Abstract
Gap genes are involved in segment determination during the early development of the fruit fly Drosophila melanogaster as well as in other insects. This review attempts to synthesize the current knowledge of the gap gene network through a comprehensive survey of the experimental literature. I focus on genetic and molecular evidence, which provides us with an almost-complete picture of the regulatory interactions responsible for trunk gap gene expression. I discuss the regulatory mechanisms involved, and highlight the remaining ambiguities and gaps in the evidence. This is followed by a brief discussion of molecular regulatory mechanisms for transcriptional regulation, as well as precision and size-regulation provided by the system. Finally, I discuss evidence on the evolution of gap gene expression from species other than Drosophila. My survey concludes that studies of the gap gene system continue to reveal interesting and important new insights into the role of gene regulatory networks in development and evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The gap gene network of the fruit fly Drosophila melanogaster is one of the most thoroughly studied developmental gene regulatory networks. There are hundreds of publications in the literature describing genetic and molecular analyses of gap genes, their expression, regulation, and their regulatory effect on downstream targets, and yet, we are far from a complete understanding of its pattern-forming and regulatory capacities, not to mention its evolutionary history.
Gap genes have attracted the interest of developmental, evolutionary, and systems biologists for three main reasons: First, they play a key role in patterning the early embryo. The gap gene system implements the most upstream regulatory layer of the segmentation gene network, which determines both the position and the identities of body segments [1, 2]. It solves a fundamental problem of embryonic patterning: how to establish discrete territories of gene expression based on regulatory input from a long-range protein gradient [3–5]. Such gradient-based patterning occurs in most multi-cellular organisms studied so far (see [6–10] for recent reviews).
Second, gap genes played a crucial role during the evolution of segment determination. While most segmented animals—arthropods, annelids, and vertebrates—add segments to their body sequentially during growth, some higher insects have evolved a mode of segment determination in which segments form by simultaneous subdivision of the embryo. This is called the long-germband mode of segment determination. It appears to have evolved many times independently [11, 12], a process which probably involved the recruitment of gap genes into the segmentation gene network [13, 14].
Finally, the gap gene network has become one of the few examples of a developmental gene network, which can be studied using data-driven mathematical modeling. Such modeling studies have allowed us to reconstruct the regulatory structure of the gap gene network in silico, to assign particular patterning functions to each regulatory interaction, and to study regulatory feedback based on gap gene cross-regulation in the intact, wild-type system [15–24]. These analyses establish the gap gene network as a model system for the quantitative study of the developmental and evolutionary dynamics of pattern-forming processes.
In this review, I summarize what is known (and what is not) about regulation of gap genes. The information presented here is predominantly based on genetic and molecular evidence. In addition, I have included evidence from selected mathematical models, if (and only if) those models closely adhere to experimental data, and provide specific biological predictions or insights into gap gene regulation. A comprehensive historical review of modeling the Drosophila blastoderm is provided elsewhere [25].
The review is structured as follows: After a brief introduction to segmentation, maternal inputs to the gap genes, and the dominant (but inaccurate) conceptual framework traditionally used to interpret pattern formation in the early Drosophila embryo, I describe phenotypes, expression, and regulation of gap genes in separate sections. This is followed by brief sections summarizing the molecular nature of gap gene regulation, the issue of precision of gap gene expression, as well as gap gene evolution. Since this is a work of reference, not all of these sections need to be read in sequence. Each section is designed to be understandable without the others. Readers interested in specific aspects of gap gene regulation are encouraged to skip ahead to those parts of the review that are relevant to them.
Segmentation genes and segment determination
The gap gene network is involved in segment determination during early embryogenesis. As mentioned above, body segments can be determined in two ways: either they are formed sequentially, by adding them to the posterior end of a growing embryo (short-germband development), or (more or less) simultaneously, by subdividing an embryo into equally sized sub-domains (long-germband development). While vertebrates, annelids, and most arthropods use the former mode of segmentation, insects show both types (including many intermediates between the two extremes; see [11, 12, 26, 27] for review).
Early insect development typically proceeds through syncytial cleavage and blastoderm stages (Fig. 1a) [11, 28, 29]. During these early stages, nuclei divide rapidly and almost simultaneously without becoming separated by cell membranes. Each nucleus is surrounded by microtubule-rich cytoplasm, with which it forms a unit called an energid. Towards the end of the cleavage stage, most energids start to migrate. After a number of cleavage divisions (nine in Drosophila), they arrive at the surface of the embryo to form the syncytial blastoderm, a peripheral layer of nuclei lying within a zone of yolk-free periplasm. At this stage, embryos are most conveniently classified by the number of nuclear divisions: cleavage cycle n, corresponds to the period between mitosis n − 1 and mitosis n [30]. These cycles become increasingly longer during the blastoderm stage (from about 10 to 50 min between cycles 10 and 14A in Drosophila; [30, 31]). The embryo becomes cellularized through invagination of cell membranes between nuclei. Subsequently gastrulation starts, during which the three germ layers (ecto-, endo-, and mesoderm) are formed. This is followed by extension and retraction of the germband. Tissue rearrangements occur mainly during and after gastrulation.
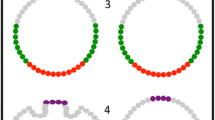
Segment determination in Drosophila. a The first 3 h of development of Drosophila melanogaster. Numbers indicate cleavage cycle number, where cycle n covers the time between mitosis n − 1 and mitosis n. The blastoderm stage lasts from 1 min into cycle 10 to the onset of gastrulation (grey background). The embryo remains syncytial (without membranes between nuclei) until cellularization occurs during cycle 14A. The cellular blastoderm stage is more or less instantaneous, since gastrulation begins immediately after cellularization is complete. Cycle 14B denotes the part of cycle 14, which occurs after the onset of gastrulation. Embryos are shown with the anterior pole to the top. b The regulatory hierarchy of the Drosophila segmentation gene network. Segment determination is based on a molecular pre-pattern established by the segmentation genes, which are active during the blastoderm stage. Different regulatory tiers of the network can be distinguished based on mutant phenotypes, epistatic interactions, and expression patterns. Maternal co-ordinate genes are expressed in broad gradients (Bcd protein distribution is shown as an example). They regulate the zygotic gap genes, expressed in broad overlapping domains (the central domain of Kr is shown). Gap genes and pair-rule genes together regulate pair-rule genes, which are expressed in 7–8 stripes (shown for Even-skipped (Eve) protein). Pair-rule genes in turn regulate segment-polarity genes whose expression in 14 stripes becomes established just before the onset of gastrulation (shown for en mRNA). These stripes constitute the segmentation pre-pattern and correspond to the positions of parasegmental boundaries later in development. Arrows indicate regulatory interactions between classes of segmentation genes. Circular arrows represent cross-regulation within a class. Embryo images are shown with anterior to the left, and dorsal up (see text for details). a is reproduced with permission from the Journal of Cell Science: http://jcs.biologists.org [30]. b Embryo images (Bcd, Kr, and Eve) are from the FlyEx database [164, 166]. The image of en is courtesy of Carlos E. Vanario-Alonso
Short- versus long-germband modes of development are reviewed in [11, 12]. In most short- and intermediate-germband insects, the blastoderm embryo occupies only a small fraction of the egg (the remainder consists of yolk and extra-embryonic tissue). A number of anterior segments become determined during the blastoderm stage, while posterior segments are added after gastrulation. In contrast, most long-germband embryos take up a large proportion of the egg, and segment determination occurs before the onset of gastrulation. No tissue growth is involved in this process. The morphological formation of segments occurs much later in development; segmental boundaries are clearly visible at the extended germband stage.
The first systematic molecular study of the process of segment determination was carried out in the fruit fly Drosophila melanogaster. Like all dipterans, Drosophila is a long-germband insect [12]. In vitro culture and transplantation experiments established that segment determination occurs at the blastoderm stage [32, 33], 1.5–3 h after egg laying (AEL) [30]. In the late 1970s, methods were developed to saturate the genome of Drosophila with mutations, and to efficiently select for segmentation phenotypes among the mutant progeny [34]. This led to the identification of several dozen genes involved in axis patterning and segmentation [35–37]. The resulting mutant phenotypes were easily classifiable into distinct groups: Mutations affecting the minor (dorso-ventral, D–V) embryonic axis rarely affected patterning along the major (antero-posterior, A–P) axis and vice versa. Zygotic mutants in A–P patterning could be further subdivided into those lacking entire regions of the embryo (gap), those missing every other segment (pair-rule), and those affecting polarity within segments (segment-polarity genes). Screens for maternal factors affecting segmentation uncovered an additional class involved in A–P patterning: the maternal co-ordinate genes [38]. These genes can be subdivided into anterior, posterior, and terminal maternal systems, depending on the regions of the embryo that are affected in the corresponding mutants.
In the decade after the initial screening efforts, segmentation genes were cloned and analyzed molecularly (reviewed in [1, 2]). They encode transcriptional or translational regulators, or proteins involved in signal transduction. Genetic analyses of their epistatic relationships revealed that these factors form a complex hierarchical network of regulatory interactions. The distinct groups of phenotypes correspond to distinct layers in the regulatory hierarchy of the network (Fig. 1b): maternal co-ordinate genes regulate gap genes; both of them jointly regulate pair-rule genes, which in turn regulate the initial expression of segment-polarity genes. In addition, all classes of segmentation genes show cross-regulation. In contrast, products of genes in the lower tiers of the network do not regulate genes in the layers above. Pair-rule genes, for example, do not regulate gap genes and so on.
At the same time, segmentation gene expression patterns were visualized by in situ hybridization and antibody staining. These studies revealed that genes in each layer of the segmentation gene network are expressed in similar patterns, which are clearly distinguishable from those of genes in other layers (Fig. 1b). The protein products of the maternal co-ordinate genes form long-range gradients along the A–P axis. Gap genes are expressed in broad, overlapping domains about 10–20 nuclei wide. The first periodic expression patterns occur at the level of the pair-rule genes, which are expressed in seven to eight stripes, each being about four nuclei wide. Segment-polarity genes show expression in 14 narrow stripes, which form a molecular pre-pattern involved in positioning the morphological segment boundaries later in development. This occurs through the formation of parasegment boundaries—tissue compartment boundaries between cells expressing distinct segment-polarity genes that no cells can cross—which are phase-shifted with regard to the morphological segmental boundaries [39–41]. At the same time, segment identity is established by the expression of homoeotic (Hox) genes during the late blastoderm stage [42]. Hox gene expression is regulated by maternal co-ordinate and gap genes.
Maternal systems, gradients, and the French Flag paradigm
Gap genes receive their initial regulatory inputs by three sub-groups of maternal co-ordinate genes. The anterior and posterior maternal systems are based on long-range gradients of maternal proteins along the A–P axis.
During oogenesis, the mRNA of bicoid (bcd) is localized to the anterior pole of the embryo by other components of the anterior system, such as the protein products of swallow (swa), exuperantia (exu), and staufen (stau) [43, 44]. After fertilization, bcd mRNA spreads further posterior, forming a gradient along the A–P axis [43, 45]. Bcd protein is thought to diffuse from its predominantly anterior source to form an exponential anterior-to-posterior gradient (Fig. 2a) [46–48]. Bcd has been shown to regulate zygotic target genes in a concentration-dependent manner [49–56]. In addition, it represses translation of the ubiquitous maternal caudal (cad) mRNA, establishing a posterior gradient of Cad protein [57–61]. This gradient spans the middle third of the embryo while Cad is present at uniformly high levels in more posterior regions (Fig. 2a).
Maternal gradients and French Flags. a–c Three maternal systems regulate the expression of gap genes: a The anterior system is based on the Bcd gradient, which regulates gap gene transcription in a concentration-dependent manner and also establishes the posterior gradient of Cad through translational repression. b The posterior system is based on the Nos gradient, whose only function is to repress the translation of maternal hb mRNA in the posterior region of the embryo to form an anterior Hb protein gradient. c The terminal system is based on Tor signaling from both terminal ends of the embryo, which induces the expression of the terminal gap genes tll and hkb at both poles of the embryo. Expression profiles are based on integrated data from the FlyEx database [164, 166], except for Nos, which is illustrated by a mirrored Bcd gradient due to the absence of quantitative Nos expression data. d Wolpert’s French Flag model: A morphogen is produced at a source (shown in green), diffuses through the tissue (without protein degradation) and is degraded at a sink (pink), at the other end of the tissue. Specific concentration thresholds in the resulting linear gradient (T 1, T 2) are detected by cells (or nuclei) in the tissue, which switch on alternative target genes (represented by blue, white, and red), which in turn lead to distinct differentiation pathways in each region of the embryo. In this model, development is seen as a two-step process: First, positional information is implemented by the morphogen gradient (step 1). Subsequently, cells in the tissue passively interpret this information (step 2). Concentration thresholds in the gradient correspond exactly to borders of downstream expression territories. e A revised French Flag, incorporating target domain shifts and increasing precision over time. New evidence shows that maternal gradients are not sufficient to determine precise downstream boundary positions on their own. Instead, cross-regulation among target genes leads to (a) shifts in boundary positions over time and (b) an observed increase in the precision with which boundaries are placed. In this model, there is no longer a precise correspondence between concentration thresholds in the gradient and the final position of target domain boundaries
The posterior system works in a similar way: mRNA of the main posterior determinant nanos (nos) diffuses and becomes trapped at the posterior pole of the embryo [62]. Only its posteriorly localized pool is actively translated [63–65]. This is thought to establish a posterior-to-anterior gradient of Nos protein (Fig. 2b). In contrast to Bcd, Nos does not function as a transcriptional regulator (and thus does not affect gap genes directly), but instead acts as a translational repressor of the uniformly distributed maternal hunchback (hb) mRNA establishing an anterior Hb protein gradient (Fig. 2b) [66–72]. Translational regulation of maternal hb is likely to be Nos’ only essential contribution to segmentation gene expression, since embryos from mothers mutant for both nos and hb are viable [68–70].
The maternal gradients of Bcd and Hb specify the position of gap domain boundaries in a concentration-dependent manner [50–52, 55, 56, 73]. In 1968, Lewis Wolpert had suggested a model—using an analogy to the French Flag—of how such positional specification can be achieved (Fig. 2d) [3, 4]. He proposed that there are specific concentration thresholds in the gradient, which can be detected by cells in the tissue. The cells thus ‘interpret’ the gradient by initiating expression of different sets of target genes, depending on whether they experience regulator concentrations above or below a given threshold. This provides a straightforward and testable hypothesis for a global patterning mechanism in which the maternal gradient imposes positional information onto its target tissue.
Wolpert repeatedly used the positioning of gap domain boundaries as an example of the French Flag mechanism [74–76]. However, other authors have criticized this proposition as not being robust, since it depends too strongly on precise measurement of gradient concentrations (see [77, 78], and the appendix of [74]). Even Wolpert himself has stressed the importance of local regulatory interactions [74, 79]. Alternative models were proposed, in which gradient-based patterning is complemented by cross-regulation among downstream targets [77, 78, 80, 81]. Current evidence indicates that such target gene cross-regulation is indeed essential for the patterning function and robustness of the gap gene network [5, 15, 16, 22, 23, 82].
Terminal gap genes and the terminal maternal system
In contrast to the long-range gradients described above, which are involved in patterning the segmented, central region of the embryo, the terminal system is based on localized signaling through the Torso (Tor) MAP-kinase cascade at both poles of the embryo (Fig. 2c; reviewed in [83]). Tor signaling acts predominantly through activation of head gap genes (discussed below) [55, 84] and the terminal gap genes tailless (tll) and huckebein (hkb) [85–87]. Activation of the latter is achieved through localized relief from constitutive repression [88–93] and depends on the strength of the Tor signal [94–98]. The expression domains of tll and hkb are missing in loss-of-function mutants of the terminal system [94, 96, 99] and are expanded centrally in gain-of-function alleles of Tor signaling [100]. Bcd and the D–V system play an important part in the regulation of the anterior tll and hkb domains [94, 101, 102]. In contrast, posterior expression of tll and hkb largely depends on the terminal system [94, 99, 103], with the notable exception of a subtle fine-tuning effect of the posterior system on the extent of tll and hkb de-repression [104]. These domains are not affected at all by any other gap genes [95, 99, 105], and therefore provide an independent, external input to the rest of the gap gene network. For this reason, their regulation will not be discussed further below.
Gap genes: phenotypes, gene structure, and protein products
This review mainly focuses on the gap genes hunchback (hb), Krüppel (Kr), knirps (kni), and giant (gt) involved in patterning of the segmented trunk (gnathal, thoracic, and abdominal) region of the embryo. Gap genes were initially defined based on their mutant phenotypes, which exhibit deletions in one or two contiguous regions of the embryo covering multiple segments [34]. Only hb has a maternal component. Embryos without zygotic hb lack the labial and all thoracic segments and show defects in the posterior abdomen [34, 36, 106, 107]. Mutants, which lack both maternal and zygotic hb, have a more severe phenotype: they have no gnathal and thoracic segments, and exhibit mirror duplications of anterior abdominal and enlargement of posterior abdominal segments [106, 108]. This phenotype can be rescued if a single copy of zygotic hb is supplied paternally [106]. Kr null mutants show deletion of thoracic and anterior abdominal segments as well as frequent mirror duplications in the abdomen [34, 35, 109–111]. kni mutant embryos show defects in the head plus all but the most posterior abdominal segments [34, 36, 112–115]. Finally, strong gt alleles show defects in the head and the fifth to seventh abdominal segments [37, 116–118]. All of these phenotypes only appear about 10–20 min after the onset of gastrulation.
Unlike the clustered Hox genes, gap genes are dispersed throughout the genome (Table 1). Each trunk gap gene is located on a different chromosome arm [34–37, 66, 118–120] (only tll and hkb map to the same arm as hb [95, 121–123]). Like other genes that are expressed during the blastoderm stage, gap genes are all unusually compact: zygotic transcripts are short (Table 1; about 1–3 kilobases (kb), with at most one or two short introns [66, 95, 101, 120, 123–125]). Such compact gene structure seems to be required for gap gene expression during the extremely short mitotic cycles of the early blastoderm stage, as the much longer maternal and late zygotic transcript of hb (about 6 kb; Table 1) and the knirps-related (knrl) gene (a duplication of kni, which seems to be functionally redundant but contains a much larger intron) only become expressed during the extended interphase of cleavage cycle 14A [114, 126].
All Drosophila gap genes encode transcription factors (Table 1): Hb, Kr, Kni, Tll, and Hkb contain zinc-finger DNA-binding domains [66, 67, 95, 120, 123, 124, 127, 128]. Kni and Tll belong to the steroid receptor superfamily [120, 123]. Gt belongs to the basic leucine zipper (bZip) family [125]. All gap proteins show predominantly nuclear sub-cellular localization [61, 67, 129–132]. The transcription factors encoded by gap genes usually act as transcriptional repressors (see, for example, [100, 133–147]), although there is evidence for activation in some specific cases [102, 134, 135, 148, 149].
Apart from being involved in segment determination, most gap genes have additional roles later in development: hb, Kr, tll, and hkb are involved in neurogenesis [105, 132, 150–155]. Kr is required for the development of the malpighian tubules and trachea [111], larval photoreceptor organs [156], muscles [157], and extraembryonic tissue [130]. kni is involved in tracheal [158], gut [159, 160], and wing-vein development [161, 162]. hkb is required for gut development [87, 95, 102].
Gap gene expression and regulation
In the blastoderm embryo of Drosophila melanogaster, the trunk gap genes hb, Kr, kni, and gt are expressed and regulated in two clearly distinguishable phases (Fig. 3) [19]: Early gap gene expression is established through strictly feed-forward regulation by maternal gradients, and each gap gene is regulated independently. At this stage, expression is highly variable; gap domain boundaries sharpen, but their positions do not shift over time [19]. During cleavage cycle 13—as gap proteins start accumulating in significant amounts—gap–gap cross-interactions begin to introduce feedback regulation to the system. These mostly repressive cross-regulatory interactions are involved in sharpening and maintaining gap domain boundaries [163], but also lead to dynamic shifts in the position of expression borders during cycle 14A [15, 16, 22–24, 61]. The regulatory logic of the system becomes much more complex at this stage as gap gene expression patterns become dependent on each other. After providing a brief description of gap gene expression patterns, I will analyze each of these two separate regulatory stages in detail.
Early versus late gap gene regulation. Gap gene regulation can be divided into two distinct phases: early regulation of gap mRNA domains is based on maternal gradients only, while late regulation of protein domains involves gap–gap cross-regulatory interactions. The position of gap domains along the major, or antero-posterior (A–P) axis of the embryo is shown schematically as colored boxes. Only the trunk region of the embryo (approx. 35–95% A–P position) is included in the diagram. Anterior is to the left, posterior to the right. Background color represents activating inputs by Bcd and Cad. Top panel: arrowheads represent activating; T-bars represent repressive inputs responsible for setting specific domain boundaries. Bottom panel: arrows and T-bars represent activating and repressive gap–gap cross-regulation, respectively. Circular arrows represent auto-activation. The thickness of the T-bars corresponds to repressive strength. Question marks indicate missing or ambiguous evidence, or other open questions regarding gap gene regulation (see text for details)
Expression patterns
Quantitative mRNA expression patterns for Kr, kni, and gt at the early blastoderm stage have been published in Jaeger et al. [19], early hb mRNA expression has been analyzed quantitatively in [56], while protein expression patterns during later cycles are described in detail in [61]. A comprehensive data set of gap protein expression patterns—at high temporal and spatial resolution—is available online from the FlyEx database (http://urchin.spbcas.ru/FlyEx) [164–166]. Additional mRNA expression patterns at lower temporal resolution are available from the Berkeley Drosophila Genome Project (BDGP) in situ database (http://www.fruitfly.org/cgi-bin/ex/insitu.pl) [167–169]. Moreover, the BDGP is developing a database of three-dimensional, quantified mRNA expression patterns in the early Drosophila embryo [170–172].
Transcription is initiated at slightly different times for each gap gene during the early blastoderm stage (Fig. 4). The earliest reported expression patterns are transient localized domains of kni and Kr, which appear during pro-, meta-, and anaphase of cleavage divisions 9 and 10 respectively (Fig. 4, inset) [19, 173]. These early domains vanish again during telo- and interphase, only to reappear during the subsequent mitosis. The function (if any) and regulation of these early domains is unknown.
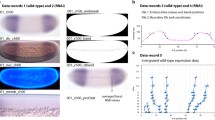
Early gap gene expression. mRNA distribution is visualized by fluorescent in situ hybridization for Kr, kni, and gt during early blastoderm stage (cycles 11–13). The inset shows transient early Kr expression during mitosis 11. Embryo images are from [19], shown with anterior to the left, dorsal up. Plots show individual one-dimensional expression profiles for each gene from the middle 10% along the dorso-ventral (D–V) axis at late cycle 13, illustrating the large embryo-to-embryo variability of the patterns at this stage. Relative mRNA concentration is plotted against position along the A–P axis (in %, where 0% is the anterior pole) (see [19, 165] for details on data quantification)
The earliest detectable expression patterns of gap genes during interphase are those of hb [56] and tll [94], which both appear during cleavage cycle 9. Some embryos initiate expression of gt during cycle 11, while most only show detectable gt expression during cycle 12 [19, 118, 131]. Kr can also be first detected during cycle 12 [19, 174]. The last gap gene to become expressed during interphase is kni. Some authors have reported its appearance during interphase of cycle 12 [113, 175] while others have only been able to detect it during mitosis 12 and early cycle 13 [19].
What all early gap mRNA domains have in common is that their initial expression is weak and appears as a dotted nuclear signal (Fig. 4) [19]. During early cycle 13, levels of transcription increase dramatically, and nuclear export leads to increasing accumulation of gap gene mRNAs in the cytoplasm, where they are translated [19]. Moreover, early expression of Kr, kni, and gt is highly variable, as positions of early gap domain boundaries at the mRNA level differ by as much as 10–15% egg length between embryos of the same age (Fig. 4, bottom row) [19]. In contrast, early expression of hb appears to be surprisingly precise at cleavage cycle 11 already [56] (see also below).
Zygotic protein products of gap genes appear later than their respective mRNA domains. Kr and Gt proteins become detectable during cycle 12, while Kni only appears during cycle 13 [61, 176]. The accumulation of zygotic Hb protein is difficult to monitor, as it is chemically indistinguishable from maternal Hb. While the maternal Hb gradient gradually transforms into its zygotic expression pattern in the anterior half of the embryo during cleavage cycles 10–13 [61, 67], at least some maternal Hb protein persists until the onset of cellularization [177]. Terminal gap gene products Tll and Hkb have only been detected in early cycle 13 ([61]; J. Jaeger, unpublished). However, the much earlier appearance of tll mRNA suggests that they may already be present before that.
Gap gene expression during the late blastoderm stage is very dynamic (Fig. 5). After their initial establishment, gap domain borders sharpen [163] and those of Kr, kni, and gt in the posterior region of the embryo shift anteriorly during cleavage cycle 14A [15, 16, 22, 23, 61], while the posterior domain of hb only appears during early cycle 14A [66, 67, 126, 178]. Similarly, the dynamics of gap gene expression changes dramatically in the anterior of the embryo during this stage. The broad and relatively uniform anterior expression of hb refines into a stripe at the position of parasegment 4 (PS4) and more irregular and weaker expression further anterior [66, 67, 126, 178]. The anterior domain of gt splits into two stripe-like domains and an additional dorsal patch of expression appears anterior to these [118, 131, 176]. The ventral, anterior domain of kni—which is not involved in segment determination [115])—expands dorsally at its posterior margin to form an L-shaped pattern during mid-cycle 14A [113, 175]. Finally, anterior and posterior domains of Kr appear, which also do not play any role in segmentation [130, 150, 179, 180].
Late gap gene expression showing dynamic shifts in gap domain positions. Protein expression patterns are shown for Hb, Kr, Kni, and Hb at eight time classes during cycle 14A (T1–T8) [61]. Plots show integrated one-dimensional expression patterns from the middle 10% along the D–V axis over time, illustrating the anterior shift in boundary position for all expression domains posterior of the central Kr domain. Relative protein concentration is plotted against position along the A–P axis (in %, where 0% is the anterior pole). Embryo images and integrated data for plots are from the FlyEx database [164, 166], shown with anterior to the left, dorsal up (see [165] for details on data quantification)
Early regulation of gap genes by maternal gradients
Since gap gene mRNAs appear before gap proteins (and do not play any role in gap–gap cross-regulation) initial regulation of localized expression must depend exclusively on maternal gradients. While gene expression in head gap domains is activated by the terminal system (see below) [55, 84], the only maternal gradients that are known to directly regulate gap gene transcription in the trunk region are the activator gradients of Bcd and Cad as well as the repressor gradient of Hb (Fig. 2a, b) [19]. Early gap gene regulation depends on a delicate balance between activation and repression (summarized in Fig. 3, top panel).
Cad activates the posterior gt domain, which is absent or very strongly reduced in embryos mutant for maternal and zygotic cad [103, 181], and—in concert with Bcd—the abdominal domain of kni, which is absent in embryos lacking both maternal Bcd and Cad [181, 182]. Expression of hb and Kr is not affected in cad mutants [103] or embryos over-expressing cad [183].
Bcd activates the anterior domains of gt and hb, which are absent in embryos from bcd mutant mothers [67, 131, 176, 181]. In the case of hb, activation occurs through Bcd binding sites in the hb regulatory region [50, 52]. The evidence is far more complicated for activation of Kr by Bcd: Early studies indicated that Kr is activated by ubiquitous maternal transcription factors [184], while Bcd was thought to repress Kr since the central Kr domain expands anteriorly in embryos from bcd mutant mothers [108, 185, 186]. However, exactly the same expansion can be seen in gt; hb double mutants. This indicates that the effect is indirect [187], as both anterior gt and hb domains are absent in a bcd mutant background [67, 176]. Later molecular studies identified a regulatory element of Kr containing multiple Bcd binding sites whose expression depends on the presence of Bcd [188, 189]. This suggests activation of Kr by Bcd. The fact that Kr expression is still present in embryos without Bcd can be explained either by an activating effect of Hb at low concentrations [108, 190, 191] or redundant activation of Kr by Cad [15] (see also below).
Maternal Hb is required for robust early expression of hb [56]. In addition, it represses Kr, kni, and the posterior domain of gt: It binds to the regulatory region of Kr [189] and Kr expression expands anteriorly in hb mutants [163, 179, 185]. The abdominal domain of kni expands anteriorly in zygotic mutants of hb; expression in its expanded domain is much stronger in embryos lacking both maternal and zygotic Hb [108]. Both abdominal kni and posterior gt domains are lacking in embryos with Hb present in the posterior region of the embryo [113, 120, 131, 175, 176, 187]. In contrast, Hb does not seem to have an effect on the anterior domain of gt. This could either be because the anterior and the posterior domains of gt are regulated by different enhancer elements, implementing different regulatory mechanisms [192–194], or because Bcd and Cad modulate the effect of Hb on gt where they are present [19] (see also below). A similar dependence on third factors has been demonstrated for the effect of Hb on the regulation of stripes 2 and 3 of the pair-rule gene even-skipped (eve), where Hb activates expression in stripe 2 due to modulation by Bcd, while it represses stripe 3 on its own [195–198].
Regulation of target genes by Bcd and Hb is concentration-dependent [50, 51, 55, 56, 73, 190]. How is this achieved at the molecular level? Two alternative explanations have been provided: Activation of some Bcd target genes depends on the number and affinity of Bcd binding sites. Regulatory elements of the head gap gene orthodenticle (otd) (see below) and hb contain a mixture of both high- and low-affinity Bcd binding sites [52, 53], while the regulatory region of kni contains a tightly spaced array of six high-affinity sites [181]. However, a more comprehensive survey of Bcd target genes found no correlation between Bcd binding site number and affinity and the position of the target gene’s boundary along the A–P axis [194]. In this case, boundary position depends on the context of the Bcd binding sites, i.e., the presence of additional binding sites for third factors—such as Hb or Kr—in a regulatory element. Such context-dependence has also been found in an equivalent survey on Hb targets [73]. The importance of genomic context is further corroborated by the fact that many homo- and heterotypic combinations of binding sites are significantly enriched in regulatory regions of segmentation genes [199].
In contrast to the concentration-dependent effect of Bcd and Hb, Cad only activates gap genes in the posterior of the embryo, where its concentration is high and constant across space (Fig. 2a) [61]. Although Cad is required for the normal expression of these genes, there is no evidence that it is actively involved in positioning any early gap domains.
The evidence presented above strongly suggests that multiple gradients are required for the placement of most gap domains. This is further supported by the fact that domains of segmentation genes—and the fate map of the embryo in general—shift less in mutants with varying doses of bcd than expected if they would depend on Bcd alone [49]. It has been proposed that regulatory synergism between maternal Hb and Bcd could account for this effect [200, 201]. However, the exact molecular nature of this synergism remains unclear.
Alternatively, the reduced shifts in bcd dosage mutants could be explained by Bcd not reaching its steady state until late during the blastoderm stage [202, 203]. However, there is currently no evidence supporting this proposition, and it has been demonstrated that the gradient of nuclear Bcd protein remains stable throughout the relevant stages of development [48, 61].
Maternal gradients can position target gene expression boundaries in two different ways: Activator gradients induce boundaries with the same polarity as the gradient itself, while repressor gradients position counter-polar boundaries (Fig. 6). Accordingly, Bcd can only set posterior boundaries of gap domains, while repression by Hb is the only available mechanism for positioning anterior borders (see Fig. 3, top panel). For example, in the abdominal domain of kni, repression by Hb positions the anterior boundary [19, 108]. Bcd appears to be responsible for establishing the posterior boundary [181], although this border is only partially developed before repression by terminal gap genes leads to the full retraction of kni from the posterior pole of the embryo during cycle 13 [19, 61].
In light of this, there is a problem for positioning early gap domain boundaries in the central and anterior region of the embryo where the concentration of Hb changes very rapidly during cycles 10–13 [19]. It is unclear how a balance between Bcd activation and Hb repression can be achieved in this region to position, for example, both boundaries of the central Kr domain. Despite the rapidly changing concentration of Hb, early boundaries of Kr remain at a constant position during cycles 12 and 13 (Fig. 4). Mathematical models of early gap gene regulation corroborate the fact that Hb repression is insufficient for placing these borders [19].
To avoid these problems, it has been suggested that Kr is repressed at high and activated at low concentrations of Hb (see Fig. 3, bottom panel). Such a concentration-dependent switch between activation and repression has been observed in assays with cell lines carrying reporter constructs that monitor the regulatory effect of transcription factors such as Kr [134] or Engrailed (En) [135]. Cells were exposed to varying levels of regulator concentration. However, it is difficult to establish whether such an effect occurs at physiologically relevant regulator concentrations. Although mathematical models incorporating such a switch can lead to a gap-like (bell-shaped) target gene expression profile [204], these models still fail to reproduce the stability of Kr boundaries over time in the presence of a rapidly changing Hb repressor gradient [19].
In summary, the evidence presented above suggests that known maternal gradients are not sufficient to account for early gap gene regulation, and we may still be missing a maternal regulator required for the establishment of early gap domain boundaries [19].
Gap gene cross-regulation and gap domain shifts
As mentioned above, gap gene regulation can be subdivided into an early (maternal-only) and a late phase (including gap-gene cross-repression). Due to its complexity, it is useful to further subdivide the latter into five separate regulatory mechanisms (Fig. 7): (a) broad activation of gap genes by maternal gradients of Bcd and Cad. (b) gap gene auto-activation. (c) Strong mutual repression between gap genes that show complementary expression patterns (hb and kni; Kr and gt). (d) Weaker, asymmetric repression between overlapping gap genes (Hb on gt; Gt on kni; Kni on Kr; Kr on hb, and Hb on Kr) and (e) repression by the terminal gap genes tll and hkb in the pole regions of the embryo. In the following sub-sections, I will discuss each of these mechanisms in turn.
The five main regulatory mechanisms for late gap gene regulation: a Gap genes are activated by maternal Bcd and Cad in broad regions of the embryo. b Auto-activation leads to intensification and sharpening of domain boundaries in specific gap domains. c Strong cross-repression between gap genes with mutually exclusive expression domains leads to the basic staggered arrangement of gap domains (alternating cushions hypothesis). d Weaker cross-repression between gap genes with overlapping domains of expression leads to anterior shifts in boundary positions over time. e Repression by terminal gap genes establishes the posterior boundaries of several gap domains and excludes gap gene expression from the un-segmented terminal regions of the embryo. Horizontal axis, background color, gap domains, and regulatory links as in Fig. 3. Colored picture elements highlight those domains involved in or affected by a specific mechanism
Late activating contributions by Bcd and Cad
I have already described that activation by Bcd plays an important role in establishing early boundaries of gap gene domains, while activation by Cad does not contribute to positional specification. During cleavage cycle 14A, both of these activating contributions continue to occur, but not even Bcd is significantly involved in the placement of domain boundaries anymore [15, 16]. Instead, activation by Bcd and Cad contributes to the maintenance of gap gene expression (Fig. 7a), until about 10–15 min before gastrulation when the Bcd gradient starts to rapidly decay [46, 61]. At the same time, Cad disappears from the abdominal region due to transcriptional repression by Hb [103, 182, 205], and its expression domain refines into a narrow posterior stripe [61], which is regulated by gap and pair-rule genes [103, 205]. This does not contradict the general rule that maternal co-ordinate genes are not regulated by gap and pair-rule genes. Late zygotic expression of cad plays a very different role than that of maternal Cad: it is involved in determining the identity of the posterior-most abdominal segment in a homoeotic-gene-like fashion [206]. The late decrease in overall maternal activation is reflected by decreasing levels of gap proteins right before the onset of gastrulation [16, 61].
Auto-regulation
Early theoretical analyses of segment determination postulated a prominent and essential role for auto-activation in gap gene regulation [80, 81]. In contrast, more recent studies suggest that auto-regulation only plays a minor part in gap gene regulation (Fig. 7b). Auto-activation by itself cannot be involved in positioning of domain boundaries as it only amplifies differences in expression levels which are already present. Instead, it contributes to sharpening and maintenance of domain borders [15]. Moreover, it does not seem to be strictly essential for correct gap gene expression (although it is clearly present in the embryo) since models of the gap gene network that lack auto-regulation show perfectly normal expression patterns [17].
Experimental support for auto-activation is strongest for hb: Early and late stages of zygotic hb expression are driven by two distinct promoters (P2 and P1, respectively), whose transcripts vary in their first exon but encode identical proteins [126, 178]. Early zygotic expression from P2 occurs in a broad anterior domain and depends on activation by Bcd (see above). Robustness, but not positioning, of this early hb expression domain also requires maternal Hb [56]. In contrast, localized late expression from P1 in its PS4 stripe depends on earlier hb expression [107, 178, 200] but not on Bcd [207]. Either maternal or early zygotic Hb on their own are sufficient for auto-activation as PS4 expression is normal in embryos lacking early zygotic expression from P2 [207], and in maternal mutants with a single paternal copy of hb [70]. PS4 expression is strongly expanded in embryos mis-expressing hb [178]. Finally, a predicted Hb binding site is present in the hb P1 promoter [208]. Note that auto-activation is not required for expression of the posterior hb domain, which is driven by both P1 and P2 promoters [107].
The evidence is less clear for auto-activation of other gap genes. The central domain of Kr is narrowed and weakened [209], and the intensification of gt domains during cycle 13 is delayed [176] in mutants of these genes expressing non-functional proteins. Moreover, recent computational studies predict that both Kr and Gt bind to some of their own regulatory elements [193].
In the case of Kr, kni, and the posterior hb domain, some authors have suggested auto-repression [178, 188, 210, 211]. Reporter assays using the two redundant Kr regulatory elements driving expression in the central domain reveal that one element—the one containing Kr binding sites—shows much weaker reporter activity than the other one [188]. In the case of kni, auto-repression is supported by the fact that reporter gene expression driven by kni regulatory elements is up-regulated in a kni mutant background [175]. Similarly, reporter gene expression in the posterior hb domain is expanded in hb mutants, and decreased in embryos over-expressing hb [178]. However, the evidence for gap gene auto-repression is weak and circumstantial, and the potential regulatory role for such negative auto-feedback remains unclear.
Repression between complementary gap genes
The basic staggered arrangement of trunk gap domains consists of two complementary pairs of expression patterns—those of hb and kni, as well as Kr and gt—which are out of phase with respect to one another (Figs. 5, 7c). This pattern is maintained and stabilized by strong mutual repression between the members of each of these complementary pairs of genes creating positive (or double-negative) regulatory feedback [187, 212]. This has been called the ‘alternating cushions’ mechanism, as one gap domain excludes—and thus buffers against—another. It is strongly supported by experimental evidence.
Repressive feedback between hb and kni is suggested by the following: The abdominal kni domain expands anteriorly in hb mutants [108, 175, 212] while kni is repressed in regions of embryos where hb is mis-expressed [73, 187, 212] or in embryos where Hb is present in the posterior region [113, 120]. Posterior expansion of kni in hb mutants has never been observed, which may be due to redundant repression by Gt and Tll in this region [15]. Very low levels of Hb are required for effective repression of kni [73, 190]. It has been suggested that this repression may be indirect, through repression of zygotic cad by Hb [182]. This is contradicted by the fact that kni expression is still observed in mutants lacking both maternal and zygotic Cad [103]. A direct interaction of Hb and kni is further supported by the fact that Hb binds to the regulatory region of kni [213] a molecular interaction that depends on co-factors of the Polycomb group [136, 139].
The effect of Kni on hb is more subtle. Only a slight expansion of the posterior hb domain can be detected in kni mutants, while the anterior hb domain remains unaffected [163, 212, 214]. Double mutants of Kr and kni, however, show complete de-repression of hb in the central region of the embryo [212]—indicating redundant repression of hb by these two factors. Furthermore, hb is repressed in regions of embryos mis-expressing kni [212, 215–217].
It has been noted that Kr and gt expression patterns are always complementary, in wild-type and various mutant backgrounds [131, 190]. In Kr mutants, both anterior and posterior domains of gt expand into the region of the central Kr domain, but do not meet in the middle [118, 125, 131, 176]. Moreover, the posterior domain of gt expands further anterior in bcd; Kr double mutants than in bcd mutants alone [190]. Finally, reporter gene expression from an enhancer driving expression in the posterior gt domain expands anteriorly in a Kr mutant background [192]. While there is only a very subtle and late effect on Kr expression in gt mutants [147, 176, 185, 218], mis-expression of gt abolishes Kr expression very effectively and the resulting embryos show a phenotype that is strikingly similar to the Kr mutant phenotype [144, 176, 187]. Moreover, the central Kr domain expands more strongly to the anterior in hb; gt double mutants than in hb mutants alone [187]. Finally, Gt has been shown to bind to multiple regulatory elements of Kr [125].
Repression between overlapping gap genes
In addition to the repressive feedback between mutually exclusive gap genes described above, there is experimental evidence for additional repressive interactions between gap genes with overlapping expression domains (Fig. 7d). For a long time, the function of these interactions remained mysterious, and they seemed to be redundant with repression between complementary gap genes. Recent studies using mathematical models of the gap gene network suggest that repressive interactions between overlapping gap genes regulate anterior shifts of gap domain boundaries during cleavage cycle 14A [15–17, 22–24]. These shifts are independent of nuclear movements [61, 171], and can cover more than 15% of the embryo’s length in the case of the posterior border of posterior gt [61].
Mathematical models allow us to identify precisely how such cross-repression can lead to boundary shifts, a task which would be extremely challenging based on traditional experimental approaches alone. Posterior of the central Kr domain, where such shifts are observed, repression from the posterior to the anterior neighbor is much stronger than the other way around. For instance, Gt represses kni, but Kni does not repress gt. This leads to an asymmetric cascade of repressive feedback with posterior dominance (Fig. 7d).
This cascade involves the following interactions, which are all supported by experimental evidence: The appearance of the posterior hb domain during early cycle 14A is made possible by Tll activation [218, 219] (which is probably indirect, via repression of kni [217]), and the absence of repression by Gt [125, 147, 176, 187]. Hb then starts to repress gt, and causes its retraction from the posterior pole [118, 176]. Gt in turn accumulates in the posterior part of the abdominal kni domain. This is possible since Kr—a strong repressor of Gt [118, 125, 131, 176]—has shifted anteriorly due to increased repression by Kni [147, 163, 179]. Gt down-regulates kni [125, 176] inducing an anterior shift in kni’s posterior border. Meanwhile, the anterior border of kni is shifting as well due to the retraction and sharpening of the anterior hb domain (Hb strongly represses kni; [108, 175, 187, 212]). Therefore, the anterior boundaries of both abdominal kni and posterior gt shift as an effect of the shift (or sharpening) of the posterior boundaries of the central Kr and the anterior hb domain.
Mathematical models suggest that this complicated chain of repressive interactions leads to the observed compaction and shift of the domains of Kr, kni, and gt in the central to posterior region of the embryo [5, 15, 16, 23, 24]. Note that such positional shifts due to gap–gap cross-repression are in direct contrast to the French Flag mechanism proposed for the gap gene system by Wolpert (Fig. 2e) [5].
In general, repression between overlapping neighbors is much weaker than that between gap genes with mutually exclusive expression patterns. This is to be expected because several nuclei express both neighboring gap genes simultaneously in each transition zone between domains. This imposes an upper limit on the strength of repression, as too strong an interaction would lead to mutual exclusion. This is probably the reason why the genetic evidence on many of these interactions remains quite ambiguous.
Repression of gt by Hb is indicated by the fact that the posterior gt domain fails to retract from the posterior pole of the embryo around mid-cycle 14A [118, 131, 176] while no gt expression can be detected in embryos over-expressing hb [131, 176, 187, 190]. As in the case of kni, repression of gt by Hb depends on co-factors of the Polycomb group [136]. In contrast, expression of the posterior hb domain is not affected in gt mutants [147, 176] or embryos mis-expressing gt [125, 187, 220].
Repression of kni by Gt has been reported by some authors, but not by others. While a posterior expansion of the abdominal kni domain was reported in one study [176], this effect was not seen in another [175]. Similarly, one study [125] reported reduced expression of abdominal kni in embryos over-expressing gt, while another [187] saw no such effect. Evidence on repression of gt by Kni is similarly ambiguous. There are slight defects of the posterior border [118, 176] and expression levels of the posterior gt domain are reduced [131, 176] in kni mutants. However, since kni is not expressed in the region of the observed defects, they are likely to be indirect.
There is little doubt that Kni represses Kr. The central Kr domain expands posteriorly into regions with reduced or lacking Kni activity in mutants [147, 163, 179, 185]. There is a Kni binding site in the Kr regulatory region, which overlaps with a Bcd activator site [221]. Repression appears to be weak, however, as mis-expression of kni fails to reduce levels of Kr in its central domain [215]. In contrast, there has been some confusion over the effect of Kr on kni. It has been proposed that Kr is required for kni activation, since expression of kni and kni reporter constructs is strongly reduced in Kr mutants [222]. However, this effect turned out to be indirect—via de-repression of gt—as kni expression is completely restored in Kr;gt double mutants [125].
Kr and hb are the only pair of overlapping gap genes that show mutual repression (Fig. 7d). Again, there is some ambiguity in the genetic evidence. While some authors have reported a posterior expansion of the anterior hb domain and its late PS4 expression in Kr mutants [107, 163, 212, 220], a quantitative study of hb expression failed to confirm this effect [214]. In any case, this interaction seems to be at least partially redundant with repression of hb by Kni, as Kr;kni double mutants show a complete de-repression of hb in the central region of the embryo [212]. Repression of Kr by Hb is suggested by an anterior expansion of the central Kr domain (or expression of corresponding Kr reporter constructs) in hb mutants [108, 163, 179, 185, 189, 223]. This expansion can be rescued by ectopic expression of hb in these mutants [216]. The interaction is probably direct, as multiple Hb binding sites have been identified in the Kr regulatory region [189]. Both of the above interactions are weak, since Kr and hb overlap across large regions of the embryo in wild-type and different mutant backgrounds [186, 195]. Moreover, Kr expression is still present in embryos over-expressing hb [187].
It has been suggested that in addition to its repressive effects, Hb can also activate Kr at low concentrations (see above and Fig. 3, bottom panel). Expression in the central Kr domain is strongly reduced in hb mutants [108, 179] and is completely absent in embryos lacking both Bcd and maternal Hb [108, 190, 191]. Reintroduction of increasing dosages of hb into the latter, leads to restoration of Kr expression in a concentration-dependent manner [190, 191]. Furthermore, there is a posterior expansion of Kr in embryos over-expressing hb [108]. However, all of these effects can be explained equally well by an indirect effect, through de-repression of kni in hb mutants, which then represses Kr [15]. Studies based on mathematical models favor this alternative mechanism and show that concentration-dependent activation of Kr by Hb is not required for correct gap gene expression [15–17, 19, 24]. At this point, both alternative explanations are equally consistent with the available evidence and expression studies in hb;kni double mutants will be required to clarify the issue.
Repression by terminal gap genes
A third layer of gap gene repression is provided by the terminal gap genes, which convey the regulatory effect of the maternal terminal system [87]. They are required to exclude trunk gap gene expression from the un-segmented pole regions of the embryo and are involved in establishing the posterior borders of the abdominal kni as well as the posterior gt and hb domains (Fig. 7e). In addition, the terminal gap gene tll may be required for activation of the posterior hb domain.
With one possible exception, the terminal gap genes have strong repressive effects on trunk gap gene expression. The evidence is quite clear, although little attention has been paid to hkb so far and its effects on Kr and kni remain to be investigated. Binding sites for Tll have been found in the regulatory regions of hb, Kr, and kni [178, 213, 221]. It represses Kr and kni in concert with the co-repressor encoded by brakeless (bks) [224]. Embryos that mis-express tll in the central region show no expression of Kr, kni, or gt [100, 175, 187, 217]. Only gt was assessed, and found to be abolished, in a similar experiment mis-expressing hkb [95]. Although, Kr expression is not affected in tll mutants alone [185, 218], embryos mutant for both tll and the posterior system show posterior expansion of the central Kr domain; this expansion extends all the way to the posterior pole if these embryos also lack hkb [87]. The abdominal domain of kni expands posteriorly both in tll and tll hkb double mutants, but it has not been established whether the expansion is larger in the latter case [99, 175, 222]. Posterior gt shows delayed retraction in tll mutants, and completely fails to retract from the posterior pole in tll hkb double mutants [99, 131, 176]. Finally, posterior hb fails to retract from the pole in hkb mutants [99, 219], while it is strongly reduced in tll and tll hkb double mutants [99, 218, 219].
In contrast to the other trunk gap genes, the posterior domain of hb is present and expanded to the anterior in embryos over-expressing tll [100, 217]. This suggests that Tll activates hb expression in its posterior domain. However, this interaction is probably indirect, since posterior hb is present in tll;kni double mutants [217]. Furthermore, it remains unclear how this activating effect overcomes translational inhibition by Nos (see above). Either, the Nos gradient has disappeared by this stage of development, or increasing amounts of hb mRNA are able to overcome translational repression by Nos. Quantitative measurements of the Nos gradient, as well as more careful studies using hb regulatory constructs will be required to resolve this issue.
Head gap genes
While head patterning is not completely independent of segment determination in the trunk [225], it involves additional head gap genes—otd, empty spiracles (ems), and buttonhead (btd) [226–232]—as well as an early gap-like expression domain of sloppy paired (slp) [233]. In contrast to the trunk gap genes, expression of the head gap genes is directly regulated by the terminal maternal system [55, 84, 227, 233–236], with additional activating contributions from Bcd [53, 226, 227, 229, 233, 234]. Although these studies indicated that Bcd activation is concentration-dependent, two more recent publications report that head gap gene expression is not seriously disrupted in embryos with a more or less uniform distribution of Bcd [55, 84]. Moreover, in contrast to the trunk, there is little evidence for gap–gap cross-regulatory interactions [233, 234, 236, 237], and head gap genes appear to act in a more or less parallel and independent manner [238, 239]. Furthermore, head gap domains—such as those of slp, btd, and the anterior domain of kni—are regulated by the maternal D–V system [175, 233, 234, 240].
Other genes with gap-like expression domains
Other genes are expressed in the blastoderm embryo in gap-like domains [167–169]. Of these, only a small number have been studied experimentally so far: nubbin (nub; also called pdm1), pdm2 [241–244] and castor (cas; also called ming) [245, 246], for example. pdm genes are regulated by gap proteins [154, 241, 244, 247], and have been shown to affect pair-rule gene expression [244]. However, in contrast to hb, Kr, kni, and gt, mutations in these genes do not lead to a gap-like phenotype and have no effect on the expression of other gap genes [244]. Therefore, they are not considered essential components of the gap gene network and will not be discussed further here.
Molecular mechanisms
So far, our discussion of gap gene regulation has remained largely at the genetic (or gene-network) level. In general, I have discussed how specific regulatory interactions (repressive or activating) affect gap gene expression without considering molecular details such as chromatin structure, or cis-regulatory elements (CREs) and the transcription factor binding sites they are composed of. Although some progress has been made towards understanding gap gene regulation at the level of regulatory sequences, our grasp of the molecular mechanisms involved is far less coherent and complete than our genetic knowledge of the system.
Zygotic gene expression before gastrulation depends on the mediator complex involved in chromatin remodeling [248]. Apart from this, very little is known about chromatin-level mechanisms of gap gene regulation and I will focus on transcriptional regulation through CREs instead.
The main conceptual problem when studying eukaryotic transcription in molecular detail is that we do not yet understand many functional and mechanistic aspects of CREs (see, for example, [249, 250]). We do not know why many of these elements are modular (i.e., located on a compact stretch of DNA), while others are dispersed across many kilobases of DNA. We cannot yet reliably predict which sets of transcription factor binding sites constitute a functional enhancer, and which ones do not. We do not have any detailed understanding how such enhancer elements interact and synergize in the regulation of whole genes. Finally, we do not have much quantitative evidence on how transcription factor occupancies at specific binding sites in a CRE affect gene expression, and whether this relationship is a simple one, as is often assumed in current assays.
For these reasons, we do not yet have a clear and satisfactory molecular understanding of the regulation of any of the gap genes discussed above. On the other hand, each of these genes can be used to illustrate some important regulatory principles that we do know about, as well as the difficulties in how to put these insights into a broader regulatory context.
The evidence presented in this section is mainly based on reporter assays in which specific stretches of regulatory sequence are combined with a heterologous promoter and a reporter gene (encoding, for example, β-galactosidase or green fluorescent protein, GFP), which are tested in transgenic animals. This is complemented by gel-shift and DNAse protection (footprinting) assays to identify specific transcription factor binding sites (see [251]). More recently, attempts have been made at determining the binding specificity of all maternal-co-ordinate and gap genes [252–254], and large-scale computational screens have been used to identify and analyze CREs (usually based on a combination of binding site cluster detection and identification of regulatory sequences which are conserved across species) [73, 192–194, 199, 211, 255, 256]. In addition, ChIP-on-chip data are now available which indicate that maternal co-ordinate and gap transcription factors bind to thousands of regulatory sequences across the entire Drosophila genome [257, 258].
As mentioned earlier, hb is transcribed from two different promoters, which vary in the first exon of their transcripts but not in the protein they encode (Fig. 8a) [126, 178]. The upstream P1 promoter has a brief open reading frame in its first exon, which has been implicated in translational regulation although its function remains unclear [126]. Maternal transcription originates exclusively from P1 [126, 178, 259]. A 1.1-kilobase (kb) region surrounding the P1 transcription start site, and containing multiple predicted GAGA factor binding sites, is necessary and sufficient to drive hb expression during oogenesis [259]. In contrast, early zygotic expression in the anterior half of the embryo is driven by the P2 promoter, which lies in the first intron of the P1 transcript [126]. A 123-bp element about 200 bp upstream of the P2 promoter is both necessary and sufficient for early anterior hb expression [50, 51]. This regulatory element contains several weak and strong binding sites for Bcd [50, 52] and Hb [260]. Late zygotic expression in the posterior hb domain and PS4 shows contributions by both P1 and P2 promoters and is under control of a regulatory element that lies 3 kb upstream of the P1 promoter [178, 208]. This element contains several predicted Kr [208] and Tll [178] binding sites. The presence of additional regulatory sequences between the upstream element and the P1 promoter is suggested by ChIP-on-chip data [257], but their function (if any) remains unknown. In summary, maternal and early zygotic hb regulation occur through entirely distinct molecular mechanisms, and hb can be considered as two independent genes encoding the same protein at these stages. In contrast, late zygotic transcription occurs through both promoters involving a shared upstream CRE. It is not clear how the switch between early and late regulation is achieved.
Molecular mechanisms of gap gene regulation. Transcripts (start site is indicated by arrow, exons by grey boxes, and introns by thin triangular lines) and protein coding sequence (black boxes), as well as cis-regulatory elements (CREs; thick black bars) involved in gap gene regulation are shown schematically for hb (a), Kr (b), kni (c), and gt (d). Solid and dashed curved arrows in a indicate early regulation by separate CREs and late regulation by a common CRE, respectively. Inset in b shows repression by competitive binding, c shows repression by interactions between CREs (kni_kd is composed of 223 and 64 bp-sub-elements; Hb-binding to the 223-bp element masks Bcd-activation in the 64-bp element in the posterior of the embryo), and d shows that strong repression of gt by Hb (required for the anterior boundary of the posterior domain) must be overcome for correct expression in the anterior domain. Genomic positions are not drawn to scale (see text for details)
In addition to expression in its central domain during the blastoderm stage, Kr shows a very complex expression pattern at later stages of development. Accordingly, its regulatory region is very complex. Kr regulatory sequences extend from 1.3 kb downstream of its transcriptional start site (including the single, short intron) up to 17 kb upstream of it [188]. Within this large region, there are specific CREs for each of the different expression domains [189, 223]. The extent of these CREs and how they interact remains controversial. There are two at least partially redundant elements (CD1, CD2) driving expression in the central domain (Fig. 8b). It remains unclear why two CREs are present and how they interact in Kr regulation. Such redundancy of CREs does not seem to be limited to Kr. Redundant CREs (called ‘sibling’ or ‘shadow’ enhancers) are now being discovered in many gene regulatory regions, including those of several other gap genes [256]. Footprint assays revealed binding sites for Bcd and Hb [189] as well as for Kni, Tll [221], and Gt [125] in both of these elements, while Kr sites are only present in CD2 [188]. In most cases, repressor sites overlap with Bcd activator sites, suggesting repression by competitive binding (see inset in Fig. 8b) [221].
Dissection of the 4.4-kb upstream region of kni has uncovered two repressive CREs that are required for setting boundaries of the abdominal kni expression domain [175, 213]. There are two discrete sub-elements responsible for transcriptional activation in the upstream region of kni (in the kni_kd element): The 64-bp element contains six binding sites for Bcd and mediates Bcd-dependent reporter expression, whereas the 223-bp element contains six Cad binding sites and mediates Cad-dependent reporter expression in the posterior part of the embryo (Fig. 8c) [181]. When these two CREs are combined, the anterior expression of the 64-bp element becomes eclipsed by Hb-mediated repression through the 223-bp element [181]. Here, in contrast to Kr, repression is achieved by interactions between CREs, rather than competitive binding of transcription factors to overlapping binding sites (see inset in Fig. 8c). The molecular mechanism for this interaction remains unclear. In addition, a CRE driving anterior kni expression (kni -5 ), as well as an intronic element driving both anterior and an imprecise, extended posterior pattern (kni +1 ) were identified using computational predictions [193, 211].
CREs for gt expression were only identified relatively recently using computational approaches. Three such elements drive reporter gene expression in the posterior (gt −3 ) and distinct anterior domains (gt −6, gt −10 ), respectively, while another element (gt −1 ) reproduces endogenous gt expression in both anterior and posterior domains (Fig. 8d) [192–194, 211, 256]. It is unknown how these elements interact, why both domain-specific and multi-domain enhancers are present, and how strong repression by Hb (required for positioning the early posterior domain) is overcome in the anterior of the embryo in the gt −1 reporter construct or in regulation of the endogenous gt gene (see inset in Fig. 8d) [19, 193].
Several recent studies based on computational modeling have attempted to predict and analyze expression of reporter constructs [211, 256, 261] or whole endogenous gap genes [204, 262] based on regulatory sequences and transcription factor concentrations. However, these predictions must be considered preliminary at best at this point. The accuracy of the predicted patterns requires further improvement: predicted boundaries are often missing or appear at a significantly different position than those measured experimentally [211, 256]. In addition, one of these studies proposes regulatory mechanisms, which are in severe conflict with the genetic evidence presented above (tll is repressed by other gap genes; Tll represses hb; Kni represses gt; Kr’s main activator is Cad) [211]. Other transcriptional models [204, 256, 261, 262] provide more plausible insights into gap gene regulation, but—in contrast to gene network models [15–20, 22, 23]—are not yet able to reproduce the dynamically shifting patterns of gap gene expression in the blastoderm embryo. This emphasizes our limited grasp of gap gene regulation at the molecular level. Further work on quantitative, dynamical models of transcriptional regulation will be required to resolve this issue.
Patterning precision and size regulation
Since segmentation gene patterns eventually determine the position of morphological body segments, they must be positioned precisely. So far, we have only considered developmental precision with regard to where, when, and how gap domain boundaries are placed, sharpened, and maintained in a ‘typical’ or ‘average’ embryo. Precise patterning, however, also requires that variability in boundary positions be minimized across embryos in a population. To achieve this, the patterning system must exhibit stability or robustness in the presence of genetic and environmental variation. Since growth rates and embryo morphology can vary across a population, the system must also be able to maintain expression domains at the same relative positions in embryos of different shape and size.
In 2002, a study by Houchmandzadeh et al. [214] found that the posterior boundary of the anterior Hb domain exhibits surprisingly little positional variability between embryos at the late blastoderm stage, while the corresponding spatial error in the maternal Bcd gradient is large. In addition, the relative position of the Hb boundary remained constant in embryos of different sizes, while no such size regulation could be detected in the Bcd gradient [214]. Similar results were reported for the pair-rule gene Eve and Bcd in a later study [263]. Furthermore, while the Bcd gradient is affected by temperature changes (as it is established by diffusion of its mRNA and/or protein), Hb precision is not [214]; in fact, hb (and eve) expression are quite unaffected if a large temperature gradient is applied across the embryo using a microfluidic device [264, 265]. Finally, the precision of Hb is maintained in mutants for all three maternal systems, other gap genes, and even in embryos lacking whole chromosome arms [214]. The only exceptions to this are certain alleles of the anterior system gene staufen (stau), which show strongly increased variability in the position of the posterior boundary of Hb [214]. This led to suggestions—based on theoretical considerations—that Hb precision could be due to transport of hb mRNA by Stau protein [266], or an unknown maternal posterior gradient which interferes with Bcd activation [267–269]. However, there is currently no experimental evidence to support either of these proposed mechanisms.
In contrast, a study using reporter constructs consisting of three concatenated Bcd binding sites found that such reporters can show sharp posterior boundaries with only very slightly increased spatial variability compared to hb [270]. Even a heterologous anterior gradient based on the yeast GAL4 transcription factor induced precise reporter gene expression [270]. None of these reporter constructs are affected by regulators other than their respective maternal protein gradients. Therefore, these results suggested that such gradients alone are capable of setting precise and sharp target gene boundaries. Precise early expression of hb at cleavage cycle 11 (before other gap proteins can be detected) provides further evidence that Bcd is sufficient to provide precise positioning [56]. In addition, some of the Bcd variability measured earlier [214] turned out to be due to methodological artifacts, and embryo-to-embryo variability measured in vivo (using a fusion of Bcd with GFB) exhibited surprisingly little spatial error in the central region of the embryo [54]. The same authors also measured the input/output ratio between Bcd and Hb protein levels in blastoderm nuclei and recovered a sigmoid distribution with very little variance, suggesting a tight correlation between concentration levels of Bcd and those of Hb. Moreover, disruption of Hb precision in stau mutants is correlated with increased variability of the Bcd gradient in these embryos [271]. Finally, there is now evidence that Bcd does exhibit size regulation within and among populations of Drosophila melanogaster [271, 272].
Yet, for many reasons, it remains highly unlikely that Bcd is indeed sufficient to establish precise positioning of gap domain boundaries. Spatial variability in the Bcd gradient is still higher than that of hb [273] or other gap domain boundaries [61] at the late blastoderm stage. Moreover, sensitivity analysis—based on the Berg-Purcell theory of bacterial chemotaxis [274]—shows that Bcd input on hb would have to be integrated over almost 2 h for it to be able to achieve the observed precision [54]. In contrast, the establishment of the anterior hb domain occurs within 20–30 min in the embryo [56, 61, 66, 67]. During this time, precision of gap gene expression increases significantly: early gap mRNA domains (with the exception of hb [56]) show very large positional variability, and only become more precise once gap–gap cross-regulation has been initiated [19, 61, 173]. At the same time, the distribution of spatial variability in the expression domains of gap genes and the pair-rule gene eve becomes increasingly de-correlated with spatial errors in the Bcd gradient (which grow steadily with lower concentrations towards the posterior of the embryo) [263]. Finally, and most significantly, none of the studies purporting to show precise regulation by Bcd take gap–gap interactions into account, although we know, for example, that at the relevant stage of development hb is repressed by Kr and Kni [212]. Such cross-regulatory interactions have been known for a long time to affect the regulation of gap domain boundaries [163], and therefore cannot be excluded from any serious analysis of patterning precision in the gap gene system.
Two recent studies confirm this and provide a mechanism for the increasing precision of gap gene expression patterns based on gap–gap cross-regulatory interactions [22, 23]. First, they show that Hb precision is reduced to that of Bcd in double mutants for Kr and kni (note that only single gap gene mutants were considered in [214], since all gap genes are on different chromosome arms; Table 1). This establishes that gap genes are important for Hb precision. Second, they use dynamical models of the gap gene network, which reproduce the observed precision of Hb (and five additional gap domain boundaries) when exposed to variation in Bcd concentration. The authors perform a numerical analysis of these models, which establishes that Kr and kni are responsible for this reduction of expression noise. They show that this is due to regulatory compensation: Since Bcd activates both hb and its repressors Kr and kni, increasing activation by Bcd is compensated by increasing repression by Kr and Kni (and vice versa) [22, 23]. Equivalent mechanisms were found for other gap domains.
While it appears that robustness of gap gene expression depends on zygotic regulatory interactions, it is the Bcd gradient that establishes size regulation. The length scale of this gradient was shown to adjust to embryo size within a wild-type laboratory population [271], and relative positions of gap gene and eve expression patterns are constant in embryos of D. melanogaster populations (collected from the wild) that differ significantly in size [272]. Genetic crosses between flies of these two populations show that this effect is entirely maternal, and is not influenced by zygotic feedback. Size regulation also occurs between certain (but not all) species of flies: While the closely related D. simulans and D. sechellia do not show scaling of gap gene patterns [272], such scaling has been found for Bcd, gap and pair-rule patterns in some very small (D. buskii) and some very large (Lucilia sericata, Calliphora vicina) fly embryos [275, 276]. Bcd proteins are of similar size between species, and gradients formed by Lucilia or Calliphora Bcd scale to the correct host embryo size if expressed in D. melanogaster [276]. Dextran injection shows that the cytoplasm of these different embryos does not impart different diffusive properties [275]. Instead, gradient scaling depends on conserved sequences in the Bcd protein required for nuclear localization and protein degradation [276]. Based on this and the observation that Bcd is rapidly imported into nuclei in embryos of D. melanogaster, it has been suggested that scaling is achieved through regulation of protein degradation [275, 276] and/or rapid nuclear import of Bcd protein [48, 276, 277].
The evolution of the gap gene network
Drosophila melanogaster is a long-germband insect. This mode of development is a derived character trait, which only occurs in some higher, holometabolous insects (these insects have a distinct larval stage with subsequent pupation, while hemimetabolous insects show gradual transformation of the larvae into the adult imago during successive moults; see Fig. 9a) [11, 12, 278]. In contrast, all other segmented animals—including vertebrates, annelid worms and most arthropods (including insects)—grow segments sequentially after gastrulation (short-germband segmentation). This ancestral, sequential mechanism is based on oscillatory temporal patterns of Notch signaling and its downstream targets, such as homologues of the pair-rule gene hairy (h). Such oscillatory patterns have been observed in vertebrates (reviewed in [26]), annelids [279], and arthropods such as spiders [280–282], centipedes [283, 284], and the cockroach Periplaneta americana—a hemimetabolous insect [285] (Fig. 9a). This may either indicate a common origin of segmentation [286, 287], or convergent co-option of the Notch signaling cascade into the segmentation process in all these phyla [288].
The evolution of the gap gene system. a A simplified phylogenetic tree for the arthropods is shown to the left (based on [323, 360, 361]) indicating relationships between taxa containing species in which gap genes have been studied in some detail. The prevalent mode of segment determination is shown in the first column (S short-, L long-germband). The presence or absence of an oscillator based on Notch-signaling is indicated in the second column. Evidence for or against gap-like expression patterns and phenotypes for the trunk gap genes hb, Kr, kni, and gt is indicated in the remaining two columns to the right (see key for abbreviations). b A simplified phylogenetic tree for the diptera (based on [362]) is shown to the left, indicating relationships between dipteran families containing species in which gap genes have been studied in some detail. The presence or absence of maternal gradients is indicated in the first column (see key for abbreviations). Only higher (cyclorrhaphan) flies have a Bcd gradient. The relative position of gap domains [from left to right in Drosophila: gt, hb (anterior), Kr, kni, gt, and hb (posterior)] and the number of pair-rule (eve) stripes before gastrulation are shown schematically to the right. There are two convergent branches, which have evolved an extreme form of long-germband development: Mosquitoes (Culicidae, top) and higher flies (Phoridae, Syrphidae, Tephritidae, and Drosophilidae, bottom) show seven eve stripes and posterior gt/hb domains before gastrulation. In contrast, midges (Psychodidae/Scatopsidae) lack posterior gt/hb and only develop 3–6 eve stripes during the blastoderm stage. Note that the posterior domains of hb and gt have swapped positions (double arrow) in mosquitoes. Question marks indicate unknown gap gene expression patterns (see main text for details)
However, Notch signaling is not involved in segment determination in holometabolous insects such as Drosophila [14], or (surprisingly) the short-germband beetle Tribolium [286] (Fig. 9a). In this latter species, the pair-rule genes themselves form an oscillatory feedback loop driving the sequential appearance of expression stripes [289]. This indicates that the gene networks governing segment determination in Tribolium—despite exhibiting short-germband dynamics—are derived compared to those in hemimetabolan short-germband insects.
Long-germband development can be seen as a heterochronic shift of segment determination to stages before gastrulation [12]. The transition from short-germband to long-germband development has occurred repeatedly during insect evolution [12] and is thought to be an adaptation to fast embryonic development [11, 290]. Some authors have suggested that this process is associated with the co-option or recruitment of gap genes into the segment determination process [13, 14, 290, 291] (the most conserved—and thus probably ancestral—role of gap genes is in head patterning and neurogenesis [154, 292–298]). In long-germband insects, gap genes provide spatially specific regulatory input for the regulation of pair-rule stripes, which replaces the regulation of such stripes by oscillatory temporal mechanisms involved in short-germband segment determination.
However, the evolutionary origins and timing of gap gene recruitment remains unclear [12, 299]. There is almost no evidence on gap gene expression and regulation outside insects (Fig. 9a). They do not play a role in segmentation of centipedes [291, 296], and hb is only expressed after segments have already formed in the crustacean Artemia fransiscana [300]. In contrast, hb is required for segmentation in the spider Achaearanea tepidariorum, where it is expressed in a complex, dynamic pattern of stripes, and leads to the loss of multiple segments upon knock-down by RNA interference (RNAi) [298]. Similarly, Kr shows a gap-like expression patterns in this species [301]. Current evidence does not allow us to distinguish whether the segmentation function of these gap genes was lost in centipedes and crustaceans, or convergently acquired in chelicerates and insects.
Somewhat more detailed evidence is available within the insects (Fig. 9). In short-germband species such as crickets and grasshoppers [292, 295, 302–304], the milkweed bug Oncopeltus fasciatus [293, 294, 305, 306] or the flour beetle Tribolium castaneum [297, 307–311], trunk gap genes are expressed in broad domains with roughly the same order along the A–P axis as in Drosophila. Small-scale mutagenesis screens in Tribolium uncovered several gap phenotypes [312, 313], one of which (the jaws mutant) is caused by a mutation in Tc-Kr [310]. Similar gap-like phenotypes have been observed in RNAi knock-down of hb, Kr and gt in Oncopeltus [293, 294, 305, 306], as well as hb in Gryllus bimaculatus [295] and Locusta migratoria [304].
In addition, mutants of the mille pattes (mlpt) gene of Tribolium also cause gap-like phenotypes [314]. This gene is not involved in segment determination in Drosophila, where it is known as tarsalless (tal) or polished rice (pri) [315, 316]. This suggests that gap genes may not only be recruited but also be lost during evolution of long-germband development. Another interesting aspect of mlpt is that it encodes a polycistronic mRNA, which codes for several, very short peptides of unknown function [314–316].
Still, there is considerable doubt that the function of gap genes is conserved in short-germband insects. RNAi knock-down of hb in the cricket Gryllus bimaculatus [295], of hb and Kr in Oncopeltus [293, 294], and of hb, Kr, kni and gt in Tribolium [297, 309–311] indicates that gap genes may be primarily involved in hox gene regulation, growth zone maintenance, or head patterning, rather than the determination of trunk segments through their effect on pair-rule genes (Fig. 9a). Moreover, RNAi knock-down of Oncopeltus gt does not affect the expression of other gap genes, despite it showing a clear gap-like phenotype, while kni knock-down does not show any phenotype at all in this species [306]. In summary, the evidence remains ambiguous, and more systematic analyses—both in terms of species and gene sampling—will be required for a better understanding of gap gene function in these insects.
In contrast, gap genes are clearly involved in segment determination in long-germband hymenopteran insects such as the parasitic wasp Nasonia vitripennis or the honeybee Apis mellifera (Fig. 9a). Nasonia mutants lacking hb, as well as Nasonia and Apis embryos exposed to Kr or gt RNAi knock-down show gap-like phenotypes [177, 317, 318]. Wild-type zygotic expression patterns of hb, Kr, kni, and gt in Nasonia, as well as Kr and gt in Apis, closely resemble those of Drosophila [103, 177, 317, 318]. Moreover, several interactions such as repression of Kr by Gt, of hb by Kr, or activation of the posterior domains of kni and gt by Cad are present in both Nasonia and Drosophila [103, 177, 317, 319].
Other aspects of gap gene expression in hymenopterans differ from Drosophila in interesting ways: Maternal gradients of the product of otd1, one of the two Nasonia orthologs of the head gap gene otd, replace Tor signaling in the terminal maternal system at both poles of the embryo [319, 320]. otd1 also activates the anterior tll domain in Apis, while the posterior domain seems to be established exclusively by mRNA localization [321]. In addition, a maternal gradient of Gt protein is present, which prevents expression of Kr in the anterior region of the embryo [317]. Maternal expression of gt is also detected in Apis, but its mRNA is not localized anteriorly as it is in Nasonia [318].
It appears that the striking similarities in gap gene expression and function between hymenopterans and Drosophila reflect convergent evolution, rather than evolutionary conservation: coleopterans (beetles) and lepidopterans (butterflies/moths)—both placed between hymenopterans and dipterans in recent phylogenies [322, 323]—show a large range of variation between long- and short-germband types of segment determination (Fig. 9a). While Tribolium is a short-germband insect (see above), other beetle species show intermediate or long-germband modes of development [324]. Unfortunately, very little is known about the gene networks involved in segment determination in these species.
The same wide range of variation was observed in those few lepidopteran species that have been studied so far (Fig. 9a): both short- and long-germband mode of development occur in the (very derived) embryos of the silkworm Bombyx mori and the tobacco hawkmoth Manduca sexta, respectively [325–329]. Consistent with this, the posterior domain of hb only appears after gastrulation in Bombyx [327], while it is present before gastrulation in Manduca [325]. The anterior domain of hb is very similar to Drosophila in both species [325, 327], and Kr expression is also conserved in Manduca [325].
Although all dipterans are long-germband insects, there are significant differences in regulatory inputs from maternal co-ordinate genes and in gap gene expression between species. Drosophila shows an extreme form of long-germband development, in which all gap domains and pair-rule stripes are formed before gastrulation (Fig. 9b). This arrangement appears to be conserved in the cyclorrhapha, the group of higher flies (Brachycera) to which Drosophila belongs: the dung fly Themira minor (family: Sepsidae) [330], the medfly Ceratitis capitata [331], the house fly Musca domestica (Muscidae) [332], various species of blowflies (Calliphoridae) [275, 276, 333], the hoverfly Episyrphus balteatus (Syrphidae) [334–336], and the hump-backed or scuttle fly Megaselia abdita (Phoridae) [334, 337, 338] all show seven pair-rule stripes before gastrulation, and gap gene expression patterns that are virtually identical to those of Drosophila (Fig. 9b).
Little functional evidence is available for gap–gap cross-regulation, but RNAi experiments have shown that many aspects of gap gene regulation by maternal factors are conserved among cyclorrhaphans: Bcd activates hb in drosophilids [208, 339], Musca (through the P2 promoter as in Drosophila) [340, 341], Megaselia (again via P2) [338, 342], and Episyrphus [336]. Anterior expression of tll in Musca involves Bcd, in concert with the dorso-ventral and terminal maternal systems [341, 343]. In Episyrphus, the terminal system activates tll and hkb, in addition to its role in regulation of the head hap gene otd [336]. Finally, Episyrphus Cad activates the posterior domains of kni and gt, as it does in Drosophila [336].
On the other hand, there are also important regulatory differences. These are evidently required in light of the fact that maternal inputs show considerable variability among cyclorrhaphan flies (Fig. 9b): otd (which encodes a homeobox transcription factor with the same affinity as Bcd) is expressed maternally in tephritid fruit flies [344], but not in Drosophila [227, 228] or Episyrphus [335]. Furthermore, while Megaselia lacks a maternal contribution to cad expression (Fig. 9b) [345], Episyrphus has no maternal hb [336], and Cad plays a much more prominent role in gap gene regulation in this fly. Episyrphus embryos exposed to cad RNAi show no trunk segments at all [335], and Cad is required not only for expression of kni and gt but also for hb and tll in the posterior of the embryo [336]. Similarly, the terminal system plays a more important role in Episyrphus than in Drosophila, as it not only regulates expression of otd, but also of cad, kni, and gt in the anterior region [336]. Finally, embryos lacking Bcd in Drosophila [346, 347] and Musca [348] show anterior deletions, but no mirror-abdomen (bicaudal) phenotypes as observed in equivalent embryos of lower cyclorrhaphan flies such as Megaselia [338, 342] and Episyrphus, [336]. This is not surprising for Episyrphus, which lacks the maternal hb contribution that maintains embryo polarity in Drosophila bcd mutants, but also suggest a comparatively minor patterning role for maternal Hb in Megaselia.
Similar to higher flies, the malaria mosquito Anopheles gambiae (Culicidae) shows seven pair-rule stripes and expression in all gap gene domains before gastrulation (Fig. 9b) [349]. However, significant differences in maternal co-ordinate and gap gene expression suggest that this form of extreme long-germband development is very probably convergent to that in higher flies. Non-cyclorrhaphan flies (including dance flies, horse flies, midges, and mosquitoes) do not have a bcd gene (Fig. 9b) [342, 350–353]. The identity of the anterior determinant—whose existence is strongly suggested by classical experiments using embryo centrifugation and UV irradiation in chironomid midges [11]—remains unknown. Neither otd nor hb are expressed maternally in Anopheles (Fig. 9b) [349] as they are in Tribolium [354]. Mosquitoes also show transient anterior localization of nos, in addition to its conserved posterior function [349, 355–357]. Moreover, gap gene expression is not entirely conserved between the two evolutionary branches, since the posterior domains of gt and hb have swapped positions in Anopheles compared to Drosophila (Fig. 9b) [349].
Expression data from basally branching dipterans such as Psychodid or Scatopsid midges corroborate the convergent nature of long-germband development in mosquitoes and higher flies. The moth midge Clogmia albipunctata (Psychodidae) only shows 6, and the phantom midge Coboldia fuscipes (Scatopsidae) only 3–5 stripes of the pair-rule gene eve before gastrulation (Fig. 9b) [334, 337, 358]. Moreover, while anterior gap gene expression is well conserved, Clogmia does not exhibit any significant posterior expression of gt, and its posterior hb domain only forms after gastrulation (Fig. 9b) [337, 358]. This reduction and delay of posterior patterning in basal dipterans suggests that both mosquitoes and higher flies have independently acquired gt expression as well as heterochronic shifts toward earlier hb and eve expression in the posterior region of the embryo.
These delays in posterior segmentation gene expression are reminiscent of (but not equivalent to) the sequential addition of segments observed during short-germband development. Although some posterior expression features only form after gastrulation in basal dipterans, there is no tissue growth involved in their establishment [12].
Another feature reminiscent of sequential segmentation is the anterior shifts in gap domain positions described above [16, 61]. These shifts are conserved among dipterans since they occur in Epysyrphus [336], as well as in Clogmia, where they are significantly more pronounced than in Drosophila [358]. Similar (although periodically repeating) traveling waves of gene expression can be observed during vertebrate somitogenesis [26] and centipede segmentation [283, 284], and are very probably also occurring in embryos of spiders [280–282] and cockroaches [285]. More detailed and comprehensive studies of gap gene expression and regulation in insects outside the Diptera will be required to reveal whether there is a true mechanistic connection between delays and shifts in posterior gap gene expression in flies and the ancestral short-germband mode of development.
Conclusions
In this review, I have attempted to provide a comprehensive overview on our current knowledge of gap gene regulation in development and evolution. For the trunk gap genes hb, Kr, kni, and gt, this knowledge is more detailed and complete than for any other developmental gene regulatory network. By now, we have a solid understanding of how regulatory interactions between maternal co-ordinate and gap genes produce the observed expression dynamics. Only minor ambiguities and gaps remain in the evidence: Does Hb affect Kr by activation and repression at different concentrations? How is translational repression by Nos overcome in the posterior hb domain? How are stable early boundaries established in light of rapid changes in hb concentration? Are we missing a posterior repressor required for the establishment of early gap domain boundaries or the control of precisely placed expression boundaries? How are head gap genes regulated? All of these remaining issues can be resolved by existing experimental and computational approaches.
On the other hand, some fundamental and intriguing questions remain: Our understanding of the molecular mechanisms underlying gap gene regulation is sketchy at best. We still cannot reliably predict expression dynamics from regulatory sequence, since it is difficult to identify those sets of transcription factor binding sites, which are essential for particular expression features. We do not understand why apparently redundant CREs are present, and how CREs interact with each other in regulation of endogenous genes. A better and quantitative understanding of eukaryotic transcription is absolutely essential to connect the genetic regulatory mechanisms—which are the focus of this review—with molecular processes at the level of the genome. Novel, experimental approaches to monitor chromatin dynamics and binding site occupancy in CREs combined with data-driven mathematical modeling of CRE interactions and function will be required to investigate these problems.
Another intriguing issue concerns gap phenotypes and their relation to underlying molecular events: Segmental deletions observed in gap mutant phenotypes most often do not coincide with the extent of the corresponding gap gene expression domains. It has been argued on theoretical grounds, that this is due to gap–gap cross-regulation, such that the absence of one gap transcription factor not only affects its own domain of expression but also those of neighboring genes [80, 81]. Furthermore, many gap gene mutants exhibit segmental duplications and inversions. In this case, it has been suggested that such phenotypes can be understood only if ratios between protein levels are considered to be relevant for positional specification, instead of absolute concentrations of individual gap proteins [359]. However, the exact mechanistic basis of these propositions remains unclear.
Finally, we do not yet have a very good understanding of the causal flow of regulatory information in complex, feedback-driven processes such as the regulation of gap domain shifts. What we do know is that this process involves interactions among all gap genes, and therefore is a network-level property of the system. A better, quantitative understanding of such properties will be required to understand the regulatory dynamics of gap gene expression, and how it influences the evolution of segment determination across different species of insects. Such an understanding can only be gained by quantitative studies combining genetic approaches with data-driven modeling of gene network dynamics.
These challenges illustrate the two central points I wanted to make in this review: First, it is undoubtedly worth taking a second, quantitative and more detailed look at biological systems that appear to have been studied to exhaustion. The more we learn about gap genes and their developmental and evolutionary context, the more interesting and important new questions we uncover. It is not mere details that remain to be discovered in these times of ‘omics’ and systems biology: Answering questions such as those described above will lead to fundamental insights and novel conceptual tools for developmental and evolutionary biology.
This leads me to the second point I am trying to make: The gap gene system—with all its biological features that have been described here, and its incomparable wealth of experimental evidence—provides a unique opportunity to study the role of gene regulatory networks in development and evolution in an integrative and quantitative manner. How do dynamic expression patterns emerge from the collective regulatory interactions within the network? What are the molecular mechanisms underlying these interactions? How do changes in regulatory mechanisms affect gene expression? Or in other words, how does random change at the level of the genome translate into non-random changes in phenotype? I have no doubt that much pioneering work to address these important issues will be based on studies of the gap gene network.
References
Akam M (1987) The molecular basis for metameric pattern in the Drosophila embryo. Development 101:1–22
Ingham PW (1988) The molecular genetics of embryonic pattern formation in Drosophila. Nature 335:25–34
Wolpert L (1968) The French Flag problem: a contribution to the discussion on pattern development and regulation. In: Waddington CH (ed) Towards a theoretical biology. Edinburgh University Press, Edinburgh, UK, pp 125–133
Wolpert L (1969) Positional information and the spatial pattern of cellular differentiation. J Theor Biol 25:1–47
Jaeger J, Reinitz J (2006) On the dynamic nature of positional information. BioEssays 28:1102–1111
Tabata T, Takei Y (2004) Morphogens, their identification and regulation. Development 131:703–712
Ashe HL, Briscoe J (2006) The interpretation of morphogen gradients. Development 133:385–394
Lander AD (2007) Morpheus unbound: reimagining the morphogen gradient. Cell 128:245–256
Ibañes M, Izpisúa Belmonte JC (2008) Theoretical and experimental approaches to understand morphogen gradients. Mol Syst Biol 4:176
Jaeger J, Irons D, Monk N (2008) Regulative feedback in pattern formation: towards a general relativistic theory of positional information. Development 135:3175–3183
Sander K (1976) Specification of the basic body pattern in insect embryogenesis. Adv Insect Physiol 12:125–238
Davis GK, Patel NH (2002) Short, long and beyond: molecular and embryological approaches to insect segmentation. Ann Rev Entomol 47:669–699
Peel A (2004) The evolution of arthropod segmentation mechanisms. BioEssays 26:1108–1116
Peel A, Chipman AD, Akam M (2005) Arthropod segmentation: beyond the Drosophila paradigm. Nat Rev Genet 6:905–916
Jaeger J, Blagov M, Kosman D, Kozlov KN, Manu, Myasnikova E, Surkova S, Vanario-Alonso CE, Samsonova M, Sharp DH, Reinitz J (2004) Dynamical analysis of regulatory interactions in the gap gene system of Drosophila melanogaster. Genetics 167:1721–1737
Jaeger J, Surkova S, Blagov M, Janssens H, Kosman D, Kozlov KN, Manu, Myasnikova E, Vanario-Alonso CE, Samsonova M, Sharp DH, Reinitz J (2004) Dynamic control of positional information in the early Drosophila embryo. Nature 430:368–371
Perkins TJ, Jaeger J, Reinitz J, Glass L (2006) Reverse engineering the gap gene network. PLoS Comp Biol 2:e51
Fomekong-Nanfack Y, Kaandorp JA, Blom J (2007) Efficient parameter estimation for spatio-temporal models of pattern formation: case study of Drosophila melanogaster. Bioinformatics 23:3356–3363
Jaeger J, Sharp DH, Reinitz J (2007) Known maternal gradients are not sufficient for the establishment of gap domains in Drosophila melanogaster. Mech Dev 124:108–128
Ashyraliyev M, Jaeger J, Blom JG (2008) Parameter estimation and determinability analysis applied to Drosophila gap gene circuits. BMC Syst Biol 2:83
Gursky VV, Kozlov KN, Samsonov AM, Reinitz J (2008) Model with asymptotically stable dynamics for Drosophila gap gene network. Biophysics 53:164–176
Manu, Surkova S, Spirov AV, Gursky V, Janssens H, Kim A-R, Radulescu O, Vanario-Alonso CE, Sharp DH, Samsonova M, Reinitz J (2009) Canalization of gene expression in the Drosophila blastoderm by gap gene cross regulation. PLoS Biol 7:e1000049
Manu, Surkova S, Spirov AV, Gursky V, Janssens H, Kim A-R, Radulescu O, Vanario-Alonso CE, Sharp DH, Samsonova M, Reinitz J (2009) Canalization of gene expression and domain shifts in the Drosophila blastoderm by dynamical attractors. PLoS Comp Biol 5:e1000303
Ashyraliyev M, Siggens K, Janssens H, Blom J, Akam M, Jaeger J (2009) Gene circuit analysis of the terminal gap gene huckebein. PLoS Comp Biol 5:e1000548
Jaeger J (2009) Modelling the Drosophila embryo. Mol BioSyst 5:1549–1568
Dequéant M-L, Pourquié O (2008) Segmental patterning of the vertebrate embryonic axis. Nat Rev Genet 9:370–382
Chipman AD (2008) Annelids step forward. Evol Dev 10:141–142
Schwalm FE (1997) Arthropods: the insects. In: Gilbert SF, Raunio AM (eds) Embryology: constructing the organism. Sinauer Associates Inc., Sunderland, MA, USA, pp 259–278
Heming BS (2003) Insect development and evolution. Cornell University Press, Ithaca, NY
Foe VE, Alberts BM (1983) Studies of nuclear and cytoplasmic behaviour during the five mitotic cycles that precede gastrulation in Drosophila embryogenesis. J Cell Sci 61:31–70
Foe VE (1989) Mitotic domains reveal early commitment of cells in Drosophila embryos. Development 107:1–22
Chan L-N, Gehring WJ (1971) Determination of blastoderm cells in Drosophila melanogaster. Proc Natl Acad Sci USA 68:2217–2221
Simcox AA, Sang JH (1983) When does determination occur in Drosophila embryos? Dev Biol 97:212–221
Nüsslein-Volhard C, Wieschaus E (1980) Mutations affecting segment number and polarity in Drosophila. Nature 287:795–801
Nüsslein-Volhard C, Wieschaus E, Kluding H (1984) Mutations affecting the pattern of the larval cuticle in Drosophila melanogaster. I. Zygotic loci on the second chromosome. Roux Arch Dev Biol 193:267–282
Jürgens G, Wieschaus E, Nüsslein-Volhard C, Kluding H (1984) Mutations affecting the pattern of the larval cuticle in Drosophila melanogaster. II. Zygotic loci on the third chromosome. Roux Arch Dev Biol 193:283–295
Wieschaus E, Nüsslein-Volhard C, Jürgens G (1984) Mutations affecting the pattern of the larval cuticle in Drosophila melanogaster. III. Zygotic loci on the x-chromosome and fourth chromosome. Roux Arch Dev Biol 1983:296–307
Nüsslein-Volhard C, Frohnhofer HG, Lehmann R (1987) Determination of anteroposterior polarity in Drosophila. Science 238:1675–1687
Lawrence PA (1981) The cellular basis of segmentation in insects. Cell 26:3–10
Martinez-Arias A, Lawrence P (1985) Parasegments and compartments in the Drosophila embryo. Nature 313:639–642
Ingham PW, Martinez-Arias A (1992) Boundaries and fields in early embryos. Cell 68:221–235
McGinnis W, Krumlauf R (1992) Homeobox genes and axial patterning. Cell 68:283–302
Frigerio G, Burri M, Bopp D, Baumgartner S, Noll M (1986) Structure of the segmentation gene paired and the Drosophila prd gene set as part of a gene network. Cell 47:735–746
Berleth T, Burri M, Thoma G, Bopp D, Richstein S, Frigerio G, Noll M, Nüsslein-Volhard C (1988) The role of localization of bicoid RNA in organizing the anterior pattern of the Drosophila embryo. EMBO J 7:1749–1756
Spirov A, Fahmy K, Schneider M, Frei E, Noll M, Baumgartner S (2009) Formation of the bicoid morphogen gradient: an mRNA gradient dictates the protein gradient. Development 136:605–614
Driever W, Nüsslein-Volhard C (1988) A gradient of bicoid protein in Drosophila embryos. Cell 54:83–93
Boring L, Weir M, Schubiger G (1993) Egg ligation alters the bcd protein gradient and segmentation gene expression in embryos of Drosophila. Mech Dev 42:97–111
Gregor T, Wieschaus EF, McGregor AP, Bialek W, Tank DW (2007) Stability and nuclear dynamics of the bicoid morphogen gradient. Cell 130:141–152
Driever W, Nüsslein-Volhard C (1988) The bicoid protein determines position in the Drosophila embryo in a concentration-dependent manner. Cell 54:95–104
Driever W, Nüsslein-Volhard C (1989) The bicoid protein is a positive regulator of hunchback transcription in the early Drosophila embryo. Nature 337:138–143
Struhl G, Struhl K, Macdonald PM (1989) The gradient morphogen bicoid is a concentration-dependent transcriptional activator. Cell 57:1259–1273
Driever W, Thoma G, Nüsslein-Volhard C (1989) Determination of spatial domains of zygotic gene expression in the Drosophila embryo by the affinity of binding sites for the bicoid morphogen. Nature 340:363–367
Gao Q, Finkelstein R (1998) Targeting gene expression to the head: the Drosophila orthodenticle gene is a direct target of the Bicoid morphogen. Development 125:4185–4193
Gregor T, Tank DW, Wieschaus EF, Bialek W (2007) Probing the limits to positional information. Cell 130:153–164
Ochoa-Espinosa A, Yu D, Tsirigos A, Struffi P, Small S (2009) Anterior-posterior postitional information in the absence of a strong Bicoid gradient. Proc Natl Acad Sci USA 106:3823–3828
Porcher A, Abu-Arish A, Huart S, Roelens B, Fradin C, Dostatni N (2010) The time to measure positional information: maternal Hunchback is required for the synchrony of the Bicoid transcriptional response at the onset of zygotic transcription. Development 137:2795–2804
Mlodzik M, Gehring WJ (1987) Expression of the caudal gene in the germ line of Drosophila: formation of an RNA and protein gradient during early embryogenesis. Cell 48:465–478
Macdonald PM, Struhl G (1986) A molecular gradient in early Drosophila embryos and its role in specifying the body pattern. Nature 324:537–545
Dubnau J, Struhl G (1996) RNA recognition and translational regulation by a homeodomain protein. Nature 379:694–649
Rivera-Pomar R, Niessing D, Schmidt-Ott U, Gehring WJ, Jäckle H (1996) RNA binding and translational suppression by bicoid. Nature 379:746–749
Surkova S, Kosman D, Kozlov K, Manu, Myasnikova E, Samsonova AA, Spirov A, Vanario-Alonso CE, Samsonova M, Reinitz J (2008) Characterization of the Drosophila segment determination morphome. Dev Biol 313:844–862
Forrest KM, Gavis ER (2003) Live imaging of endogenous RNA reveals a diffusion and entrapment mechanism for nanos mRNA localization in Drosophila. Curr Biol 13:1159–1168
Wang C, Lehmann R (1991) Nanos is the localized posterior determinant in Drosophila. Cell 66:637–647
Lehmann R, Nüsslein-Volhard C (1991) The maternal gene nanos has a central role in posterior pattern formation of the Drosophila embryo. Development 112:679–691
Dahanukar A, Wharton RP (1996) The Nanos gradient in Drosophila embryos is generated by translational regulation. Genes Dev 10:2610–2620
Tautz D, Lehmann R, Schnürch H, Schuh R, Seifert E, Kienlin A, Jones K, Jäckle H (1987) Finger protein of novel structure encoded by hunchback, a second member of the gap class of Drosophila segmentation genes. Nature 327:383–389
Tautz D (1988) Regulation of the Drosophila segmentation gene hunchback by two maternal morphogenetic centres. Nature 332:281–284
Hülskamp M, Schröder C, Pfeifle C, Jäckle H, Tautz D (1989) Posterior segmentation of the Drosophila embryo in the absence of a maternal posterior organizer gene. Nature 338:629–632
Irish V, Lehmann R, Akam M (1989) The Drosophila posterior-group gene nanos functions by repressing hunchback activity. Nature 338:646–648
Struhl G (1989) Differing strategies for organizing anterior and posterior body pattern in Drosophila embryos. Nature 338:741–744
Wharton RP, Struhl G (1991) RNA regulatory elements mediate control of Drosophila body pattern by the posterior morphogen nanos. Cell 67:955–967
Murata Y, Wharton RP (1995) Binding of Pumilio to maternal hunchback mRNA is required for posterior patterning in Drosophila embryos. Cell 80:747–756
Yu D, Small S (2008) Precise registration of gene expression boundaries by a repressive morphogen in Drosophila. Curr Biol 18:868–876
Wolpert L (1989) Positional information revisited. Development (Suppl) 107:3–12
Wolpert L (1994) Positional information and pattern formation in development. Dev Genet 15:485–490
Wolpert L (1996) One hundred years of positional information. Trends Genet 12:359–364
Meinhardt H (1977) A model of pattern formation in insect embryogenesis. J Cell Sci 23:117–139
Meinhardt H (1978) Space-dependent cell determination under the control of a morphogen gradient. J Theor Biol 74:307–321
Kerszberg M, Wolpert L (2007) Specifying positional information in the embryo: looking beyond morphogens. Cell 130:205–209
Meinhardt H (1986) Hierarchical inductions of cell states: a model for segmentation in Drosophila. J Cell Sci (Suppl) 4:357–381
Meinhardt H (1988) Models for maternally supplied positional information and the activation of segmentation genes in Drosophila embryogenesis. Development (Suppl) 104:95–110
Isalan M, Lemerle C, Serrano L (2005) Engineering gene networks to emulate Drosophila embryonic pattern formation. PLoS Biol 3:e64
Furriols M, Casanova J (2003) In and out of Torso RTK signaling. EMBO J 22:1947–1952
Löhr U, Chung H-R, Beller M, Jäckle H (2009) Antagonistic action of Bicoid and the repressor Capicua determines the spatial limits of Drosophila head gene expression domains. Proc Natl Acad Sci USA 106:21695–21700
Klingler M, Erdélyi M, Szabad J, Nüsslein-Volhard C (1988) Function of torso in determining the terminal anlagen of the Drosophila embryo. Nature 335:275–277
Strecker TR, Halsell SR, Fisher WW, Lipshitz HD (1988) Reciprocal effects of hyper- and hypoactivity mutations in the Drosophila pattern gene torso. Science 243:1062–1066
Weigel D, Jürgens G, Klingler M, Jäckle H (1990) Two gap genes mediate maternal terminal pattern information in Drosophila. Science 248:495–498
Liaw GJ, Rudolph KM, Huang JD, Dubnicoff T, Courey AJ, Lengyel JA (1995) The torso response element binds GAGA and NTF-1/ELF-1, and regulates tailless by relief of repression. Genes Dev 9:3163–3176
Paroush Z, Wainwright SM, Ish-Horowicz D (1997) Torso signalling regulates terminal patterning in Drosophila by antagonising Groucho-mediated repression. Development 124:3827–3834
Häder T, Wainwright D, Shandala T, Saint R, Taubert H, Brönner G, Jäckle H (1999) Receptor tyrosine kinase signaling regulates different modes of Groucho-dependent control of Dorsal. Curr Biol 10:51–54
Jiménez G, Guichet A, Ephrussi A, Casanova J (2000) Relief of gene repression by Torso RTK signaling: role of capicua in Drosophila terminal and dorsoventral patterning. Genes Dev 14:224–231
Chen YJ, Chiang CS, Weng LC, Lengyel JA, Liaw GJ (2002) Tramtrack69 is required for the early repression of tailless expression. Mech Dev 116:75–83
Chen Y-C, Lin SI, Chen Y-K, Chiang C-S, Liaw G-J (2009) The Torso signaling pathway modulates a dual transcriptional switch to regulate tailless expression. Nucl Acid Res 37:1061–1072
Pignoni F, Steingrimsson E, Lengyel JA (1992) bicoid and the terminal system activate tailless expression in the early Drosophila embryo. Development 115:239–251
Brönner G, Chu-LaGraff Q, Doe CQ, Cohen B, Weigel D, Taubert H, Jäckle H (1994) Sp1/Egr-like zinc-finger protein required for endoderm specification and germ-layer formation in Drosophila. Nature 369:664–668
Furriols M, Sprenger F, Casanova J (1996) Variation in the number of activated torso receptors correlates with differential gene expression. Development 122:2313–2317
Greenwood S, Struhl G (1997) Different levels of Ras activity can specify distinct transcriptional and morphological consequences in early Drosophila embryos. Development 124:4879–4886
Ghiglione C, Perrimon N, Perkins LA (1999) Quantitative variations in the level of MAPK activity control patterning of the embryonic termini in Drosophila. Dev Biol 205:181–193
Brönner G, Jäckle H (1991) Control and function of terminal gap gene activity in the posterior pole region of the Drosophila embryo. Mech Dev 35:205–211
Steingrimsson E, Pignoni F, Liaw GJ, Lengyel JA (1991) Dual role of the Drosophila pattern gene tailless in embryonic termini. Science 254:418–421
Liaw GJ, Lengyel JA (1992) Control of tailless expression by bicoid, dorsal and synergistically interacting terminal system regulatory elements. Mech Dev 40:47–61
Reuter R, Leptin M (1994) Interacting functions of snail, twist and huckebein during the early development of germ layers in Drosophila. Development 120:1137–1150
Olesnicky EC, Brent AE, Tonnes L, Walker M, Pultz MA, Leaf D, Desplan C (2006) A caudal mRNA gradient controls posterior development in the wasp Nasonia. Development 133:3973–3982
Cinnamon E, Gur-Wahnon D, Helman A, St Johnston D, Jiménez G, Paroush Z (2004) Capicua integrates input from two maternal systems in Drosophila terminal patterning. EMBO J 23:4571–4582
Rudolph KM, Liaw GJ, Daniel A, Green P, Courey AJ, Hartenstein V, Lengyel JA (1997) Complex regulatory region mediating tailless expression in early embryonic patterning and brain development. Development 124:4297–4308
Lehmann R, Nüsslein-Volhard C (1987) hunchback, a gene required for segmentation of an anterior and posterior region of the Drosophila embryo. Dev Biol 119:402–417
Hülskamp M, Lukowitz W, Beermann A, Glaser G, Tautz D (1994) Differential regulation of target genes by different alleles of the segmentation gene hunchback in Drosophila. Genetics 138:125–134
Hülskamp M, Pfeifle C, Tautz D (1990) A morphogenetic gradient of hunchback protein organizes the expression of the gap genes Krüppel and knirps in the early Drosophila embryo. Nature 346:577–580
Gloor H (1950) Schädigungsmuster eines Letalfaktors (Kr) von Drosophila melanogaster. Arch Jul Klaus Stiftung 25:28–44
Gloor H (1954) Phänotypus der heterozygoten bei der unvollständig dominanten, homozygot letalen Mutante Kr (=Krüppel) von Drosophila melanogaster. Arch Jul Klaus Stiftung 29:277–287
Wieschaus E, Nüsslein-Volhard C, Kluding H (1984) Krüppel, a gene whose activity is required early in the zygotic genome for normal embryonic segmentation. Dev Biol 104:172–186
Lehmann R (1988) Phenotypic comparison between maternal and zygotic genes controlling the segmental pattern of the Drosophila embryo. Development 104(Suppl):17–27
Rothe M, Nauber U, Jäckle H (1989) Three hormone receptor-like Drosophila genes encode an identical DNA-binding finger. EMBO J 8:3087–3094
Rothe M, Pehl M, Taubert H, Jäckle H (1992) Loss of gene function through rapid mitotic cycles in the Drosophila embryo. Nature 359:156–159
González-Gaitán M, Rothe M, Wimmer EA, Taubert H, Jäckle H (1994) Redundant functions of the genes knirps and knirps-related for the establishment of anterior Drosophila head structures. Proc Natl Acad Sci USA 91:8567–8571
Gergen JP, Wieschaus E (1986) Localized requirements for gene activity in segmentation of Drosophila embryos: analysis of armadillo, fused, giant and unpaired mutations in mosaic embryos. Roux Arch Dev Biol 195:49–62
Petschek JP, Perrimon N, Mahowald AP (1987) Region-specific defects in l(1)giant embryos of Drosophila melanogaster. Dev Biol 119:175–189
Mohler J, Eldon ED, Pirrotta V (1989) A novel spatial transcription pattern associated with the segmentation gene, giant, of Drosophila. EMBO J 8:1539–1548
Preiss A, Rosenberg UB, Kienlin A, Seifert E, Jäckle H (1985) Molecular genetics of Krüppel, a gene required for segmentation of the Drosophila embryo. Nature 313:27–32
Nauber U, Pankratz MJ, Kienlin A, Seifert E, Klemm U, Jäckle H (1988) Abdominal segmentation of the Drosophila embryo requires a hormone receptor-like protein encoded by the gap gene knirps. Nature 336:489–492
Strecker TR, Kongsuwan K, Lengyel JA, Merriam JR (1986) The zygotic mutant tailless affects the anterior and posterior ectodermal regions of the Drosophila embryo. Dev Biol 113:64–76
Strecker TR, Merriam JR, Lengyel JA (1988) Graded requirement for the zygotic terminal gene, tailless, in the brain and tail region of the Drosophila embryo. Development 102:721–734
Pignoni F, Baldarelli RM, Steingrimsson E, Diaz RJ, Patapoutian A, Merriam JR, Lengyel JA (1990) The Drosophila gene tailless is expressed at the embryonic termini and is a member of the steroid receptor superfamily. Cell 62:151–163
Rosenberg UB, Schröder C, Preiss A, Kienlin A, Côté S, Riede I, Jäckle H (1986) Structural homology of the product of the Drosophila Krüppel gene with Xenopus transcription factor IIIa. Nature 319:336–339
Capovilla M, Eldon ED, Pirrotta V (1992) The giant gene of Drosophila encodes a b-Zip DNA-binding protein that regulates the expression of other segmentation gap genes. Development 114:99–112
Schröder C, Tautz D, Seifert E, Jäckle H (1988) Differential regulation of the two transcripts from the Drosophila gap segmentation gene hunchback. EMBO J 7:2881–2887
Bender M, Horikami S, Cribbs D, Kaufman TC (1988) Identification and expression of the gap segmentation gene hunchback in Drosophila melanogaster. Dev Genet 9:715–732
Liaw GJ, Steingrimsson E, Pignoni F, Courey AJ, Lengyel JA (1993) Characterization of downstream elements in a Raf-1 pathway. Proc Natl Acad Sci USA 90:858–862
Ollo R, Maniatis T (1987) Drosophila Krüppel gene product produced in a baculovirus expression system is a nuclear phosphoprotein that binds to DNA. Proc Natl Acad Sci USA 84:5700–5704
Gaul U, Seifert E, Schuh R, Jäckle H (1987) Analysis of Krüppel protein distribution during early Drosophila development reveals posttranscriptional regulation. Cell 50:639–647
Kraut R, Levine M (1991) Spatial regulation of the gap gene giant during Drosophila development. Development 111:601–609
McDonald JA, Doe CQ (1997) Establishing neuroblast-specific gene expression in the Drosophila CNS: huckebein is activated by Wingless and Hedgehog and repressed by Engrailed and Gooseberry. Development 124:1079–1087
Licht JD, Grossel MD, Figge J, Hansen UM (1990) Drosophila Krüppel protein is a transcriptional repressor. Nature 346:76–79
Sauer F, Jäckle H (1991) Concentration-dependent transcriptional activation or repression by Krüppel from a single binding site. Nature 353:563–566
Zuo P, Stanojevic D, Colgan J, Han K, Levine M, Manley JL (1991) Activation and repression of transcription by the gap proteins hunchback and Krüppel in cultured Drosophila cells. Genes Dev 5:254–264
Pelegri F, Lehmann R (1994) A role of Polycomb group genes in the regulation of gap gene expression in Drosophila. Genetics 136:1341–1353
Gray S, Levine M (1996) Short-range transcriptional repressors mediate both quenching and direct repression within complex loci in Drosophila. Genes Dev 10:700–710
Arnosti DN, Gray S, Barolo S, Zhou J, Levine M (1996) The gap protein knirps mediates both quenching and direct repression in the Drosophila embryo. EMBO J 15:3659–3666
Kehle J, Beuchle D, Treuheit S, Christen B, Kennison JA, Bienz M, Müller J (1998) dMi-2, a hunchback-interacting protein that functions in Polycomb repression. Science 282:1897–1900
Nibu Y, Zhang H, Bajor E, Barolo S, Small S, Levine M (1998) dCTBP mediates transcriptional repression by Knirps, Krüppel and Snail in the Drosophila embryo. EMBO J 17:7009–7020
Nibu Y, Zhang H, Levine M (1998) Interaction of short-range repressors with Drosophila CTBP in the embryo. Science 280:101–104
Hewitt GF, Strunk B, Margulies C, Priputin T, Wang XD, Amey R, Pabst B, Kosman D, Reinitz J, Arnosti DN (1999) Transcriptional repression by the Drosophila giant protein: cis element positioning provides an alternative means of interpreting an effector gradient. Development 126:1201–1210
Keller SA, Mao Y, Struffi P, Margulies C, Yurk CE, Anderson AR, Amey RL, Moore S, Ebels JM, Foley K, Corado M, Arnosti DN (2000) dCTBP-dependent and -independent repression activities of the Drosophila Knirps protein. Mol Cell Biol 20:7247–7258
Nibu Y, Zhang H, Levine M (2001) Local action of long-range repressors in the Drosophila embryo. EMBO J 20:2246–2253
Nibu Y, Levine M (2001) CTBP-dependent activities of the short-range Giant repressor in the Drosophila embryo. Proc Natl Acad Sci USA 98:6204–6208
Ryu JR, Olson LK, Arnosti DN (2001) Cell-type specificity of short-range transcriptional repressors. Proc Natl Acad Sci USA 98:12960–12965
Strunk B, Struffi P, Wright K, Pabst B, Thomas J, Qin L, Arnosti DN (2001) Role of CTBP in transcriptional repression by the Drosophila giant protein. Dev Biol 239:229–240
Licht JD, Hanna-Rose W, Reddy JC, English MA, Ro M, Grossel MD, Shaknovich R, Hansen UM (1994) Mapping and mutagenesis of the amino-terminal transcriptional repression domain of the Drosophila Krüppel protein. Mol Cell Biol 14:4057–4066
Brönner G, Jäckle H (1996) Regulation and function of the terminal gap gene huckebein in the Drosophila blastoderm. Intl J Dev Biol 40:157–165
González-Gaitán M, Jäckle H (1995) Invagination centers within the Drosophila stomatogastric nervous system anlage are positioned by Notch-mediated signaling which is spatially controlled through wingless. Development 121:2313–2325
Romani S, Jimenez F, Hoch M, Patel NH, Taubert H, Jäckle H (1996) Krüppel, a Drosophila segmentation gene, participates in the specification of neurons and glial cells. Mech Dev 60:95–107
Lundell MJ, Chu-LaGraff Q, Doe CQ, Hirsh J (1996) The engrailed and huckebein genes are essential for development of serotonin neurons in the Drosophila CNS. Mol Cell Neurosci 7:46–61
Younossi-Hartenstein A, Green P, Liaw GJ, Rudolph KM, Lengyel JA, Hartenstein V (1997) Control of early neurogenesis of the Drosophila brain by the head gap genes tll, otd, ems and btd. Dev Biol 182:270–283
Isshiki T, Pearson B, Holbrook S, Doe CQ (2001) Drosophila neuroblasts sequentially express transcription factors which specify the temporal identity of their neuronal progeny. Cell 106:511–521
Novotny T, Eiselt R, Urban J (2002) Hunchback is required for the specification of the early sublineage of neuroblast 7–3 in the Drosophila central nervous system. Development 129:1027–1036
Schmucker D, Taubert H, Jäckle H (1992) Formation of the Drosophila larval photoreceptor organ and its neuronal differentiation require continuous Krüppel gene activity. Neuron 9:1025–1039
Ruiz-Gomez M, Romani S, Hartmann C, Jäckle H, Bate M (1997) Specific muscle identities are regulated by Krüppel during Drosophila embryogenesis. Development 124:3407–3414
Chen CK, Kühnlein RP, Eulenberg KG, Vincent S, Affolter M, Schuh R (1998) The transcription factors Knirps and Knirps-related control cell migration and branch morphogenesis during Drosophila tracheal development. Development 125:4959–4968
Fuss B, Meissner T, Bauer R, Lehmann C, Eckardt F, Hoch M (2001) Control of endoreduplication domains in the Drosophila gut by the knirps and knirps-related genes. Mech Dev 100:15–23
Fuss B, Hoch M (2002) Notch signaling controls cell fate specification along the dorsoventral axis of the Drosophila gut. Curr Biol 12:171–179
Lunde K, Biehs B, Nauber U, Bier E (1998) The knirps and knirps-related genes organize development of the second wing vein in Drosophila. Development 125:4145–4154
Lunde K, Trimble JL, Guichard A, Guss KA, Nauber U, Bier E (2003) Activation of the knirps locus links patterning to morphogenesis of the second wing vein in Drosophila. Development 130:235–248
Jäckle H, Tautz D, Schuh R, Seifert E, Lehmann R (1986) Cross-regulatory interactions among the gap genes of Drosophila. Nature 324:668–670
Poustelnikova E, Pisarev A, Blagov M, Samsonova M, Reinitz J (2004) A database for management of gene expression data in situ. Bioinformatics 20:2212–2221
Surkova S, Myasnikova E, Janssens H, Kozlov KN, Samsonova AA, Reinitz J, Samsonova M (2008) Pipeline for acquisition of quantitative data on segmentation gene expression from confocal images. Fly 2:1–9
Pisarev A, Poustelnikova E, Samsonova M, Reinitz J (2009) FlyEx, the quantitative atlas on segmentation gene expression at cellular resolution. Nucl Acid Res 37:D560–D566
Tomancak P, Beaton A, Weiszmann R, Kwan E, Shu S, Lewis SE, Richards S, Ashburner M, Hartenstein V, Celniker SE, Rubin GM (2002) Systematic determination of patterns of gene expression during Drosophila embryogenesis. Genome Biol 3:0088
Tomancak P, Berman BP, Beaton A, Weiszmann R, Kwan E, Hartenstein V, Celniker SE, Rubin GM (2007) Global analysis of patterns of gene expression during Drosophila embryogenesis. Genome Biol 8:R145
Frise E, Hammonds AS, Celniker S (2010) Systematic image-driven analysis of the spatial Drosophila embryonic expression landscape. Mol Syst Biol 6:345
Luengo Hendriks CL, Keranen SV, Fowlkes CC, Simirenko L, Weber GH, DePace AH, Henriquez C, Kaszuba DW, Hamann B, Eisen MB, Malik J, Sudar D, Biggin MD, Knowles DW (2006) Three-dimensional morphology and gene expression in the Drosophila blastoderm at cellular resolution I: data acquisition pipeline. Genome Biol 7:R123
Keranen SV, Fowlkes CC, Luengo Hendriks CL, Sudar D, Knowles DW, Malik J, Biggin MD (2006) Three-dimensional morphology and gene expression in the Drosophila blastoderm at cellular resolution II: dynamics. Genome Biol 7:R124
Fowlkes CC, Hendriks CL, Keranen SV, Weber GH, Rubel O, Huang MY, Chatoor S, DePace AH, Simirenko L, Henriquez C, Beaton A, Weiszmann R, Celniker S, Hamann B, Knowles DW, Biggin MD, Eisen MB, Malik J (2008) A quantitative spatiotemporal atlas of gene expression in the Drosophila blastoderm. Cell 133:364–374
Pritchard DK, Schubiger G (1996) Activation of transcription in Drosophila embryos is a gradual process mediated by the nucleocytoplasmic ratio. Genes Dev 10:1131–1142
Knipple DC, Seifert E, Rosenberg UB, Preiss A, Jäckle H (1985) Spatial and temporal patterns of Krüppel gene expression in early Drosophila embryos. Nature 317:40–44
Rothe M, Wimmer EA, Pankratz MJ, González-Gaitán M, Jäckle H (1994) Identical transacting factor requirement for knirps and knirps-related gene expression in the anterior but not in the posterior region of the Drosophila embryo. Mech Dev 46:169–181
Eldon ED, Pirrotta V (1991) Interactions of the Drosophila gap gene giant with maternal and zygotic pattern-forming genes. Development 111:367–378
Pultz MA, Westendorf L, Gale SD, Hawkins K, Lynch J, Pitt JN, Reeves NL, Yao JCY, Small S, Desplan C, Leaf DS (2005) A major role for zygotic hunchback in patterning the Nasonia embryo. Development 132:3705–3715
Margolis JS, Borowsky ML, Steingrimsson E, Shim CW, Lengyel JA, Posakony JW (1995) Posterior stripe expression of hunchback is driven from two promoters by a common enhancer element. Development 121:3067–3077
Harding K, Levine M (1988) Gap genes define the limits of Antennapedia and bithorax gene expression during early development in Drosophila. EMBO J 7:205–214
Gaul U, Weigel D (1990) Regulation of Krüppel expression in the anlage of the malpighian tubules in the Drosophila embryo. Mech Dev 33:57–67
Rivera-Pomar R, Lu X, Perrimon N, Taubert H, Jäckle H (1995) Activation of posterior gap gene expression in the Drosophila blastoderm. Nature 376:253–256
Schulz C, Tautz D (1995) Zygotic caudal regulation by hunchback and its role in abdominal segment formation of the Drosophila embryo. Development 121:1023–1028
Mlodzik M, Gibson G, Gehring WJ (1990) Effects of ectopic expression of caudal during Drosophila development. Development 109:271–277
Kerrigan LA, Croston GE, Lira LM, Kadonaga JT (1991) Sequence-specific transcriptional antirepression of the Drosophila Krüppel gene by the GAGA factor. J Biol Chem 266:574–582
Gaul U, Jäckle H (1987) Pole region-dependent repression of the Drosophila gap gene Krüppel by maternal gene products. Cell 51:549–555
Gaul U, Jäckle H (1989) Analysis of maternal effect mutant combinations elucidates regulation and function of the overlap of hunchback and Krüppel gene expression in the Drosophila blastoderm embryo. Development 107:651–662
Kraut R, Levine M (1991) Mutually repressive interactions between the gap genes giant and Krüppel define middle body regions of the Drosophila embryo. Development 111:611–621
Hoch M, Schröder C, Seifert E, Jäckle H (1990) Cis-acting control elements for Krüppel expression in the Drosophila embryo. EMBO J 9:2587–2595
Hoch M, Seifert E, Jäckle H (1991) Gene expression mediated by cis-acting sequences of the Krüppel gene in response to the Drosophila morphogens bicoid and hunchback. EMBO J 10:2267–2278
Struhl G, Johnston P, Lawrence PA (1992) Control of Drosophila body pattern by the hunchback morphogen gradient. Cell 69:237–249
Schulz C, Tautz D (1994) Autonomous concentration-dependent activation and repression of Krüppel by hunchback in the Drosophila embryo. Development 120:3043–3049
Berman BP, Nibu Y, Pfeiffer BD, Tomancak P, Celniker SE, Levine M, Rubin GM, Eisen MB (2002) Exploiting transcription factor binding site clustering to identify cis-regulatory modules involved in pattern formation in the Drosophila genome. Proc Natl Acad Sci USA 99:757–762
Schroeder MD, Pearce M, Fak J, Fan HQ, Unnerstall U, Emberly E, Rajewsky N, Siggia ED, Gaul U (2004) Transcriptional control in the segementation gene network of Drosophila. PLoS Biol 2:e271
Ochoa-Espinosa A, Yucel G, Kaplan L, Pare A, Pura N, Oberstein A, Papatsenko D, Small S (2005) The role of binding site cluster strength in Bicoid-dependent patterning in Drosophila. Proc Natl Acad Sci USA 102:4960–4965
Stanojevic D, Hoey T, Levine M (1989) Sequence-specific DNA-binding activities of the gap proteins encoded by hunchback and Krüppel in Drosophila. Nature 341:331–335
Stanojevic D, Hoey T, Levine M (1991) Regulation of a segmentation stripe by overlapping activators and repressors in the Drosophila embryo. Science 254:1385–1387
Small S, Blair A, Levine M (1992) Regulation of even-skipped stripe 2 in the Drosophila embryo. EMBO J 11:4047–4057
Small S, Blair A, Levine H (1996) Regulation of two pair-rule stripes by a single enhancer in the Drosophila embryo. Dev Biol 175:314–324
Papatsenko D, Goltsev Y, Levine M (2009) Organization of developmental enhancers in the Drosophila embryo. Nucl Acid Res 37:5665–5677
Simpson-Brose M, Treisman J, Desplan C (1994) Synergy between the hunchback and bicoid morphogens is required for anterior patterning in Drosophila. Cell 78:855–865
Reinitz J, Mjolsness E, Sharp DH (1995) Cooperative control of positional information in Drosophila by bicoid and maternal hunchback. J Exp Zool 271:47–56
Bergmann S, Sandler O, Sberro H, Shnider S, Schejter E, Shilo B-Z, Barkai N (2007) Pre-steady-state decoding of the Bicoid morphogen gradient. PLoS Biol 5:e46
Morton de Lachapelle A, Bergmann S (2010) Precision and scaling in morphogen gradient read-out. Mol Syst Biol 6:351
Papatsenko D, Levine M (2008) Dual regulation by the Hunchback gradient in the Drosophila embryo. Proc Natl Acad Sci USA 105:2901–2906
Mlodzik M, Gehring WJ (1987) Hierarchy of the genetic interactions that specify the anteroposterior segmentation pattern of the Drosophila embryo as monitored by caudal protein expression. Development 101:421–435
Moreno E, Morata G (1999) caudal is the hox gene that specifies the most posterior Drosophila segment. Nature 400:873–877
Wimmer EA, Carleton A, Harjes P, Turner T, Desplan C (2000) Bicoid-independent formation of thoracic segments in Drosophila. Science 287:2476–2479
Lukowitz W, Schröder C, Glaser G, Hülskamp M, Tautz D (1994) Regulatory and coding regions of the segmentation gene hunchback are functionally conserved between Drosophila virilis and Drosophila melanogaster. Mech Dev 45:105–115
Warrior R, Levine M (1990) Dose-dependent regulation of pair-rule stripes by gap proteins and the initiation of segment polarity. Development 110:759–767
Howard K (1990) The blastoderm prepattern. Semin Cell Biol 1:161–172
Segal E, Raveh-Sadka T, Schroeder M, Unnerstall U, Gaul U (2008) Predicting expression patterns from regulatory sequence in Drosophila segmentation. Nature 451:535–540
Clyde DE, Corado MSG, Wu X, Paré A, Papatsenko D, Small S (2003) A self-organizing system of repressor gradients establishes segmental complexity in Drosophila. Nature 426:849–853
Pankratz MJ, Busch M, Hoch M, Seifert E, Jäckle H (1992) Spatial control of the gap gene knirps in the Drosophila embryo by posterior morphogen system. Science 255:986–989
Houchmandzadeh B, Wieschaus E, Leibler S (2002) Establishment of developmental precision and proportions in the early Drosophila embryo. Nature 415:798–802
Kosman D, Small S (1997) Concentration-dependent patterning by an ectopic expression domain of the Drosophila gap gene knirps. Development 124:1343–1354
Wu X, Vasisht V, Kosman D, Reinitz J, Small S (2001) Thoracic patterning by the Drosophila gap gene hunchback. Dev Biol 237:79–92
Morán E, Jiménez G (2006) The Tailless nuclear receptor acts as a dedicated repressor in the early Drosophila embryo. Mol Cell Biol 26:3446–3454
Reinitz J, Levine M (1990) Control of the initiation of homeotic gene expression by the gap genes giant and tailless in Drosophila. Dev Biol 140:57–72
Casanova J (1990) Pattern formation under the control of the terminal system in the Drosophila embryo. Development 110:621–628
Wu X, Vakani R, Small S (1998) Two distinct mechanisms for differential positioning of gene expression borders involving the Drosophila gap protein giant. Development 125:3765–3774
Hoch M, Gerwin N, Taubert H, Jäckle H (1992) Competition for overlapping sites in the regulatory region of the Drosophila gene Krüppel. Science 256:94–97
Pankratz MJ, Hoch M, Seifert E, Jäckle H (1989) Krüppel requirement for knirps enhancement reflects overlapping gap gene activities in the Drosophila embryo. Nature 341:337–340
Jacob Y, Sather S, Martin JR, Ollo R (1991) Analysis of Krüppel control elements reveals that localized expression results from the interaction of multiple subelements. Proc Natl Acad Sci USA 88:5912–5916
Haecker A, Qi D, Lilja T, Moussian B, Andrioli LP, Luschnig S, Mannervik M (2007) Drosophila Brakeless interacts with Atrophin and is required for Tailless-mediated transcriptional repression in early embryos. PLoS Biol 5:e145
Crozatier M, Valle D, Dubois L, Ibnsouda S, Vincent A (1999) Head versus trunk patterning in the Drosophila embryo; collier requirement for formation of the intercalary segment. Development 126:4385–4394
Dalton D, Chadwick R, McGinnis W (1989) Expression and embryonic function of empty spiracles: a Drosophila homeo box gene with two patterning functions on the anterior-posterior axis of the embryo. Genes Dev 3:1940–1956
Finkelstein R, Perrimon N (1990) The orthodenticle gene is regulated by bicoid and torso and specifies Drosophila head development. Nature 346:485–488
Finkelstein R, Smouse D, Capaci TM, Spradling AC, Perrimon N (1990) The orthodenticle gene encodes a novel homeo domain protein involved in the development of the Drosophila nervous system and ocellar visual structures. Genes Dev 4:1516–1527
Walldorf U, Gehring WJ (1992) empty spiracles, a gap gene containing a homeobox involved in Drosophila head development. EMBO J 11:2247–2259
Wieschaus E, Perrimon N, Finkelstein R (1992) orthodenticle activity is required for the development of medial structures in the larval and adult epidermis of Drosophila. Development 115:801–811
Wimmer EA, Jäckle H, Pfeifle C, Cohen SM (1993) A Drosophila homologue of human Sp1 is a head-specific segmentation gene. Nature 366:690–694
Wimmer EA, Frommer G, Purnell BA, Jäckle H (1996) buttonhead and D-SpI: a novel Drosophila gene pair. Mech Dev 59:53–62
Grossniklaus U, Cadigan KM, Gehring WJ (1994) Three maternal coordinate systems cooperate in the patterning of the Drosophila head. Development 120:3155–3171
Wimmer EA, Simpson-Brose M, Cohen SM, Desplan C, Jäckle H (1995) Trans- and cis-acting requirements for blastodermal expression of the head gap gene buttonhead. Mech Dev 53:235–245
Gao Q, Wang Y, Finkelstein R (1996) orthodenticle regulation during embryonic head development in Drosophila. Mech Dev 56:3–15
Hartmann B, Reichert H, Walldorf U (2001) Interaction of gap genes in the Drosophila head: tailless regulates expression of empty spiracles in early embryonic patterning and brain development. Mech Dev 109:161–172
Cohen SM, Jürgens G (1990) Mediation of Drosophila head development by gap-like segmentation genes. Nature 346:482–485
Wimmer EA, Cohen SM, Jäckle H, Desplan C (1997) buttonhead does not contribute to a combinatorial code proposed for Drosophila head development. Development 124:1509–1517
Gallitano-Mendel A, Finkelstein R (1998) Ectopic orthodenticle expression alters segment polarity gene expression but not head segment identity in the Drosophila embryo. Dev Biol 199:125–137
Zeitlinger J, Zinzen RP, Stark A, Kellis M, Zhang H, Young RA, Levine M (2007) Whole-genome ChIP-chip analysis of Dorsal, Twist, and Snail suggests integration of diverse patterning processes in the Drosophila embryo. Genes Dev 21:385–390
Lloyd A, Sakonju S (1991) Characterization of two Drosophila POU domain genes, related to Oct-1 and Oct-2, and the regulation of their expression patterns. Mech Dev 36:87–102
Dick T, Yang X, Yeo S, Chia W (1991) Two closely linked Drosophila POU domain genes are expressed in neuroblasts and sensory elements. Proc Natl Acad Sci USA 88:7645–7649
Billin AN, Cockerill KA, Poole SJ (1991) Isolation of a family of Drosophila POU domain genes expressed in early development. Mech Dev 34:75–84
Cockerill KA, Billin AN, Poole SJ (1993) Regulation of expression domains and effects of ectopic expression reveal gap gene-like properties of the linked pdm genes of Drosophila. Mech Dev 41:139–153
Cui X, Doe CQ (1992) ming is expressed in neuroblast sublineages and regulates gene expression in the Drosophila central nervous system. Development 116:943–952
Mellerick DM, Kassis JA, Zhang S-D, Odenwald WF (1992) castor encodes a novel zinc-finger protein required for the development of a subset of CNS neurons in Drosophila. Neuron 9:789–803
Kambadur R, Koizumi K, Stivers C, Nagle J, Poole SJ, Odenwald WF (1998) Regulation of POU genes by castor and hunchback establishes layered compartments in the Drosophila CNS. Genes Dev 12:246–260
Bosveld F, van Hoek S, Sibon OCM (2008) Establishment of cell fate during early Drosophila embryogenesis requires transcriptional mediator subunit dMED31. Dev Biol 313:802–813
Reinitz J, Hou S, Sharp DH (2003) Transcriptional control in Drosophila. ComPlexUs 1:54–64
Janssens H, Hou S, Jaeger J, Kim AR, Myasnikova E, Sharp D, Reinitz J (2006) Quantitative and predictive model of transcriptional control of the Drosophila melanogaster even skipped gene. Nat Genet 38:1159–1165
Bergman CM, Carlson JW, Celniker SE (2005) Drosophila DNAse I footprint database: a systematic genome annotation of transcription factor binding sites in the fruitfly, Drosophila melanogaster. Bioinformatics 21:1747–1749
Adryan B, Teichmann S (2006) FlyTF: a systematic review of site-specific transcription factors in the fruit fly Drosophila melanogaster. Bioinformatics 22:1532–1533
Noyes MB, Meng X, Wakabayashi A, Sinha S, Brodsky MH, Wolfe SA (2008) A systematic characterization of factors that regulate Drosophila segmentation via a bacterial one-hybrid system. Nucl Acid Res 36:2547–2560
Noyes MB, Christensen RG, Wakabayashi A, Stormo GD, Brodsky MH, Wolfe SA (2008) Analysis of homeodomain specificities allows the family-wide prediction of preferred recognition sites. Cell 133:1277–1289
Berman BP, Pfeiffer BD, Laverty TR, Salzberg SL, Rubin GM, Eisen MB, Celniker SE (2004) Computational identification of developmental enhancers: conservation and function of transcription factor binding-site clusters in Drosophila melanogaster and Drosophila pseudoobscura. Genome Biol 5:R61
Kazemian M, Blatti C, Richards A, McCutchan M, Wakabayashi-Ito N, Hammonds AS, Celniker SE, Kumar S, Wolfe SA, Brodsky MH, Sinha S (2010) Quantitative analysis of the Drosophila segmentation regulatory network using pattern generating potentials. PLoS Biol 8:e1000456
Li X-Y, MacArthur S, Bourgon R, Nix D, Pollard DA, Iyer VN, Hechmer A, Simirenko L, Stapleton M, Luengo Hendriks CL, Chu HC, Ogawa N, Inwood W, Sementchenko V, Beaton A, Weiszmann R, Celniker S, Knowles DW, Gingeras T, Speed TP, Eisen MB, Biggin MD (2008) Transcription factors bind thousands of active and inactive regions in the Drosophila blastoderm. PLoS Biol 6:e27
MacArthur S, Li X-Y, Li J, Brown JB, Chu HC, Zeng L, Grondona BP, Hechmer A, Simirenko L, Keranen SV, Knowles DW, Stapleton M, Bickel P, Biggin MD, Eisen MB (2009) Developmental roles of 21 Drosophila transcription factors are determined by quantitative differences in binding to an overlapping set of thousands of genomic regions. Genome Biol 10:R80
Margolis JS, Borowsky ML, Shim CW, Posakony JW (1994) A small region surrounding the distal promoter of the hunchback gene directs maternal expression. Dev Biol 163:381–388
Treisman J, Desplan C (1989) The products of the Drosophila gap genes hunchback and Krüppel bind to the hunchback promoters. Nature 341:335–337
Fakhouri WD, Ay A, Sayal R, Dresch J, Dayringer E, Arnosti DN (2010) Deciphering a transcriptional regulatory code: modeling short-range repression in the Drosophila embryo. Mol Syst Biol 6:341
Zinzen RP, Papatsenko D (2007) Enhancer responses to similarly distributed antagonistic gradients in development. PLoS Comp Biol 3:e84
Holloway DM, Harrison LG, Kosman D, Vanario-Alonso CE, Spirov AV (2006) Analysis of pattern precision shows that Drosophila segmentation develops substantial independence from gradients of maternal gene products. Dev Dyn 235:2949–2960
Lucchetta EM, Lee JH, Fu LA, Patel NH, Ismagilov RF (2005) Dynamics of Drosophila embryonic patterning network perturbed in space and time using microfluidics. Nature 434:1134–1138
Lucchetta EM, Vincent ME, Ismagilov RF (2008) A precise Bicoid gradient is nonessential during cycles 11–13 for precise patterning in the Drosophila blastoderm. PLoS ONE 3:e3651
Aegerter-Wilmsen T, Aegerter CM, Bisseling T (2005) Model for the robust establishment of precise proportions in the early Drosophila embryo. J Theor Biol 234:13–19
Howard M, ten Wolde PR (2005) Finding the center reliably: robust patterns of developmental gene expression. Phys Rev Lett 95:208103
Houchmandzadeh B, Wieschaus E, Leibler S (2005) Precise domain specification in the developing Drosophila embryo. Phys Rev E 72:061920
McHale P, Rappel W-J, Levine H (2006) Embryonic pattern scaling achieved by oppositely directed morphogen gradients. Phys Biol 3:107–120
Crauk O, Dostatni N (2005) Bicoid determines sharp and precise target gene expression in the Drosophila embryo. Curr Biol 15:1888–1898
He F, Wen Y, Deng J, Lin X, Lu LJ, Jiao R, Ma J (2008) Probing intrinsic properties of a robust morphogen gradient in Drosophila. Dev Cell 15:558–567
Lott SE, Kreitman M, Palsson A, Alekseeva E, Ludwig MZ (2007) Canalization of segmentation and its evolution in Drosophila. Proc Natl Acad Sci USA 104:10926–10931
Reinitz J (2007) A ten per cent solution. Nature 448:418–419
Berg HC, Purcell EM (1977) Physics of chemoreception. Biophys J 20:193–219
Gregor T, Bialek W, van Steveninck RR, Tank DW, Wieschaus EF (2005) Diffusion and scaling during early embryonic pattern formation. Proc Natl Acad Sci USA 102:18403–18407
Gregor T, McGregor AP, Wieschaus E (2008) Shape and function of the Bicoid morphogen gradient in dipteran species with different sized embryos. Dev Biol 316:350–358
Coppey M, Berezhkovskii AM, Kim Y, Boettiger AN, Shvartsman SY (2007) Modeling the Bicoid gradient: diffusion and reversible nuclear trapping of a stable protein. Dev Biol 312:623–630
Mito T, Nakamura T, Noji S (2010) Evolution of insect development: to the hemimetabolous paradigm. Curr Opin Genet Dev 20:335–361
Rivera AS, Weisblat DA (2009) And lophotrochozoa makes three: Notch/Hes signaling in annelid segmentation. Dev Genes Evol 219:37–43
Stollewerk A, Schoppmeier M, Damen WGM (2003) Involvement of Notch and delta genes in spider segmentation. Nature 423:863–865
Schoppmeier M, Damen WGM (2005) suppressor of hairless and presenilin phenotypes imply involvement of canonical Notch-signaling in segmentation of the spider Cupiennius salei. Dev Biol 280:211–224
Oda H, Nishimura O, Hirao Y, Tarui H, Agata K, Akiyama-Oda Y (2007) Progressive activation of Delta-Notch signaling from around the blastopore is required to set up a functional caudal lobe in the spider Achaearanea tipidariorum. Development 134:2195–2205
Chipman AD, Arthur W, Akam M (2004) A double segment periodicity underlies segment generation in centipede development. Curr Biol 14:1250–1255
Chipman AD, Akam M (2008) The segmentation cascade in the centipede Strigamia maritima: involvement of the Notch pathway and pair-rule gene homologues. Dev Biol 319:160–169
Pueyo JI, Lanfear R, Couso JP (2008) Ancestral Notch-mediated segmentation revealed in the cockroach Periplaneta americana. Proc Natl Acad Sci USA 105:16614–16619
Tautz D (2004) Segmentation. Dev Cell 7:301–312
Couso JP (2009) Segmentation, metamerism and the Cambrian explosion. Intl J Dev Biol 53:1305–1316
Chipman AD (2010) Parallel evolution of segmentation by co-option of ancestral gene regulatory networks. BioEssays 32:60–70
Choe CP, Miller SC, Brown SJ (2006) A pair-rule gene circuit defines segments sequentially in the short-germ insect Tribolium castaneum. Proc Natl Acad Sci USA 103:6560–6564
Peel AD (2008) The evolution of developmental gene networks: lessons from comparative studies on holometabolous insects. Phil Trans Roy Soc Lond B 363:1539–1547
Chipman AD (2008) Thoughts and speculations on the ancestral arthropod segmentation pathway. In: Minelli A, Fusco G (eds) Evolving pathways. Cambridge University Press, Cambridge, UK, pp 343–358
Patel NH, Hayward DC, Lall S, Pirkl NR, DiPietro D, Ball EE (2001) Grasshopper hunchback expression reveals conserved and novel aspects of axis formation and segmentation. Development 128:3459–3472
Liu PZ, Kaufman TC (2004) hunchback is required for suppression of abdominal identity, and for proper germband growth and segmentation in the intermediate germband insect Oncopeltus fasciatus. Development 131:1515–1527
Liu PZ, Kaufman TC (2004) Krüppel is a gap gene in the intermediate germband insect Oncopeltus fasciatus and is required for development of both blastoderm and germband-derived segment. Development 131:4567–4579
Mito T, Sarashina I, Zhang H, Iwahashi A, Okamoto H, Miyawaki K, Shinmyo Y, Ohuchi H, Noji S (2005) Non-canonical functions of hunchback in segment patterning of the intermediate germ cricket Gryllus bimaculatus. Development 132:2069–2079
Chipman AD, Stollewerk A (2006) Specification of neural precursor identity in the geophilomorph centipede Strigamia maritima. Dev Biol 290:337–350
Cerny AC, Grossmann D, Bucher G, Klingler M (2008) The Tribolium ortholog of knirps and knirps-related is crucial for head segmentation but plays a minor role during abdominal patterning. Dev Biol 321:284–294
Schwager EE, Pechmann M, Feitosa NM, McGregor AP, Damen WGM (2009) hunchback functions as a segmentation gene in the spider Achaearanea tepidariorum. Curr Biol 19:1333–1340
Damen WGM (2007) Evolutionary conservation and divergence of the segmentation process in arthropods. Dev Dyn 236:1379–1391
Kontarakis Z, Copf T, Averof M (2006) Expression of hunchback during trunk segmentation in the branchiopod crustacean Artemia franciscana. Dev Genes Evol 216:89–93
McGregor AP, Pechmann M, Schwager EE, Feitosa NM, Kruck S, Aranda M, Damen WGM (2008) Wnt8 is required for growth-zone establishment and development of opisthosomal segments in a spider. Curr Biol 18:1619–1623
Mito T, Okamoto H, Shinahara W, Shinmyo Y, Miyawaki K, Ohuchi H, Noji S (2006) Krüppel acts as a gap gene regulating expression of hunchback and even-skipped in the intermediate germ cricket Gryllus bimaculatus. Dev Biol 294:471–481
Shinmyo Y, Mito T, Uda T, Nakamura T, Miyawaki K, Ohuchi H, Noji S (2006) brachyenteron is necessary for morphogenesis of the posterior gut but not for anteroposterior axial elongation from the posterior growth zone in the intermediate-germband cricket Gryllus bimaculatus. Development 133:4539–4547
He Z-B, Cao Y-Q, Yin Y-P, Wang Z-K, Chen B, Peng G-X, Xia Y-X (2006) Role of hunchback in segment patterning of Locusta migratoria manilensis revealed by parental RNAi. Dev Growth Diff 48:439–445
Liu PZ, Patel NH (2010) giant is a bone fide gap gene in the intermediate germband insect, Oncopeltus fasciatus. Development 137:835–844
Ben-David J, Chipman AD (2010) Mutual regulatory interactions of the trunk gap genes during blastoderm patterning in the hemipteran Oncopeltus fasciatus. Dev Biol 346:140–149
Sommer RJ, Tautz D (1993) Involvement of an orthologue of the Drosophila pair-rule gene hairy in segment formation of the short germ-band embryo of Tribolium (Coleoptera). Nature 361:400–4001
Wolff C, Sommer R, Schröder R, Glaser G, Tautz D (1995) Conserved and divergent expression aspects of the Drosophila segmentation gene hunchback in the short germ band embryo of the flour beetle Tribolium. Development 121:4227–4236
Bucher G, Klingler M (2004) Divergent segmentation mechanism in the short germ insect Tribolium revealed by giant expression and function. Development 131:1729–1740
Cerny AC, Bucher G, Schröder R, Klingler M (2005) Breakdown of abdominal patterning in the Tribolium Krüppel mutant jaws. Development 132:5353–5363
Marques-Souza H, Aranda M, Tautz D (2008) Delimiting the conserved features of hunchback function for the trunk organization of insects. Development 135:881–888
Sulston IA, Anderson KV (1996) Embryonic patterning mutants in Tribolium castaneum. Development 122:805–814
Maderspacher F, Bucher G, Klingler M (1998) Pair-rule and gap gene mutants in the flour beetle Tribolium castaneum. Dev Genes Evol 208:558–568
Savard J, Marques-Souza H, Aranda M, Tautz D (2006) A segmentation gene in Tribolium produces a polycistronic mRNA that codes for multiple conserved peptides. Cell 126:559–569
Galindo MI, Pueyo JI, Fouix S, Bishop SA, Couso JP (2007) Peptides encoded by short ORFs control development and define a new eukaryotic gene family. PLoS Biol 5:e106
Kondo T, Hashimoto Y, Kato K, Inagaki S, Hayashi S, Kageyama Y (2007) Small peptide regulators of actin-based cell morphogenesis encoded by a polycistronic mRNA. Nat Cell Biol 9:660–665
Brent AE, Yucel G, Small S, Desplan C (2007) Permissive and instructive anterior patterning rely on mRNA localization in the wasp embryo. Science 315:1841–1843
Wilson MJ, Havler M, Dearden PK (2010) giant, Krüppel, and caudal act as gap genes with extensive roles in patterning the honeybee embryo. Dev Biol 339:200–211
Lynch JA, Brent AE, Leaf DS, Pultz MA, Desplan C (2006) Localized maternal orthodenticle patterns anterior and posterior in the long germ wasp Nasonia. Nature 439:728–732
Lynch JA, Olesnicky EC, Desplan C (2006) Regulation and function of tailless in the long germ wasp Nasonia vitripennis. Dev Genes Evol 216:493–498
Wilson MJ, Dearden PK (2009) Tailless patterning functions are conserved in the honeybee even in the absence of Torso signaling. Dev Biol 335:276–287
Savard J, Tautz D, Richards S, Weinstock GM, Gibbs RA, Werren JH, Tettelin H, Lercher MJ (2006) Phylogenomic analysis reveals bees and wasps (Hymenoptera) at the base of the radiation of holometabolous insects. Genome Res 16:1334–1338
Wiegmann BM, Trautwein MD, Kim J-W, Cassel BK, Bertone MA, Winterton SL, Yeates DK (2009) Single-copy nuclear genes resolve the phylogeny of the holometabolous insects. BMC Biol 7:34
Patel NH, Condron BG, Zinn K (1994) Pair-rule expression patterns of even-skipped are found in both short- and long-germ beetles. Nature 367:429–434
Kraft R, Jäckle H (1994) Drosophila mode of metamerization in the embryogenesis of the lepidopteran insect Manduca sexta. Proc Natl Acad Sci USA 91:6634–6638
Nagy L, Riddiford L, Kiguchi K (1994) Morphogenesis in the early embryo of the lepidopteran Bombyx mori. Dev Biol 165:137–151
Xu X, Xu P-X, Amanai K, Suzuki Y (1997) Double-segment defining role of even-skipped homologs along the evolution of insect pattern formation. Dev Growth Diff 39:515–522
Liu W, Yang F, Jia S, Miao X, Huang Y (2008) Cloning and characterization of Bmrunt from the silkworm Bombyx mori during embryonic development. Arch Insect Biochem Physiol 69:47–59
Nakao H (2010) Characterization of Bombyx embryo segmentation process: expression profiles of engrailed, even-skipped, caudal, and wnt1/wingless homologues. J Exp Zool 314B:224–231
Hare EE, Peterson BK, Iyer VN, Meier R, Eisen MB (2008) Sepsid even-skipped enhancers are functionally conserved in Drosophila despite lack of sequence conservation. PLoS Genet 4:e1000106
Peterson BK, Hare EE, Iyer VN, Storage S, Conner L, Papaj DR, Kurashima R, Jang E, Eisen MB (2009) Big genomes facilitate the comparative identification of regulatory elements. PLoS ONE 4:e4688
Sommer R, Tautz D (1991) Segmentation gene expression in the housefly Musca domestica. Development 113:419–430
McGregor AP, Shaw PJ, Dover GA (2001) Sequence and expression of the hunchback gene in Lucilia sericata: a comparison with other dipterans. Dev Genes Evol 211:315–318
Bullock SL, Stauber M, Prell A, Hughes JR, Ish-Horowicz D, Schmidt-Ott U (2004) Differential cytoplasmic mRNA localisation adjusts pair-rule transcription factor activity to cytoarchitecture in dipteran evolution. Development 131:4251–4261
Lemke S, Schmidt-Ott U (2009) Evidence for a composite anterior determinant in the hover fly Episyrphus balteatus (Syrphidae), a cyclorrhaphan fly with an anterodorsal serosa anlage. Development 136:117–127
Lemke S, Busch SE, Antonopoulos DA, Meyer F, Domanus MH, Schmidt-Ott U (2010) Maternal activation of gap genes in the hover fly Episyrphus. Development 137:1709–1719
Rohr KB, Tautz D, Sander K (1999) Segmentation gene expression in the mothmidge Clogmia albipunctata (Diptera, Psychodidae) and other primitive dipterans. Dev Genes Evol 209:145–154
Stauber M, Taubert H, Schmidt-Ott U (2000) Function of bicoid and hunchback homologs in the basal cyclorrhaphan fly Megaselia (Phoridae). Proc Natl Acad Sci USA 97:10844–10849
Schröder R, Sander K (1993) A comparison of transplantable bicoid activity and partial bicoid homeobox sequences in several Drosophila and blowfly species (Calliphoridae). Roux Arch Dev Biol 203:34–43
Bonneton F, Shaw PJ, Fazakerley C, Shi M, Dover GA (1997) Comparison of bicoid-dependent regulation of hunchback between Musca domestica and Drosophila melanogaster. Mech Dev 66:143–156
Shaw PJ, Wratten NS, McGregor AP, Dover GA (2002) Coevolution in bicoid-dependent promoters and the inception of regulatory incompatibilities among species of higher diptera. Evol Dev 4:265–277
Lemke S, Stauber M, Shaw PJ, Rafiqi AM, Prell A, Schmidt-Ott U (2008) bicoid occurrence and Bicoid-dependent hunchback regulation in lower cyclorrhaphan flies. Evol Dev 10:413–420
Wratten NS, McGregor AP, Shaw PJ, Dover GA (2006) Evolutionary and functional analysis of the tailless enhancer in Musca domestica and Drosophila melanogaster. Evol Dev 8:6–15
Schetelig MF, Schmid BGM, Zimowska G, Wimmer EA (2008) Plasticity in mRNA expression and localization of orthodenticle within higher diptera. Evol Dev 10:700–704
Stauber M, Lemke S, Schmidt-Ott U (2008) Expression and regulation of caudal in the lower cyclorrhaphan fly Megaselia. Dev Genes Evol 218:81–87
Frohnhöfer HG, Nüsslein-Volhard C (1986) Organization of anterior pattern in the Drosophila embryo by the maternal gene bicoid. Nature 324:120–125
Frohnhöfer HG, Nüsslein-Volhard C (1987) Maternal genes required for the anterior localization of bicoid activity in the embryo of Drosophila. Genes Dev 1:880–890
Shaw PJ, Salameh A, McGregor AP, Bala S, Dover GA (2001) Divergent structure and function of the bicoid gene in muscoidea fly species. Evol Dev 3:251–262
Goltsev Y, Hsiong W, Lanzaro G, Levine M (2004) Different combinations of gap repressors for common stripes in Anopheles and Drosophila embryos. Dev Biol 275:435–446
Stauber M, Jäckle H, Schmidt-Ott U (1999) The anterior determinant bicoid of Drosophila is a derived hox class 3 gene. Proc Natl Acad Sci USA 96:3786–3789
Stauber M, Prell A, Schmidt-Ott U (2002) A single hox3 gene with composite bicoid and zerknüllt expression characteristics in non-cyclorrhaphan flies. Proc Natl Acad Sci USA 99:274–279
Holt RA, Subramanian GM, Halpern A, Sutton GG, Charlab R, Nusskern DR, Wincker P, Clark AG, Ribeiro JM, Wides R, Salzberg SL, Loftus B, Yandell M, Majoros WH, Rusch DB, Lai Z, Kraft CL, Abril JF, Anthouard V, Arensburger P, Atkinson PW, Baden H, Berardinis Vd, Baldwin D, Benes V, Biedler J, Blass C, Bolanos R, Boscus D, Barnstead M, Cai S, Center A, Chaturverdi K, Christophides GK, Chrystal MA, Clamp M, Cravchik A, Curwen V, Dana A, Delcher A, Dew I, Evans CA, Flanigan M, Grundschober-Freimoser A, Friedli L, Gu Z, Guan P, Guigo R, Hillenmeyer ME, Hladun SL, Hogan JR, Hong YS, Hoover J, Jaillon O, Ke Z, Kodira C, Kokoza E, Koutsos A, Letunic I, Levitsky A, Liang Y, Lin JJ, Lobo NF, Lopez JR, Malek JA, McIntosh TC, Meister S, Miller J, Mobarry C, Mongin E, Murphy SD, O’Brochta DA, Pfannkoch C, Qi R, Regier MA, Remington K, Shao H, Sharakhova MV, Sitter CD, Shetty J, Smith TJ, Strong R, Sun J, Thomasova D, Ton LQ, Topalis P, Tu Z, Unger MF, Walenz B, Wang A, Wang J, Wang M, Wang X, Woodford KJ, Wortman JR, Wu M, Yao A, Zdobnov EM, Zhang H, Zhao Q et al (2002) The genome sequence of the malaria mosquito Anopheles gambiae. Science 298:129–149
Nene V, Wortman JR, Lawson D, Haas B, Kodira C, Tu ZJ, Loftus B, Xi Z, Megy K, Grabherr M, Ren Q, Zdobnov EM, Lobo NF, Campbell KS, Brown SE, Bonaldo MF, Zhu J, Sinkins SP, Hogenkamp DG, Amedeo P, Arensburger P, Atkinson PW, Bidwell S, Biedler J, Birney E, Bruggner RV, Costas J, Coy MR, Crabtree J, Crawford M, Debruyn B, Decaprio D, Eiglmeier K, Eisenstadt E, El-Dorry H, Gelbart WM, Gomes SL, Hammond M, Hannick LI, Hogan JR, Holmes MH, Jaffe D, Johnston JS, Kennedy RC, Koo H, Kravitz S, Kriventseva EV, Kulp D, Labutti K, Lee E, Li S, Lovin DD, Mao C, Mauceli E, Menck CF, Miller JR, Montgomery P, Mori A, Nascimento AL, Naveira HF, Nusbaum C, O’Leary S, Orvis J, Pertea M, Quesneville H, Reidenbach KR, Rogers YH, Roth CW, Schneider JR, Schatz M, Shumway M, Stanke M, Stinson EO, Tubio JM, Vanzee JP, Verjovski-Almeida S, Werner D, White O, Wyder S, Zeng Q, Zhao Q, Zhao Y, Hill CA, Raikhel AS, Soares MB, Knudson DL, Lee NH, Galagan J, Salzberg SL, Paulsen IT, Dimopoulos G, Collins FH, Birren B, Fraser-Liggett CM, Severson DW (2007) Genome sequence of Aedes aegypti, a major arbovirus vector. Science 316:1718–1723
Schröder R (2003) The genes orthodenticle and hunchback substitute for bicoid in the beetle Tribolium. Nature 422:621–625
Curtis D, Apfeld J, Lehmann R (1995) nanos is an evolutionarily conserved organizer of anterior-posterior polarity. Development 121:1899–1910
Calvo E, Walter M, Adelman ZN, Jimenez A, Onal S, Marinotti O, James AA (2005) Nanos (nos) genes of the vector mosquitoes, Anopheles gambiae, Anopheles stephensi and Aedes aegypti. Insect Biochem Mol Biol 35:789–798
Juhn J, Marinotti O, Calvo E, James AA (2008) Gene structure and expression of nanos (nos) and oskar (osk) orthologues in the vector mosquito, Culex quinquefasciatus. Insect Mol Biol 17:545–552
García-Solache MA, Jaeger J, Akam M (2010) A systematic analysis of the gap gene system in the moth midge Clogmia albipunctata. Dev Biol 344:306–318
Kauffman SA, Goodwin BC (1990) Spatial harmonics and pattern specification in early Drosophila development. Part II. The four colour wheels model. J Theor Biol 144:321–345
Grimaldi D, Engel MS (2005) Evolution of the insects. Cambridge University Press, Cambridge, UK
Caravas J, Friedrich M (2010) Of mites and millipedes: recent progress in resolving the base of the arthropod tree. BioEssays 32:488–495
Yeates DK, Wiegmann BM (1999) Congruence and controversy: toward a higher-level phylogeny of diptera. Ann Rev Entomol 44:397–428
Acknowledgments
The literature reviewed here is large and complex. I have done my best to include citations to all relevant primary sources and apologize to authors of any significant contribution to the study of gap genes which I may have forgotten to mention. I would like to thank John Reinitz, Manu, Svetlana Surkova, Urs Schmidt-Ott, and members of my laboratory for discussion and helpful comments on the manuscript. I am especially grateful to Hilde Janssens for help with the figures, and Carlos E. Vanario-Alonso for the embryo image of en in Fig. 1b. Finally, I would like to thank two anonymous reviewers for their detailed and constructive criticism of this paper.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Jaeger, J. The gap gene network. Cell. Mol. Life Sci. 68, 243–274 (2011). https://doi.org/10.1007/s00018-010-0536-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-010-0536-y