Abstract
The insulin-like growth factor-1 (IGF-1) signaling pathway is crucial for the regulation of growth and development. The correct processing of the IGF-1Ea prohormone (proIGF-1Ea) and the IGF-1 receptor (IGF-1R) peptide precursor requires proper N-glycosylation. Deficiencies of N-linked glycosylation lead to a clinically heterogeneous group of inherited diseases called Congenital Disorders of Glycosylation (CDG). The impact of N-glycosylation defects on IGF-1/IGF-1R signaling components is largely unknown. In this study, using dermal fibroblasts from patients with different CDG [PMM2-CDG (n = 7); ALG3-CDG (n = 2); ALG8-CDG (n = 1); GMPPB-CDG (n = 1)], we analyzed the glycosylation pattern of the proIGF-1Ea, IGF-1 secretion efficiency and IGF-1R signaling activity. ALG3-CDG, ALG8-CDG, GMPPB-CDG and some PMM2-CDG fibroblasts showed hypoglycosylation of the proIGF-1Ea and lower IGF-1 secretion when compared with control (CTR). Lower IGF-1 serum concentration was observed in ALG3-CDG, ALG8-CDG and in some patients with PMM2-CDG, supporting our in vitro data. Furthermore, reduced IGF-1R expression level was observed in ALG3-CDG, ALG8-CDG and in some PMM2-CDG fibroblasts. IGF-1-induced IGF-1R activation was lower in most PMM2-CDG fibroblasts and was associated with decreased ERK1/2 phosphorylation as compared to CTR. In general, CDG fibroblasts showed a slight upregulation of Endoplasmic Reticulum (ER) stress genes compared with CTR, uncovering mild ER stress in CDG cells. ER-stress-related gene expression negatively correlated with fibroblasts IGF-1 secretion. This study provides new evidence of a direct link between N-glycosylation defects found in CDG and the impairment of IGF-1/IGF-1R signaling components. Further studies are warranted to determine the clinical consequences of reduced systemic IGF-1 availability and local activity in patients with CDG.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The insulin-like growth factor-1 (IGF-1) signaling pathway is fundamental for growth regulation, especially in childhood, while also playing a central role in development and metabolic homeostasis. In the bloodstream, IGF-1 circulates primarily as part of a 150 kDa ternary complex with two additional proteins, the IGF binding protein-3 (IGFBP-3) and the acid-labile subunit (ALS). This system extends the half-life of circulating IGF-1 from 10 to 12 min to about 15 h, preserving stable serum levels of IGF-1 [1, 2]. Although growth hormone (GH) is a major regulator of hepatic IGF-1 production, IGF-1 is also regulated and secreted by other organs in an autocrine/paracrine manner [3]. Tissue-specific IGF-1 knockout models demonstrated the role of circulating IGF-1 in regulation of body size and tissue growth and revealed autocrine actions in specific tissues such as skeletal muscle [4], bone [5] and nervous system [6]. Thus, the activity of IGF-1 is due to a combination of local expression together with IGF-1 delivered to the tissue from the circulation where high levels are maintained [7].
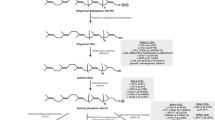
The function of all the IGF-1 system components is affected by their glycosylation status [8, 9]. The mature IGF-1 is a non-glycosylated polypeptide of 7.6 kD derived from post-translational cleavage of the C terminal E peptide from the IGF-1 prohormone (proIGF-1E) [9]. In human, the most ubiquitously expressed prohormone, named proIGF-1Ea, is glycosylated (N-glycosylation site on residue 92 of the Ea petide) and proIGF-1Ea N-glycosylation ensures proper secretion of the mature IGF-1 [9, 10]. Both IGFBP-3 and ALS proteins are glycosylated. IGFBP-3 contains three N-glycosylation sites [1] and ALS is a soluble glycoprotein of 85 kDa [11]. Previous studies have demonstrated that N-glycosylation prolongs the half-life of IGFBP-3 [12], increases the affinity of ALS to the IGFBP-3/IGF-1 complex [13] and thus is required for the correct formation and stabilization of the 150 kDa ternary complex [2]. The actions of IGF-1 are regulated by the tyrosine kinase receptor IGF-1R. This receptor is synthesized as a single polypeptide precursor (proreceptor), which undergoes proteolytic cleavage into α (130 KDa) and β (97 kDa) chains that form a tetramer (α-β-β-α). Each α and β subunit contains 11 and 5N-glycosylation sites, respectively [14]; therefore, the α-β-β-α tetramer structure may enclose 32 glycosylation positions [15]. Several studies demonstrated that proper N-glycosylation of IGF-1R proreceptor is required for correct IGF-1R maturation and transport to the cell surface [15,16,17].
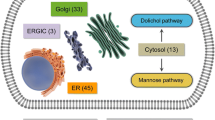
Defects in either IGF-1 or its receptor can result in poor pre- and post-natal growth [18, 19]. Beside growth failure, patients with IGF-1R haploinsufficiency often exhibit additional clinical features including skeletal malformation, intellectual disability, cardiac defects and facial dysmorphisms [19, 20]. In contrast, much less is known about the impact of N-glycosylation genetic defects, such as those found in Congenital Disorders of Glycosylation (CDG), on the IGF-1 system. CDG are a large family of rare inborn errors of metabolism caused by defective glycoprotein and glycolipid glycan synthesis and attachment. They comprise a broad spectrum of clinical manifestations, because glycosylation occurs in every cell and involves all organs and tissues [21]. Major clinical manifestations include nearly always neurological involvement (such as developmental/intellectual disability, hypotonia, epilepsy, stroke-like episodes, polyneuropathy) besides other organ involvement [21]. Nearly 160 CDG have been described and their number increases exponentially [22]. Glycosylation comprises a large number of biochemical pathways, including N-linked and O-linked pathways, glycosylphosphatidylinositol (GPI) anchor synthesis and glycolipid glycosylation. It occurs in several organelles: the cytoplasm, the endoplasmic reticulum (ER) and ER–Golgi intermediate compartment (ERGIC), the Golgi and vesicular network and the plasma and sarcolemmal membranes [23]. Most CDG are ultrarare disorders, those affecting N-glycosylation are the most common. Several specific therapeutic strategies are under evaluation for these diseases [24, 25]. PMM2-CDG is the most common N-glycosylation defect with more than 900 reported patients worldwide [21].
N-linked glycosylation disorders are usually identified by analyzing the serum transferrin isoforms by isoelectric focusing (IEF) or mass spectrometry [23]. On the basis of the transferrin IEF profiles (type 1 or type 2 pattern), N-glycosylation defects are classified as CDG-I or CDG-II. CDG-I results from defects in the glycan assembly in the cytoplasm and ER and is hallmarked by the absence of one or more N-glycans on glycoproteins [23]. CDG type-II is Golgi-related and characterized by defective glycans [23]. Hypoglycosylation leads to protein misfolding and a mild increase of ER-stress markers in patients’ fibroblasts [26].
Growing evidence suggests that insufficient levels and/or activity of growth factors might contribute to some of the clinical manifestations in CDG [8, 27,28,29]. The endocrine data from PMM2-affected children showed that circulating IGF-1 levels were in the low or low normal range in infants and children below the age of 9 years (n = 15 median, 33 µg/L; range, 12–92 µg/L; normal, 77–135 µg/L) and in the low-to-normal IGF-1 levels in adolescents (n = 8; median, 267 µg/L; range, 107–437 µg/L; normal, 350–504 µg/L) [29]. Similarly, Miller et al. [8] reported reduced serum levels of ALS, IGFBP-3, IGF-1 and of ternary complex formation in children with PMM2-CDG (n = 12; age = 8.1 ± 6.6 years) compared with age-matched controls, despite normal or elevated levels of GH. These features were also found in the recently developed mouse model of PMM2-CDG [28].
The molecular basis of the IGF-1 system deficiency found in PMM2-CDG patients is currently unknown.
In this study, we investigated whether glycosylation is directly involved in the function of the IGF-1 system. We hypothesize that primary fibroblasts from patients with CDG show aberrant proIGF-1Ea N-glycosylation and impaired activation of the IGF-1R signaling pathways.
Materials and methods
CDG fibroblasts
Primary fibroblasts derived from patients with CDG caused by different N-glycosylation defects were studied. ALG3-CDG (α-1,3-mannosyltransferase deficiency), PGAP2-CDG (post-GPI attachment to proteins 2 deficiency), ALG8-CDG (α-1,3-glucosyltransferase 2 deficiency), GMPPB-CDG (GDP-mannose pyrophosphorylase B) and control (CTR) fibroblasts from age-matched volunteers were provided by the Neuroscience Department of the Meyer Children’s Hospital (Florence, Italy). PMM2-CDG (phosphomannomutase-2 deficiency) fibroblasts were obtained from the Giannina Gaslini Institute-Telethon Network of Genetic Biobanks (Genoa, Italy) [30]. Clinical features of these patients have been previously published in part [31] and are described in the Supplementary Table 1. Fibroblasts were cultured in Dulbecco’s Modified Eagle Medium (DMEM, high glucose) supplemented with 10% heat-inactivated fetal bovine serum (FBS), 2 mM glutamine, penicillin (100 U/mL) and streptomycin (100 μg/mL) and maintained in a 5% CO2 atmosphere at 37 °C. For IGF-1R activation, after 24 h of serum-free culture in 12-well plates, cells were stimulated with IGF-1 (100 ng/mL) (Cat #I3769-50UG, Sigma-Aldrich) for 30 or 60 min before lysis.
For IGF-1 gene overexpression, fibroblasts were transfected using the Human Dermal Fibroblast Nucleofector™ Kit (Cat #VPD1001; Lonza) using the Nucleofector™ Device (Lonza) following the instruction manual. Briefly, 1 × 106 fibroblasts were resuspended in the nucleofector solution, combined with 2.5 µg of plasmid construct containing the class 1 IGF-1Ea isoform [32], transferred to the cuvette supplied and finally transfected with the U-23 program of the Nucleofector™ Device. After electroporation, cells were mixed with 500 µl of DMEM and immediately plated into a 6-well plate. The efficiency of transfection was estimated by GFP-dependent fluorescence and by qPCR for total IGF-1 mRNA levels at 48 h after transfection, as previously described [32]. The fibroblasts' concentration and cell viability were determined by the LUNA-II™ Automated Cell Counter (Logos Biosystems, Twin Helix) with trypan blue staining.
ELISA assay
Quantitative determination of IGF-1 concentrations in transfected fibroblasts supernatants was performed by a commercially available ELISA kit following the manufacturer’s instructions (Human IGF-I/IGF-1 DuoSet ELISA Cat #DY291-05; R&D Systems). The data were acquired at a wavelength of 450 nm using Model 680 microplate reader (Bio-Rad Laboratories).
Western blot and lectin blot analyses
Fibroblasts were processed for western blot analyses as previously reported [32]. The protein samples (20–40 µg total proteins) were electrophoresed with 10% SDS-PAGE and then transferred to nitrocellulose or PVDF membranes (Bio-Rad Laboratories) for immunoblotting. Primary antibodies against phospho-IGF-1 Receptor β (1:2000; Cat #3024 Cell Signaling Technology); IGF-1 Receptor β (1:2000; Cat #3027 Cell Signaling Technology), phospho-Akt (Ser473) (1:2000; Cat #9271 Cell Signaling Technology), Akt (1:2000; Cat #9272 Cell Signaling Technology), phospho-p44/42 (ERK1/2) (1:2000; Cat #9101 Cell Signaling Technology), p44/42 (ERK1/2) (1:2000; Cat #9102 Cell Signaling Technology) and IGF-1 (1:2000; Cat #500P11 Peprotech) were incubated overnight at 4 °C. For lectin blotting, membranes were probed with biotinylated Concanavalin A (ConA, 1:1000; Cat #B-1005-5) and Phaseolus vulgaris leucoagglutinin (PHA-L, 1:200; Cat #B-1115-2) lectins (Vector laboratories, D.B.A. Italia) at room temperature while shaking for 1 h. After washes, the membranes were incubated with the appropriate horseradish peroxidase (HRP)-conjugated secondary antibody (against primary antibodies) or streptavidin–HRP (for biotinylated lectins) at room temperature for 1 h and were then washed three times. Blots were developed using Clarity Western ECL Substrate (Bio-Rad Laboratories) and were quantified using the ChemiDoc MP (Bio-Rad Laboratories) equipped with Image Lab software.
RNA extraction, cDNA synthesis and quantitative real-time polymerase chain reaction
The total RNA was extracted and genes of interest analyzed by real-time qPCR as previously described [33]. Briefly, the expression of ER-stress-related genes was monitored by qPCR using TB Green Premix Ex TaqII Mastermix (Takara Bio Europe, France), in a RotorGene 6000 instrument (Corbett life science, Sydney, Australia). As template, 2 ng of total RNA used for cDNA synthesis was used in each PCR tube and a non-template control was included for each primer pair reaction as negative control. All amplification reactions were performed in duplicate. The amplification conditions were: 95 °C for 10 min, 95 °C for 10 s and 60 °C for 50 s for 40 cycles. At the end of each run, a melting curve analysis from 65 °C to 95 °C was performed to exclude the presence of non-specific products or primer dimers. The data were normalized against the reference gene GAPDH (glyceraldehyde-3-phosphate dehydrogenase). The relative expression levels were calculated using the 2−ΔΔCt method [34].
Statistical analyses
The data are represented as mean ± SEM of at least three independent experiments. Statistical analyses were performed using the t test or one-way ANOVA as appropriate, followed by Tukey’s Multiple Comparison post hoc test. Correlation between fibroblasts IGF-1 secretion, densitometric quantification of proIGF-1Ea and IGF-1R immunoreactive bands, lectin binding and ER-stress-related genes was analyzed using Pearson’s correlation. The statistical tests were performed using SPSS (IBM SPSS Statistics for Windows, Version 20.0, IBM Corp.) and GraphPad Prism version 5 (GraphPad Software, Inc., La Jolla, CA, USA). A p value ≤ 0.05 was considered statistically significant.
Results
proIGF-1Ea N-glycosylation pattern and mature IGF-1 secretion in CDG fibroblasts
To evaluate the expression level of IGF-1 in CTR and CDG fibroblasts, we conducted preliminary analyses on the IGF-1 mRNA and protein quantity. IGF-1 mRNA quantification by qPCR showed that all fibroblasts had detectable levels of IGF-1 mRNA (mean CT value 30.5); however, the IGF-1 protein level was too low to be detected by western blot or ELISA. We also tried to quantify the IGF-1 secretion in the fibroblast supernatants after concentration using 3 kDa Spin Columns (by about 20-fold; Amicon® Ultra Merck Millipore), but the IGF-1 concentration still remained below the ELISA sensitivity (93.8 pg/mL). Subsequently, to assess the expression pattern and the N-glycosylation status of the IGF-1 protein, we transiently transfected fibroblasts with a plasmid encoding the IGF-1Ea isoform or an empty vector [32] by electroporation.
As shown in Fig. 1A, two distinct bands, likely representing glycosylated (~ 17 kDa) and unglycosylated (~ 12 kDa) proIGF-1Ea, were detected in CTR and PGAP2-CDG fibroblasts. In contrast, the glycosylation pattern of proIGF-1Ea differed in other CDG fibroblasts. In particular, the two fibroblasts from ALG3-CDG presented a distinct band of about 14 kDa, while ALG8-CDG and GMPPB-CDG revealed a marked accumulation of 12 kDa proteins. It is likely that all these bands represent underglycosylated forms of the proIGF-1Ea. Among PMM2-CDG fibroblasts analyzed, there was a general preservation of proIGF-1Ea N-glycosylation pattern, however the IGF-1 bands intensity in PMM2_p1-CDG, PMM2_p6-CDG and PMM2_p7-CDG fibroblasts were lower compared to CTR. Densitometric analysis of western blot bands further confirmed a reduction of glycosylated proIGF-1Ea band in most CDG fibroblasts compared to CTR (Fig. 1B), while the intensities of un/under-glycosylated bands did not change (ANOVA, p = 0.101). Subsequently, we quantified the IGF-1 protein level in the cell culture media of IGF-1Ea-transfected fibroblasts (Fig. 1C). We observed that both ALG3-CDG fibroblasts derived from two different patients, GMPPB-CDG, PMM2_p1-CDG and PMM2_p6-CDG showed a decreased IGF-1 secretion as compared to CTR. The secretion pattern of IGF-1 positively correlated with the glycosylated proIGF-1Ea quantity (r = 0.612, p = 0.020), corroborating the previous finding that N-glycosylation is required for proper proIGF-1Ea processing and IGF-1 secretion [9, 10]. IGF-1 serum level was also quantified from some available patients (Supplementary Table 1). Low serum IGF-1 was found in ALG3-CDG (52 ng/mL, normal 95–312 ng/mL) and ALG8-CDG patients (80 ng/mL, normal, 95–460 ng/mL), while normal levels were observed in PGAP2-CDG (116 ng/mL, normal 47–231 ng/mL). Among PMM2-CDG patients, decreased IGF-1serum levels were observed in patients 2 (64 ng/ml, normal 99–238 ng/ml) and 4 (80 ng/ml, normal 99–238 ng/ml) while it was in the low normal or normal range in patients 1 (101 ng/ml, normal 82–214 ng/ml) and 7 (361 ng/mL, normal 119–395 ng/mL), respectively.
IGF-1Ea prohormone expression patterns and IGF-1 secretion. Representative western blot showing the IGF-1Ea prohormone expression patterns (A) and relative protein expression level (B) of IGF-1Ea transfected fibroblasts (mean ± SEM of three technical replicates for each fibroblast subtype). Quantification of IGF-1 level in fibroblast supernatants measured by ELISA (C) (mean ± SEM of three technical replicates for each fibroblast subtype). Gly_proIGF-1Ea: glycosylated IGF-1Ea prohormone; un/under gly_proIGF-1Ea: un/underglycosylated IGF-1Ea prohormone. *significantly different from CTR fibroblasts; ***p < 0.001, **p < 0.01, *p < 0.05
IGF-1R expression and IGF-1R pathway activation in CDG fibroblasts
As shown in Fig. 2A, ALG3-CDG fibroblasts from two different patients and ALG8-CDG cells revealed reduced IGF-1R expression levels compared to CTR fibroblasts. PMM2-CDG fibroblasts from different patients presented significant variability in the IGF-1R expression levels. IGF-1R was reduced in about 50% of PMM2-CDG cells tested. Reduced levels of IGF-1R found in CDG fibroblasts were not followed by an IGF-1R proreceptor accumulation (Fig. 2B and C). Altogether, CDG fibroblasts showed reduced IGF-1R expression compared with CTR (Fig. 2D). To further investigate the possible differences in the two main study CDG subtypes, we performed a post hoc subgroup analysis for ALG-CDG (n = 3) and PMM2-CDG (n = 7) showing reduced IGF-1R levels both in ALG-CDG (p = 0.003) and PMM2-CDG (p = 0.02). To evaluate if IGF-1R deficiency observed in PMM2-CDG was associated with reduced IGF-1R pathway activation, fibroblasts were treated with recombinant IGF-1 (100 ng/ml) for 30 and 60 min and the level of IGF-1R (Fig. 3A), Akt (Fig. 3B) and ERK1/2 (Fig. 3C) phosphorylation was quantified by western blotting. The IGF-1-induced activation of IGF-1R and ERK1/2 was reduced in most PMM2-CDG fibroblasts, while Akt phosphorylation level was more heterogeneous. Notably, in some PMM2-CDG, the decreased response to IGF-1 treatment was mainly due to higher basal IGF-1R and ERK1/2 activation (e.g., PMM2_p2-CDG; PMM2_p3-CDG; PMM2_p5-CDG; PMM2_p6-CDG), instead of general inhibition of IGF-1R and ERK1/2 activity (Fig. 3D). When PMM2-CDG fibroblasts derived from different patients were pooled together (n = 7), both IGF-1R and ERK1/2 IGF-1-induced activation were reduced compared to CTR (Fig. 3E). Preliminary results obtained in some ALG-CDG fibroblasts showed IGF-1 pathway downregulation (not shown) (Fig. 4).
IGF-1R protein levels in CDG fibroblasts. Relative expression level of IGF-1R (A) and IGF-1R proreceptor (B) in different CDG quantified by western blot (mean ± SEM of three technical replicates for each fibroblast subtype). Representative western blot showing expression levels of IGF-1R proreceptor (~ 200 kDa), IGF-1R (~ 97 kDa) and GAPDH (~ 36 kDa) (C). Relative expression levels of IGF-1R in CTR (mean of three technical replicates for each of five biological replicates) and CDG- fibroblasts (mean of three technical replicates for each of twelve biological replicates) quantified by western blot (D). *significantly different from CTR fibroblasts; ***p < 0.0001, **p < 0.001, *p < 0.05
IGF-1R signaling pathway activation. Relative expression levels of phosphorylated IGF-1R (A), Akt (B) and ERK1/2 (C) in PMM2-CDG fibroblasts obtained from different patients (mean ± SEM of three technical replicates for each fibroblast subtype). Representative western blot showing expression levels of phosphorylated and total IGF-1R, Akt, ERK1/2 and GAPDH (D). IGF-1R, Akt and ERK1/2 phosphorylation level in CTR (mean of three technical replicates for each of five biological replicates) and PMM2-CDG (mean of three technical replicates for each of seven biological replicates) fibroblasts (E). *significantly different from CTR fibroblasts; #significantly different from recombinant IGF-1 untreated cells *** and ###p < 0.0001, ##**p < 0.001, *p < 0.01; #p < 0.05
Lectin-binding analysis. Concanavalin A (ConA) and Phaseolus vulgaris leucoagglutinin (PHA-L) binding to CTR (mean of three technical replicates for each of five biological replicates) and CDG (mean of three technical replicates for each of twelve biological replicates) fibroblasts. Con A and PHA-L recognize high mannose and complex type N-glycans, respectively. *significantly different from CTR fibroblasts, *p < 0.0001
Lectin binding analysis and ER-stress-related markers in CDG fibroblasts
Lectin blotting with ConA and PHA-L, which recognize high mannose and beta 1,6 branched oligosaccharides complex type N-glycans, respectively, showed a non-significant decrement trend of PHA-L binding in CDG- compared to CTR- fibroblasts (p = 0.29), while ConA reactivity did not change (p = 0.43). Subsequently, qPCR was carried out to quantify the expression level of selected ER-stress-related genes: CHOP/DDIT3, sXBP1, uXBP1, MAP1LC3B, HSPA5, CEBPB, CHAC1 and ATF4. As shown in Fig. 5, CHOP/DDIT3, uXBP1 and ATF4 mRNA levels were higher in CDG compared to CTR fibroblasts. The post hoc subgroup analysis showed that ATF4 mRNA expression was higher in ALG-CDG as compared to PMM2-CDG and CTR fibroblasts (p = 0.008), while CHOP/DDIT3 and uXBP1 mRNA levels did not differ between ALG-CDG and PMM2-CDG. The Pearson’s correlation coefficient showed that fibroblast IGF-1 secretion was directly associated with PHA-L binding (r = 0.516, p = 0.05) and inversely correlated with CHOP/DDIT3 (r = − 0.747, p = 0.002), uXBP1 (r = − 0.715, p = 0.004), CEBPB (r = − 0.548, p = 0.043) and ATF4 (r = − 0.735, p = 0.003) mRNA expression.
ER-stress markers' mRNA expression in CDG fibroblasts. Quantification of the expression level of selected ER-stress-related genes: CHOP/DDIT3, sXBP1, uXBP1, HSPA5, CEBPB, CHAC1, MAP1LC3B and ATF4 in CTR (mean of three technical replicates for each of five biological replicates) and CDG (mean of three technical replicates for each of 12 biological replicates) fibroblasts. *significantly different from CTR fibroblasts, **p < 0.01, *p < 0.05
Discussion
In this study, we provide new evidence for IGF-1 system impairment in CDG fibroblasts due to defective N-glycosylation of proIGF-1Ea and IGF-1R, which partially disrupts IGF-1 signaling pathway activation. We previously demonstrated that, under physiological conditions, intracellular IGF-1 is mainly expressed as a heavily N-glycosylated prohormone of 17–22 kDa [10]. Here, using CTR and CDG fibroblasts transiently transfected with the IGF-1Ea isoform, we monitored the N-glycosylation pattern of proIGF-1Ea and IGF-1 secretion. Two distinct bands, likely representing glycosylated and unglycosylated proIGF-1Ea, were found in CTR, PGAP2-CDG and most PMM2-CDG fibroblasts by western blotting. Conversely, a unique band was present in fibroblasts derived from ALG3-CDG, ALG8-CDG and GMPPB-CDG. In both ALG3-CDG, we found an abnormal band of about 14 kDa, while in ALG8-CDG and in GMPPB-CDG, we found a marked accumulation of a 12 KDa isoform. These bands probably represent hypoglycosylated isoforms of the proIGF-1Ea. To evaluate if aberrant proIGF-1Ea glycosylation impaired IGF-1 secretion, we quantified the IGF-1 protein level in cell culture supernatants and found that most CDG fibroblasts released less IGF-1 in the culture media compared to CTR. These data corroborate our previous findings that N-glycosylation of the proIGF-1Ea is essential for correct IGF-1 secretion [10]. Normal proIGF-1Ea N-glycosylation and IGF-1 secretion in PGAP2-CDG fibroblasts was expected, since PGAP2 is not involved in N-glycosylation but in GPI-anchor synthesis of GPI-anchored proteins. The IGF-1 serum level was evaluated in seven patients, whose fibroblasts have been included in this work (ALG3-CDG, ALG8-CDG, PGAP2-CDG and PMM2_p1,_p2,_p4,_p7-CDG). ALG3-CDG and ALG8-CDG patients showed low IGF-1 serum levels, while low-to-normal IGF-1 serum levels were found in studied patients with PMM2-CDG. These data remain provisional, due to the low number of patients tested, but they are in line with the published data on reduced circulating IGF-1 levels in CDG patients [8, 29].
Because the effects of IGF-1 are mediated principally through the IGF-1R, we subsequently analyzed the IGF-1R levels in primary fibroblasts from different CDG. Previous studies have shown that N-glycosylation contributes to ligand binding, kinase activity and turnover of the IGF-1R [35]. Accordingly, we found that IGF-1R expression and activity were reduced in primary fibroblasts from different CDG. Levels of IGF-1R were particularly low in ALG3-CDG, ALG-8-CDG and in some of the PMM2-CDG fibroblast cell lines tested. We did not find an accumulation of the IGF-1 proreceptor in these cells, suggesting that defective N-glycosylation causes IGF-1R proreceptor instability and degradation [27]. In PMM2-CDG, we also analyzed the IGF-1-induced IGF-1R pathway activation. Most PMM2-CDG fibroblasts showed reduced IGF-1R and ERK1/2 phosphorylation compared to CTR, while the Akt activation did not differ significantly probably due to the high variability of the Akt response among PMM2-CDG fibroblasts.
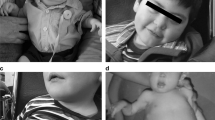
Altogether, these data corroborate and expand previous observations demonstrating an IGF-1 system deficiency in CDG [8, 27,28,29]. In particular, our results suggest that the glycosylation defect found in CDG might impair the IGF-1 system not only indirectly, by destabilizing the ternary complex [8], but also directly due to proIGF-1Ea hypoglycosylation and IGF-1 secretory defect. Furthermore, local growth-promoting effects of IGF-1 might be compromised by the reduced IGF-1R proreceptor glycosylation and related impairment of IGF-1 signal transduction. We believe that these findings are relevant in the context of CDG since both systemic IGF-1 production and local bioactivity are essential to support normal growth during development [5, 7]. Patients with genetic IGF-1R defects, show symptoms (i.e., pre and postnatal growth retardation, microcephaly, cardiac defects and dysmorphic features) that significantly overlap with those presented in various CDG types [19, 20]. For example, the majority of CDG patients analyzed in the present study showed growth failure and microcephaly (Supplementary Table 1), which are also clinical hallmarks of IGF-1/IGF-1R defects [36,37,38]. Further studies are needed, also taking advantage of the recently developed CDG animal models [28], to evaluate the impact of glycosylation defects on each component of the IGF-1 system. This, in turn, will help to clarify the association between clinical CDG features and IGF-1/IGF-1R signaling abnormalities. Interestingly, a case study showed that treatment with recombinant IGF-1 prompted linear growth in a child with PMM2-CDG [39], suggesting the potential of IGF-1 therapy in the improvement of clinical outcome of CDG patients. However, taking into account our data together with previous findings [27], the residual IGF-1R activity should be measured before starting a recombinant IGF-1 therapy.
Although typical clinical features of CDG were present in patients whose fibroblasts we studied, clinical manifestations and severity have a wide spectrum, which is probably related to the specific gene variants and/or enzyme residual activity [30 and Supplementary Table 1]. Global quantitative protein glycosylation performed with lectins that have specific affinity for different types of N-glycans (ConA and PHA-L) failed to highlight difference between different CDG fibroblast subtypes. We found a modest, and not significant, decrease of PHA-L reactivity, indicating reduced levels of β1–6 branching structures and complex N-glycans. A positive association between IGF-1 secretion and PHA-L binding was found. In addition, the analysis of ER-stress-related gene expression showed a slight increase of CHOP/DDIT3, uXBP1 and ATF4 mRNA level in CDG compared to CTR fibroblasts, which negatively correlate with fibroblasts IGF-1 secretion. A correlation between ER stress and IGF-1 system alterations has been proposed in several neurodegenerative diseases that share as common feature the accumulation of misfolded proteins [40]. Although further studies are needed, the broad modification of protein N-glycosylation patterns and the mild ER-stress in CDG fibroblasts might also have contributed to the IGF-1/IGF-1R impaired signaling observed in CDG fibroblasts [40, 41].
Conclusions
In this study, we have shown that primary fibroblasts from CDG patients have reduced levels of proIGF-1Ea and IGF-1R proreceptor glycosylation. The decreased glycosylation of IGF-1/IGF-1R signaling pathway components reduced the IGF-1 bioactivity in CDG fibroblasts. Thus, our molecular data provide new insight into CDG pathogenesis and suggest that both IGF-1 production (circulating/local) and bioactivity might be compromised in CDG due to reduced N-glycosylation.
These findings pave the way to future studies focused on the impact of CDG on the different IGF-1 system components, which include IGF-1, IGFBPs and the IGF-1R. More efforts will be needed to clarify the correlation between reduced IGF-1/IGF-1R signaling in CDG patients and their clinical manifestations.
Availability of data and materials
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- Akt:
-
RAC-alpha serine/threonine-protein kinase
- ALG3:
-
Dol-P-Man:Man(5)GlcNAc(2)-PP-Dol alpha-1,3-mannosyltransferase
- ALG8:
-
Dolichyl pyrophosphate Glc1Man9GlcNAc2 alpha-1,3-glucosyltransferase
- ALS:
-
Acid-labile subunit
- ATF4:
-
Activating transcription factor 4
- CDG:
-
Congenital Disorder(s) of Glycosylation
- CEBPB:
-
CCAAT enhancer binding protein beta
- CHAC1:
-
ChaC glutathione-specific gamma-glutamylcyclotransferase 1
- CHOP/DDIT3:
-
DNA damage inducible transcript 3
- ConA:
-
Concanavalin A
- CTR:
-
Control
- DMEM:
-
Dulbecco’s Modified Eagle Medium
- ER:
-
Endoplasmic reticulum
- ERGIC:
-
ER–Golgi intermediate compartment
- ERK:
-
Mitogen-activated protein kinase
- FBS:
-
Fetal bovine serum
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- GH:
-
Growth hormone
- GMPPB:
-
GDP-mannose pyrophosphorylase
- GPI:
-
Glycosylphosphatidylinositol
- HRP:
-
Horseradish peroxidase
- HSPA5:
-
Heat shock protein family A (Hsp70) member 5
- IEF:
-
Isoelectric focusing
- IGF-1:
-
Insulin-like growth factor-1
- IGF1-R:
-
Insulin-like growth factor-1 receptor
- IGFBP-3:
-
IGF binding protein-3
- MAP1LC3B:
-
Microtubule associated protein 1 light chain 3 beta
- PGAP2:
-
Post-GPI attachment to proteins factor 2
- PHA-L:
-
Phaseolus vulgaris Leucoagglutinin
- PMM2:
-
Phosphomannomutase 2
- proIGF-1E:
-
IGF-1 prohormone
- sXBP1:
-
Spliced X-box binding protein 1
- uXBP1:
-
Unspliced X-box binding protein 1
References
Collett-Solberg PF, Cohen P (1996) The role of the insulin-like growth factor binding proteins and the IGFBP proteases in modulating IGF action. Endocrinol Metab Clin North Am 25:591–614. https://doi.org/10.1016/s0889-8529(05)70342-x
Boisclair YR, Rhoads RP, Ueki I, Wang J, Ooi GT (2001) The acid-labile subunit (ALS) of the 150 kDa IGF-binding protein complex: an important but forgotten component of the circulating IGF system. J Endocrinol 170:63–70. https://doi.org/10.1677/joe.0.1700063
Ohlsson C, Mohan S, Sjögren K, Tivesten A, Isgaard J, Isaksson O et al (2009) The role of liver-derived insulin-like growth factor-I. Endocrine Rev. https://doi.org/10.1210/er.2009-0010
Vassilakos G, Lei H, Yang Y, Puglise J, Matheny M, Durzynska J et al (2019) Deletion of muscle IGF-I transiently impairs growth and progressively disrupts glucose homeostasis in male mice. The FASEB J. https://doi.org/10.1096/fj.201800459R
Yakar S, Werner H, Rosen CJ (2018) Insulin-like growth factors: actions on the skeleton. J Mol Endocrinol 61:T115–T137. https://doi.org/10.1530/JME-17-0298
Lewitt MS, Boyd GW (2019) The role of insulin-like growth factors and insulin-like growth factor-binding proteins in the nervous system. Biochem Insights. https://doi.org/10.1177/1178626419842176
LeRoith D, Holly JMP, Forbes BE (2021) Insulin-like growth factors: ligands, binding proteins, and receptors. Mol Metab. https://doi.org/10.1016/j.molmet.2021.101245
Miller BS, Khosravi MJ, Patterson MC, Conover CA (2009) IGF system in children with congenital disorders of glycosylation. Clin Endocrinol (Oxf) 70:892–897. https://doi.org/10.1111/j.1365-2265.2009.03531.x
Philippou A, Maridaki M, Pneumaticos S, Koutsilieris M (2014) The complexity of the IGF1 gene splicing, posttranslational modification and bioactivity. Mol Med. https://doi.org/10.2119/molmed.2014.00011
Annibalini G, Contarelli S, de Santi M, Saltarelli R, di Patria L, Guescini M et al (2018) The intrinsically disordered E-domains regulate the IGF-1 prohormones stability, subcellular localisation and secretion. Sci Rep 8:8919. https://doi.org/10.1038/s41598-018-27233-3
Janosi JB, Ramsland PA, Mott MR, Firth SM, Baxter RC, Delhanty PJ (1999) The acid-labile subunit of the serum insulin-like growth factor-binding protein complexes. Structural determination by molecular modeling and electron microscopy. J Biol Chem 274:23328–23332. https://doi.org/10.1074/jbc.274.33.23328
Firth SM, McDougall F, McLachlan AJ, Baxter RC (2002) Impaired blockade of insulin-like growth factor I (IGF-I)-induced hypoglycemia by IGF binding protein-3 analog with reduced ternary complex-forming ability. Endocrinology 143:1669–1676. https://doi.org/10.1210/endo.143.5.8764
Janosi JBM, Firth SM, Bond JJ, Baxter RC, Delhanty PJD (1999) N-linked glycosylation and sialylation of the acid-labile subunit. J Biol Chem 274:5292–5298. https://doi.org/10.1074/jbc.274.9.5292
Dricu A, Carlberg M, Wang M, Larsson O (1997) Inhibition of N-linked glycosylation using tunicamycin causes cell death in malignant cells: role of down-regulation of the insulin-like growth factor 1 receptor in induction of apoptosis. Cancer Res 57:543–548
Carlberg M, Dricu A, Blegen H, Wang M, Hjertman M, Zickert P et al (1996) Mevalonic acid is limiting for N-linked glycosylation and translocation of the insulin-like growth factor-1 receptor to the cell surface. Evidence for a new link between 3-hydroxy-3-methylglutaryl-coenzyme a reductase and cell growth. J Biol Chem 271:17453–17462. https://doi.org/10.1074/jbc.271.29.17453
Siddals KW, Marshman E, Westwood M, Gibson JM (2004) Abrogation of insulin-like growth factor-I (IGF-I) and insulin action by mevalonic acid depletion: synergy between protein prenylation and receptor glycosylation pathways. J Biol Chem 279:38353–38359. https://doi.org/10.1074/jbc.M404838200
Dricu A, Kanter L, Wang M, Nilsson G, Hjertman M, Wejde J et al (1999) Expression of the insulin-like growth factor 1 receptor (IGF-1R) in breast cancer cells: evidence for a regulatory role of dolichyl phosphate in the transition from an intracellular to an extracellular IGF-1 pathway. Glycobiology 9:571–579. https://doi.org/10.1093/glycob/9.6.571
Klammt J, Pfäffle R, Werner H, Kiess W (2008) IGF signaling defects as causes of growth failure and IUGR. Trends Endocrinol Metab 19:197–205. https://doi.org/10.1016/j.tem.2008.03.003
Ester WA, van Duyvenvoorde HA, de Wit CC, Broekman AJ, Ruivenkamp CA, Govaerts LC et al (2009) Two short children born small for gestational age with insulin-like growth factor 1 receptor haploinsufficiency illustrate the heterogeneity of its phenotype. J Clin Endocrinol Metab 94:4717–4727. https://doi.org/10.1210/jc.2008-1502
Veenma DC, Eussen HJ, Govaerts LC, de Kort SW, Odink RJ, Wouters CH et al (2010) Phenotype-genotype correlation in a familial IGF1R microdeletion case. J Med Genet 47:492–498. https://doi.org/10.1136/jmg.2009.070730
Ondruskova N, Cechova A, Hansikova H, Honzik T, Jaeken J (2021) Congenital disorders of glycosylation: still “hot” in 2020. Biochim Biophys Acta Gen Subj 1865:129751. https://doi.org/10.1016/j.bbagen.2020.129751
Péanne R, de Lonlay P, Foulquier F, Kornak U, Lefeber DJ, Morava E et al (2018) Congenital disorders of glycosylation (CDG): Quo vadis? Eur J Med Genet 61:643–663. https://doi.org/10.1016/j.ejmg.2017.10.012
Chang IJ, He M, Lam CT (2018) Congenital disorders of glycosylation. Ann Transl Med 6:477. https://doi.org/10.21037/atm.2018.10.45
Ng BG, Freeze HH (2018) Perspectives on glycosylation and its congenital disorders. Trends Genet 34:466–476. https://doi.org/10.1016/j.tig.2018.03.002
Verheijen J, Tahata S, Kozicz T, Witters P, Morava E (2020) Therapeutic approaches in congenital disorders of glycosylation (CDG) involving N-linked glycosylation: an update. Genet Med 22:268–279. https://doi.org/10.1038/s41436-019-0647-2
Lecca MR, Wagner U, Patrignani A, Berger EG, Hennet T (2005) Genome-wide analysis of the unfolded protein response in fibroblasts from congenital disorders of glycosylation type-I patients. FASEB J 19:240–242. https://doi.org/10.1096/fj.04-2397fje
Dhaunsi GS (2017) Receptor-mediated selective impairment of insulin-like growth factor-1 activity in congenital disorders of glycosylation patients. Pediatr Res 81:526–530. https://doi.org/10.1038/pr.2016.96
Chan B, Clasquin M, Smolen GA, Histen G, Powe J, Chen Y et al (2016) A mouse model of a human congenital disorder of glycosylation caused by loss of PMM2. Hum Mol Genet 25:2182–2193. https://doi.org/10.1093/hmg/ddw085
de Zegher F, Jaeken J (1995) Endocrinology of the carbohydrate-deficient glycoprotein syndrome type 1 from birth through adolescence. Pediatric Res. https://doi.org/10.1203/00006450-199504000-00003
Filocamo M, Mazzotti R, Corsolini F, Stroppiano M, Stroppiana G, Grossi S et al (2014) Cell line and DNA biobank from patients affected by genetic diseases. Open J Bioresour 1:e2. https://doi.org/10.5334/ojb.ab
Barone R, Carrozzi M, Parini R, Battini R, Martinelli D, Elia M et al (2015) A nationwide survey of PMM2-CDG in Italy: high frequency of a mild neurological variant associated with the L32R mutation. J Neurol 262:154–164. https://doi.org/10.1007/s00415-014-7549-7
de Santi M, Annibalini G, Barbieri E, Villarini A, Vallorani L, Contarelli S et al (2016) Human IGF1 pro-forms induce breast cancer cell proliferation via the IGF1 receptor. Cell Oncol (Dordr) 39:149–159. https://doi.org/10.1007/s13402-015-0263-3
Galluzzi L, Diotallevi A, de Santi M, Ceccarelli M, Vitale F, Brandi G et al (2016) Leishmania infantum induces mild unfolded protein response in infected macrophages. PLoS ONE 11:e0168339. https://doi.org/10.1371/journal.pone.0168339
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29:e45. https://doi.org/10.1093/nar/29.9.e45
Ferreira IG, Pucci M, Venturi G, Malagolini N, Chiricolo M, Dall’Olio F (2018) Glycosylation as a main regulator of growth and death factor receptors signaling. Int J Mol Sci. https://doi.org/10.3390/ijms19020580
Walenkamp MJE, Wit JM (2019) A homozygous mutation in the highly conserved Tyr60 of the mature IGF1 peptide broadens the spectrum of IGF1 deficiency. Eur J Endocrinol 181:C29-33. https://doi.org/10.1530/EJE-19-0801
Walenkamp MJE, Robers JML, Wit JM, Zandwijken GRJ, van Duyvenvoorde HA, Oostdijk W et al (2019) Phenotypic features and response to GH treatment of patients with a molecular defect of the IGF-1 receptor. J Clin Endocrinol Metab 104:3157–3171. https://doi.org/10.1210/jc.2018-02065
Gkourogianni A, Andrade AC, Jonsson B, Segerlund E, Werner-Sperker A, Horemuzova E et al (2020) Pre- and postnatal growth failure with microcephaly due to two novel heterozygous IGF1R mutations and response to growth hormone treatment. Acta Paediatr 109:2067–2074. https://doi.org/10.1111/apa.15218
Miller BS, Duffy MM, Addo OY, Sarafoglou K (2013) rhIGF-1 therapy for growth failure and IGF-1 deficiency in congenital disorder of glycosylation Ia (PMM2 Deficiency). J Investig Med High Impact Case Rep 1:232470961350331. https://doi.org/10.1177/2324709613503316
Garcia-Huerta P, Troncoso-Escudero P, Jerez C, Hetz C, Vidal RL (2016) The intersection between growth factors, autophagy and ER stress: a new target to treat neurodegenerative diseases? Brain Res 1649:173–180. https://doi.org/10.1016/j.brainres.2016.02.052
Xia W, Wang Y, Zhang Y, Ge X, Lv P, Cheng J et al (2020) Endoplasmic reticulum stress induces growth retardation by inhibiting growth hormone IGF-I axis. Growth Horm IGF Res 55:101341. https://doi.org/10.1016/j.ghir.2020.101341
Acknowledgements
“The Cell Line and DNA Biobank from Patients Affected by Genetic Diseases”, member of the Telethon Network of Genetic Biobanks (project no. GTB18001), funded by Telethon Italy, provided us with specimens”. Thanks to the Professor Timothy Bloom for a critical reading of this work. We gratefully acknowledge AMMeC (Associazione Malattie Metaboliche Congenite, Italy), for technical assistance and collaboration to the Molecular and Cell Biology Laboratory of Neurometabolic Diseases at the Meyer Children’s Hospital of Florence, Italy.
Funding
The laboratory part of this research did not receive any specific Grant from funding agencies in the public, commercial or not-for-profit sectors. Patients were studied in the context of the Tuscany Region Call for Health 2018 (Grant DECODE-EE).
Author information
Authors and Affiliations
Contributions
LDP, GA and EB conceived and designed the study. LDP, GA, RS and MB performed cell culture experiments, western blot and lectin blot analyses and ELISA assay. LG and AD performed the quantification of ER-stress-related genes. GA, AM, LF and EB analyzed the data and drafted the manuscript. MAD, RB, RG, JJ, VS and EB provided overall direction to the project and revised the manuscript. All the authors contributed to manuscript revision, read and approved the submitted version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Code availability
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Di Patria, L., Annibalini, G., Morrone, A. et al. Defective IGF-1 prohormone N-glycosylation and reduced IGF-1 receptor signaling activation in congenital disorders of glycosylation. Cell. Mol. Life Sci. 79, 150 (2022). https://doi.org/10.1007/s00018-022-04180-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04180-x