Abstract
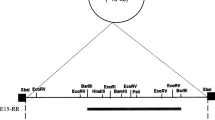
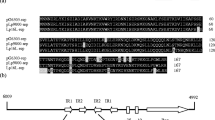
The replication region of pUCL22, the lactose-protease plasmid ofLactococcus lactis subsp.lactis (Lc. lactis) CNRZ270, was isolated on an 18-kbBamHI fragment, by cloning into pUCB300, anEscherichia coli vector encoding Emr, and selecting for Emr inLc. lactis MG1614. Subcloning and deletion analysis localized the replication region, namedRep22, on a 2.3-kb fragment. Replicons based on this region followed a theta-type mechanism of replication inLc. lactis. An internal 1251-bpDraI fragment ofRep22 used as a probe hybridized with numerous plasmids in bacteria from the generaLactococcus, Lactobacillus, andLeuconostoc. In some strains, two or three coresident plasmids hybridized with the probe in stringent conditions. It appears, therefore, that this family of theta-type replicons is widely distributed in lactic acid bacteria and contains several incompatibility groups.
Similar content being viewed by others
Literature Cited
Anderson DG, McKay LL (1983) Simple and rapid method for isolating large plasmid DNa from lactic streptococci. Appl Environ Microbiol 46:549–552
Anderson DG, McKay LL (1977) Plasmids, loss of lactose metabolism and appearance of partial and full lactosefermenting revertants inStreptococcus cremoris B1. J Bacteriol 129:367–377
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res 7:1513–1519
Brantl S, Behnke D (1992) The amount of RepR protein determines the copy number of plasmid pIP501 inBacillus subtilis. J Bacteriol 174:5475–5478
Brewer BJ, Fangman WL (1987) The localization of replication origins on ARS plasmids inS. cerevisiae. Cell 51:463–471
Bruand C, Ehrlich SD, Jannière L (1991) Unidirectional theta replication of the structurally stableEnterococcus faecalis plasmid pAMβ1. EMBO J 10:2171–2177
Courvalin P, Carlier C (1986) Transposable multiple antibiotic resistance inStreptococcus pneumoniae. Mol Gen Genet 205:291–297
de Man JC, Rogosa M, Sharpe ME (1960) A medium for the cultivation of lactobacilli. J Appl Bacteriol 23:130–135
de Vos WM (1986) Gene cloning in lactic streptococci. Neth Milk Dairy J 40:141–154
Efstathiou JD, McKay LL (1976) Plasmids inStreptococcus lactis: evidence that lactose metabolism and proteinase activity are plasmid linked. Appl. Environ Microbiol 32:38–44
Gasson MJ (1983) Plasmid complements ofStreptococcus lactis NCDO712 and other streptococci after protoplast-induced curing. J Bacteriol 154:1–9
Hintermann G, Fischer HM, Crameri R, Hutter R (1981) Simple procedure for distinguishing CCC, OC and L forms of plasmid DNA by agarose gel electrophoresis. Plasmid 5:371–373
Holo H, Nes IF (1989) High-frequency transformation, by electroporation, ofLactococcus lactis subsp.cremoris grown with glycine in osmotically stabilized media. Appl Environ Microbiol 55:3119–3123
Kempler GM, McKay LL (1979) Genetic evidence for plasmid-linked lactose metabolism inStreptococcus lactis subsp.diacetylactis. Appl Environ Microbiol 37:1041–1043
Kuhl SA, Larsen LD, McKay LL (1979) Plasmid profile of lactose negatives and proteinase deficient mutants ofS. lactis C10, ML3 and M18. Appl Environ Microbiol 37:1193–1195
Mandel M, Higa A (1970) Calcium-dependent bacteriophage DNA infection. J Mol Biol 53:159–162
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory
Novel M, Huang DC, Novel G (1988) Transposition ofStreptococcus lactis subsp.lactis Z270 lactose plasmide to pVA797: demonstration of an insertion sequence and its relationship to an inverted repeat sequence isolated by selfannealing. Biochimie 70:543–551
Novick RP (1987) Plasmid incompatibility. Microbiol Rev 51:381–395
Polzin KM, McKay LL (1991) Identification, DNA sequence, and distribution of IS981, a new high-copy-number insertion sequence in Lactococci. Appl Environ Microbiol 57:734–743
Ramos, P, Novel M, Lemosquet M, Novel G (1983) Fragmentation du plasmide lactose-protéase chez des dérivés lactose-négatifs deStreptococcus lactis etS. lactis ssp.diacetylactis. Ann. Microbiol. (Inst. Pasteur) 134B:387–399
Southern EM (1975) Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol 98:503–517
te Riele H, Michel B, Ehrlich SD (1986) Are single-stranded circles intermediates in plasmid DNA replication? EMBO J 5:631–637
Terzaghi BE, Sandine WE (1975) Improved medium for lactic streptococci and their bacteriophages. Appl Environ Microbiol 29:807–813
Trieu-Cuot P, Poyart-Salmero C, Carlier C, Courvalin P (1990) Nucleotide sequence of the erythromycin resistance gene of the conjugative transposon Tn1545. Nucleic Acids Res 18:3660
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–119
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Frère, J., Benachour, A., Novel, M. et al. Identification of the theta-type minimal replicon of theLactococcus lactis subsp.lactis CNRZ270 lactose protease plasmid pUCL22. Current Microbiology 27, 97–102 (1993). https://doi.org/10.1007/BF01570865
Issue Date:
DOI: https://doi.org/10.1007/BF01570865