Abstract
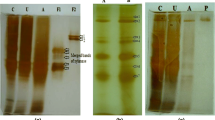
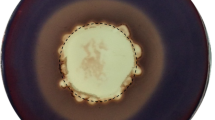
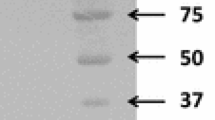
Spirochetes capable of degrading xylan or cellulose have not been commonly isolated, nor have their polysaccharolytic activities been characterized.Spirochaeta thermophila strain RI 19.B1 is xylanolytic and grows well at 65°C with oatspelt (OX), birchwood (BX), corncob (CCX-A) xylans, or glucuronoxylan (MGX) as the energy source. All xylans were extensively degraded and utilized during growth. About 72–82% of the initial hexuronic acids and 57–79% of initial pentoses disappeared during growth.S. thermophila possessed xylanase, xylosidase, and arabinofuranosidase enzyme activities. Low levels of these activities were detected with growth on glucose, but high expression of these activities occurred during growth on OX. All three activities were cell-associated and were more stable in cells than cell extracts. Xylan-degrading activities were measured with cells or cell extracts exposed (60 min) to a variety of temperatures (65°–85°C) and pHs (5.0–8.0). More than 50% loss of activities occurred at temperatures above 75°C. Although pH stability was affected by buffer, the optimal range was pH 6.5–7.5. These temperature and pH profiles for xylan-degrading activities coincide with those found for the growth ofS. thermophila.
Similar content being viewed by others
Literature Cited
Aksenova HY, Rainey FA, Jassen PH, Zavarzin GA, Morgan HW (1992)Spirochaeta thermophila sp. nov., an obligately anaerobic, polysaccharolytic, extremely thermophilic bacterium. Int J Syst Bacteriol 42:175–177
Ashwell G (1957) Colorimetric analysis of sugars. Methods Enzymol 3:73–105
Borneman WC, Ljungdahl LG, Hartley RD, Akin DE (1992) Purification and partial characterization of two feruloyl esterases from the anaerobic fungus Neocallimastix strain MC-2. Appl Environ Microbiol 58:3762–3766
Bradford MM (1976) A rapid and sensitive method for quantitation of microgram quantities of proteins utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bryant MP (1952) The isolation and characteristics of a spirochete from the bovine rumen. J Bacteriol 64:325–335
Cwyk WM, Canale-Parola E (1979)Treponema succinifaciens sp. nov.m an anaerobic spirochete from the swine intestine. Arch Microbiol 122:231–239
Hespell RB (1992) Fermentation of xylans byButyrivibrio fibrisolvens andThermoanaerobacter strain B6a: utilization of uronic acids and xylanolyytic activities. Curr Microbiol 25:189–195
Hespell RB, O'Bryan-Shah PJ (1988) Esterase activities inButyrivibrio fibrisolvens strains. Appl Environ Microbiol 54:1917–1922
Hespell RB, Whitehead TR (1990) Physiology and genetics of xylan degradation by gastrointestinal tract bacteria. J Dairy Sci 73:3013–3022
Hespell RB, Wolf R, Bothast RJ (1988) Fermentation of xylans byButyrivibrio fibrisolvens and other ruminal bacterial species. Appl Environ Microbiol 53:2849–2853
Janssen PH, Morgan HW (1992) Glucose metabolism bySpirochaeta thermophila. J Bacteriol 174:2449–2453
McDermid KP, Forsberg CW, MacKenzie CR (1990) Purification and properties of an acetylxylan esterase fromFibrobacter succinogenes S85. Appl Environ Microbiol 56:3805–3810
Paster BJ, Canale-Parola E (1982) Physiological diversity of rumen spirochetes. Appl Environ Microbiol 43:686–693
Paster BJ, Canale-Parola E (1985)Treponema saccharophilum sp. nov., a large pectinolytic spirochete from the bovine rumen. Appl Environ Microbiol 50:212–219
Paster BJ, Stackebrandt E, Hespell RB, Hahn CM, Woese CR (1984) The phylogeny of the spirochetes. Syst Appl Microbiol 5:337–351
Paster BJ, Dehirst FE, Weisburg WG, Tordoff LA, Fraser GJ, Hespell RB, Stanton TB, Zablen L, Mandelico L, Woese CR (1991) Phylogenetic analysis of the spirochetes. J Bacteriol 173:6101–6109
Rainey FA, Jansson PH, Wild DJC, Morgan HW (1991) Isolation and characterization of an obligately anaerobic, polysaccharolytic, extremely thermophilic member of the genusSpirochaeta. Arch Microbiol 155:396–401
Salyers AA, Balascio JR, Palmer JK (1982) Breakdown of xylan by enzymes from human colonic bacteria. J Food Biochem 6:39–55
Schneider WC (1957) Determination of nucleic acids by pentose analysis. Methods Enzymol 3:374–381
Scott RW (1979) Colorimetric determination of hexuronic acids in plant materials. Anal Chem 51:936–941
Shao W, Wiegel J (1992) Purification and characterization of a thermostable β-xylosidase fromThermoanaerobacter ethanolicus. J Bacteriol 174:5848–5853
Stack RJ (1988) Neutral sugar composition of extracellular polysaccharides produced by strains ofButyrivibrio fibrisolvens. Appl Environ Microbiol 54:878–883
Stanton TB, Canale-Parola E (1980)Treponema bryantii sp. nov., a spirochete that interacts with cellulolytic bacteria. Arch Microbiol 127:145–156
Thomson JA (1993) Molecular biology of xylan degradation. FEMS Microbiol Rev 104:65–82
Whitehead TR, Hespell RB (1990) Heterologus expression of theBacteroides ruminicola xylanase gene inBacteroides fragilis andBacteroides uniformis. FEMS Microbiol Lett 66:61–66
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hespell, R.B. Xylanolytic activities ofSpirochaeta thermophila . Current Microbiology 29, 343–347 (1994). https://doi.org/10.1007/BF01570227
Issue Date:
DOI: https://doi.org/10.1007/BF01570227