regulate c-MYC target genes and cellular DNA damage response. Cell Rep. 25, 1485–1500 e1484 (2018).
Zhang, X. et al. TET2 suppresses VHL deficiency-driven clear cell renal cell carcinoma by inhibiting HIF signaling. Cancer Res. 82, 2097–2109 (2022).
Gu, T. P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Guo, F. et al. Active and passive demethylation of male and female pronuclear DNA in the mammalian zygote. Cell Stem Cell 15, 447–459 (2014).
Shen, L. et al. Tet3 and DNA replication mediate demethylation of both the maternal and paternal genomes in mouse zygotes. Cell Stem Cell 15, 459–471 (2014).
Ladstatter, S. & Tachibana-Konwalski, K. A surveillance mechanism ensures repair of DNA lesions during zygotic reprogramming. Cell 167, 1774–1787 e1713 (2016).
Bogdanovic, O. et al. Active DNA demethylation at enhancers during the vertebrate phylotypic period. Nat. Genet. 48, 417–426 (2016).
Weng, Y. L. et al. An intrinsic epigenetic barrier for functional axon regeneration. Neuron 94, 337–346 e336 (2017).
Yu, H. et al. Tet3 regulates synaptic transmission and homeostatic plasticity via DNA oxidation and repair. Nat. Neurosci. 18, 836–843 (2015).
Reizel, Y. et al. Postnatal DNA demethylation and its role in tissue maturation. Nat. Commun. 9, 2040 (2018).
Beck, D. B. et al. Delineation of a human Mendelian disorder of the DNA demethylation machinery: TET3 deficiency. Am. J. Hum. Genet. 106, 234–245 (2020).
Antunes, C. et al. Tet3 ablation in adult brain neurons increases anxiety-like behavior and regulates cognitive function in mice. Mol. Psychiatry 26, 1445–1457 (2021).
Da, L. et al. Hepatic TET3 contributes to type-2 diabetes by inducing the HNF4alpha fetal isoform. Nat. Commun. 11, 342 (2020).
Chen, B. et al. Maternal inheritance of glucose intolerance via oocyte TET3 insufficiency. Nature 605, 761–766 (2022).
Bian, C. & Yu, X. PGC7 suppresses TET3 for protecting DNA methylation. Nucleic Acids Res. 42, 2893–2905 (2014).
Nakamura, T. et al. PGC7 binds histone H3K9me2 to protect against conversion of 5mC to 5hmC in early embryos. Nature 486, 415–419 (2012).
Piccolo, F. M. et al. Different roles for Tet1 and Tet2 proteins in reprogramming-mediated erasure of imprints induced by EGC fusion. Mol. Cell 49, 1023–1033 (2013).
Hackett, J. A. et al. Germline DNA demethylation dynamics and imprint erasure through 5-hydroxymethylcytosine. Science 339, 448–452 (2013).
Marina, R. J. et al. TET-catalyzed oxidation of intragenic 5-methylcytosine regulates CTCF-dependent alternative splicing. EMBO J. 35, 335–355 (2016).
Finley, L. W. S. et al. Pluripotency transcription factors and Tet1/2 maintain Brd4-independent stem cell identity. Nat. Cell Biol. 20, 565–574 (2018).
Lu, Y. et al. Reprogramming to recover youthful epigenetic information and restore vision. Nature 588, 124–129 (2020).
Kusuyama, J. et al. Placental superoxide dismutase 3 mediates benefits of maternal exercise on offspring health. Cell Metab. 33, 939–956 e938 (2021).
Yang, R. et al. Hydrogen sulfide promotes Tet1- and Tet2-mediated Foxp3 demethylation to drive regulatory T cell differentiation and maintain immune homeostasis. Immunity 43, 251–263 (2015).
Yang, R. et al. Tet1 and Tet2 maintain mesenchymal stem cell homeostasis via demethylation of the P2rX7 promoter. Nat. Commun. 9, 2143 (2018).
Zhu, X. et al. Role of Tet1/3 genes and chromatin remodeling genes in cerebellar circuit formation. Neuron 89, 100–112 (2016).
Issuree, P. D. et al. Stage-specific epigenetic regulation of CD4 expression by coordinated enhancer elements during T cell development. Nat. Commun. 9, 3594 (2018).
Yue, X., Lio, C. J., Samaniego-Castruita, D., Li, X. & Rao, A. Loss of TET2 and TET3 in regulatory T cells unleashes effector function. Nat. Commun. 10, 2011 (2019).
Yue, X. et al. Control of Foxp3 stability through modulation of TET activity. J. Exp. Med. 213, 377–397 (2016).
Tanaka, S. et al. Tet2 and Tet3 in B cells are required to repress CD86 and prevent autoimmunity. Nat. Immunol. 21, 950–961 (2020).
Fang, S. et al. Tet inactivation disrupts YY1 binding and long-range chromatin interactions during embryonic heart development. Nat. Commun. 10, 4297 (2019).
Tsagaratou, A. et al. TET proteins regulate the lineage specification and TCR-mediated expansion of iNKT cells. Nat. Immunol. 18, 45–53 (2017).
Yan, H. et al. Distinct roles for TET family proteins in regulating human erythropoiesis. Blood 129, 2002–2012 (2017).
An, J. et al. Acute loss of TET function results in aggressive myeloid cancer in mice. Nat. Commun. 6, 10071 (2015).
Shukla, V. et al. TET deficiency perturbs mature B cell homeostasis and promotes oncogenesis associated with accumulation of G-quadruplex and R-loop structures. Nat. Immunol. 23, 99–108 (2022).
Hu, X. et al. Tet and TDG mediate DNA demethylation essential for mesenchymal-to-epithelial transition in somatic cell reprogramming. Cell Stem Cell 14, 512–522 (2014).
Lu, F., Liu, Y., Jiang, L., Yamaguchi, S. & Zhang, Y. Role of Tet proteins in enhancer activity and telomere elongation. Genes Dev. 28, 2103–2119 (2014).
Dai, H. Q. et al. TET-mediated DNA demethylation controls gastrulation by regulating Lefty-Nodal signalling. Nature 538, 528–532 (2016).
Verma, N. et al. TET proteins safeguard bivalent promoters from de novo methylation in human embryonic stem cells. Nat. Genet. 50, 83–95 (2018).
Ginno, P. A. et al. A genome-scale map of DNA methylation turnover identifies site-specific dependencies of DNMT and TET activity. Nat. Commun. 11, 2680 (2020).
Putiri, E. L. et al. Distinct and overlapping control of 5-methylcytosine and 5-hydroxymethylcytosine by the TET proteins in human cancer cells. Genome Biol. 15, R81 (2014).
Ko, M. et al. Ten-Eleven-Translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc. Natl Acad. Sci. USA 108, 14566–14571 (2011).
Dawlaty, M. M. et al. Combined deficiency of Tet1 and Tet2 causes epigenetic abnormalities but is compatible with postnatal development. Dev. Cell 24, 310–323 (2013).
Zhao, H. & Chen, T. Tet family of 5-methylcytosine dioxygenases in mammalian development. J. Hum. Genet. 58, 421–427 (2013).
Mellen, M., Ayata, P., Dewell, S., Kriaucionis, S. & Heintz, N. MeCP2 binds to 5hmC enriched within active genes and accessible chromatin in the nervous system. Cell 151, 1417–1430 (2012).
Spruijt, C. G. et al. Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 152, 1146–1159 (2013).
Wossidlo, M. et al. 5-Hydroxymethylcytosine in the mammalian zygote is linked with epigenetic reprogramming. Nat. Commun. 2, 241 (2011).
Kaji, K. et al. The NuRD component Mbd3 is required for pluripotency of embryonic stem cells. Nat. Cell Biol. 8, 285–292 (2006).
Yildirim, O. et al. Mbd3/NURD complex regulates expression of 5-hydroxymethylcytosine marked genes in embryonic stem cells. Cell 147, 1498–1510 (2011).
Serandour, A. A. et al. Dynamic hydroxymethylation of deoxyribonucleic acid marks differentiation-associated enhancers. Nucleic Acids Res. 40, 8255–8265 (2012).
Hill, P. W. S. et al. Epigenetic reprogramming enables the transition from primordial germ cell to gonocyte. Nature 555, 392–396 (2018).
Feng, J. et al. Role of Tet1 and 5-hydroxymethylcytosine in cocaine action. Nat. Neurosci. 18, 536–544 (2015).
Smeriglio, P. et al. Inhibition of TET1 prevents the development of osteoarthritis and reveals the 5hmC landscape that orchestrates pathogenesis. Sci. Transl. Med. 12, eaax2332 (2020).
Amouroux, R. et al. De novo DNA methylation drives 5hmC accumulation in mouse zygotes. Nat. Cell Biol. 18, 225–233 (2016).
Mellen, M., Ayata, P. & Heintz, N. 5-hydroxymethylcytosine accumulation in postmitotic neurons results in functional demethylation of expressed genes. Proc. Natl Acad. Sci. USA 114, E7812–E7821 (2017).
Xie, D. et al. TET3 epigenetically controls feeding and stress response behaviors via AGRP neurons. J. Clin. Invest. 132, e162365 (2022).
Wu, F. et al. Spurious transcription causing innate immune responses is prevented by 5-hydroxymethylcytosine. Nat. Genet. 55, 100–111 (2023).
Kaas, G. A. et al. TET1 controls CNS 5-methylcytosine hydroxylation, active DNA demethylation, gene transcription, and memory formation. Neuron 79, 1086–1093 (2013).
Damal Villivalam, S. et al. TET1 is a beige adipocyte-selective epigenetic suppressor of thermogenesis. Nat. Commun. 11, 4313 (2020).
Chrysanthou, S. et al. The DNA dioxygenase Tet1 regulates H3K27 modification and embryonic stem cell biology independent of its catalytic activity. Nucleic Acids Res. 50, 3169–3189 (2022).
Stolz, P. et al. TET1 regulates gene expression and repression of endogenous retroviruses independent of DNA demethylation. Nucleic Acids Res. 50, 8491–8511 (2022).
Montalban-Loro, R. et al. TET3 prevents terminal differentiation of adult NSCs by a non-catalytic action at Snrpn. Nat. Commun. 10, 1726 (2019).
Zhang, Q. et al. Tet2 is required to resolve inflammation by recruiting Hdac2 to specifically repress IL-6. Nature 525, 389–393 (2015).
Ito, K. et al. Non-catalytic roles of Tet2 are essential to regulate hematopoietic stem and progenitor cell homeostasis. Cell Rep. 28, 2480–2490 e2484 (2019).
Fu, L. et al. Tet-mediated formation of 5-hydroxymethylcytosine in RNA. J. Am. Chem. Soc. 136, 11582–11585 (2014).
Delatte, B. et al. RNA biochemistry. Transcriptome-wide distribution and function of RNA hydroxymethylcytosine. Science 351, 282–285 (2016).
Shen, Q. et al. Tet2 promotes pathogen infection-induced myelopoiesis through mRNA oxidation. Nature 554, 123–127 (2018).
Guallar, D. et al. RNA-dependent chromatin targeting of TET2 for endogenous retrovirus control in pluripotent stem cells. Nat. Genet. 50, 443–451 (2018).
Huang, X. et al. A TET1-PSPC1-Neat1 molecular axis modulates PRC2 functions in controlling stem cell bivalency. Cell Rep. 39, 110928 (2022).
Lan, J. et al. Functional role of Tet-mediated RNA hydroxymethylcytosine in mouse ES cells and during differentiation. Nat. Commun. 11, 4956 (2020).
He, C. et al. High-resolution mapping of RNA-binding regions in the nuclear proteome of embryonic stem cells. Mol. Cell 64, 416–430 (2016).
He, C. et al. TET2 chemically modifies tRNAs and regulates tRNA fragment levels. Nat. Struct. Mol. Biol. 28, 62–70 (2021).
Shen, H. et al. TET-mediated 5-methylcytosine oxidation in tRNA promotes translation. J. Biol. Chem. 296, 100087 (2021).
Pfaffeneder, T. et al. Tet oxidizes thymine to 5-hydroxymethyluracil in mouse embryonic stem cell DNA. Nat. Chem. Biol. 10, 574–581 (2014).
Neri, F. et al. TET1 is controlled by pluripotency-associated factors in ESCs and downmodulated by PRC2 in differentiated cells and tissues. Nucleic Acids Res. 43, 6814–6826 (2015).
Etchegaray, J. P. et al. The histone deacetylase SIRT6 controls embryonic stem cell fate via TET-mediated production of 5-hydroxymethylcytosine. Nat. Cell Biol. 17, 545–557 (2015).
Yang, Y. A. et al. FOXA1 potentiates lineage-specific enhancer activation through modulating TET1 expression and function. Nucleic Acids Res. 44, 8153–8164 (2016).
Jiang, X. et al. Targeted inhibition of STAT/TET1 axis as a therapeutic strategy for acute myeloid leukemia. Nat. Commun. 8, 2099 (2017).
Tovy, A. et al. p53 is essential for DNA methylation homeostasis in naive embryonic stem cells, and its loss promotes clonal heterogeneity. Genes Dev. 31, 959–972 (2017).
Collignon, E. et al. Immunity drives TET1 regulation in cancer through NF-kappaB. Sci. Adv. 4, eaap7309 (2018).
Cui, Q. et al. Downregulation of TLX induces TET3 expression and inhibits glioblastoma stem cell self-renewal and tumorigenesis. Nat. Commun. 7, 10637 (2016).
Jacobsen, A. et al. Analysis of microRNA-target interactions across diverse cancer types. Nat. Struct. Mol. Biol. 20, 1325–1332 (2013).
Tu, J. et al. MicroRNA-29b/Tet1 regulatory axis epigenetically modulates mesendoderm differentiation in mouse embryonic stem cells. Nucleic Acids Res. 43, 7805–7822 (2015).
Zhu, J. et al. Stem cell-homing hydrogel-based miR-29b-5p delivery promotes cartilage regeneration by suppressing senescence in an osteoarthritis rat model. Sci. Adv. 8, eabk0011 (2022).
Chuang, K. H. et al. MicroRNA-494 is a master epigenetic regulator of multiple invasion-suppressor microRNAs by targeting ten eleven translocation 1 in invasive human hepatocellular carcinoma tumors. Hepatology 62, 466–480 (2015).
Li, H. et al. MicroRNA-191 acts as a tumor promoter by modulating the TET1-p53 pathway in intrahepatic cholangiocarcinoma. Hepatology 66, 136–151 (2017).
Song, S. J. et al. The oncogenic microRNA miR-22 targets the TET2 tumor suppressor to promote hematopoietic stem cell self-renewal and transformation. Cell Stem Cell 13, 87–101 (2013).
Zhang, P., Huang, B., Xu, X. & Sessa, W. C. Ten-eleven translocation (Tet) and thymine DNA glycosylase (TDG), components of the demethylation pathway, are direct targets of miRNA-29a. Biochem. Biophys. Res. Commun. 437, 368–373 (2013).
Cheng, J. et al. An extensive network of TET2-targeting microRNAs regulates malignant hematopoiesis. Cell Rep. 5, 471–481 (2013).
Jiang, S., Yan, W., Wang, S. E. & Baltimore, D. Dual mechanisms of posttranscriptional regulation of Tet2 by Let-7 microRNA in macrophages. Proc. Natl Acad. Sci. USA 116, 12416–12421 (2019).
Ma, Q., Dasgupta, C., Shen, G., Li, Y. & Zhang, L. MicroRNA-210 downregulates TET2 and contributes to inflammatory response in neonatal hypoxic-ischemic brain injury. J. Neuroinflammation 18, 6 (2021).
Li, N. et al. miR-144-3p suppresses osteogenic differentiation of BMSCs from patients with aplastic anemia through repression of TET2. Mol. Ther. Nucleic Acids 19, 619–626 (2020).
Scherm, M. G. et al. miRNA142-3p targets Tet2 and impairs Treg differentiation and stability in models of type 1 diabetes. Nat. Commun. 10, 5697 (2019).
Fu, X. et al. MicroRNA-26a targets ten eleven translocation enzymes and is regulated during pancreatic cell differentiation. Proc. Natl Acad. Sci. USA 110, 17892–17897 (2013).
Lopez-Bertoni, H. et al. Sox2 induces glioblastoma cell stemness and tumor propagation by repressing TET2 and deregulating 5hmC and 5mC DNA modifications. Signal Transduct. Target Ther. 7, 37 (2022).
Selimoglu-Buet, D. et al. A miR-150/TET3 pathway regulates the generation of mouse and human non-classical monocyte subset. Nat. Commun. 9, 5455 (2018).
Bauer, C. et al. Phosphorylation of TET proteins is regulated via O-GlcNAcylation by the O-linked N-acetylglucosamine transferase (OGT). J. Biol. Chem. 290, 4801–4812 (2015).
Zhang, Y. W. et al. Acetylation enhances TET2 function in protecting against abnormal DNA methylation during oxidative stress. Mol. Cell 65, 323–335 (2017).
Wu, D. et al. Glucose-regulated phosphorylation of TET2 by AMPK reveals a pathway linking diabetes to cancer. Nature 559, 637–641 (2018).
Nakagawa, T. et al. CRL4(VprBP) E3 ligase promotes monoubiquitylation and chromatin binding of TET dioxygenases. Mol. Cell 57, 247–260 (2015).
Chen, L. L. et al. USP15 suppresses tumor immunity via deubiquitylation and inactivation of TET2. Sci. Adv. 6, eabc9730 (2020).
Rao, V. K. et al. Phosphorylation of Tet3 by cdk5 is critical for robust activation of BRN2 during neuronal differentiation. Nucleic Acids Res. 48, 1225–1238 (2020).
Lv, L. et al. Vpr targets TET2 for degradation by CRL4(VprBP) E3 ligase to sustain IL-6 expression and enhance HIV-1 replication. Mol. Cell 70, 961–970 e965 (2018).
Wang, Y. & Zhang, Y. Regulation of TET protein stability by calpains. Cell Rep. 6, 278–284 (2014).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Shim, E. H. et al. L-2-Hydroxyglutarate: an epigenetic modifier and putative oncometabolite in renal cancer. Cancer Discov. 4, 1290–1298 (2014).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of alpha-ketoglutarate-dependent dioxygenases. Cancer Cell 19, 17–30 (2011).
Xiao, M. et al. Inhibition of alpha-KG-dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 26, 1326–1338 (2012).
Chen, L. L. et al. Itaconate inhibits TET DNA dioxygenases to dampen inflammatory responses. Nat. Cell Biol. 24, 353–363 (2022).
Traube, F. R. et al. Redirected nuclear glutamate dehydrogenase supplies Tet3 with alpha-ketoglutarate in neurons. Nat. Commun. 12, 4100 (2021).
Blaschke, K. et al. Vitamin C induces Tet-dependent DNA demethylation and a blastocyst-like state in ES cells. Nature 500, 222–226 (2013).
Hore, T. A. et al. Retinol and ascorbate drive erasure of epigenetic memory and enhance reprogramming to naive pluripotency by complementary mechanisms. Proc. Natl Acad. Sci. USA 113, 12202–12207 (2016).
Yin, R. et al. Ascorbic acid enhances Tet-mediated 5-methylcytosine oxidation and promotes DNA demethylation in mammals. J. Am. Chem. Soc. 135, 10396–10403 (2013).
Ostriker, A. C. et al. TET2 protects against vascular smooth muscle cell apoptosis and intimal thickening in transplant vasculopathy. Circulation 144, 455–470 (2021).
Eyres, M. et al. TET2 drives 5hmc marking of GATA6 and epigenetically defines pancreatic ductal adenocarcinoma transcriptional subtypes. Gastroenterology 161, 653–668 e616 (2021).
Gustafson, C. B. et al. Epigenetic reprogramming of melanoma cells by vitamin C treatment. Clin. Epigenetics 7, 51 (2015).
Peng, D. et al. Vitamin C increases 5-hydroxymethylcytosine level and inhibits the growth of bladder cancer. Clin. Epigenetics 10, 94 (2018).
Agathocleous, M. et al. Ascorbate regulates haematopoietic stem cell function and leukaemogenesis. Nature 549, 476–481 (2017).
Chen, H. Y. et al. Epigenetic remodeling by vitamin C potentiates plasma cell differentiation. Elife 11, e73754 (2022).
DiTroia, S. P. et al. Maternal vitamin C regulates reprogramming of DNA methylation and germline development. Nature 573, 271–275 (2019).
Cimmino, L. et al. Restoration of TET2 function blocks aberrant self-renewal and leukemia progression. Cell 170, 1079–1095 e1020 (2017).
Thienpont, B. et al. Tumour hypoxia causes DNA hypermethylation by reducing TET activity. Nature 537, 63–68 (2016).
Burr, S. et al. Oxygen gradients can determine epigenetic asymmetry and cellular differentiation via differential regulation of Tet activity in embryonic stem cells. Nucleic Acids Res. 46, 1210–1226 (2018).
Zhao, B. et al. Redox-active quinones induces genome-wide DNA methylation changes by an iron-mediated and Tet-dependent mechanism. Nucleic Acids Res. 42, 1593–1605 (2014).
Singh, A. K. et al. Selective targeting of TET catalytic domain promotes somatic cell reprogramming. Proc. Natl Acad. Sci. USA 117, 3621–3626 (2020).
Chua, G. N. L. et al. Cytosine-based TET enzyme inhibitors. ACS Med. Chem. Lett. 10, 180–185 (2019).
Guan, Y. et al. A therapeutic strategy for preferential targeting of TET2 mutant and TET-dioxygenase deficient cells in myeloid neoplasms. Blood Cancer Discov. 2, 146–161 (2021).
Yin, R., Mo, J., Dai, J. & Wang, H. Nickel(II) inhibits Tet-mediated 5-methylcytosine oxidation by high affinity displacement of the cofactor Iron(II). ACS Chem. Biol. 12, 1494–1498 (2017).
Sun, J. et al. SIRT1 activation disrupts maintenance of myelodysplastic syndrome stem and progenitor cells by restoring TET2 function. Cell Stem Cell 23, 355–369 e359 (2018).
Morinishi, L., Kochanowski, K., Levine, R. L., Wu, L. F. & Altschuler, S. J. Loss of TET2 affects proliferation and drug sensitivity through altered dynamics of cell-state transitions. Cell Syst. 11, 86–94 e85 (2020).
Bensberg, M. et al. TET2 as a tumor suppressor and therapeutic target in T-cell acute lymphoblastic leukemia. Proc. Natl Acad. Sci. USA 118, e2110758118 (2021).
Maekawa, T., Miyake, T., Tani, M. & Uemoto, S. Diverse antitumor effects of ascorbic acid on cancer cells and the tumor microenvironment. Front. Oncol. 12, 981547 (2022).
Chen, Q. et al. Pharmacologic ascorbic acid concentrations selectively kill cancer cells: action as a pro-drug to deliver hydrogen peroxide to tissues. Proc. Natl Acad. Sci. USA 102, 13604–13609 (2005).
Chen, Q. et al. Ascorbate in pharmacologic concentrations selectively generates ascorbate radical and hydrogen peroxide in extracellular fluid in vivo. Proc. Natl Acad. Sci. USA 104, 8749–8754 (2007).
Gao, P. et al. HIF-dependent antitumorigenic effect of antioxidants in vivo. Cancer Cell 12, 230–238 (2007).
Polireddy, K. et al. High dose parenteral ascorbate inhibited pancreatic cancer growth and metastasis: mechanisms and a phase I/IIa study. Sci. Rep. 7, 17188 (2017).
Gerecke, C., Schumacher, F., Berndzen, A., Homann, T. & Kleuser, B. Vitamin C in combination with inhibition of mutant IDH1 synergistically activates TET enzymes and epigenetically modulates gene silencing in colon cancer cells. Epigenetics 15, 307–322 (2020).
Monti, D. A. et al. Phase I evaluation of intravenous ascorbic acid in combination with gemcitabine and erlotinib in patients with metastatic pancreatic cancer. PLoS ONE 7, e29794 (2012).
Stephenson, C. M., Levin, R. D., Spector, T. & Lis, C. G. Phase I clinical trial to evaluate the safety, tolerability, and pharmacokinetics of high-dose intravenous ascorbic acid in patients with advanced cancer. Cancer Chemother. Pharm. 72, 139–146 (2013).
Hoffer, L. J. et al. High-dose intravenous vitamin C combined with cytotoxic chemotherapy in patients with advanced cancer: a phase I-II clinical trial. PLoS ONE 10, e0120228 (2015).
Nauman, G., Gray, J. C., Parkinson, R., Levine, M. & Paller, C. J. Systematic review of intravenous ascorbate in cancer clinical trials. Antioxidants 7, 89 (2018).
Mussa, A. et al. High-dose vitamin C for cancer therapy. Pharmaceuticals 15, 711 (2022).
Koh, K. P. et al. Tet1 and Tet2 regulate 5-hydroxymethylcytosine production and cell lineage specification in mouse embryonic stem cells. Cell Stem Cell 8, 200–213 (2011).
Ficz, G. et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473, 398–402 (2011).
Chen, K. et al. Loss of 5-hydroxymethylcytosine is linked to gene body hypermethylation in kidney cancer. Cell Res. 26, 103–118 (2016).
Vasanthakumar, A. & Godley, L. A. 5-hydroxymethylcytosine in cancer: significance in diagnosis and therapy. Cancer Genet. 208, 167–177 (2015).
Sjostrom, M. et al. The 5-hydroxymethylcytosine landscape of prostate cancer. Cancer Res. 82, 3888–3902 (2022).
Zhang, Z. et al. Values of 5mC, 5hmC, and TET2 for identifying the presence and progression of breast precancerous lesion. J. Clin. Lab Anal. 34, e23162 (2020).
Fernandez, A. F. et al. Loss of 5hmC identifies a new type of aberrant DNA hypermethylation in glioma. Hum. Mol. Genet. 27, 3046–3059 (2018).
Jin, S. G. et al. 5-Hydroxymethylcytosine is strongly depleted in human cancers but its levels do not correlate with IDH1 mutations. Cancer Res. 71, 7360–7365 (2011).
Uribe-Lewis, S. et al. 5-hydroxymethylcytosine marks promoters in colon that resist DNA hypermethylation in cancer. Genome Biol. 16, 69 (2015).
Lian, C. G. et al. Loss of 5-hydroxymethylcytosine is an epigenetic hallmark of melanoma. Cell 150, 1135–1146 (2012).
Killian, J. K. et al. Succinate dehydrogenase mutation underlies global epigenomic divergence in gastrointestinal stromal tumor. Cancer Discov. 3, 648–657 (2013).
Li, W. et al. 5-Hydroxymethylcytosine signatures in circulating cell-free DNA as diagnostic biomarkers for human cancers. Cell Res. 27, 1243–1257 (2017).
Song, C. X. et al. 5-Hydroxymethylcytosine signatures in cell-free DNA provide information about tumor types and stages. Cell Res. 27, 1231–1242 (2017).
Guler, G. D. et al. Detection of early stage pancreatic cancer using 5-hydroxymethylcytosine signatures in circulating cell free DNA. Nat. Commun. 11, 5270 (2020).
Hu, X. et al. Integrated 5-hydroxymethylcytosine and fragmentation signatures as enhanced biomarkers in lung cancer. Clin. Epigenetics 14, 15 (2022).
Zhang, J. et al. 5-Hydroxymethylome in circulating cell-free DNA as a potential biomarker for non-small-cell lung cancer. Genomics Proteom. Bioinform. 16, 187–199 (2018).
Thomson, J. P. & Meehan, R. R. The application of genome-wide 5-hydroxymethylcytosine studies in cancer research. Epigenomics 9, 77–91 (2017).
Gilat, N. et al. Single-molecule quantification of 5-hydroxymethylcytosine for diagnosis of blood and colon cancers. Clin. Epigenetics 9, 70 (2017).
Cai, J. et al. Genome-wide mapping of 5-hydroxymethylcytosines in circulating cell-free DNA as a non-invasive approach for early detection of hepatocellular carcinoma. Gut 68, 2195–2205 (2019).
Dong, C. et al. 5-Hydroxymethylcytosine signatures in circulating cell-free DNA as diagnostic and predictive biomarkers for coronary artery disease. Clin. Epigenetics 12, 17 (2020).
Chen, H. Y. et al. 5-Hydroxymethylcytosine profiles of cfDNA are highly predictive of R-CHOP treatment response in diffuse large B cell lymphoma patients. Clin. Epigenetics 13, 33 (2021).
Stroud, H., Feng, S., Morey Kinney, S., Pradhan, S. & Jacobsen, S. E. 5-Hydroxymethylcytosine is associated with enhancers and gene bodies in human embryonic stem cells. Genome Biol. 12, R54 (2011).
Wu, H. et al. Genome-wide analysis of 5-hydroxymethylcytosine distribution reveals its dual function in transcriptional regulation in mouse embryonic stem cells. Genes Dev. 25, 679–684 (2011).
Jin, S. G., Wu, X., Li, A. X. & Pfeifer, G. P. Genomic mapping of 5-hydroxymethylcytosine in the human brain. Nucleic Acids Res. 39, 5015–5024 (2011).
Robertson, A. B. et al. A novel method for the efficient and selective identification of 5-hydroxymethylcytosine in genomic DNA. Nucleic Acids Res. 39, e55 (2011).
Robertson, A. B., Dahl, J. A., Ougland, R. & Klungland, A. Pull-down of 5-hydroxymethylcytosine DNA using JBP1-coated magnetic beads. Nat. Protoc. 7, 340–350 (2012).
Huang, Y., Pastor, W. A., Zepeda-Martinez, J. A. & Rao, A. The anti-CMS technique for genome-wide mapping of 5-hydroxymethylcytosine. Nat. Protoc. 7, 1897–1908 (2012).
Li, J. et al. Aberrant DNA hydroxymethylation reshapes transcription factor binding in myeloid neoplasms. Clin. Epigenetics 14, 81 (2022).
Song, C. X. et al. Selective chemical labeling reveals the genome-wide distribution of 5-hydroxymethylcytosine. Nat. Biotechnol. 29, 68–72 (2011).
Yu, M. et al. Base-resolution analysis of 5-hydroxymethylcytosine in the mammalian genome. Cell 149, 1368–1380 (2012).
Booth, M. J. et al. Quantitative sequencing of 5-methylcytosine and 5-hydroxymethylcytosine at single-base resolution. Science 336, 934–937 (2012).
Zeng, H. et al. Bisulfite-free, nanoscale analysis of 5-hydroxymethylcytosine at single base resolution. J. Am. Chem. Soc. 140, 13190–13194 (2018).
Liu, Y. et al. Subtraction-free and bisulfite-free specific sequencing of 5-methylcytosine and its oxidized derivatives at base resolution. Nat. Commun. 12, 618 (2021).
Liu, Y. et al. Bisulfite-free direct detection of 5-methylcytosine and 5-hydroxymethylcytosine at base resolution. Nat. Biotechnol. 37, 424–429 (2019).
Hu, L. et al. Jump-seq: genome-wide capture and amplification of 5-hydroxymethylcytosine sites. J. Am. Chem. Soc. 141, 8694–8697 (2019).
Schutsky, E. K. et al. Nondestructive, base-resolution sequencing of 5-hydroxymethylcytosine using a DNA deaminase. Nat. Biotechnol. 36, 1083–1090 (2018).
Wang, Y., Zhang, X., Wu, F., Chen, Z. & Zhou, X. Bisulfite-free, single base-resolution analysis of 5-hydroxymethylcytosine in genomic DNA by chemical-mediated mismatch. Chem. Sci. 10, 447–452 (2019).
Wang, Y., Zhao, Y., Bollas, A., Wang, Y. & Au, K. F. Nanopore sequencing technology, bioinformatics and applications. Nat. Biotechnol. 39, 1348–1365 (2021).
Laszlo, A. H. et al. Detection and mapping of 5-methylcytosine and 5-hydroxymethylcytosine with nanopore MspA. Proc. Natl Acad. Sci. USA 110, 18904–18909 (2013).
Huber, S. M. et al. Formation and abundance of 5-hydroxymethylcytosine in RNA. Chembiochem 16, 752–755 (2015).
Huber, S. M., van Delft, P., Tanpure, A., Miska, E. A. & Balasubramanian, S. 2'-O-methyl-5-hydroxymethylcytidine: a second oxidative derivative of 5-methylcytidine in RNA. J. Am. Chem. Soc. 139, 1766–1769 (2017).
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Yu, C. et al. CRL4 complex regulates mammalian oocyte survival and reprogramming by activation of TET proteins. Science 342, 1518–1521 (2013).
Hon, G. C. et al. 5mC oxidation by Tet2 modulates enhancer activity and timing of transcriptome reprogramming during differentiation. Mol. Cell 56, 286–297 (2014).
Javierre, B. M., Esteller, M. & Ballestar, E. Epigenetic connections between autoimmune disorders and haematological malignancies. Trends Immunol. 29, 616–623 (2008).
Jones, P. A. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat. Rev. Genet. 13, 484–492 (2012).
Nishiyama, A. & Nakanishi, M. Navigating the DNA methylation landscape of cancer. Trends Genet. 37, 1012–1027 (2021).
de Groote, M. L., Verschure, P. J. & Rots, M. G. Epigenetic editing: targeted rewriting of epigenetic marks to modulate expression of selected target genes. Nucleic Acids Res. 40, 10596–10613 (2012).
Voigt, P. & Reinberg, D. Epigenome editing. Nat. Biotechnol. 31, 1097–1099 (2013).
Maeder, M. L. et al. Targeted DNA demethylation and activation of endogenous genes using programmable TALE-TET1 fusion proteins. Nat. Biotechnol. 31, 1137–1142 (2013).
Ou, K. et al. Targeted demethylation at the CDKN1C/p57 locus induces human beta cell replication. J. Clin. Invest. 129, 209–214 (2019).
Xu, X. et al. A CRISPR-based approach for targeted DNA demethylation. Cell Discov. 2, 16009 (2016).
Amabile, A. et al. Inheritable silencing of endogenous genes by hit-and-run targeted epigenetic editing. Cell 167, 219–232 e214 (2016).
Liu, X. S. et al. Editing DNA methylation in the mammalian genome. Cell 167, 233–247 e217 (2016).
Morita, S. et al. Targeted DNA demethylation in vivo using dCas9-peptide repeat and scFv-TET1 catalytic domain fusions. Nat. Biotechnol. 34, 1060–1065 (2016).
Josipovic, G. et al. Antagonistic and synergistic epigenetic modulation using orthologous CRISPR/dCas9-based modular system. Nucleic Acids Res. 47, 9637–9657 (2019).
Taghbalout, A. et al. Enhanced CRISPR-based DNA demethylation by Casilio-ME-mediated RNA-guided coupling of methylcytosine oxidation and DNA repair pathways. Nat. Commun. 10, 4296 (2019).
Chan, W. F. et al. Activation of stably silenced genes by recruitment of a synthetic de-methylating module. Nat. Commun. 13, 5582 (2022).
Baumann, V. et al. Targeted removal of epigenetic barriers during transcriptional reprogramming. Nat. Commun. 10, 2119 (2019).
Godecke, N. et al. Controlled re-activation of epigenetically silenced Tet promoter-driven transgene expression by targeted demethylation. Nucleic Acids Res. 45, e147 (2017).
Chen, H. et al. Induced DNA demethylation by targeting Ten-Eleven Translocation 2 to the human ICAM-1 promoter. Nucleic Acids Res. 42, 1563–1574 (2014).
Xu, X. et al. High-fidelity CRISPR/Cas9- based gene-specific hydroxymethylation rescues gene expression and attenuates renal fibrosis. Nat. Commun. 9, 3509 (2018).
Devesa-Guerra, I. et al. DNA methylation editing by CRISPR-guided excision of 5-methylcytosine. J. Mol. Biol. 432, 2204–2216 (2020).
Sapozhnikov, D. M. & Szyf, M. Unraveling the functional role of DNA demethylation at specific promoters by targeted steric blockage of DNA methyltransferase with CRISPR/dCas9. Nat. Commun. 12, 5711 (2021).
Horii, T. et al. Successful generation of epigenetic disease model mice by targeted demethylation of the epigenome. Genome Biol. 21, 77 (2020).
Liu, X. S. et al. Rescue of fragile X syndrome neurons by DNA methylation editing of the FMR1 gene. Cell 172, 979–992 e976 (2018).
Qian, J. et al. Multiplex epigenome editing of MECP2 to rescue Rett syndrome neurons. Sci. Transl. Med. 15, eadd4666 (2023).
Fujiki, K. et al. PPARgamma-induced PARylation promotes local DNA demethylation by production of 5-hydroxymethylcytosine. Nat. Commun. 4, 2262 (2013).
Fidalgo, M. et al. Zfp281 coordinates opposing functions of Tet1 and Tet2 in pluripotent states. Cell Stem Cell 19, 355–369 (2016).
Sun, Z. et al. EGR1 recruits TET1 to shape the brain methylome during development and upon neuronal activity. Nat. Commun. 10, 3892 (2019).
Kim, H. et al. Ten-eleven translocation protein 1 modulates medulloblastoma progression. Genome Biol. 22, 125 (2021).
Wu, B. K., Mei, S. C., Chen, E. H., Zheng, Y. & Pan, D. YAP induces an oncogenic transcriptional program through TET1-mediated epigenetic remodeling in liver growth and tumorigenesis. Nat. Genet. 54, 1202–1213 (2022).
Li, J. et al. TET1 dioxygenase is required for FOXA2-associated chromatin remodeling in pancreatic beta-cell differentiation. Nat. Commun. 13, 3907 (2022).
Jiang, X. et al. miR-22 has a potent anti-tumour role with therapeutic potential in acute myeloid leukaemia. Nat. Commun. 7, 11452 (2016).
Zhong, J. et al. TET1 modulates H4K16 acetylation by controlling auto-acetylation of hMOF to affect gene regulation and DNA repair function. Nucleic Acids Res. 45, 672–684 (2017).
Zhang, C. et al. The hepatic AMPK-TET1-SIRT1 axis regulates glucose homeostasis. Elife 10, e70672 (2021).
Zeng, Y. et al. Lin28A binds active promoters and recruits Tet1 to regulate gene expression. Mol. Cell 61, 153–160 (2016).
Li, W. et al. Nono deficiency compromises TET1 chromatin association and impedes neuronal differentiation of mouse embryonic stem cells. Nucleic Acids Res. 48, 4827–4838 (2020).
Zhou, T. et al. Lupus enhancer risk variant causes dysregulation of IRF8 through cooperative lncRNA and DNA methylation machinery. Nat. Commun. 13, 1855 (2022).
Wei, T. et al. An HDAC2-TET1 switch at distinct chromatin regions significantly promotes the maturation of pre-iPS to iPS cells. Nucleic Acids Res. 43, 5409–5422 (2015).
Zhu, F. et al. Sin3a-Tet1 interaction activates gene transcription and is required for embryonic stem cell pluripotency. Nucleic Acids Res. 46, 6026–6040 (2018).
Arab, K. et al. GADD45A binds R-loops and recruits TET1 to CpG island promoters. Nat. Genet. 51, 217–223 (2019).
Wu, L. et al. Hepatic Gadd45beta promotes hyperglycemia and glucose intolerance through DNA demethylation of PGC-1alpha. J. Exp. Med. 218, e20201475 (2021).
Neri, F. et al. Genome-wide analysis identifies a functional association of Tet1 and Polycomb repressive complex 2 in mouse embryonic stem cells. Genome Biol. 14, R91 (2013).
Dixon, G. et al. QSER1 protects DNA methylation valleys from de novo methylation. Science 372, eabd0875 (2021).
Okashita, N. et al. PRDM14 promotes active DNA demethylation through the ten-eleven translocation (TET)-mediated base excision repair pathway in embryonic stem cells. Development 141, 269–280 (2014).
Lio, C. W. et al. Tet2 and Tet3 cooperate with B-lineage transcription factors to regulate DNA modification and chromatin accessibility. Elife 5, e18290 (2016).
Xiong, J. et al. Cooperative action between SALL4A and TET proteins in stepwise oxidation of 5-methylcytosine. Mol. Cell 64, 913–925 (2016).
Suzuki, T. et al. RUNX1 regulates site specificity of DNA demethylation by recruitment of DNA demethylation machineries in hematopoietic cells. Blood Adv. 1, 1699–1711 (2017).
Lio, C. J. et al. TET enzymes augment activation-induced deaminase (AID) expression via 5-hydroxymethylcytosine modifications at the Aicda superenhancer. Sci. Immunol. 4, eaau7523 (2019).
Cheng, Z. L. et al. The Zscan4-Tet2 transcription nexus regulates metabolic rewiring and enhances proteostasis to promote reprogramming. Cell Rep. 32, 107877 (2020).
Mendes, K. et al. The epigenetic pioneer EGR2 initiates DNA demethylation in differentiating monocytes at both stable and transient binding sites. Nat. Commun. 12, 1556 (2021).
Morante-Palacios, O. et al. Coordinated glucocorticoid receptor and MAFB action induces tolerogenesis and epigenome remodeling in dendritic cells. Nucleic Acids Res. 50, 108–126 (2022).
Kim, M. R., Wu, M. J., Zhang, Y., Yang, J. Y. & Chang, C. J. TET2 directs mammary luminal cell differentiation and endocrine response. Nat. Commun. 11, 4642 (2020).
Wang, L. et al. TET enzymes regulate skeletal development through increasing chromatin accessibility of RUNX2 target genes. Nat. Commun. 13, 4709 (2022).
Deplus, R. et al. TET2 and TET3 regulate GlcNAcylation and H3K4 methylation through OGT and SET1/COMPASS. EMBO J. 32, 645–655 (2013).
Vella, P. et al. Tet proteins connect the O-linked N-acetylglucosamine transferase Ogt to chromatin in embryonic stem cells. Mol. Cell 49, 645–656 (2013).
Chen, Q., Chen, Y., Bian, C., Fujiki, R. & Yu, X. TET2 promotes histone O-GlcNAcylation during gene transcription. Nature 493, 561–564 (2013).
Jeong, J. J. et al. Cytokine-regulated phosphorylation and activation of TET2 by JAK2 in hematopoiesis. Cancer Discov. 9, 778–795 (2019).
Ravichandran, M. et al. Rinf regulates pluripotency network genes and Tet enzymes in embryonic stem cells. Cell Rep. 28, 1993–2003 e1995 (2019).
Wang, X. et al. PROSER1 mediates TET2 O-GlcNAcylation to regulate DNA demethylation on UTX-dependent enhancers and CpG islands. Life Sci. Alliance 5, e202101228 (2022).
Guan, W. et al. Methylcytosine dioxygenase TET3 interacts with thyroid hormone nuclear receptors and stabilizes their association to chromatin. Proc. Natl Acad. Sci. USA 114, 8229–8234 (2017).
Thakur, A. et al. Hepatocyte nuclear factor 4-alpha is essential for the active epigenetic state at enhancers in mouse liver. Hepatology 70, 1360–1376 (2019).
Xue, S. et al. TET3 inhibits type I IFN production independent of DNA demethylation. Cell Rep. 16, 1096–1105 (2016).
Zhang, Q. et al. Differential regulation of the ten-eleven translocation (TET) family of dioxygenases by O-linked beta-N-acetylglucosamine transferase (OGT). J. Biol. Chem. 289, 5986–5996 (2014).
Huang, Z. et al. The chromosomal protein SMCHD1 regulates DNA methylation and the 2c-like state of embryonic stem cells by antagonizing TET proteins. Sci. Adv. 7, eabb9149 (2021).
Hatanaka, Y. et al. Histone H3 methylated at arginine 17 is essential for reprogramming the paternal genome in zygotes. Cell Rep. 20, 2756–2765 (2017).
Moran-Crusio, K. et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 20, 11–24 (2011).
Zhao, Z. et al. Combined loss of Tet1 and Tet2 promotes B cell, but not myeloid malignancies, in mice. Cell Rep. 13, 1692–1704 (2015).
Kang, J. et al. Simultaneous deletion of the methylcytosine oxidases Tet1 and Tet3 increases transcriptome variability in early embryogenesis. Proc. Natl Acad. Sci. USA 112, E4236–E4245 (2015).
Huisman, C. et al. Re-expression of selected epigenetically silenced candidate tumor suppressor genes in cervical cancer by TET2-directed demethylation. Mol. Ther. 24, 536–547 (2016).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32201229), Shanghai Pujiang Program (21PJ1409100), GuangCi Professorship Program of Ruijin Hospital Shanghai Jiao Tong University School of Medicine, Nanning Yongjiang Plan Program, the Science and Technology Commission of Shanghai Municipality(20JC1410100)and Innovative Research Team of High-Level Local Universities in Shanghai (SHSMU-ZDCX20211801). We thank Nanning Jiuzhouyuan Biotechnology Co. Ltd. and Shanghai Frontiers Science Center of Cellular Homeostasis and Human Diseases.
Author information
Authors and Affiliations
Contributions
X.Z. and Y.Z. wrote the manuscript. X.W. and C.W. designed and revised the manuscript. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, X., Zhang, Y., Wang, C. et al. TET (Ten-eleven translocation) family proteins: structure, biological functions and applications. Sig Transduct Target Ther 8, 297 (2023). https://doi.org/10.1038/s41392-023-01537-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-023-01537-x
- Springer Nature Limited
This article is cited by
-
High estrogen during ovarian stimulation induced loss of maternal imprinted methylation that is essential for placental development via overexpression of TET2 in mouse oocytes
Cell Communication and Signaling (2024)
-
Wnt/β-catenin signaling in the development and therapeutic resistance of non-small cell lung cancer
Journal of Translational Medicine (2024)
-
A prognostic signature based on genes associated with m6A/m5C/m1A/m7G modifications and its immunological characteristics in clear cell renal cell carcinoma
Scientific Reports (2024)
-
Epigenetic regulation of DNA repair gene program by Hippo/YAP1-TET1 axis mediates sorafenib resistance in HCC
Cellular and Molecular Life Sciences (2024)