Abstract
The human immunity-related GTPase M (IRGM) is a GTP-binding protein that regulates selective autophagy including xenophagy and mitophagy. IRGM impacts autophagy by (1) affecting mitochondrial fusion and fission, (2) promoting the co-assembly of ULK1 and Beclin 1, (3) enhancing Beclin 1 interacting partners (AMBRA1, ATG14L1, and UVRAG), (4) interacting with other key proteins (ATG16L1, p62, NOD2, cGAS, TLR3, and RIG-I), and (5) regulating lysosomal biogenesis. IRGM also negatively regulates NLRP3 inflammasome formation and therefore, maturation of the important pro-inflammatory cytokine IL-1β, impacting inflammation and pyroptosis. Ultimately, this affords protection against chronic inflammatory diseases. Importantly, ten IRGM polymorphisms (rs4859843, rs4859846, rs4958842, rs4958847, rs1000113, rs10051924, rs10065172, rs11747270, rs13361189, and rs72553867) have been associated with human inflammatory disorders including cancer, which suggests that these genetic variants are functionally relevant to the autophagic and inflammatory responses. The current review contextualizes IRGM, its modulation of autophagy, and inflammation, and emphasizes the role of IRGM as a cross point of immunity and tumorigenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Autophagy is a conserved intracellular process involving the digestion of damaged cytosolic cellular components. The autophagic machinery may be co-opted by the innate immune system for selective digestion of microbial factors, a processed termed xenophagy. Autophagy is regulated by three upstream pathways—the class I PI3K pathway, the AMPK pathway, and the class III PI3K pathway [1]—and the overall process comprises three stages—initiation, maturation, and degradation (Fig. 1). Autophagy is executed by autophagy-related gene (ATG) proteins, which facilitate the process by forming essential complexes with other molecules (Fig. 1).
The autophagy pathway. mTOR is the fulcrum for autophagic initiation, inhibiting autophagy during cellular metabolic satiety. [1] Inhibition of mTOR triggers autophagy. The activated ULK complex translocates from the cytosol to the endoplasmic reticulum (ER), initiating phagophore formation. [1] The PI3K complex is recruited to the ER and phosphatidylinositol 3-phosphate (PI3P) is generated. [2] PI3P-binding proteins (WIPI1, WIPI2, DFCP1, and Alfy) are recruited to nucleation sites, expanding the phagophore. [2, 3] Phagophore maturation requires two ubiquitin-like conjugation complexes, ATG12-ATG5-ATG16L1 and MAP1LC3 A/B/C-GABARAPs, to form the autophagosome. [1] Specific cargo, such as virulence factors, adhere to the luminal LC3. On completion, the ATG12-ATG5-ATG16L1 complex dissociates from the autophagosome, and this autophagosome fuses with the lysosome to form the autolysosome. [1, 4] The contents are then degraded through lysosomal hydrolases. [4] mTOR, mammalian target of rapamycin; LC3, microtubule-associated protein light chain 3; ULK, Unc-51 like autophagy activating kinase; ULK complex—made of ULK1/2, ATG13, FIP200, and ATG101
What is IRGM?
The immunity-related-GTPases (IRGs), also known as p47 GTPases, perform a pivotal function in innate immunity. The murine IRG gene family comprises 23 genes as tandem clusters on three chromosomes [5]. Murine models demonstrate that one of these genes, Irgm1, influences autophagic flux at the maturation phase when localised to the lysosomal compartment [6]. Murine Irgm1 modulates autophagy by assisting with autophagosome formation [7] and preventing lysosomal deacidification [7]. Irgm1 lysosomal localisation is IFN-γ-induced [8] during bacterial infections such as Salmonella enterica serovar typhimurium infection [9].
Interestingly, this family is reduced to only three genes in humans. This is possibly the consequence of host–pathogen coevolution driven by competition between IRG resistance proteins and pathogen virulence factors as it has been suggested by seminal studies conducted on Chlamydia muridarum [10] and C. trachomatis [11].
The three identifiable IRG genes in humans are IRGC, IRGQ, and IRGM. IRGC and IRGQ are located in chromosome 19 and are not involved in human immunity [5]. IRGM, which is located in chromosome 5q33.1, is the mammalian ortholog of murine Irgm1 and has a role in immunity, providing protection against intracellular pathogens [12]. Bekpen et al. [13] identified the process of ancestral Irgm1 pseudogenation and subsequent reactivation via insertion of the endogenous retroviral element 9 (EVR9) in human lineages. Some murine Irgm1 autophagy-related functions are performed similarly by IRGM. Of note, IRGM, in association with ATG8, translocates Stx17, an SNARE component, to the autophagosome for lysosomal fusion [14]. In addition, human IRGM functions upstream of autophagic initiation and throughout the autophagic process. Importantly, human IRGM is not IFN-γ-dependant, lacking a γ-activated sequence (GAS) [5]; however, recent evidence suggests it does act as a master negative regulator of cellular interferon responses [13]. There are four IRGM isoforms (IRGMa, IRGMb, IRGMc, and IRGMd) with distinct functions. Isoforms IRGMa and IRGMc lack the C-terminal G5 (SAK) motif, which is present in IRGMb and IRGMd [12]. IRGMd isoform becomes embedded within the mitochondrial membrane via cardiolipin, depolarising the membrane, inducing Bax–Bak-dependant cell death by increasing mitochondrial fission [12, 15]. Isoforms IRGMa and IRGMc also exert the same effect at high concentrations, albeit with different kinetic profiles [15]. Tian et al. [16] demonstrated that concentrations of IRGMb, IRGMc, and IRGMd were higher in cancerous tissue compared to paracancerous tissue in melanoma patients, while IRGMa concentrations were relatively lower. In addition, IRGMb was shown to increase melanoma cell survival via increasing autophagic flux, in an HMGB1-dependant manner [16]. This study also established IRGMb as a notable independent risk factor for melanoma progression [16]. Given that generation of alternatively spliced isoforms is frequently associated with drug resistance in cancer therapy, further studies are required to determine the role of IRGM isoforms in diverse tumours.
IRGM in autophagy
IRGM modulates selective autophagy including xenophagy and mitophagy, both directly and indirectly. IRGM affects autophagy indirectly through mitochondrial fusion and fission, by modulating mitochondrial membrane collapse [15]. IRGM localisation to the mitochondrial membrane via cardiolipin facilitates mitophagy [15, 17]. IRGM negatively regulates mitofilin stability during mitochondrial depolarisation, resulting in PINK1-Parkin-dependant ubiquitination and subsequent clearance of faulty mitochondria [17]. IRGM also stabilises AMPK in its Thr-172 phosphorylated form [18], which is required for AMPK activation and further induction of autophagy via phosphorylation of the ULK1 complex and Beclin 1 (Fig. 2) [19, 20].
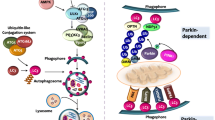
IRGM regulates autophagy. IRGM has been shown to be a potent autophagy regulator via five mechanisms: (1) pattern recognition receptors, including NOD2, are triggered upon bacterial infection via PAMPs. Activated NOD2 enhances IRGM binding to ATG16L1, to form a tripartite complex that induces autophagy. (2) IRGM activates AMPK, which in turn phosphorylates ULK1 and Beclin 1 to induce autophagy. (3) IRGM influences the composition of the Beclin 1 complex, by competing with the negative regulators Bcl2 and Rubicon, to trigger autophagy. (4) By binding to ATG8, IRGM induces Stx17 recruitment and stimulates autophagosome-lysosome fusion. (5) Finally, it induces TFEB translocation and lysosomal biogenesis by interacting with calcineurin. Additionally, IRGM isoforms mediate mitochondrial fission by facilitating mitochondrial depolarisation via cardiolipin, potentiating cell death. Furthermore, IRGM prevents type-1 interferon response by sequestering nucleic acid-sensing PRR and inducing their proteasomal degradation via SQSTM1-associated polyubiquitination. PAMPs, pathogen-associated molecular patterns; NOD2, nucleotide-binding oligomerization domain-containing protein 2; AMPK, 5’ AMP activated protein kinase, ATG16L1, autophagy-related gene 16-like 1; ULK1, unc-51 like autophagy activating kinase 1; Bcl2, B-cell lymphoma 2, ATG8, autophagy-related gene 8; Stx17, syntaxin 17; TFEB, transcription factor EB; Bax, Bcl2 associated X; Bak, Bcl2 homologous antagonist killer; TLR3, toll-like receptor 3; cGAS, cyclic GMP-AMP synthase; RIG-I, retinoic acid-inducible gene 1
Chauhan et al. [18] demonstrated that IRGM also physically interacts with essential autophagy proteins including ULK1, Beclin 1, ATG14L, and ATG16L1. During microbial insult, IRGM interacts with the pattern recognition receptor (PRR) nucleotide-binding oligomerization domain protein 2 (NOD2) in response to pathogen-associated molecular patterns (PAMPs). NOD2 enhances the K63-linked polyubiquitination of IRGM, which helps IRGM interact with the core autophagic protein ATG16L1, and thus, induces a xenophagic response (Fig. 2) [18]. IRGM also links p62/SQSTM1 to nucleic acid-sensing PPRs including cyclic GMP–AMP synthase (cGAS), Toll-like receptor (TLR) 3, and retinoic acid-inducible gene (RIG-I), enhancing their polyubiquitination and culminating in their proteasomal degradation [21]. Furthermore, IRGM was shown to determine the composition of the Beclin 1 complex, by enhancing the Beclin 1 interacting partners, AMBRA1, ATG14L1, and UVRAG (Fig. 2) [18]. Beclin 1 has two negative regulators, Bcl2 and Rubicon, that bind to its BH3, CCD, and ECD domains. ATG14L1, an enabling regulator, also has a binding site for Beclin 1’s CCD domain. IRGM and ATG14L1 simultaneously bind to Beclin 1 and compete with the negative regulators, thus, initiating autophagy [18]. In addition, absence of IRGM causes proteasomal degradation of the proteins ULK1, ATG14L1, AMBRA1, and ATG16L1, thus indicating a role of IRGM in degradative ubiquitination [22].
Finally, IRGM, in concert with mammalian ATG8 proteins, regulate lysosomal biogenesis, a fundamental process for any autophagic pathway. Kumar et al. [23] elucidated that IRGM’s interaction with transcription factor EB (TFEB) at domain of unknown function (DUF) 3371 and calcineurin PPP3CB causes dephosphorylation of TFEB by PPP3CB, resulting in TFEB nuclear translocation and stimulation of lysosomal biogenesis. The study also demonstrated that IRGM inactivates mTOR, the phosphorylating agent of TFEB, under nutrient starvation, and bacterial and viral infection [23].
IRGM in cancer
The role of autophagy in cancer is context-dependent, acting as both a promoter and suppressor of tumorigenesis [24]. Evidence that mutations in ATG genes are associated with cancer was first provided when a monoallelic deletion in BECN1, an essential autophagy gene encoding Beclin 1, was shown to result in tumorigenesis in breast cancer [25]. This was later observed in 40–75% cases of breast, ovary, and prostate cancers [26, 27]. Studies on other important autophagy genes (ATG2B, ATG3, ATG5, ATG9, ATG12, and ATG16L1) have also been conducted and suggest that autophagy plays a key role in tumorigenesis [28, 29].
IRGM appears to be a fulcrum between immunity and tumorigenesis. Studies on melanoma [16, 30], hepatocellular carcinoma [31], glioma [32], and gastric cancer [33] have shown that IRGM can promote carcinogenesis. In human glioma cell lines, overexpression of IRGM was linked to increased cell colony formation, cell proliferation, and Akt activation [32]. Xu et al. [34] also established that highly expressed IRGM in glioma cells promoted IL-8 production and M2 macrophage polarization via p62/TRAF6/NK-κB signalling. In gastric cancer, mRNA and protein levels of IRGM were shown to be significantly upregulated in the peripheral blood of cancer patients compared to healthy controls, and these levels were higher in stage IV than in stage I cancer patients [33]. Furthermore, there were significant differences in IRGM expression in patients presenting with melanoma, wherein the highest expression occurred in metastatic tumours, followed by primary tumours, and finally, benign adjacent nevus tissue [16]. A similar pattern was observed according to disease stage, where patients in stages III–IV of melanoma showed significantly higher expression of IRGM mRNA and protein levels, compared to patients in stages I and II [16]. In hepatocellular carcinoma (HCC), overexpression of the metallocarboxypeptidase AGBL2, an independent prognostic biomarker that promotes HCC cell survival and proliferation, enhanced autophagy and inhibited apoptosis via IRGM up-regulation [31].
Given the plethora of IRGM-related mechanisms that impact autophagy, a pathway that plays a complex role in cancer per se, it is pivotal to establish the precise mechanisms by which IRGM can promote carcinogenesis.
IRGM in other inflammatory conditions
Inflammation is a hallmark of cancer. In fact, many chronic inflammatory disorders including inflammatory bowel diseases (IBD) (e.g., Crohn’s disease (CD) and ulcerative colitis) [35] and autoimmune diseases (e.g., primary Sjogren syndrome, rheumatoid arthritis, systemic lupus erythematosus, and systemic sclerosis) [36] are well known to significantly increase the risk of cancer. Recent studies demonstrate that IRGM modulates anti-inflammatory processes, particularly in the context of IBD [37]. Irgm1 deficient murine models have exhibited functional abnormalities in intestinal Paneth cells and hyper-inflammation in colon and ileum, upon dextran sodium sulphate exposure [38]. Mehto et al. [39] have revealed that IRGM is a negative regulator of the NLRP3 inflammasome, suppressing inflammation and providing protection against inflammatory diseases including CD. IRGM has been shown to control inflammation by interacting with SQSTM1/p62 and mediating p62-dependent selective autophagy of NLRP3 and ASC [39]. In addition, IRGM appears to block NLPR3 and ASC oligomerization, hindering inflammasome assembly [39]. Thus, IRGM restricts inflammasome activity and protects from pyroptosis (Fig. 3) [39]. Furthermore, by inhibiting inflammasome activation, Irgm1 has been shown to negatively regulate cellular inflammation in immune and intestinal epithelial cells in a CD murine model [39]. Knockdown of IRGM has resulted in morphology changes of dendritic cells, leading to hyperstability of the immunologic synapse as well as increased T-cell activation [37]. This mechanism might explain the loss of immune tolerance in the intestine and increased adaptive immunity in CD patients who carry ATG16L1 and IRGM risk alleles [40].
IRGM and inflammation. In inflammatory diseases, IRGM controls inflammation by negatively regulating the NLRP3 inflammasome. A Upon sensing-specific stimuli, NLRP3 and ASC oligomerize to form a caspase-1 activating scaffold. B IRGM physically complexes with NLRP3 inflammasome components and obstructs inflammasome activity. C IRGM interacts with SQSTM/p62 and mediates p62-dependent selective autophagy of NLRP3 and ASC and, thus, restricts inflammasome activity. D Thus, IRGM suppresses inflammation and provides protection against inflammatory diseases. E Inflammatory conditions that have been associated with IRGM. NLR, NOD-like receptors; ASC, apoptosis-associated speck-like protein containing CARD; CD, Crohn’s disease; GIT, gastrointestinal; NAFLD, non-alcoholic fatty liver disease; VAT, visceral adipose tissue. Adapted from Mehto et al. [36].
Recently, Jena et al. [21] demonstrated that IRGM prevents translocation of IFN-α/β transcription factors IRF3 and IRF7 via proteasomal degradation of nucleic acid-sensing PRRs and mitophagy of hyperpolarised mitochondria. Hyperpolarised mitochondria yield mtROS and cause mtDNA soiling, which leads to RIG-I, TLR3, and cGAS activation and concludes with IRF3/7 phosphorylation and nuclear translocation [21]. Furthermore, in vivo Irgm1−/− murine models present with mitochondria-dependant type-1 interferonopathy that is tissue specific [41]. An aberrant type-1 IFN response results in increased apoptosis and contributes to human autoimmune disease, such as systemic lupus erythematosus [21, 41]. Indeed, Irgm1-null murine models exhibit IFN-1-dependant autoimmune syndrome, which is comparable to Sjogren’s syndrome via defective mtDNA clearance [42]. Thus, by modulating mitochondrial fission [15, 41] and polyubiquitination of PRRs, IRGM prevents type-1 interferonopathy [21]. Advancing our knowledge of IRGM as a master switch of type-1 IFN responses will be beneficial for generating therapeutics against autoimmune disorders.
However, IRGM has also been shown to support immunopathogenesis. In mouse and human intestinal epithelial cell lines, it was found that IRGM can modulate necroptosis and release damage associated molecular patterns to induce gastrointestinal inflammation [43]. Furthermore, Fang et al. [44] have demonstrated that Irgm1 deficient murine models lead to macrophage apoptosis rescue by preventing ROS accumulation and phosphorylation of JNK/p38/ERK in the MAPK pathway. Irgm1 imparts a pro-inflammatory M1 macrophage phenotype by stabilising M1-associated transcription factors Irf5 and Irf8 [45]. In addition, Irgm1-haplodeficient mice demonstrated reduced iNOS activity in M1 macrophages and reduced M1 polarization [45].
IRGM during microbial insult
Infectious diseases represent the third leading cause of cancer worldwide. In fact, 15.4% of cancers are attributable to carcinogenic infections [46]. Helicobacter pylori, high-risk human papillomavirus (HPV), hepatitis B virus (HBV), and hepatitis C virus (HCV) account for 90% of infection-related cancers worldwide [46].
Human IRGM has been extensively investigated as a xenophagy inducer during bacterial infection [12, 47, 48]. H. pylori-infected patients harbouring IRGM rs13361189 demonstrated a remarkably increased risk of gastric cancer development [49]. Considering that H. pylori induces IRGM downregulation in a strain-dependant manner, IRGM polymorphisms and microbial suppression act synergistically to prevent xenophagic clearance [49]. The deletion or knock-down of the murine ortholog Irgm1 results in increased susceptibility to both intracellular and extracellular bacteria, including Citrobacter rodentium [50], S. typhimurium [9], Mycobacterium tuberculosis [12], Listeria monocytogenes [51], and C. trachomatis [52]. Numerous mechanisms for increased susceptibility in murine Irgm1−/− models have been proposed: failure of monocyte maturation upon lamina propia infiltration and apoptosis [50], abolishment of macrophage movement and adhesion [9], and loss of intracellular bacterial restriction mechanisms [52]. In vitro models using M. leprae and M. tuberculosis corroborate murine models, identifying increased IRGM expression upon infection [12, 53]. Interestingly, impaired IRGM expression results in persistent replication of adherent-invasive Escherichia coli (AIEC) in epithelial cells and macrophages, which has been implicated in CD pathogenesis, with further increased production of IL-6 and TNF-α [47]. The inability of macrophage-mediated AIEC clearance has been further demonstrated in CD patient-derived macrophages harbouring IRGM rs10065172 [54].
IRGM is also a fundamental negative regulator of type-1 IFN response against viral pathogens [21, 55]. IRGM plays a key role in replication of HCV, an important risk factor for hepatocellular carcinoma, by regulating Golgi fragmentation and leading to co-localization of Golgi vesicles with replicating HCV [48]. Furthermore, autophagosome formation is stimulated in HIV-infected and HCV-infected HeLa cells, via IRGM interaction with HIV-NEF and HCV-NS3 [56]. This may aid in viral survival by autophagic degradation of nucleic acid-sensing proteins RIG-I and cGAS [21]. Epithelial and monocytic cells lines with abolished IRGM demonstrate enhanced antiviral properties, including MHC-I presentation and PKR stress granule formation, and are resistant to ZIKV and SARS-CoV-2 infection [55].
While the interaction of IRGM and fungi remains understudied, Rosentul et al. [57] investigated the effects of HIV+patient-derived peripheral blood mononuclear cells cytokine stimulation by Candida albicans blastoconidia, demonstrating patients harbouring IRGM rs13361189 had increased IL-8 levels compared to patients not harbouring this variant. IRGM also plays a pivotal role in limiting parasitic protozoan proliferation, including Trypanosoma cruzi and Toxoplasma Gondii, by ensuring macrophage maturation and secluding protozoan vacuoles to the lysosome [51, 58].
In the era of the microbiome, the limited number of studies investigating the potential impact of IRGM on whole microbial communities needs to be addressed. This would be particularly pertinent in gastrointestinal disorders. To date, the most relevant study investigating the impact of IRGM genetic variants on gut dysbiosis [59] demonstrated that several IBD risk alleles, including IRGM rs11741861, are associated with a decreased abundance of the genus Roseburia in healthy individuals (FDR = 0.017).
IRGM germline variants associated with disease
IRGM polymorphisms have been investigated in relation to cancer including gastric cancer, renal cell carcinoma, and glioma (Table 1) [49, 60,61,62]. Of these, consistent associations have been found between IRGM rs4958847 and rs13361189 and gastric cancer; we [49] showed that rs4958847 decreases the risk of gastric cancer in ethnic Han Chinese populations, while Burada et al. [60] showed comparable results in Caucasian populations. Interestingly, both studies reported borderline associations between IRGM rs13361189 and gastric carcinogenesis. IRGM rs13361189 is in perfect linkage disequilibrium with a 20-kb deletion located immediately upstream of the IRGM promoter gene [63]. This deletion is replaced with seven nucleotides, causing IRGM segregation in the population with two distinct upstream sequences and alters IRGM regulation, which subsequently affects autophagy [63]. Importantly, IRGM rs13361189 appears to increase the risk of other types of cancer in Chinese populations, as evidenced by Ge et al. [61] who demonstrated an increased risk of glioma in subjects harbouring this polymorphism.
Several studies establishing links between IRGM polymorphisms and other human inflammatory diseases have been conducted (Table 1) The most established associations have been with CD and tuberculosis. An early meta-analysis by Li et al. [64], including 5183 CD patients and 5571 healthy controls, showed a significant association between rs13361189 and CD, but not rs4958847 and rs10065172. However, a meta-analysis by Lu et al. [65], comprising a much larger study sample size (20590 IBD cases and 27670 controls), has demonstrated that these three IRGM polymorphisms (rs13361189, rs4958847, and rs10065172) significantly increase the risk of CD. In addition, subgroup analyses by ethnicity conducted in both meta-analyses [64, 65] revealed significant associations between these IRGM polymorphisms and an increased risk of CD among Caucasians but not among Asian populations. Recently, Ajayi et al. [66] demonstrated that subjects harbouring IRGM polymorphisms (rs13361189 and rs10065172) present with reduced IRGM expression in their serum and terminal ileum, indicating that these disease-associated SNP also affect IRGM expression, not only protein activity. Importantly, IRGM polymorphisms also appear to exacerbate symptoms (rs4958847) [67] and complicate prognoses (rs4958847 and rs13361189) [68, 69] following treatment of CD.
A meta-analysis by Xie et al. [70], investigating the association between IRGM polymorphisms status and the risk of tuberculosis, comprising 3780 patients with tuberculosis and 4835 controls, reported a decreased risk of this disease in the presence of IRGM rs10065172, rs4958842, rs4859843, and rs4859846. The capacity for IRGM polymorphisms to affect tuberculosis development appears to be species-dependant, as the variant rs9637876 (−261TT) results in significant protection from M. tuberculosis, but not M. africanum or M. bovis [71].
Other inflammatory diseases that have been associated with IRGM polymorphisms include non-alcohol fatty liver disease (rs4958847, rs13361189 and rs10065172) [72, 73], visceral adipose tissue (rs4958847 and rs13361189) [72], autoimmune thyroid disorders (rs10065172, rs4958847, and rs13361189) [74], leprosy (rs13361189) [75], chronic periodontitis (rs11747270) [76], ankylosing spondylitis (rs4958846) [77], ulcerative colitis (rs13361189 and rs4958847) [78], and sepsis (rs10065172) [79].
Conclusions
Over the past decade, autophagy has evolved from a purely homeostatic tool to a complex axis of immunological, inflammatory, and carcinogenic processes, spurred by several seminal studies. In healthy tissue, autophagy can function as a survival mechanism, maintaining viability during stress by sequestering damaged organelles or intracellular pathogens. IRGM, being the only molecule that has been shown to regulate autophagy upon infection, thus, becomes greatly important. IRGM modulates autophagy by promoting the co-assembly of ULK1 and Beclin 1, and by its interaction with key proteins such as ATG16L1, NOD2, and p62. IRGM also affects mitochondrial fusion and fission, regulates lysosomal biogenesis, and restricts NLRP3 inflammasome activity. Consistently, IRGM polymorphisms have been associated with human diseases including cancer (gastric cancer, renal cell carcinoma, and glioma), infection (tuberculosis and leprosy), autoimmunity (autoimmune thyroid disorders) and inflammatory disorders (CD, non-alcohol fatty liver disease, and chronic periodontitis). Given the intersection that occurs among autophagy, inflammation, and tumorigenesis, understanding the functional relevance of IRGM could facilitate the translation of new therapies for cancer and other prevalent inflammatory disorders.
References
Yu L, Chen Y, Tooze SA. Autophagy pathway: cellular and molecular mechanisms. Autophagy. 2018;14(2):207–15. https://doi.org/10.1080/15548627.2017.1378838.
Axe EL, Walker SA, Manifava M, Chandra P, Roderick HL, Habermann A, et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J Cell Biol. 2008;182:685–701. https://doi.org/10.1083/jcb.200803137.
Biazik J, Ylä-Anttila P, Vihinen H, Jokitalo E, Eskelinen EL. Ultrastructural relationship of the phagophore with surrounding organelles. Autophagy. 2015;11(3):439–51. https://doi.org/10.1080/15548627.2015.1017178.
Dunn WA Jr. Autophagy and related mechanisms of lysosome-mediated protein degradation. Trends Cell Biol. 1994;4(4):139–43. https://doi.org/10.1016/0962-8924(94)90069-8.
Bekpen C, Hunn JP, Rohde C, Parvanova I, Guethlein L, Dunn DM, et al. The interferon-inducible p47 (IRG) GTPases in vertebrates: loss of the cell autonomous resistance mechanism in the human lineage. Genome Biol. 2005;6(11):R92. https://doi.org/10.1186/gb-2005-6-11-r92.
Maric-Biresev J, Hunn JP, Krut O, Helms JB, Martens S, Howard JC. Loss of the interferon-γ-inducible regulatory immunity-related GTPase (IRG), Irgm1, causes activation of effector IRG proteins on lysosomes, damaging lysosomal function and predicting the dramatic susceptibility of Irgm1-deficient mice to infection. BMC Biol. 2016;14(1):33. https://doi.org/10.1186/s12915-016-0255-4.
Traver MK, Henry SC, Cantillana V, Oliver T, Hunn JP, Howard JC, et al. Immunity-related GTPase M (IRGM) proteins influence the localization of guanylate-binding protein 2 (GBP2) by modulating macroautophagy. J Biol Chem. 2011;286(35):30471–80. https://doi.org/10.1074/jbc.M111.251967.
Zhao YO, Könen-Waisman S, Taylor GA, Martens S, Howard JC. Localisation and mislocalisation of the interferon-inducible immunity-related GTPase, Irgm1 (LRG-47) in mouse cells. PLoS ONE. 2010;5(1):e8648. https://doi.org/10.1371/journal.pone.0008648.
Henry SC, Daniell X, Indaram M, Whitesides JF, Sempowski GD, Howell D, et al. Impaired macrophage function underscores susceptibility to <em>salmonella</em> in mice lacking Irgm1 (LRG-47). J Immunol. 2007;179(10):6963. https://doi.org/10.4049/jimmunol.179.10.6963.
Coers J, Bernstein-Hanley I, Grotsky D, Parvanova I, Howard JC, Taylor GA, et al. Chlamydia muridarum evades growth restriction by the IFN-gamma-inducible host resistance factor Irgb10. J Immunol. 2008;180(9):6237–45. https://doi.org/10.4049/jimmunol.180.9.6237.
Bernstein-Hanley I, Coers J, Balsara ZR, Taylor GA, Starnbach MN, Dietrich WF. The p47 GTPases Igtp and Irgb10 map to the chlamydia trachomatis susceptibility locus Ctrq-3 and mediate cellular resistance in mice. Proc Natl Acad Sci USA. 2006;103(38):14092–7. https://doi.org/10.1073/pnas.0603338103.
Singh SB, Davis AS, Taylor GA, Deretic V. Human IRGM induces autophagy to eliminate intracellular mycobacteria. Science. 2006;313(5792):1438–41. https://doi.org/10.1126/science.1129577.
Bekpen C, Marques-Bonet T, Alkan C, Antonacci F, Leogrande MB, Ventura M, et al. Death and resurrection of the human IRGM gene. PLoS Genet. 2009;5(3): e1000403. https://doi.org/10.1371/journal.pgen.1000403.
Kumar S, Jain A, Farzam F, Jia J, Gu Y, Choi SW, et al. Mechanism of Stx17 recruitment to autophagosomes via IRGM and mammalian Atg8 proteins. J Cell Biol. 2018;217(3):997–1013. https://doi.org/10.1083/jcb.201708039.
Singh SB, Ornatowski W, Vergne I, Naylor J, Delgado M, Roberts E, et al. Human IRGM regulates autophagy and cell-autonomous immunity functions through mitochondria. Nat Cell Biol. 2010;12:1154–65. https://doi.org/10.1038/ncb2119.
Tian L, Meng H, Dong X, Li X, Shi Z, Li H, et al. IRGM promotes melanoma cell survival through autophagy and is a promising prognostic biomarker for clinical application. Mol Ther Oncolytics. 2021;20:187–98. https://doi.org/10.1016/j.omto.2020.12.005.
Guo X, Zhang W, Wang C, Zhang B, Li R, Zhang L, et al. IRGM promotes the PINK1-mediated mitophagy through the degradation of Mitofilin in SH-SY5Y cells. FASEB J. 2020;34(11):14768–79. https://doi.org/10.1096/fj.202000943RR.
Chauhan S, Mandell MA, Deretic V. IRGM governs the core autophagy machinery to conduct antimicrobial defense. Mol Cell. 2015;58:507–21. https://doi.org/10.1016/j.molcel.2015.03.020.
Egan D, Kim J, Shaw RJ, Guan KL. The autophagy initiating kinase ULK1 is regulated via opposing phosphorylation by AMPK and mTOR. Autophagy. 2011;7(6):643–4. https://doi.org/10.4161/auto.7.6.15123.
Kim J, Kim YC, Fang C, Russell RC, Kim JH, Fan W, et al. Differential regulation of distinct Vps34 complexes by AMPK in nutrient stress and autophagy. Cell. 2013;152:290–303. https://doi.org/10.1016/j.cell.2012.12.016.
Jena KK, Mehto S, Nath P, Chauhan NR, Sahu R, Dhar K, et al. Autoimmunity gene IRGM suppresses cGAS-STING and RIG-I-MAVS signaling to control interferon response. EMBO Rep. 2020. https://doi.org/10.15252/embr.202050051.
Antonioli M, Di Rienzo M, Piacentini M, Fimia GM. Emerging mechanisms in initiating and terminating autophagy. Trends Biochem Sci. 2017;42(1):28–41. https://doi.org/10.1016/j.tibs.2016.09.008.
Kumar S, Jain A, Choi SW, da Silva GPD, Allers L, Mudd MH, et al. Mammalian Atg8 proteins and the autophagy factor IRGM control mTOR and TFEB at a regulatory node critical for responses to pathogens. Nat Cell Biol. 2020;22(8):973–85. https://doi.org/10.1038/s41556-020-0549-1.
Bhutia SK, Mukhopadhyay S, Sinha N, Das DN, Panda PK, Patra SK, et al. Autophagy: cancer’s friend or foe? Adv Cancer Res. 2013;118:61–95. https://doi.org/10.1016/b978-0-12-407173-5.00003-0.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H, et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature. 1999;402(6762):672–6. https://doi.org/10.1038/45257.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H, et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature. 1999;402:672–6. https://doi.org/10.1038/45257.
Aita VM, Liang XH, Murty VV, Pincus DL, Yu W, Cayanis E, et al. Cloning and genomic organization of beclin 1, a candidate tumor suppressor gene on chromosome 17q21. Genomics. 1999;59(1):59–65. https://doi.org/10.1006/geno.1999.5851.
Takamura A, Komatsu M, Hara T, Sakamoto A, Kishi C, Waguri S, et al. Autophagy-deficient mice develop multiple liver tumors. Genes Dev. 2011;25(8):795–800. https://doi.org/10.1101/gad.2016211.
Kang MR, Kim MS, Oh JE, Kim YR, Song SY, Kim SS, et al. Frameshift mutations of autophagy-related genes ATG2B, ATG5, ATG9B and ATG12 in gastric and colorectal cancers with microsatellite instability. J Pathol. 2009;217(5):702–6. https://doi.org/10.1002/path.2509.
Dong H, Tian L, Li R, Pei C, Fu Y, Dong X, et al. IFNg-induced Irgm1 promotes tumorigenesis of melanoma via dual regulation of apoptosis and Bif-1-dependent autophagy. Oncogene. 2015;34:5363–71. https://doi.org/10.1038/onc.2014.459.
Wang LL, Jin XH, Cai MY, Li HG, Chen JW, Wang FW, et al. AGBL2 promotes cancer cell growth through IRGM-regulated autophagy and enhanced aurora a activity in hepatocellular carcinoma. Cancer Lett. 2018;414:71–80. https://doi.org/10.1016/j.canlet.2017.11.003.
Xu Y, Liu R, Liao C, Liu J, Zhao H, Li Z, et al. High expression of immunity-related GTPase family M protein in glioma promotes cell proliferation and autophagy protein expression. Pathol Res Pract. 2019;215(1):90–6. https://doi.org/10.1016/j.prp.2018.10.004.
Song Z, Guo C, Zhu L, Shen P, Wang H, Guo C, et al. Elevated expression of immunity-related GTPase family M in gastric cancer. Tumour Biol. 2015;36(7):5591–6. https://doi.org/10.1007/s13277-015-3229-1.
Xu Y, Liao C, Liu R, Liu J, Chen Z, Zhao H, et al. IRGM promotes glioma M2 macrophage polarization through p62/TRAF6/NF-κB pathway mediated IL-8 production. Cell Biol Int. 2019;43(2):125–35. https://doi.org/10.1002/cbin.11061.
Axelrad JE, Lichtiger S, Yajnik V. Inflammatory bowel disease and cancer: the role of inflammation, immunosuppression, and cancer treatment. World J Gastroenterol. 2016;22(20):4794–801. https://doi.org/10.3748/wjg.v22.i20.4794.
Giat E, Ehrenfeld M, Shoenfeld Y. Cancer and autoimmune diseases. Autoimmun Rev. 2017;16(10):1049–57. https://doi.org/10.1016/j.autrev.2017.07.022.
Wildenberg ME, Vos AC, Wolfkamp SC, Duijvestein M, Verhaar AP, Te Velde AA, et al. Autophagy attenuates the adaptive immune response by destabilizing the immunologic synapse. Gastroenterology. 2012;142(7):1493-503.e6. https://doi.org/10.1053/j.gastro.2012.02.034.
Liu B, Gulati AS, Cantillana V, Henry SC, Schmidt EA, Daniell X, et al. Irgm1-deficient mice exhibit Paneth cell abnormalities and increased susceptibility to acute intestinal inflammation. Am J Physiol Gastrointest Liver Physiol. 2013;305(8):G573–84. https://doi.org/10.1152/ajpgi.00071.2013.
Mehto S, Jena KK, Nath P, Chauhan S, Kolapalli SP, Das SK, et al. The crohn’s disease risk factor IRGM limits NLRP3 inflammasome activation by impeding its assembly and by mediating its selective autophagy. Mol Cell. 2019;73:429-45.e7. https://doi.org/10.1016/j.molcel.2018.11.018.
Brest P, Lapaquette P, Souidi M, Lebrigand K, Cesaro A, Vouret-Craviari V, et al. A synonymous variant in IRGM alters a binding site for miR-196 and causes deregulation of IRGM-dependent xenophagy in Crohn’s disease. Nat Genet. 2011;43(3):242–5. https://doi.org/10.1038/ng.762.
Rai P, Janardhan KS, Meacham J, Madenspacher JH, Lin W-C, Karmaus PWF, et al. IRGM1 links mitochondrial quality control to autoimmunity. Nat Immunol. 2021;22(3):312–21. https://doi.org/10.1038/s41590-020-00859-0.
Azzam KM, Madenspacher JH, Cain DW, Lai L, Gowdy KM, Rai P, et al. Irgm1 coordinately regulates autoimmunity and host defense at select mucosal surfaces. JCI Insight. 2017;2(16): e91914. https://doi.org/10.1172/jci.insight.91914.
Roy S, Esmaeilniakooshkghazi A, Patnaik S, Wang Y, George SP, Ahrorov A, et al. Villin-1 and gelsolin regulate changes in actin dynamics that affect cell survival signaling pathways and intestinal inflammation. Gastroenterology. 2018;154(5):1405-20.e2. https://doi.org/10.1053/j.gastro.2017.12.016.
Fang S, Sun S, Cai H, Zou X, Wang S, Hao X, et al. <i>IRGM/Irgm1</i> facilitates macrophage apoptosis through ROS generation and MAPK signal transduction: <i>Irgm1</i><sup>+/-</sup> mice display increases atherosclerotic plaque stability. Theranostics. 2021;11(19):9358–75. https://doi.org/10.7150/thno.62797.
Fang S, Xu Y, Zhang Y, Tian J, Li J, Li Z, et al. Irgm1 promotes M1 but not M2 macrophage polarization in atherosclerosis pathogenesis and development. Atherosclerosis. 2016;251:282–90. https://doi.org/10.1016/j.atherosclerosis.2016.07.011.
Plummer M, de Martel C, Vignat J, Ferlay J, Bray F, Franceschi S. Global burden of cancers attributable to infections in 2012: a synthetic analysis. Lancet Glob Health. 2016;4(9):e609–16. https://doi.org/10.1016/S2214-109X(16)30143-7.
Lapaquette P, Bringer MA, Darfeuille-Michaud A. Defects in autophagy favour adherent-invasive Escherichia coli persistence within macrophages leading to increased pro-inflammatory response. Cell Microbiol. 2012;14(6):791–807. https://doi.org/10.1111/j.1462-5822.2012.01768.x.
Hansen MD, Johnsen IB, Stiberg KA, Sherstova T, Wakita T, Richard GM, et al. Hepatitis C virus triggers Golgi fragmentation and autophagy through the immunity-related GTPase M. Proc Natl Acad Sci. 2017;114(17):E3462. https://doi.org/10.1073/pnas.1616683114.
Castaño-Rodríguez N, Kaakoush NO, Goh KL, Fock KM, Mitchell HM. Autophagy in Helicobacter pylori infection and related gastric cancer. Helicobacter. 2015;20(5):353–69. https://doi.org/10.1111/hel.12211.
Taylor GA, Huang H-I, Fee BE, Youssef N, Jewell ML, Cantillana V, et al. Irgm1-deficiency leads to myeloid dysfunction in colon lamina propria and susceptibility to the intestinal pathogen citrobacter rodentium. PLoS Pathog. 2020;16(5):e1008553. https://doi.org/10.1371/journal.ppat.1008553.
Collazo CM, Yap GS, Sempowski GD, Lusby KC, Tessarollo L, Vande Woude GF, et al. Inactivation of LRG-47 and IRG-47 reveals a family of interferon gamma-inducible genes with essential, pathogen-specific roles in resistance to infection. J Exp Med. 2001;194(2):181–8. https://doi.org/10.1084/jem.194.2.181.
Coers J, Gondek DC, Olive AJ, Rohlfing A, Taylor GA, Starnbach MN. Compensatory T cell responses in IRG-deficient mice prevent sustained Chlamydia trachomatis infections. PLoS Pathog. 2011;7(6):e1001346. https://doi.org/10.1371/journal.ppat.1001346.
Yang D, Chen J, Zhang L, Cha Z, Han S, Shi W, et al. Mycobacterium leprae upregulates IRGM expression in monocytes and monocyte-derived macrophages. Inflammation. 2014;37(4):1028–34. https://doi.org/10.1007/s10753-014-9825-1.
Buisson A, Douadi C, Ouchchane L, Goutte M, Hugot J-P, Dubois A, et al. Macrophages inability to mediate adherent-invasive E coli. replication is linked to autophagy in crohn’s disease patients. Cells. 2019;8(11):1394.
Nath P, Chauhan NR, Jena KK, Datey A, Kumar ND, Mehto S, et al. Inhibition of IRGM establishes a robust antiviral immune state to restrict pathogenic viruses. EMBO Rep. 2021. https://doi.org/10.15252/embr.202152948.
Grégoire IP, Richetta C, Meyniel-Schicklin L, Borel S, Pradezynski F, Diaz O, et al. IRGM is a common target of RNA viruses that subvert the autophagy network. PLoS pathog. 2011;7(12):e1002422. https://doi.org/10.1371/journal.ppat.1002422.
Rosentul DC, Plantinga TS, Farcas M, Oosting M, Hamza OJM, Scott WK, et al. Role of autophagy genetic variants for the risk of Candida infections. Med Mycol. 2014;52:333–41. https://doi.org/10.1093/mmy/myt035.
Santiago HC, Feng CG, Bafica A, Roffe E, Arantes RM, Cheever A, et al. Mice deficient in LRG-47 display enhanced susceptibility to <em>Trypanosoma cruzi</em> infection associated with defective hemopoiesis and intracellular control of parasite growth. J Immunol. 2005;175(12):8165. https://doi.org/10.4049/jimmunol.175.12.8165.
Imhann F, Vich Vila A, Bonder MJ, Fu J, Gevers D, Visschedijk MC, et al. Interplay of host genetics and gut microbiota underlying the onset and clinical presentation of inflammatory bowel disease. Gut. 2018;67(1):108–19. https://doi.org/10.1136/gutjnl-2016-312135.
Burada F, Plantinga TS, Ioana M, Rosentul D, Angelescu C, Joosten LA, et al. IRGM gene polymorphisms and risk of gastric cancer. J Dig Dis. 2012;13(7):360–5. https://doi.org/10.1111/j.1751-2980.2012.00602.x.
Ge J, Li L, Jin Q, Liu YC, Zhao L, Song H-H. Functional IRGM polymorphism is associated with language impairment in glioma and upregulates cytokine expressions. Tumor Biol. 2014;35(8):8343–8. https://doi.org/10.1007/s13277-014-2091-x.
Santoni M, Piva F, De Giorgi UGO, Mosca A, Basso U, Santini D, et al. Autophagic gene polymorphisms in liquid biopsies and outcome of patients with metastatic clear cell renal cell carcinoma. Anticancer Res. 2018;38(10):5773. https://doi.org/10.21873/anticanres.12916.
McCarroll SA, Huett A, Kuballa P, Chilewski SD, Landry A, Goyette P, et al. Deletion polymorphism upstream of IRGM associated with altered IRGM expression and Crohn’s disease. Nat Genet. 2008;40(9):1107–12. https://doi.org/10.1038/ng.215.
Li Y, Feng ST, Yao Y, Yang L, Xing Y, Wang Y, et al. Correlation between IRGM genetic polymorphisms and Crohn’s disease risk: a meta-analysis of case-control studies. Genet Mol Res. 2014;13(4):10741–53. https://doi.org/10.4238/2014.December.18.15.
Lu XC, Tao Y, Wu C, Zhao PL, Li K, Zheng JY, et al. Association between variants of the autophagy related gene—IRGM and susceptibility to crohn’s disease and ulcerative colitis: a meta-analysis. PLoS ONE. 2013;8(11):e80602. https://doi.org/10.1371/journal.pone.0080602.
Ajayi TA, Innes CL, Grimm SA, Rai P, Finethy R, Coers J, et al. Crohn’s disease IRGM risk alleles are associated with altered gene expression in human tissues. Am J Physiol Gastrointest Liver Physiol. 2018. https://doi.org/10.1152/ajpgi.00196.2018.-Crohn.
Latiano A, Palmieri O, Cucchiara S, Castro M, D’Incà R, Guariso G, et al. Polymorphism of the IRGM gene might predispose to fistulizing behavior in Crohn’s disease. Am J Gastroenterol. 2009;104(1):110–6. https://doi.org/10.1038/ajg.2008.3.
Sehgal R, Berg A, Polinski JI, Hegarty JP, Lin Z, McKenna KJ, et al. Mutations in IRGM are associated with more frequent need for surgery in patients with ileocolonic Crohn’s disease. Dis Colon Rectum. 2012;55(2):115–21. https://doi.org/10.1097/DCR.0b013e31823ccea8.
Kline BP, Weaver T, Brinton DL Jr, Deiling S, Yochum GS, Berg AS, et al. Clinical and genetic factors associated with complications after crohn’s ileocolectomy. Dis Colon Rectum. 2020;63(3):357–64. https://doi.org/10.1097/dcr.0000000000001574.
Xie H, Li C, Zhang M, Zhong N, Chen L. Association between IRGM polymorphisms and tuberculosis risk: a meta-analysis. Medicine (Baltimore). 2017;96(43):e8189. https://doi.org/10.1097/md.0000000000008189.
Intemann CD, Thye T, Niemann S, Browne ENL, Amanua Chinbuah M, Enimil A, et al. Autophagy gene variant IRGM −261T contributes to protection from tuberculosis caused by mycobacterium tuberculosis but not by M africanum strains. PLOS Pathog. 2009;5(9):e1000577. https://doi.org/10.1371/journal.ppat.1000577.
Simon TG, Deng X, Liu C-T, Chung RT, Long MT. The immunity-related GTPase M rs13361189 variant does not increase the risk for prevalent or incident steatosis; results from the framingham heart study. Liver Int. 2019;39(6):1022–6. https://doi.org/10.1111/liv.14039.
Simon TG, Van Der Sloot KWJ, Chin SB, Joshi AD, Lochhead P, Ananthakrishnan AN, et al. IRGM gene variants modify the relationship between visceral adipose tissue and NAFLD in patients with crohn’s disease. Inflamm Bowel Dis. 2018;24(10):2247–57. https://doi.org/10.1093/ibd/izy128.
Yao Q-M, Zhu Y-F, Wang W, Song Z-Y, Shao X-Q, Li L, et al. Polymorphisms in autophagy-related gene IRGM are associated with susceptibility to autoimmune thyroid diseases. Biomed Res Int. 2018. https://doi.org/10.1155/2018/7959707.
Yang D, Chen J, Shi C, Jing Z, Song N. Autophagy gene polymorphism is associated with susceptibility to leprosy by affecting inflammatory cytokines. Inflammation. 2014;37(2):593–8. https://doi.org/10.1007/s10753-013-9773-1.
Folwaczny M, Tsekeri E, Glas J. A haplotypic variant at the IRGM locus and rs11747270 are related to the susceptibility for chronic periodontitis. Inflamm Res. 2018;67(2):129–38. https://doi.org/10.1007/s00011-017-1101-z.
Xia Q, Wang M, Yang X, Li X, Zhang X, Xu S, et al. Autophagy-related IRGM genes confer susceptibility to ankylosing spondylitis in a Chinese female population: a case-control study. Genes Immun. 2017;18(1):42–7. https://doi.org/10.1038/gene.2016.48.
Palomino-Morales RJ, Oliver J, Gómez-García M, López-Nevot MA, Rodrigo L, Nieto A, et al. Association of ATG16L1 and IRGM genes polymorphisms with inflammatory bowel disease: a meta-analysis approach. Genes Immun. 2009;10:356–64. https://doi.org/10.1038/gene.2009.25.
Kimura T, Watanabe E, Sakamoto T, Takasu O, Ikeda T, Ikeda K, et al. Autophagy-related IRGM polymorphism is associated with mortality of patients with severe sepsis. PLoS ONE. 2014;9(3):e91522. https://doi.org/10.1371/journal.pone.0091522.
Kee BP, Ng JG, Ng CC, Hilmi I, Goh KL, Chua KH. Genetic polymorphisms of ATG16L1 and IRGM genes in Malaysian patients with Crohn’s disease. J Dig Dis. 2020;21(1):29–37. https://doi.org/10.1111/1751-2980.12829.
Teimoori-Toolabi L, Samadpoor S, Mehrtash A, Ghadir M, Vahedi H. Among autophagy genes, ATG16L1 but not IRGM is associated with Crohn’s disease in Iranians. Gene. 2018;675:176–84. https://doi.org/10.1016/j.gene.2018.06.074.
Pranculienė G, Steponaitienė R, Skiecevičienė J, Kučinskienė R, Kiudelis G, Adamonis K, et al. Associations between NOD2, IRGM and ORMDL3 polymorphisms and pediatric-onset inflammatory bowel disease in the Lithuanian population. Medicina (Kaunas). 2016;52(6):325–30. https://doi.org/10.1016/j.medici.2016.11.006.
Na SY, Park SS, Seo JK. Genetic polymorphisms in autophagy-associated genes in korean children with early-onset crohn disease. J Pediatr Gastroenterol Nutr. 2015;61(3):285–91. https://doi.org/10.1097/mpg.0000000000000796.
Lin YC, Chang PF, Lin HF, Liu K, Chang MH, Ni YH. Variants in the autophagy-related gene IRGM confer susceptibility to non-alcoholic fatty liver disease by modulating lipophagy. J Hepatol. 2016;65(6):1209–16. https://doi.org/10.1016/j.jhep.2016.06.029.
Lu Y, Li Q, Peng J, Zhu Y, Wang F, Wang C, et al. Association of autophagy-related IRGM polymorphisms with latent versus active tuberculosis infection in a Chinese population. Tuberculosis. 2016;97:47–51. https://doi.org/10.1016/j.tube.2016.01.001.
Yuan L, Ke Z, Ma J, Guo Y, Li Y. IRGM gene polymorphisms and haplotypes associate with susceptibility of pulmonary tuberculosis in Chinese hubei han population. Tuberculosis (Edinb). 2016;96:58–64. https://doi.org/10.1016/j.tube.2015.10.014.
Song JH, Kim SY, Chung KS, Moon CM, Kim SW, Kim EY, et al. Association between genetic variants in the IRGM gene and tuberculosis in a Korean population. Infection. 2014;42(4):655–60. https://doi.org/10.1007/s15010-014-0604-6.
Bahari G, Hashemi M, Taheri M, Naderi M, Eskandari-Nasab E, Atabaki M. Association of irgm polymorphisms and susceptibility to pulmonary tuberculosis in Zahedan Southeast Iran. Scientific World J. 2012;2012:950801. https://doi.org/10.1100/2012/950801.
King KY, Lew JD, Ha NP, Lin JS, Ma X, Graviss EA, et al. Polymorphic allele of human IRGM1 is associated with susceptibility to tuberculosis in African Americans. PLoS ONE. 2011;6(1):e16317. https://doi.org/10.1371/journal.pone.0016317.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. NCR is supported by a Cancer Institute NSW Early Career Fellowship (2019/ECF1082) and a UNSW Scientia Fellowship.
Author information
Authors and Affiliations
Contributions
Conceptualisation and design: NCR. Literature search: ABG and DK. Draft manuscript: ABG and DK. Critically revised the work and editing: NCR. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interests
The authors have no conflict of interests to declare that are relevant to the content of this article.
Additional information
Responsible Editor: John Di Battista.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Goswami, A.B., Karadarević, D. & Castaño-Rodríguez, N. Immunity-related GTPase IRGM at the intersection of autophagy, inflammation, and tumorigenesis. Inflamm. Res. 71, 785–795 (2022). https://doi.org/10.1007/s00011-022-01595-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-022-01595-x