Abstract.
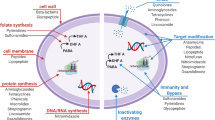
The DD-peptidase enzymes (penicillin-binding proteins) catalyze the final transpeptidation reaction of bacterial cell wall (peptidoglycan) biosynthesis. Although there is now much structural information available about these enzymes, studies of their activity as enzymes lag. It is now established that representatives of two low-molecular-mass classes of DD-peptidases recognize elements of peptidoglycan structure and rapidly react with substrates and inhibitors incorporating these elements. No members of other DD-peptidase classes, including the high-molecular-mass enzymes, essential for bacterial growth, appear to interact strongly with any particular elements of peptidoglycan structure. Rational design of inhibitors for these enzymes is therefore challenging.
Similar content being viewed by others
Author information
Authors and Affiliations
Corresponding author
Additional information
Received 30 December 2007; received after revision 19 February 2008; accepted 25 February 2008
Rights and permissions
About this article
Cite this article
Pratt, R.F. Substrate specificity of bacterial DD-peptidases (penicillin-binding proteins). Cell. Mol. Life Sci. 65, 2138–2155 (2008). https://doi.org/10.1007/s00018-008-7591-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-008-7591-7