Abstract
KDEL receptor cycles between the ER and the Golgi to retrieve ER-resident chaperones that get leaked to the secretory pathway during protein export from the ER. Recent studies have shown that a fraction of KDEL receptor may reside in the plasma membrane and function as a putative cell surface receptor. However, the trafficking itinerary and mechanism of cell surface expressed KDEL receptor remains largely unknown. In this study, we used N-terminally Halo-tagged KDEL receptor to investigate its endocytosis from the plasma membrane and trafficking itinerary of the endocytosed receptor through the endolysosomal compartments. Our results indicate that surface-expressed KDEL receptor undergoes highly complex recycling pathways via the Golgi and peri-nuclear recycling endosomes that are positive for Rab11 and Rab14, respectively. Unexpectedly, KDEL receptor appears to preferentially utilize clathrin-mediated endocytic pathway as well as clathrin-dependent transport carriers for export from the trans-Golgi network. Taken together, we suggest that KDEL receptor may be a bona fide cell surface receptor with a complex, yet well-defined trafficking itinerary through the endolysosomal compartments.







Similar content being viewed by others
References
Munro S, Pelham HR (1987) A C-terminal signal prevents secretion of luminal ER proteins. Cell 48:899–907
Lewis MJ, Pelham HR (1990) A human homologue of the yeast HDEL receptor. Nature 348:162–163
Lewis MJ, Sweet DJ, Pelham HR (1990) The ERD2 gene determines the specificity of the luminal ER protein retention system. Cell 61:1359–1363
Pulvirenti T et al (2008) A traffic-activated Golgi-based signalling circuit coordinates the secretory pathway. Nat Cell Biol 10:912–922
Capitani M, Sallese M (2009) The KDEL receptor: new functions for an old protein. FEBS Lett 583:3863–3871
Giannotta M et al (2012) The KDEL receptor couples to Galphaq/11 to activate Src kinases and regulate transport through the Golgi. EMBO J 31:2869–2881
Solis GP et al (2017) Golgi-resident Galphao promotes protrusive membrane dynamics. Cell 170:939–955 e924
Cancino J et al (2014) Control systems of membrane transport at the interface between the endoplasmic reticulum and the Golgi. Dev Cell 30:280–294
Brauer P et al (2019) Structural basis for pH-dependent retrieval of ER proteins from the Golgi by the KDEL receptor. Science 363:1103–1107
Lewis MJ, Pelham HR (1992) Ligand-induced redistribution of a human KDEL receptor from the Golgi complex to the endoplasmic reticulum. Cell 68:353–364
Townsley FM, Wilson DW, Pelham HR (1993) Mutational analysis of the human KDEL receptor: distinct structural requirements for Golgi retention, ligand binding and retrograde transport. EMBO J 12:2821–2829
Wilson DW, Lewis MJ, Pelham HR (1993) pH-dependent binding of KDEL to its receptor in vitro. J Biol Chem 268:7465–7468
Aoe T et al (1997) The KDEL receptor, ERD2, regulates intracellular traffic by recruiting a GTPase-activating protein for ARF1. EMBO J 16:7305–7316
Griffiths G et al (1994) Localization of the Lys, Asp, Glu, Leu tetrapeptide receptor to the Golgi complex and the intermediate compartment in mammalian cells. J Cell Biol 127:1557–1574
Raykhel I et al (2007) A molecular specificity code for the three mammalian KDEL receptors. J Cell Biol 179:1193–1204
Henderson MJ, Richie CT, Airavaara M, Wang Y, Harvey BK (2013) Mesencephalic astrocyte-derived neurotrophic factor (MANF) secretion and cell surface binding are modulated by KDEL receptors. J Biol Chem 288:4209–4225
Lindahl M et al (2014) MANF is indispensable for the proliferation and survival of pancreatic beta cells. Cell Rep 7:366–375
Becker B et al (2016) Cargo binding promotes KDEL receptor clustering at the mammalian cell surface. Sci Rep 6:28940
Neves J et al (2016) Immune modulation by MANF promotes tissue repair and regenerative success in the retina. Science 353:aaf3646
Norisada J, Hirata Y, Amaya F, Kiuchi K, Oh-hashi K (2016) A comparative analysis of the molecular features of MANF and CDNF. PLoS ONE 11:e0146923
Lindahl M, Saarma M, Lindholm P (2017) Unconventional neurotrophic factors CDNF and MANF: structure, physiological functions and therapeutic potential. Neurobiol Dis 97:90–102
Borner GH et al (2012) Multivariate proteomic profiling identifies novel accessory proteins of coated vesicles. J Cell Biol 197:141–160
Borner GH, Fielding AB (2014) Isolating HeLa cell fractions enriched for clathrin-coated vesicles. Cold Spring Harb Protoc 2014:1184–1187
Linford A et al (2012) Rab14 and its exchange factor FAM116 link endocytic recycling and adherens junction stability in migrating cells. Dev Cell 22:952–966
Koike S, Jahn R (2019) SNAREs define targeting specificity of trafficking vesicles by combinatorial interaction with tethering factors. Nat Commun 10:1608
Dores MR, Schnell JD, Maldonado-Baez L, Wendland B, Hicke L (2010) The function of yeast epsin and Ede1 ubiquitin-binding domains during receptor internalization. Traffic 11:151–160
Dores MR, Trejo J (2019) Endo-lysosomal sorting of G-protein-coupled receptors by ubiquitin: diverse pathways for G-protein-coupled receptor destruction and beyond. Traffic 20:101–109
Hari J et al (1987) The receptor for insulin-like growth factor II mediates an insulin-like response. EMBO J 6:3367–3371
Morgan DO et al (1987) Insulin-like growth factor II receptor as a multifunctional binding protein. Nature 329:301–307
Roth RA et al (1987) Interactions of the receptor for insulin-like growth factor II with mannose-6-phosphate and antibodies to the mannose-6-phosphate receptor. Biochem Biophys Res Commun 149:600–606
Yamamoto K et al (2003) The KDEL receptor modulates the endoplasmic reticulum stress response through mitogen-activated protein kinase signaling cascades. J Biol Chem 278:34525–34532
Wang P, Li B, Zhou L, Fei E, Wang G (2011) The KDEL receptor induces autophagy to promote the clearance of neurodegenerative disease-related proteins. Neuroscience 190:43–55
Tapia D et al (2019) KDEL receptor regulates secretion by lysosome relocation- and autophagy-dependent modulation of lipid-droplet turnover. Nat Commun 10:735
Funding
This project was sponsored by Shanghai Pujiang Program (16PJ1407 to YQ). Financial support of this study was provided exclusively by the ShanghaiTech University.
Author information
Authors and Affiliations
Contributions
IL, XY, YQ designed the study. JJ, XY, YQ, LZ, BG, SJ, YW performed the experiments. JJ, XY, YQ and IL wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
18_2020_3570_MOESM1_ESM.pdf
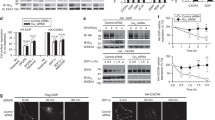
Supplementary file1. Supplementary Fig. 1(A) KDELR1 knockdown efficiency in MCF7 KDELR KD cells was determined by quantitative real-time PCR in three independent experiments and summarized by histogram. KDELR2 and KDELR3 mRNA levels were also determined as controls. Y axis indicates the relative mRNA level of KDEL receptor in MCF7 KDELR KD cells compared to MCF7 control cells. **: P < 0.01 (B) KDELR1 protein knockdown efficiency by siRNA was determined. KDELR1-3Flag-mCherry knock-in cells were transfected with scramble or KDELR1 siRNA for 72 h before lysed and analyzed by western blotting with anti-mCherry and anti-GAPDH antibodies. Supplementary Fig. 2(H) Confocal images of overexpressed Halo-KDELR or Halo-KDELR H12A in the Golgi region of HeLa cells. Endogenous GM130 was stained as a Golgi marker. Scale bar: 1 µm. Bar graph summarize the co-localization of Halo-KDELRs with GM130 (n = 15 cells). ns not significant. Statistical analysis was performed using Students’ t test. Supplementary Table 1. Summary of the Pearson coefficient values (mean ± SD) in the graphs (PDF 134 kb)
Rights and permissions
About this article
Cite this article
Jia, J., Yue, X., Zhu, L. et al. KDEL receptor is a cell surface receptor that cycles between the plasma membrane and the Golgi via clathrin-mediated transport carriers. Cell. Mol. Life Sci. 78, 1085–1100 (2021). https://doi.org/10.1007/s00018-020-03570-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-020-03570-3