Abstract
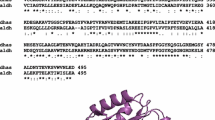
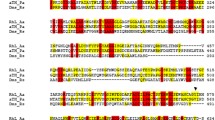
Unlike the parent wild-type strain, theKlebsiella pneumoniae mutant strain MAO4 has a 4-HBA+ phenotype. The capacity of this mutant to take up and metabolize 4-hydroxybenzoate (4-HBA) relies on the expression of a permease and an NADPH-linked monooxygenase (4-HBA-3-hydroxylase). Both enzymes are normally expressed at basal levels, and only the presence of 4-HBA in the media enhances their activities. Strikingly, when theAcinetobacter calcoaceticus pobA gene encoding 4-hydroxybenzoate-3-hydroxylase was expressed in hydroxybenzoateK. pneumoniae wild-type, the bacteria were unable to grow on 4-HBA, suggesting that the main difference between the wild-type and the mutant strain is the capability of the latter to take up 4-HBA. 4-HBA-3-hydroxylase was purified to homogeneity by affinity, gel-filtration, and anion-exchange chromatography. The native enzyme, which appeared to be a dimer of identical subunits, had an apparent molecular mass of 80 kDa and a pI of 4.6. Steady-state kinetics were analyzed; the initial velocity patterns were consistent with a concerted substitution mechanism. The purified enzyme had 362 amino acid residues, and a tyrosine seemed to be involved in substrate activation.
Similar content being viewed by others
Abbreviations
- DPC :
-
Diethylpyrocarbonate
References
Allende JL, Suárez M, Gallego M, Garrido-Pertierra A (1993) 4-Hydroxybenzoate uptake inKlebsiella pneumoniae is driven by electrical potential. Arch Biochem Biophys 300:142–147
Anderson JJ, Dagley S (1980) Catabolism of aromatic acids inTrichosporon cutaneum. J Bacteriol 143:534–543
Averhoff B, Gregg-Jolly L, Elsemore D, Ornston LN (1992) Genetic analysis of supraoperonic clustering by use of natural transformation inAcinetobacter calcoaceticus. J Bacteriol 174: 200–20
Blanke SR, Hager P (1990) Chemical modification of chloroperoxidase with diethyl pyrocarbonate. J Biol Chem 265:12454–12461
Dalziel K (1969) The interpretation of kinetic data for enzyme catalysed reactions involving three substrates. Biochem J. 114: 547–556.
Deschamps AM, Richard C, Lebeault JM (1983) Bacteriology and nutrition of environmental strains ofKlebsiella pneumoniae involved in wood and bark decay. Ann Microbiol (Inst Pasteur) 134:89–196
DiMarco AA, Averhoff BA, Kim EE, Ornston LN (1993) Evolutionary divergence ofpobA, the structural gene encodingp-hydroxybenzoate hydroxylase in anAcinetobacter calcoaceticus strain well-suited for genetic analysis. Gene 125:25–33
Dixon RA (1984) The genetic complexity of nitrogen fixation. J Gen Microbiol 130:2745–2755
Edman P, Begg G (1967) A protein sequenator. Eur J Biochem 1: 80–91
Entsch B, Ballou DP (1989) Purification, properties and oxygen reactivity ofp-hydroxybenzoate hydroxylase fromPseudomonas aeruginosa. Biochim Biophys Acta 999:313–332
Entsch B, Nan Y, Weaich K, Scott KF (1988) Sequence and organization ofpobA, the gene coding forp-hydroxybenzoate hydroxylase, an inducible enzyme fromPseudomonas aeruginosa. Gene 7:279–29
Entsch B, Palfey BA, Ballou DP, Massey V (1991) Catalytic function of tyrosine residues inpara-hydroxybenzoate hydroxylase as determined by the study of site-directed mutants. J Biol Chem 266:17341–17349
Fuji T, Kaneda T (1985) Purification and properties of NADH/NADPH dependentp-hydroxybenzoate hydroxylase fromCorynebacterium cyclohexanicum. Eur J Biochem 147:97–104
Groenewegen PEJ, Driessen AJM, Konings WN, De Bont JAM (1990) Energy-dependent uptake of 4-chlorobenzoate inCoryneform bacterium NTB-1. J Bacteriol 172:419–423
Hanahan D (1983) Studies on transformation ofEscherichia coli with plasmid DNA. J Mol Biol 166:557–561
Harayama S, Kok M, Neidle EL (1992) Functional and evolutionary relationships among diverse oxygenases. Annu Rev Microbiol 46:565–560
Hareland WA, Crawford RL, Chapman PJ, Dagley S (1975) Metabolic function and properties of 4-hydroxyphenylacetic acid 1-hydroxylase fromPseudomonas acidovorans. J Bacteriol 121: 272–285
Hofsteenge J, Vereijken JM, Weijer WJ, Beintema JJ, Wierenga RK, Drenth J (1983) Primary and tertiary structure studies ofp-hydroxybenzoate hydroxylase fromPseudomonas fluorescens. Eur J Biochem 113:141–150
Husain M, Massey V (1979) Kinetic studies on the reaction ofp-hydroxybenzoate hydroxylase. J Biol Chem 254:6657–6666
Marshall T, Williams KM (1986) Phenol enhancement of the Coomasie Blue protein dye binding assay. Biotechniques 4: 308–10
Müller F, Voordouw G, Van Berkel WJH, Steennis PJ, Visser S, Van Rooijen PJ (1979) A study ofp-hydroxybenzoate hydroxylase fromPseudomonas fluorescens. Improved purification, relative molecular mass and amino acid composition. Eur J Biochem 101:235–244
Nielsen BL, Brown LR (1984) The basis for colored silver-protein complex formation in stained polyacrylamide gels. Anal Biochem 141:311–315
Spencer RL, Wold F (1969) A new convenient method for estimation of total cystine-cysteine in proteins. Anal Biochem 32: 185–190
Streicher S, Gurney E, Valentine RC (1971) Transduction of the nitrogen-fixation genes inKlebsiella pneumoniae Proc Natl Acad Sci USA 68:1174–1177
Suárez M, Gibello A, Allende JL, Martín M, Ferrer E, Garrido-Pertierra A (1991) Degradation of 3- and 4-hydroxybenzoate byKlebsiella pneumoniae Appl Microbiol Biotechnol 34:677–682
Takeuchi M, Asano N, Kaneda J, Masui K (1986) Chemical modification of histidine residues in 3-ketovalidoxylamine A C-N lyase. J Biochem 99:1571–1577
Van Berkel W, Westphal A, Eschrich K, Eppink M, De Kok A (1992) Substitution of Arg214 at the substrate-binding site ofp-hydroxybenzoate hydroxylase fromPseudomonas fluorescens. Eur J Biochem 210:411–419
Warburg O, Christian W (1972) Biochemical procedures. In: Dawson RMC, Elliot WH, Jones KM (eds) Data for biochemical research. Oxford University Press, London, pp 537–539
Wilson M (1977) Modification of histidine residues in proteins by diethylpyrocarbonate. In: Hirs CHW, Timasheff SN (eds) Methods in enzymology, vol 47. Academic Press, New York, pp 431–432
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Suárez, M., Martín, M., Ferrer, E. et al. Purification and characterization of 4-hydroxybenzoate 3-hydroxylase from aKlebsiella pneumoniae mutant strain. Arch. Microbiol. 164, 70–77 (1995). https://doi.org/10.1007/BF02568737
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02568737