Abstract
The effectiveness of many DNA-damaging chemotherapeutic drugs depends on their ability to form monoadducts, intrastrand crosslinks and/or interstrand crosslinks (ICLs) that interfere with transcription and replication. The ERCC1–XPF endonuclease plays a critical role in removal of these lesions by incising DNA either as part of nucleotide excision repair (NER) or interstrand crosslink repair (ICLR). Engagement of ERCC1–XPF in NER is well characterized and is facilitated by binding to the XPA protein. However, ERCC1–XPF recruitment to ICLs is less well understood. Moreover, specific mutations in XPF have been found to disrupt its function in ICLR but not in NER, but whether this involves differences in lesion targeting is unknown. Here, we imaged GFP-tagged ERCC1, XPF and ICLR-defective XPF mutants to investigate how in human cells ERCC1–XPF is localized to different types of psoralen-induced DNA lesions, repaired by either NER or ICLR. Our results confirm its dependence on XPA in NER and furthermore show that its engagement in ICLR is dependent on FANCD2. Interestingly, we find that two ICLR-defective XPF mutants (R689S and S786F) are less well recruited to ICLs. These studies highlight the differential mechanisms that regulate ERCC1–XPF activity in DNA repair.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The heterodimeric ERCC1–XPF complex (also called ERCC1–ERCC4) is a structure-specific endonuclease that plays an essential role in multiple DNA repair pathways by incising DNA repair intermediate fork structures at the junction between single-stranded and double-stranded DNA. ERCC1–XPF excises bulky DNA lesions in nucleotide excision repair (NER) [1], unhooks interstrand crosslinks (ICLs), i.e., covalent linkages between two bases on opposite DNA strands, in interstrand crosslink repair (ICLR) [2,3,4], removes 3′ flaps of DNA intermediate structures in specific double-strand break (DBS) repair pathways [5] and cleaves 3′ overhangs from uncapped telomeres to shorten them [6]. Hereditary mutations in ERCC1–XPF cause several distinct human syndromes characterized by either (skin) cancer proneness, i.e., xeroderma pigmentosum (XP), developmental abnormalities and accelerated aging, i.e., XP combined with Cockayne syndrome (XPCS complex), or bone marrow failure and chromosome fragility, i.e., Fanconi anemia (FA) [1, 7,8,9,10]. The severe and pleiotropic symptoms associated with these syndromes highlight the fundamental role of the complex in promoting health by its essential role in different DNA repair processes.
The ERCC1–XPF complex was initially identified as an essential endonuclease in NER [1]. This DNA repair pathway removes many different types of DNA lesions that disrupt base-pairing, including photolesions induced by UV light and monoadducts and intrastrand crosslinks induced by widely used chemotherapeutic drugs such as mitomycin C (MMC), cisplatin and psoralens [11, 12]. NER is a multistep process consisting of the coordinated, sequential assembly of multiple repair enzymes guided by direct protein–protein and protein–DNA interactions. NER is initiated by two different damage sensing mechanisms. In global genome NER (GG-NER), lesions located anywhere in the genome are recognized by the UV-DDB and XPC–RAD23B–CETN2 complexes. In transcription-coupled NER, lesions located in the transcribed strand are sensed by stalling of elongating RNA polymerase II, which leads to the recruitment of the CSA, CSB and UVSSA proteins. Lesion recognition in both subpathways is followed by recruitment of the transcription factor TFIIH, which unwinds the DNA and verifies the presence of damage. TFIIH function is stimulated by the DNA damage-binding protein XPA [13], which together with the single-stranded DNA-binding protein RPA coordinates the positioning of the endonucleases ERCC1–XPF and XPG. XPG is first recruited through an interaction with TFIIH [14]. Next, ERCC1–XPF is recruited through a direct interaction between XPA and an XPA-binding region located within the central domain of ERCC1 [15,16,17,18,19]. XPF is the actual nuclease of this complex and incises the DNA 5′ to the lesion [20], after which XPG incises the DNA 3′ of the lesion. Following excision, the resulting 22–30 nt gap is filled by DNA synthesis and ligation.
In addition to NER, ERCC1–XPF is essential for the removal of ICLs. These are rare DNA modifications induced by either endogenous aldehydes [21] or by chemotherapeutic drugs such as MMC, cisplatin and psoralens [22]. ICLs are highly cytotoxic as they obstruct DNA strand separation, required for both replication and transcription. However, despite many studies focused on understanding how ICLs are removed from the genome, the molecular mechanism(s) of ICLR is not yet entirely clear. ICLR is a multistep, sequential assembly of repair enzymes and involves distinct subpathways depending on the cell cycle phase and type of ICL. Most ICLs are repaired during S phase, which requires the concerted action of different DNA repair systems, including the Fanconi anemia (FA) and homologous recombination (HR) repair pathways, NER and translesion synthesis (TLS). This replication-dependent ICLR is proposed to be initiated when two replication forks converge on an ICL. This leads to the helicase heterodimer FANCM/FAAP24-dependent recruitment of the FA-core complex and monoubiquitylation of the FANCD2–FANCI heterodimer [23, 24]. Experiments using Xenopus laevis egg extract and in vitro-modeled ICL-containing replication structures suggest that ERCC1–XPF is then recruited, depending on its interaction with the scaffold protein SLX4 that stimulates its function [2, 25, 26]. ERCC1–XPF incises the lagging strand to unhook the ICL, possibly together with another endonuclease or together with the exonuclease SNM1A that digests past the ICL [27]. The resulting single-stranded gap is filled by TLS and is used as homology template for repair of the remaining double-strand break by HR [28, 29]. The unhooked crosslink is likely repaired by NER. Certain types of ICLs, such as, for instance, induced by psoralens, appear to be preferentially unhooked by DNA glycosylase NEIL3-mediated cleavage of one of the two N-glycosyl bonds forming the crosslink, which circumvents the need for FA factors and incision of the DNA backbone [30]. Other forms of ICLR probably exist that are independent of replication, such as in slowly or nonreplicating cells, but these are only poorly understood. Replication-independent ICLR is proposed to involve GG-NER and/or TC-NER factors, likely depending on the type of crosslinked chemical, including ERCC1–XPF to unhook the ICL [31,32,33,34,35]. TLS then fills the resulting gap while a second round of NER removes the remaining ICL from the opposite DNA strand [22, 28].
Understanding how ERCC1–XPF is recruited to sites of DNA damage is necessary to understand how ERCC1–XPF protects against different types of DNA damage, including those induced by commonly used chemotherapeutics, how it helps to prevent cancer and how its inherited deficiency can cause different diseases. Previous imaging studies by us and others have highlighted the importance of XPA in regulating ERCC1–XPF recruitment to UV damage and have shown that patient-derived mutations within XPF can differentially affect its subcellular localization or recruitment to UV-induced DNA damage, explaining phenotypic differences observed in diseases associated with NER dysfunction, such as XP and XPCS complex [16, 17, 36, 37]. However, XPA is not implicated in replication-dependent ICLR and not much is known about how ERCC1–XPF is recruited to ICLs in human cells. Here we used imaging of wild-type and ICLR-defective XPF to investigate how ERCC1–XPF is distinctly recruited to psoralen–DNA crosslinks repaired by NER or by ICLR. Our results substantiate that FANCD2 is required for recruitment of the complex to ICLs and suggest that XPF ICLR-defective mutants are unable to efficiently associate with ICLs.
Methods
Cell culture and siRNA
U2OS cells expressing ERCC1–GFP, XPF–GFP and XPF-C236R, XPF-R689S and XPF-S786F were previously described [37, 38]. To knockout XPA, U2OS cells expressing ERCC1–GFP or XPF–GFP were transfected with pLentiCRISPR-V2 plasmid [39] encoding the sgRNA GGCGGCTTTAGAGCAACCCG, targeting the first exon of the XPA gene. Transfected cells were selected with puromycin and individual XPA KO clones were screened by immunoblot and sequencing and analyzed using Tracking Indels by DEcomposition as described [40]. All cell lines were cultured in standard condition: DMEM/F10 supplemented with 10% fetal calf serum (FCS) and 1% penicillin–streptomycin (PS) at 37 °C and 5% CO2. For siRNA treatment, cells were transfected using RNAiMax (Invitrogen) with control siRNA (Dharmacon, D-001210-05), siRNA targeting XPA (Dharmacon, MJAWM-000011) or FANCD2 (Dharmacon, D-016376-02), 48 h before UVC or 8-MOP treatment.
Immunoprecipitation
Immunoprecipitation of chromatin-enriched nuclear extracts for mass spectrometry was performed as described [41]. ERCC1–GFP cells were seeded in 14-cm culture dishes and irradiated with 20 J/m2 UVC (254 nm lamp, Philips) or left untreated. After 1 h, cells were harvested by scraping in 3 ml cold PBS containing protease inhibitor cocktail (Roche), centrifuged for 10 min at 1500 rpm and washed again with PBS. Cell pellet was then incubated in HEPES buffer (10 mM HEPES, pH 7.6, 1.5 mM MgCl2, 10 mM KCl, 0.5 mM DTT, protease inhibitor cocktail) for 10 min on ice. Dounce homogenizer with a type A pestle was used to isolate the nuclei prior to centrifugation at 3000 rpm for 10 min at 4 °C. Next, cell pellets were resuspended in HEPES buffer (100 mM HEPES, pH 7.6, 1.5 mM MgCl2, 150 mM NaCl, 0.5 mM DTT, 25% glycerol, protease inhibitor cocktail) and subsequently dounced using a type B pestle. 25 U micrococcal nuclease (MNase, Sigma) was used to digest chromatin extracts for 1 h at 4 °C to obtain mononucleosomal size material by centrifugation at 15,000 rpm for 15 min.
Immunoprecipitation of benzonase-treated lysates was performed as described [42]. Briefly, cells were lysed using IP buffer (30 mM Tris, pH 7.5, 150 mM NaCl, 2 mM MgCl2, 0.5% Triton X-100, protease inhibitor cocktail (Roche)) supplemented with 250 U/mL Benzonase® nuclease.
In both cases, GFP-Trap®_A beads (Chromotek) were used to immunoprecipitate GFP-tagged proteins. Cell lysate and immunoprecipitated samples were analyzed by immunoblot or mass spectrometry. For mass spectrometry, beads from UVC- and mock-treated cells were combined in a 1:1 ratio and label swapping was used to validate the biological findings and exclude contaminants.
Immunoblot
Protein lysates were prepared by scraping cells in 2 × sample buffer (125 mM Tris–HCl, pH 6.8, 20% glycerol, 10% 2-β-mercaptoethanol, 4% SDS, 0.01% bromophenol blue) and boiled at 98 °C for 5 min. Proteins were separated by SDS-PAGE and transferred to a PVDF membrane (0.45 µm, Merck Millipore). Subsequently, membranes were blocked in 2% BSA and incubated with primary antibodies and secondary antibodies conjugated with CF IRDye 680 and 770 (Sigma) for 1 h or overnight. Primary antibodies used were anti-XPA (sc-853, Santa Cruz Biotechnology), anti-SNF2H (ab3749, Abcam), anti-Tubulin (T6074, Sigma), anti-FANCD2 (nb100-316, Novus Biologicals), anti-GFP (ab290, Abcam), anti-SLX4 (NBP1-28680, Novus Biologicals), anti-ERCC1 (ab129267, Abcam), anti-H2B (07-371, Millipore), and anti-RPA70 (2267, Cell signaling). Secondary antibodies were visualized using the Odyssey CLx Infrared Imaging System (LI-COR Biosciences).
Clonogenic survival assays
To analyze the UV sensitivity of the ERCC1–GFP and XPF–GFP XPA-deficient cells, we seeded 500 cells in six-well plates in triplicate. After 24 h, cells were irradiated with the indicated doses of UV (254 nm UVC lamp, Philips). 5–7 days after irradiation, cells were fixed and stained with 50% methanol, 7% acetic acid, and 0.1% Brilliant Blue R (Sigma). Colonies were counted using the integrated colony counter GelCount (Oxford Optronix). The number of colonies in the non-treated cells was set to 100% and the number of colonies after UV treatment was normalized to the number of colonies in untreated samples and plotted as percentage survival.
UV, psoralen treatment and immunofluorescence
To analyze the localization of proteins to LUD, cells seeded on coverslips were irradiated with 60 J/m2 (254 nm UVC lamp, Philips) through an 8-µm microporous filter (Millipore) and fixed with 2% paraformaldehyde supplemented with 0.1% Triton X-100, 1 h after irradiation. Next, cells were permeabilized with 0.1% Triton X-100 for 20 min, incubated with 0.07 M NaOH for 5 min (to visualize CPDs) and washed in PBS containing 0.15% glycine and 0.5% BSA before proceeding with immunofluorescence. Analysis of protein localization to psoralen-induced DNA damage was performed as described [43, 44]. Briefly, cells were seeded on coverslips and incubated with 50 µM 8-methoxypsoralen (8-MOP, Sigma) for 2 h. 8-MOP was locally activated in stripes along the cell nuclei by a 355-nm UVA laser coupled to a PALM laser dissection microscope (Zeiss) through a 40 × 0.60 NA Korr LD Plan Neofluar objective. Cells were then fixed with 2% paraformaldehyde supplemented with 0.1% Triton X-100 and washed in PBS containing 0.15% glycine and 0.5% BSA before proceeding with immunofluorescence. For immunofluorescence, cells were stained with primary antibodies for 2 h and then incubated with secondary antibodies conjugated to ALEXA fluorochromes 488, 555 and 633 (Invitrogen) for 1 h. Primary antibodies used were against: GFP (ab290, Abcam), CPD (CAC-NM-DND-001; Cosmo Bio), XPA (sc-853, Santa Cruz Biotechnology), FANCD2 (nb100-316, Novus Biologicals), SLX4 (NBP1-28680, Novus Biologicals), γH2AX (ab11174, Abcam or 05-636, Millipore), XPF (sc-136153, Santa Cruz Biotechnology). DAPI Vectashield (Vector Laboratories) was used to mount the coverslips. Cells were imaged using an LSM700 microscope equipped with a 40 × Plan-apochromat 1.3 NA oil immersion lens (Carl Zeiss). The software Image J was used to quantify fluorescence intensity. Fold accumulation was calculated as the ratio between fluorescence intensity at the site of damage and fluorescence intensity in the cell nucleus, measured in at least 60 cells per condition in two or more separate experiments. Statistical analysis of the differences in GFP intensity at sites of local damage was performed using an unpaired two-tailed Student’s t test.
SILAC-based mass spectrometry analysis
For SILAC, cells were cultured in DMEM containing 10% dialyzed FBS (Gibco), 10% GlutaMAX (Life Technologies), penicillin/streptomycin (Life Technologies), unlabeled l-arginine–HCl and l-lysine–HCl (“light”) or 13C6,15N4l–arginine–HCl and 13C6,15N2l–lysine–2HCl (“heavy”) (Cambridge Isotope Laboratories). For the “Forward” experiment, immunoprecipitation extracts derived from “light”-labeled mock-treated cells were mixed with extracts derived from “heavy” UVC-treated (20 J/m2) cells; for the “Reverse” experiment, extracts derived from “light”-labeled UVC-treated (20 J/m2) cells were mixed with extracts derived from “heavy” mock-treated cells. After protein separation, SDS-PAGE gel was cut into 2-mm slices and subjected to in-gel reduction with dithiothreitol, alkylation with iodoacetamide (98%; D4, Cambridge Isotope Laboratories) and digested with trypsin (sequencing grade; Promega). Samples were analyzed by mass spectrometry using an Orbitrap Fusion™ Tribrid™ mass spectrometer and EASY-nLC™ 1000 (Thermo). MaxQuant software was used to analyze the data.
Results
ERCC1–XPF recruitment to sites of UV damage is XPA dependent
To investigate how the activity of ERCC1–XPF in response to ICLs is regulated in cells, we studied how the complex is recruited to sites of locally induced psoralen-DNA crosslinks independently of NER. ERCC1–XPF binding to damaged DNA during NER critically depends on its interaction with XPA [15,16,17,18,19]. Therefore, we generated XPA-knockout (KO) cell lines by transfecting a plasmid encoding Cas9 and an sgRNA targeting exon 1 of XPA in previously generated U2OS cells expressing ERCC1–GFP [38] and U2OS XPF KO cells expressing XPF–GFP [37]. Sequencing and Tracking Indels by DEcomposition analysis [40] revealed multiple indel mutations in the XPA gene of a selected ERCC1–GFP-expressing and a selected XPF–GFP-expressing XPA KO clone (data not shown). The absence of XPA in these clones was confirmed by immunoblot (Fig. 1a, b), while strong UV hypersensitivity (Fig. 1c, d) showed that these cells indeed lack functional NER.
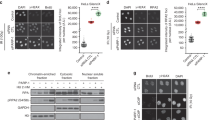
Generation of XPA knockout in ERCC1–GFP- and XPF–GFP-expressing cells. a Immunoblot showing XPA expression levels in XPA-proficient and XPA-knockout (XPA KO) U2OS cells expressing GFP-tagged ERCC1. SNF2H was used as loading control. b Immunoblot showing XPA expression levels in XPA-proficient and XPA-knockout (XPA KO) U2OS cells expressing GFP-tagged XPF. Tubulin was used as loading control. c Clonogenic UV survival assays of wild-type U2OS cells without any transgene (U2OS) and XPA-proficient and XPA KO U2OS cells expressing ERCC1–GFP. Results are plotted as average of three independent experiments, each performed in triplicate. d Clonogenic UV survival assays of XPA-proficient and XPA KO U2OS cells expressing XPF–GFP. Results are plotted as average of three independent experiments, each performed in triplicate. In c and d error bars represent the SEM
To show the validity of these cells in discerning between ERCC1–XPF activities in NER and in ICLR, we tested ERCC1–GFP and XPF–GFP recruitment to local UV damage (LUD) in XPA-proficient and XPA KO cells by immunofluorescence, after UVC irradiation through an 8-µm microporous filter (Fig. 2a–c). While both ERCC1 and XPF were clearly recruited to LUD in XPA-proficient cells, we could not observe clear recruitment in XPA KO cells. Depletion of FANCD2 in XPA-proficient and XPA KO cells expressing XPF–GFP, by a 48-h siRNA treatment (Fig. S1A), did not reduce XPF–GFP LUD recruitment as compared to control siRNA-treated cells (Fig. 2b, c). Our data confirm that ERCC1–XPF engagement in NER is completely dependent on XPA [45] and not regulated by FANCD2, indicating that the XPA KO cells can be used to study ERCC1–XPF activity outside NER.
XPF recruitment to LUD is XPA dependent. a Immunofluorescence images showing LUD recruitment of ERCC1–GFP (top panel) or XPF–GFP (lower panel) and endogenous XPA in XPA-proficient and XPA KO U2OS cells, 1 h after 60 J/m2 UVC irradiation through an 8-µm microporous filter. Cells were immunostained against XPA and CPD, as damage marker. The GFP signal was not amplified using antibody staining. Scale bar: 5 µm. b Immunofluorescence images showing XPF–GFP LUD recruitment 1 h after 60 J/m2 UVC through an 8-µm microporous filter in XPA-proficient and XPA KO U2OS cells treated with nontargeting siRNA (sictrl) or siRNA targeting FANCD2 (siFANCD2). Cells were immunostained against GFP and CPD, which was used as damage marker. Scale bar: 5 µm. c Quantification of XPF–GFP LUD recruitment in XPA-proficient and XPA KO cells, as determined by immunofluorescence experiments shown in b. The fold accumulation at sites of local damage was calculated over the nuclear background and plotted as average of at least 100 cells per condition from two independent experiments. Statistical significant difference (p < 0.05) is indicated by asterisk, n.s. non-significant. d Scatter plot of the log2 SILAC ratio of the forward and reverse SILAC ERCC1–GFP immunoprecipitations experiments, comparing ERCC1–GFP interactors with and without UVC treatment (20 J/m2, 1 h). UV-specific interactors are depicted in the upper right quadrant. In b error bars represent the SEM
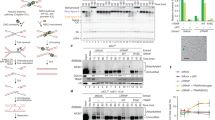
To determine whether any additional, as-of-yet unidentified factors regulate ERCC1–XPF recruitment to UV damage, we combined ERCC1–GFP immunoprecipitation with stable isotope labeling by amino acids in cell culture (SILAC)-based quantitative proteomics (including label swapping, i.e., “forward” and “reverse” replicate experiments). To this end, SILAC-labeled ERCC1–GFP-expressing U2OS cells were mock-treated or exposed to UVC (20 J/m2) and 1 h after treatment chromatin-enriched nuclear extracts were subjected to GFP-mediated immunoprecipitation and mass spectrometry analysis. Interestingly, and as expected, this showed that after UV irradiation, ERCC1–XPF interacts with multiple components of the TFIIH complex, RPA and XPG (Fig. 2d and Table 1). Also XPA was identified as an interactor of ERCC1, but only in one of the replicate experiments and, therefore, not depicted in Fig. 2d. However, no additional new factors were identified, suggesting that no major or essential unknown factors exist that regulate ERCC1–XPF involvement in NER. Moreover, SLX4 was identified as a constitutive interaction partner of the endonuclease complex, in line with previous observations [2, 25, 26]. However, the interaction ratio of this partner did not change upon UV irradiation, in line with its specific role in ICLR.
ICLR and NER core factors localize to psoralen-induced DNA damage
To analyze ERCC1–XPF recruitment to ICLs, we combined the use of the psoralen 8-methoxypsoralen (8-MOP) with UVA laser microirradiation along a user-defined track across the cell nucleus [44, 46], to activate the 8-MOP and induce local psoralen–DNA crosslinks. UVA activation of intercalated 8-MOP generates both monoadducts as well as ICLs in DNA [47,48,49,50,51]. The efficiency of this method to induce ICLs was shown by the clear recruitment of endogenous FANCD2 and SLX4 to local psoralen adducts, marked by γH2AX co-staining, as visualized by immunofluorescence in U2OS cells (Fig. 3a, 8-MOP + laser). Importantly, we did not observe recruitment of FANCD2 or SLX4 or induction of γH2AX in cells that were only UVA microirradiated but not treated with 8-MOP (laser), indicating that the applied UVA laser by itself did not generate DNA damage that leads to γH2AX signaling or is recognized by FANCD2. Furthermore, we observed clear recruitment of XPA to local psoralen adducts, as marked by γH2AX (Fig. 3b), indicative of targeting of the core NER machinery to psoralen adducts and suggesting the induction of NER substrate lesions. As monoadducts formed by psoralen are known NER substrates [52, 53], XPA is likely recruited to psoralen monoadducts as part of the NER machinery. Since NER is also implicated in replication-dependent and -independent ICLR [31, 32], XPA may be recruited to psoralen-ICLs as well. Moreover, also endogenous XPF as well as ERCC1–GFP and XPF–GFP were clearly recruited to psoralen adducts. Because of the induction of multiple types of DNA damage, this accumulation likely reflects the engagement of ERCC1–XPF in both active NER as well as ICLR.
ICLR and NER core factors are recruited to psoralen-induced DNA damage. a Immunofluorescence images showing recruitment of endogenous FANCD2 and SLX4 to sites of local UVA laser microirradiation in untreated (laser) or psoralen-treated (8-MOP + laser, 50 µM 8-MOP, 2 h) U2OS cells. Cells were immunostained against FANCD2, SLX4 and γH2AX, which was used as damage marker. Scale bar: 5 µm. b Immunofluorescence images showing recruitment to sites of local UVA laser microirradiation in psoralen-treated U2OS cells (8-MOP + laser, 50 µM 8-MOP, 2 h) of endogenous XPA and XPF (left panels) and in psoralen-treated U2OS cells expressing ERCC–GFP or XPF–GFP (right panels). Cells were immunostained against XPA, XPF, GFP and γH2AX, which was used as damage marker. Scale bar: 5 µm
ERCC1–XPF recruitment to psoralen adducts is XPA and FANCD2 dependent
To test whether ERCC1–XPF recruitment to ICLs is dependent on FANCD2, we studied the recruitment of the complex to local psoralen adducts in ERCC1–GFP- and XPF–GFP-expressing XPA-proficient and -KO cells, depleted of FANCD2 by siRNA (Fig. 4). In XPA-proficient cells, XPF–GFP recruitment to psoralen adducts was reduced after siFANCD2 treatment compared to control siRNA-treated cells (Fig. 4a, b). Also, ERCC1–GFP recruitment was slightly reduced in siFANCD2-treated cells compared to control siRNA-treated cells (Fig. 4c, d). This was less pronounced than for XPF, possibly because of competition with endogenous ERCC1 that is still present in these cells (and which we did not visualize). These observations suggest that ERCC1–XPF recruitment to psoralen adducts is partially dependent on FANCD2. Furthermore, the absence of XPA also induced a strong reduction in ERCC1–GFP and XPF–GFP localization to psoralen adducts that was further reduced by additional FANCD2 depletion (Fig. 4). Our data, therefore, suggest that, in response to psoralen adducts, ERCC1–XPF is recruited both by XPA to participate in NER and by FANCD2 to function in ICLR. These observations in human cells are in accordance with chromatin immunoprecipitation (ChIP) studies in Xenopus egg extracts suggesting that FANCD2 preceeds and is needed for ERCC1–XPF recruitment to an ICL [2].
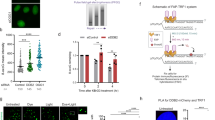
ERCC1–XPF recruitment to psoralen-induced DNA damage is XPA and FANCD2 dependent. a Immunofluorescence images showing XPF–GFP recruitment to sites of local UVA laser microirradiation in psoralen-treated (50 µM 8-MOP, 2 h) XPA-proficient and XPA KO U2OS cells treated with nontargeting siRNA (sictrl) or siRNA targeting FANCD2 (siFANCD2). Cells were immunostained against GFP and γH2AX, which was used as damage marker. Scale bar: 5 µm. b Quantification of XPF–GFP recruitment to sites of local UVA laser microirradiation in psoralen-treated XPA-proficient and XPA KO U2OS cells, as determined by immunofluorescence experiments shown in a. The fold accumulation at sites of local damage was calculated over the nuclear background and plotted as average of at least 80 cells per condition from three independent experiments. Statistically significant difference (p < 0.05) is indicated by asterisk. c Immunofluorescence images showing ERCC1–GFP recruitment to sites of local UVA laser microirradiation in psoralen-treated (50 µM 8-MOP, 2 h) XPA-proficient and XPA KO U2OS cells treated with nontargeting siRNA (sictrl) or siRNA targeting FANCD2 (siFANCD2). Cells were stained against GFP and γH2AX, which was used as damage marker. Scale bar: 5 µm. d Quantification of ERCC1–GFP recruitment to sites of local UVA laser microirradiation in psoralen-treated XPA-proficient and XPA KO U2OS cells, as determined by immunofluorescence experiments shown in c. The fold accumulation at sites of local damage was calculated over the nuclear background and plotted as average of at least 100 cells per condition from two independent experiments. Statistically significant difference (p < 0.05) is indicated by asterisk. In b and d, error bars represent the SEM
The stronger reduction in ERCC1–XPF recruitment due to XPA KO as compared to siFANCD2 could be because of incomplete FANCD2 depletion by siRNA. However, considering that we obtained efficient FANCD2 knockdown (Fig. S1a) this is likely better explained by the observation that psoralen-ICLs are preferentially repaired via unhooking by the DNA glycosylase NEIL3, which is independent of FANCD2–FANCI [30]. Moreover, XPA, but not FANCD2, is implicated in replication-independent ICLR as well [31, 33, 35]. Knockout of XPA will, therefore, also impair ERCC1–XPF engagement in this pathway. Importantly, this implies that even though in our experiments we did not distinguish between different cell cycle phases of the cells under study (which were rapidly dividing U2OS cells), the FANCD2-dependent recruitment of ERCC1–XPF reflects its engagement in replication-dependent ICLR. Finally, we observed residual ERCC1–XPF recruitment even in the absence of both XPA and FANCD2. It is thus possible that the ERCC1–XPF complex can still be targeted to psoralen adducts, albeit less efficient, even in the absence of both of these factors. Alternatively, it could be that this residual targeting reflects a function of the complex in yet another DNA repair pathway, possibly the repair of DSBs [5].
R689S and S786F mutant XPF recruitment to psoralen adducts is XPA dependent
Several of the XPF mutations reported in patients have been described to disrupt ICLR. Among these are C236R, located in the SF2 helicase-like domain and associated with XPCS complex with FA features [7]; R689S, located in the nuclease domain and associated with FA [8]; S786F, also located in the nuclease domain and identified in a breast cancer [54]. Cells carrying the R689S or S786F mutant XPF alleles are hypersensitive to MMC but not to UV irradiation, suggesting that the R689S and S786F mutations only impair XPF function in ICLR [8, 37, 54]. Conversely, C236R renders cells hypersensitive to both MMC and UV irradiation, indicating that this mutation impairs XPF function in both ICLR and NER [7, 37]. Moreover, previous incision assays and ChIP experiments in Xenopus egg extracts showed that all three mutations impair the unhooking of a sequence-specific cisplatin–DNA ICL by XPF but not its recruitment to this lesion [55]. Therefore, we determined whether in human cells these mutations affect ERCC1–XPF recruitment to psoralen adducts.
We previously generated U2OS XPF KO cell lines that stably expressed GFP-tagged XPF mutants carrying the amino acid substitutions C236R (XPF–C236R), R689S (XPF–R689S) and S786F (XPF–S786F) [37]. To study their ability to be recruited to psoralen adducts repaired by NER or repaired by ICLR, we treated these cells with control, XPA and FANCD2 siRNA (Fig. S1a, B) and applied local UVA laser microirradiation after 8-MOP treatment. XPF–C236R was clearly recruited to sites of damage (Fig. 5a, b), which was reduced after FANCD2 and, even more, after XPA depletion. These results indicate that this XPF mutant is recruited to psoralen adducts as part of both the NER and the replication-dependent ICLR machinery, similarly as wild-type XPF (Fig. 4a, b). Conversely, XPF–S786F and, even more so, XPF–R689S were recruited less efficiently to psoralen adducts. Strikingly, their recruitment was only strongly reduced after XPA depletion but not significantly after FANCD2 depletion. This suggests that ERCC1–XPF carrying R689S or S786F can still be normally recruited as part of the NER machinery, in line with previous immunofluorescence and fluorescence recovery after photobleaching experiments that showed efficient binding of these mutants to UV-induced DNA damage [37]. However, the absence of a significant dependency on FANCD2 suggests that these mutants cannot efficiently be recruited to or stably associate with psoralen–DNA ICLs, which is in line with the MMC hypersensitivity of cells expressing these mutants [8, 37, 54]. Interestingly, the less efficient recruitment to psoralen adducts of XPF–R689S compared to XPF–S786F may indicate that this mutation has a more deleterious effect on XPF function in ICLR. This is in line with the stronger MMC hypersensitivity observed in cells expressing XPF with this mutation [37] and the stronger nuclease and ICL-unhooking defect found with purified Xenopus XPF carrying this mutation [55].
R689S and S786F mutant XPF is recruited to psoralen adducts in an XPA-dependent and FANCD2-independent manner. a Immunofluorescence images showing XPF recruitment to sites of local UVA laser microirradiation in psoralen-treated (50 µM 8-MOP, 2 h) U2OS XPF KO cells expressing GFP-tagged XPF carrying the amino acid mutations C236R, R689S or S786F, treated with nontargeting siRNA (sictrl) or siRNA targeting XPA (siXPA) or FANCD2 (siFANCD2). Cells were immunostained against GFP and γH2AX, which was used as damage marker. Scale bar: 5 µm. b Quantification of mutant XPF–GFP recruitment to sites of local UVA laser microirradiation in psoralen-treated U2OS XPF KO cells expressing GFP-tagged XPF carrying the amino acid mutations C236R, R689S or S786F, as determined by immunofluorescence experiments shown in a. The fold accumulation at sites of local damage was calculated over the nuclear background and plotted as average of at least 60 cells per condition from three independent experiments. Statistical significant difference (p < 0.05) compared to sicntrl of each cell line is indicated by asterisk. c Immunoblot analysis of cell lysate (input) and GFP immunoprecipitation samples (elute) from wild-type U2OS cells (U2OS) and XPF KO U2OS cells expressing GFP-tagged wild-type XPF (XPF–GFP) or XPF carrying the amino acid mutations R689S or S786F. Samples were analyzed with antibodies against GFP, ERCC1, SLX4, FANCD2, RPA70 and H2B (as loading control). Cells were mock treated (−) or incubated with 10 µg/ml MMC for 1 h before lysis. In the FANCD2 immunoblot, the asterisk indicates aspecific staining. In b error bars represent the SEM
In an effort to understand why XPF–R689S and XPF–S786F are inefficiently recruited, we tested whether these XPF mutants showed reduced interaction with known or hypothesized XPF-interacting proteins. To this end, we performed GFP immunoprecipitation experiments on benzonase-treated lysate (i.e., to avoid isolation of DNA-mediated interactions/associations) from cells expressing GFP-tagged wild-type, R689S or S786F mutant XPF that was either mock treated or exposed to MMC. Importantly, we found that the mutant XPF proteins interacted normally with ERCC1 (Fig. 5c), which is in line with previous observations that these mutants stabilize, and thus bind ERCC1 [8, 37, 55]. Moreover, treatment with the crosslinking agent MMC did not alter the ratio of the interaction between XPF and ERCC1. It is currently unclear through which protein interactions ERCC1–XPF is recruited to ICLs, except that this depends on binding to its partner SLX4 [2, 25, 26], which was suggested to interact with mono-ubiquitylated FANCD2 [56]. Therefore, we tested whether these interactions were affected due to the XPF mutations. Strikingly, we did not observe any change in the interaction of XPF with SLX4, neither upon MMC exposure nor in the presence of the two XPF mutations. Moreover, we were unable to co-immunoprecipitate FANCD2 with XPF, suggesting that these proteins do not interact directly. Because in vitro ERCC1–XPF activity in ICLR is stimulated by RPA [27, 57], we also tested whether RPA directly interacts with wild-type or R689S or S786F mutant XPF. However, GFP-tagged XPF was unable to co-immunoprecipitate RPA70, the largest subunit of the heterotrimeric RPA complex, suggesting that stimulation of XPF activity by RPA is not via a direct physical interaction between the proteins. In summary, it remains unclear how exactly ERCC1–XPF, in complex with SLX4, is recruited to ICLs and how specific mutations in XPF could affect this. We observed that, even in the presence of both FANCD2 and XPA, recruitment of XPF–S786F and XPF–R689S is impaired. It was previously proposed that R689S and S786F affect the positioning of the XPF nuclease domain around the ICL rather than the actual catalytic activity itself [55, 58]. It is, therefore, possible that this incorrect positioning impairs a stable association of ERCC1–XPF with DNA at the lesion site, leading to inefficient visual recruitment at ICL sites as observed in our immunofluorescence experiments.
Discussion
In this study, we show that in human cells, ERCC1–XPF is involved in the repair of the multiple types of lesions induced by psoralen–DNA crosslinking, as part of the NER machinery via XPA, and as part of replication-dependent ICLR via FANCD2. Our results highlight and help to better understand the multifaceted activity and regulation of ERCC1–XPF in DNA repair, which is essential for a better comprehension of how the multiple DNA repair pathways in which ERCC1–XPF is involved tightly collaborate to protect against cancer and promote health. Moreover, our data encourage to seriously consider the activation of multiple DNA repair pathways when evaluating the (side) effects of using DNA crosslinking agents as therapeutic drugs.
It would be interesting to further investigate the kinetics of ERCC1–XPF recruitment to ICLs, to determine how this changes over time in accord with unhooking and repair of the lesions. Moreover, considering the different mechanisms involved in replication-dependent and replication-independent ICLR and the different activities that NER might have in both pathways, it would be useful to study the roles of ERCC1–XPF in these two different mechanisms by investigating how the complex is recruited to ICLs specifically in S-phase and non-S-phase cells. Additionally, the interaction and interdependency of ERCC1–XPF with other ICLR, NER and HR factors should be determined in a cell cycle phase-specific manner, to precisely elucidate the multistep processes that lead to ICL removal. Possibly, ICLR factors that we did not investigate or unknown factors influence the recruitment of ERCC1–XPF and could help explain why the recruitment of XPF–R689S and XPF–S786F to ICLs is inefficient.
References
Sijbers AM, De Laat WL, Ariza RR et al (1996) Xeroderma pigmentosum group F caused by a defect in a structure-specific DNA repair endonuclease. Cell 86:811–822. https://doi.org/10.1016/S0092-8674(00)80155-5
Klein Douwel D, Boonen RACM, Long DT et al (2014) XPF–ERCC1 acts in unhooking DNA interstrand crosslinks in cooperation with FANCD2 and FANCP/SLX4. Mol Cell 54:460–471. https://doi.org/10.1016/j.molcel.2014.03.015
Wang AT, Sengerová B, Cattell E et al (2011) Human SNM1a and XPF–ERCC1 collaborate to initiate DNA interstrand cross-link repair. Genes Dev 25:1859–1870. https://doi.org/10.1101/gad.15699211
Bhagwat N, Olsen AL, Wang AT et al (2009) XPF–ERCC1 participates in the Fanconi anemia pathway of cross-link repair. Mol Cell Biol 29:6427–6437. https://doi.org/10.1128/MCB.00086-09
Ahmad A, Robinson AR, Duensing A et al (2008) ERCC1–XPF endonuclease facilitates DNA double-strand break repair. Mol Cell Biol 28:5082–5092. https://doi.org/10.1128/MCB.00293-08
Muñoz P, Blanco R, Flores JM, Blasco MA (2005) XPF nuclease-dependent telomere loss and increased DNA damage in mice overexpressing TRF2 result in premature aging and cancer. Nat Genet 37:1063–1071. https://doi.org/10.1038/ng1633
Kashiyama K, Nakazawa Y, Pilz DT et al (2013) Malfunction of nuclease ERCC1–XPF results in diverse clinical manifestations and causes Cockayne syndrome, xeroderma pigmentosum, and Fanconi anemia. Am J Hum Genet 92:807–819. https://doi.org/10.1016/j.ajhg.2013.04.007
Bogliolo M, Schuster B, Stoepker C et al (2013) Mutations in ERCC4, encoding the DNA-repair endonuclease XPF, cause Fanconi anemia. Am J Hum Genet 92:800–806. https://doi.org/10.1016/j.ajhg.2013.04.002
Niedernhofer LJ, Garinis GA, Raams A et al (2006) A new progeroid syndrome reveals that genotoxic stress suppresses the somatotroph axis. Nature 444:1038–1043. https://doi.org/10.1038/nature05456
Jaspers NGJ, Raams A, Silengo MC et al (2007) First reported patient with human ERCC1 deficiency has cerebro-oculo-facio-skeletal syndrome with a mild defect in nucleotide excision repair and severe developmental failure. Am J Hum Genet 80:457–466. https://doi.org/10.1086/512486
Marteijn JA, Lans H, Vermeulen W, Hoeijmakers JHJ (2014) Understanding nucleotide excision repair and its roles in cancer and ageing. Nat Rev Mol Cell Biol 15:465–481. https://doi.org/10.1038/nrm3822
Schärer OD (2013) Nucleotide excision repair in eukaryotes. Cold Spring Harb Perspect Biol 5:a012609–a012609. https://doi.org/10.1101/cshperspect.a012609
Sugasawa K, Akagi J-I, Nishi R et al (2009) Two-step recognition of DNA damage for mammalian nucleotide excision repair: directional binding of the XPC complex and DNA strand scanning. Mol Cell 36:642–653. https://doi.org/10.1016/j.molcel.2009.09.035
Staresincic L, Fagbemi AF, Enzlin JH et al (2009) Coordination of dual incision and repair synthesis in human nucleotide excision repair. EMBO J 28:1111–1120. https://doi.org/10.1038/emboj.2009.49
Tsodikov OV, Ivanov D, Orelli B et al (2007) Structural basis for the recruitment of ERCC1–XPF to nucleotide excision repair complexes by XPA. EMBO J 26:4768–4776. https://doi.org/10.1038/sj.emboj.7601894
Orelli B, McClendon TB, Tsodikov OV et al (2010) The XPA-binding domain of ERCC1 is required for nucleotide excision repair but not other DNA repair pathways. J Biol Chem 285:3705–3712. https://doi.org/10.1074/jbc.M109.067538
Volker M, Moné MJ, Karmakar P et al (2001) Sequential assembly of the nucleotide excision repair factors in vivo. Mol Cell 8:213–224. https://doi.org/10.1016/S1097-2765(01)00281-7
Li L, Elledge SJ, Peterson CA et al (1994) Specific association between the human DNA repair proteins XPA and ERCC1. Proc Natl Acad Sci USA 91:5012–5016
Park CH, Sancar A (1994) Formation of a ternary complex by human XPA, ERCC1, and ERCC4(XPF) excision repair proteins. Proc Natl Acad Sci USA 91:5017–5021. https://doi.org/10.1073/pnas.91.11.5017
Enzlin JH, Schärer OD (2002) The active site of the DNA repair endonuclease XPF–ERCC1 forms a highly conserved nuclease motif. EMBO J 21:2045–2053. https://doi.org/10.1093/emboj/21.8.2045
Pontel LB, Rosado IV, Burgos-Barragan G et al (2015) Endogenous formaldehyde is a hematopoietic stem cell genotoxin and metabolic carcinogen. Mol Cell 60:177–188. https://doi.org/10.1016/j.molcel.2015.08.020
Clauson C, Scharer OD, Niedernhofer L (2013) Advances in understanding the complex mechanisms of DNA interstrand cross-link repair. Cold Spring Harb Perspect Biol 5:a012732–a012732. https://doi.org/10.1101/cshperspect.a012732
Smogorzewska A, Matsuoka S, Vinciguerra P et al (2007) Identification of the FANCI protein, a monoubiquitinated FANCD2 paralog required for DNA repair. Cell 129:289–301. https://doi.org/10.1016/j.cell.2007.03.009
Knipscheer P, Räschle M, Smogorzewska A et al (2009) The fanconi anemia pathway promotes replication-dependent DNA interstrand cross-link repair. Science (80-) 326:1698–1701. https://doi.org/10.1126/science.1182372
Muñoz IM, Hain K, Déclais AC et al (2009) Coordination of structure-specific nucleases by human SLX4/BTBD12 is required for DNA repair. Mol Cell 35:116–127. https://doi.org/10.1016/j.molcel.2009.06.020
Hodskinson MRG, Silhan J, Crossan GP et al (2014) Mouse SLX4 is a tumor suppressor that stimulates the activity of the nuclease XPF–ERCC1 in DNA crosslink repair. Mol Cell 54:472–484. https://doi.org/10.1016/j.molcel.2014.03.014
Abdullah UB, McGouran JF, Brolih S et al (2017) RPA activates the XPF–ERCC1 endonuclease to initiate processing of DNA interstrand crosslinks. EMBO J 36:2047–2060. https://doi.org/10.15252/embj.201796664
Roy U, Schärer OD (2016) Involvement of translesion synthesis DNA polymerases in DNA interstrand crosslink repair. DNA Repair (Amst). 44:33–41
Long DT, Räschle M, Joukov V, Walter JC (2011) Mechanism of RAD51-dependent DNA interstrand cross-link repair. Science (80-) 333:84–87. https://doi.org/10.1126/science.1204258
Semlow DR, Zhang J, Budzowska M et al (2016) Replication-dependent unhooking of DNA interstrand cross-links by the NEIL3 glycosylase. Cell 167:498–511.e14. https://doi.org/10.1016/j.cell.2016.09.008
Wang X, Peterson CA, Zheng H et al (2001) Involvement of nucleotide excision repair in a recombination-independent and error-prone pathway of DNA interstrand cross-link repair. Mol Cell Biol 21:713–720. https://doi.org/10.1128/MCB.21.3.713-720.2001
Sarkar S, Davies AA, Ulrich HD, McHugh PJ (2006) DNA interstrand crosslink repair during G1 involves nucleotide excision repair and DNA polymerase zeta. EMBO J 25:1285–1294. https://doi.org/10.1038/sj.emboj.7600993
Enoiu M, Jiricny J, Schärer OD (2012) Repair of cisplatin-induced DNA interstrand crosslinks by a replication-independent pathway involving transcription-coupled repair and translesion synthesis. Nucleic Acids Res 40:8953–8964. https://doi.org/10.1093/nar/gks670
Zheng H, Wang X, Warren AJ et al (2003) Nucleotide excision repair- and polymerase -mediated error-prone removal of mitomycin C interstrand cross-links. Mol Cell Biol 23:754–761. https://doi.org/10.1128/MCB.23.2.754-761.2003
Muniandy PA, Thapa D, Thazhathveetil AK et al (2009) Repair of laser-localized DNA interstrand cross-links in G1 phase mammalian cells. J Biol Chem 284:27908–27917. https://doi.org/10.1074/jbc.M109.029025
Ahmad A, Enzlin JH, Bhagwat NR et al (2010) Mislocalization of XPF–ERCC1 nuclease contributes to reduced DNA repair in XP-F patients. PLoS Genet 6:e1000871. https://doi.org/10.1371/journal.pgen.1000871
Sabatella M, Theil AF, Ribeiro-Silva C et al (2018) Repair protein persistence at DNA lesions characterizes XPF defect with Cockayne syndrome features. Nucleic Acids Res 46:9563–9577. https://doi.org/10.1093/nar/gky774
Van Cuijk L, Van Belle GJ, Turkyilmaz Y et al (2015) SUMO and ubiquitin-dependent XPC exchange drives nucleotide excision repair. Nat Commun 6:7499. https://doi.org/10.1038/ncomms8499
Sanjana NE, Shalem O, Zhang F (2014) Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods 11:783–784
Brinkman EK, Chen T, Amendola M, Van Steensel B (2014) Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res 42:e168. https://doi.org/10.1093/nar/gku936
Aydin ÖZ, Marteijn JA, Ribeiro-Silva C et al (2014) Human ISWI complexes are targeted by SMARCA5 ATPase and SLIDE domains to help resolve lesion-stalled transcription. Nucleic Acids Res 42:8473–8485. https://doi.org/10.1093/nar/gku565
Pines A, Dijk M, Makowski M et al (2018) TRiC controls transcription resumption after UV damage by regulating Cockayne syndrome protein A. Nat Commun 9:1040. https://doi.org/10.1038/s41467-018-03484-6
Lachaud C, Castor D, Hain K et al (2014) Distinct functional roles for the two SLX4 ubiquitin-binding UBZ domains mutated in Fanconi anemia. J Cell Sci 127:2811–2817. https://doi.org/10.1242/jcs.146167
Slyskova J, Sabatella M, Ribeiro-Silva C et al (2018) Base and nucleotide excision repair facilitate resolution of platinum drugs-induced transcription blockage. Nucleic Acids Res 46:9537–9549. https://doi.org/10.1093/nar/gky764
Sugitani N, Sivley RM, Perry KE et al (2016) XPA: a key scaffold for human nucleotide excision repair. DNA Repair (Amst) 44:123–135. https://doi.org/10.1016/j.dnarep.2016.05.018
Thazhathveetil AK, Liu ST, Indig FE, Seidman MM (2007) Psoralen conjugates for visualization of genomic interstrand cross-links localized by laser photoactivation. Bioconjugate Chem 18:431–437. https://doi.org/10.1021/bc060309t
Tessman JW, Isaacs ST, Hearst JE (1985) Photochemistry of the furan-side 8-methoxypsoralen-thymidine monoadduct inside the DNA helix. Conversion to diadduct and to pyrone-side monoadduct. Biochemistry 24:1669–1676. https://doi.org/10.1021/bi00328a015
Sage E, Moustacchi E (1987) Sequence context effects on 8-methoxypsoralen photobinding to defined DNA fragments. Biochemistry 26:3307–3314. https://doi.org/10.1021/bi00386a010
Hearst JE, Isaacs ST, Kanne D et al (1984) The reaction of the psoralens with deoxyribonucleic acid. Q Rev Biophys 17:1–44
Cao H, Hearst JE, Corash L, Wang Y (2008) LC–MS/MS for the detection of DNA interstrand cross-links formed by 8-methoxypsoralen and UVA irradiation in human cells. Anal Chem 80:2932–2938
Lai C, Cao H, Hearst JE et al (2008) Quantitative analysis of DNA interstrand cross-links and monoadducts formed in human cells induced by psoralens and UVA irradiation. Anal Chem. https://doi.org/10.1021/ac801520m
Wood RD, Robins P, Lindahl T (1988) Complementation of the xeroderma pigmentosum DNA repair defect in cell-free extracts. Cell 53:97–106. https://doi.org/10.1016/0092-8674(88)90491-6
Kaye J, Smith CA, Hanawalt PC (1980) DNA repair in human cells containing photoadducts of 8-methoxypsoralen or angelicin. Cancer Res 40:696–702
Osorio A, Bogliolo M, Fernández V et al (2013) Evaluation of rare variants in the New Fanconi Anemia Gene ERCC4 (FANCQ) as familial breast/ovarian cancer susceptibility alleles. Hum Mutat 34:1615–1618. https://doi.org/10.1002/humu.22438
Klein Douwel D, Hoogenboom WS, Boonen RA, Knipscheer P (2017) Recruitment and positioning determine the specific role of the XPF–ERCC1 endonuclease in interstrand crosslink repair. EMBO J 36:2034–2046. https://doi.org/10.15252/embj.201695223
Yamamoto KN, Kobayashi S, Tsuda M et al (2011) Involvement of SLX4 in interstrand cross-link repair is regulated by the Fanconi anemia pathway. Proc Natl Acad Sci 108:6492–6496. https://doi.org/10.1073/pnas.1018487108
Mu D, Bessho T, Nechev LV et al (2000) DNA interstrand cross-links induce futile repair synthesis in mammalian cell extracts. Mol Cell Biol 20:2446–2454. https://doi.org/10.1128/MCB.20.7.2446-2454.2000
Su Y, Orelli B, Madireddy A et al (2012) Multiple DNA binding domains mediate the function of the ERCC1–XPF protein in nucleotide excision repair. J Biol Chem 287:21846–21855. https://doi.org/10.1074/jbc.M111.337899
Acknowledgements
The authors would like to thank Dr. Christophe Lachaud and Dr. John Rouse for help with setting up the technique to locally induce psoralen–DNA crosslinks, the Erasmus Proteomics Center for mass spectrometry and the Erasmus MC Optical Imaging Center for support with microscopes. This work was supported by the Marie Curie Initial Training Network ‘aDDRess’ funded by the European Commission 7th Framework Programme (Grant 316390), a European Research Council Advanced Grant (Grant 340988-ERC-ID), an EMBO long-term fellowship (ALTF 663-2014) and the Dutch Cancer Society (KWF Grant 10506). This work is part of the Oncode Institute which is partly financed by the Dutch Cancer Society.
Author information
Authors and Affiliations
Contributions
MS, AP and HL performed all experiments and analyzed data. JS contributed reagents. MS, HL and WV designed experiments and wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Sabatella, M., Pines, A., Slyskova, J. et al. ERCC1–XPF targeting to psoralen–DNA crosslinks depends on XPA and FANCD2. Cell. Mol. Life Sci. 77, 2005–2016 (2020). https://doi.org/10.1007/s00018-019-03264-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-019-03264-5