Abstract
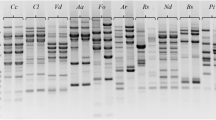
DNA polymorphisms generated by the random amplified polymorphic DNA polymerase chain reaction (RAPD PCR) were used to analyse 41 isolates investigated in the EuropeanFusarium sambucinum Project (EFSP). Employing ten arbitrary (10-mer) oligonucleotides and simple repeat sequences (M13, (GACA)4) as single primers, informative banding patterns typical for identifying European populations ofFusarium sambucinum Fuckel s. str.,F. torulosum (Berk. & Curt.) Nirenberg andF. venenatum Nirenberg were obtained.
Similar content being viewed by others
References
Saiki RK, Scharf S, Faloona F, Mullis KB, Horn GT, Erlich HA, Amheim H. Enzymatic amplification of β-globin genomic sequences and restriction site analysis of diagnosis of sickle cell anemia. Science 1985; 230: 1350–1354.
Mullis KB, Faloona FA. Specific synthesis of DNA in vitro via a polymerase-catalysed chain reaction. Methods Enzymol 1987; 155: 335–350.
Saiki RK, Gelfand DH, Stoffel S, Scharf SJ, Higuchi R, Horn GT, Mullis KB, Ehrlich HA. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 1988; 239: 487–491.
Welsh J, McClelland M. Fingerprinting genomes using PCR with arbitrary primers. Nucl Acids Res 1990; 18: 7213–7218.
Williams JGK, Kubelik AR, Livak J, Rafalski JA, Tingey SV. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucl Acids Res 1990; 18: 6531–6535.
Welsh J, Petersen C, McClelland M. Polymorphisms generated by arbitrarily primed PCR in the mouse: Application to strain identification and genetic mapping. Nucl Acids Res 1991; 19: 303–306.
Caetano-Anollés G, Bassam BJ, Gresshoff PM. DNA amplification fingerprinting using very short arbitrary oligonucleotide primers. Bio/Technol 1991; 9: 553–557.
Zilberstein A, Pnini-Cohen S, Eyal Z, Shuster S, Hillel J, Lavi U. Application of DNA fingerprinting for dctecting genetical variation among isolates of the wheat pathogenMycosphaerella graminicola. In: Chet I, ed. Biotechnology in plant diseases control. Wiley-Liss, 1993: 341–353.
Schönian G, Meusel O, Tietz H-J, Meyer W, Gräser Y, Tausch I, Presber W, Mitchel TG. Identification of clinical strains ofCandida albicans by DNA fingerprinting with the polymerase chain reaction. Mycoses 1993; 36: 171–179.
Caetano-Anollés G, Bassam BJ, Gresshoff PM. DNA amplification fingerprinting: A strategy for genome analysis. Plant Mol Biol Rep 1991; 9: 294–307.
Crowhurst RN, Hawthorne BT, Rikkerink EHA, Templeton MD. Differentiation ofFusarium solani f. sp.cucurbitae raccs 1 and 2 by random amplified polymorphic DNA. Curr Genet 1991; 20: 391–396.
Hering O: Rassendifferenzierung beiFusarium oxysporum f. sp.vasinfectum mittels RAPD-PCR (Abstract). Phytomedizin 1993; 23: 51–52.
Hamelin RC, Oulette GB, Bernier L. Identification ofGremmeniella abietina races with random amplified polymorphic DNA Markers. Appl Environ Microbiol 1993; 59: 1752–1755.
Goodwin PH, Annis SL. Rapid identification of genetic variation and pathotype ofLeptosphaeria maculans by random amplified polymorphic DNA assay. Appl Environ Microbiol 1991; 57: 2482–2486.
Schäfer C, Wöstemeyer J. Random primer dependent PCR differentiates aggressive from non-aggressive isolates of the oilseed rape pathogenPhoma lingam (Leptosphaeria maculans). J Phytopathol 1992; 136: 124–136.
Sreenivasaprasad S, Brown AE, Mills PR. DNA sequence variation and interrelationships amongCollectotrichum species causing strawberry anthracnose. Physiol Mol Plant Pathol 1992; 41: 265–281.
Bayman P, Cotty PJ. Genetic diversity inAspergillus flavus: Association with aflatoxin production and morphology. Can J Bot 1993; 71: 23–31.
Leung H, Loomis P, Shi Y. Differentiation of the wheat bunt fungi by random amplification of polymorphic DNA (Abstract). Phytopathology 1991; 81: 1190.
Nirenberg HI. Morphological differentiation ofFusarium sambucinum Fuckel s. str.,F. torulosum (Berk & Curt) Nirenberg comb. nov. andF. venenatum Nirenberg sp. nov. Mycopathologia 1995; 129: 131–141 (this issue).
Nirenberg HI. Recent advances in the taxonomy ofFusarium. Stud Mycol 1990; 32: 91–101.
Huey B, Hall J. Hypervariable DNA fingerprinting inEscherichia coli: Minisatellite probes from bacteriophage M13. J Bacteriol 1989; 171: 2528–2532.
Ali S, Müller CR, Epplen JT. DNA fingerprinting by oligonucleotide probes specific for simple repeats. Hum Genet 1986; 74: 239–243.
Weising K, Weigand F, Driesel AJ, Kahl G, Zischler H, Epplen JT. Polymorphic simple GATA/GACA repeats in plant genomes. Nucl Acids Res 1989; 17: 10128.
Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: A laboratory manual, 2. ed. New York: Cold Spring Harbor Laboratory, 1989.
Booth C. The genus Fusarium. Kew: Commonwealth Mycological Institute, 1971: 168, 189.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hering, O., Nirenberg, H.I. Differentiation ofFusarium sambucinum Fuckel sensu lato and related species by RAPD PCR. Mycopathologia 129, 159–164 (1995). https://doi.org/10.1007/BF01103341
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01103341