Abstract
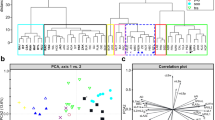
Genetic relatedness inChaenomeles was studied by RAPD analysis in 42 plants representing accessions of three wild species and one hybrid taxon. Amplification with 17 primers yielded a total of 156 polymorphic RAPD bands. Estimates of genetic relatedness suggest thatC. cathayensis andC. japonica are the most distantly related species, and that the former is comparatively homogeneous.Chaenomeles speciosa, which may have arisen through hybridization betweenC. cathayensis andC. japonica, takes an intermediate position between these two species. Analysis of diagnostic bands demonstrate that neitherC. speciosa norC. ×superba has any bands that do not occur in at least one ofC. cathayensis orC. japonica. Moreover,C. speciosa and the partly overlapping taxonC. ×superba are comparatively heterogeneous, which is also in accordance with a hybrid origin. Intraspecific variation was studied mainly inC. japonica; plants obtained from different sources of material formed well separated groups in the cluster analysis.
Similar content being viewed by others
References
Debener, T., Bartels, C., Mattiesch, L., 1996: RAPD analysis of genetic variation between a group of rose cultivars and selected wild rose species. — Molec. Breed.2: 321–327.
Doyle, J. J., Doyle, J. L., 1987: A rapid DNA isolation procedure for small quantities of fresh leaf tissue. — Phytochem. Bull.19: 11–15.
Dunemann, F., 1994: Molecular classification ofMalus with RAPD markers. — InSchmidt, H., Kellerhals, M., (Eds): Progress in temperate fruit breeding, pp. 295–300. — Dordrecht: Kluwer.
Gogorcena, Y., Parfitt, D. E., 1994: Evaluation of RAPD marker consistency for detection of polymorphism in apricot. — Sci. Hort.59: 163–167.
Graham, J., McNicol, R. J., 1995: An examination of the ability of RAPD markers to determine the relationships within and betweenRubus species. — Theor. Appl. Genet.90: 1128–1132.
Halldén, C., Hansen, M., Nilsson, N.-O., Hjerdin, A., Säll, T., 1996: Competition as a source of errors in RAPD analysis. — Theor. Appl. Genet.93: 1185–1192.
Harris, S. A., 1995: Systematics and randomly amplified polymorphic DNA in the genusLeucaena (Leguminosae, Mimosoideae). — Pl. Syst. Evol.197: 195–208.
Kuhnlein, U., Zadworny, D., Dawe, Y., Fairfull, R. W., Gavora, J. S., 1990: Assessment of inbreeding by DNA fingerprinting: development of a calibration curve using defined strains of chickens. — Genetics125: 161–165.
Millan, T., Osuna, F. F., Cobos, S., Torres, A. M., Cubero, J. I., 1996: Using RAPDs to study phylogenetic relationships inRosa. — Theor. Appl. Genet.92: 273–277.
Norusis, M. J., 1990: SPSS® base system user's guide. — Chicago: SPSS Inc.
Novy, R. G., Vorsa, N., 1996: Evidence for RAPD heteroduplex formation in cranberry: implications for pedigree and genetic relatedness studies and a source for co-dominant RAPD markers. — Theor. Appl. Genet.92: 840–849.
Phipps, J. B., Robertson, K. R., Smith, P. G., Rohrer, J. R., 1990: A checklist of the subfamilyMaloideae (Rosaceae). — Canad. J. Bot.68: 2209–2269.
Pitra, C., Curson, A., Nürnberg, P., Krawczak, M., Brown, S., 1996: An assessment of inbreeding in Asian wild horse (Equus przewalskii Poliakov 1881) populations using DNA fingerprinting. — Arch. Tierzücht. Dummerstort.39: 589–596.
Rumpunen, K., 1996:Chaenomeles — a novel source for pectic substances, organic acids and aromatic compounds. — InGarcia-Viguear, C., Cataner, M., Gil, M. I., Ferreres, F., Tomas-Barberan, F. A., (Eds): Current trends in fruit and vegetable phytochemistry, pp. 271–176. — Madrid: Consejo Superior Investigaciones Cientificas.
-Kviklys, D., Kaufmane, E., Garkava, L., 1998: BreedingChaenomeles — a new aromatic fruit crop. — Acta Horticult. (in press).
Saghai-Maroof, M. A., Soliman, K. M., Jorgensen, R. A., Allard, R. W., 1984: Ribosomal DNA spacer-polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. — Proc. Natl. Acad. Sci. USA81: 8014–8018.
Skroch, P., Nienhuis, J., 1995: Impact of scoring error and reproducibility of RAPD data on RAPD based estimates of genetic distance. — Theor. Appl. Genet.91: 1086–1091.
Spooner, D. M., Tivang, J., Nienhuis, J., Miller, J. T., Douches, D. S., Contreras, M. A., 1996: Comparison of four molecular markers in measuring relationships among the wild potato relativesSolanum sectionEutuberosum (subgenusPotatoe). — Theor. Appl. Genet.92: 532–540.
Thormann, C. E., Ferreira, M. E., Camargo, L. E. A., Tivang, J. G., Osborn, T. C., 1994: Comparison of RFLP and RAPD markers to estimating genetic relationships within and among cruciferous species. — Theor. Appl. Genet.88: 973–980.
Tivang, J., Skroch, P. W., Nienhuis, J., De Vos, N., 1996: Randomly amplified polymorphic DNA (RAPD) variation among and within artichoke (Cynara scolymus L.) cultivars and breeding populations. — J. Amer. Soc. Hort. Sci.121: 783–788.
Weber, C., 1963: Cultivars in the genusChaenomeles. — Arnoldia23: 17–75.
—, 1964: The genusChaenomeles (Rosaceae). — J. Arnold Arbor.45: 161–205, 302–345.
Weising, K., Nybom, H., Wolff, K., Meyer, W., 1994: DNA fingerprinting in plants and fungi. — Boca Raton: CRC Publishers.
Williams, J. G. K., Kubelik, A. R., Livak, K. J., Rafalski, J. A., Tingey, S. V., 1990: DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. — Nucl. Acids Res.18: 6531–6535.
Yü, T. T., Kuan, K. C., 1963: Taxa nova Rosacearum sinicarum (I). — Acta Phytotax. Sin.8: 234.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Bartish, I.V., Rumpunen, K. & Nybom, H. Genetic diversity inChaenomeles (Rosaceae) revealed by RAPD analysis. Pl Syst Evol 214, 131–145 (1999). https://doi.org/10.1007/BF00985735
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00985735