Abstract
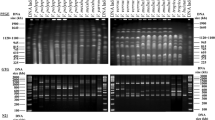
The taxonomy of the fungal genus Chrysosporium is mainly based on morphological features. In our current studies we have found several Chrysosporium species which showed intermediate morphological characteristics between several species. For this reason, we have carried out an analysis of the mitochondrial DNA restriction fragments of these strains that have permit us to classify each isolate strain in a species.
Similar content being viewed by others
References
Oorschot van, CAN. A revision of Chrysosporium and allied genera. Stud Mycol 1980; 20: 1–89.
Sigler L, Guarro J, Punsola L. New keratinophilic species of Chrysosporium. Can J Bot 1986; 64: 1212–1215.
Cano J, Guarro J. Studies on keratinophilic fungi. III. Chrysosporium siglerae sp. nov. Mycotaxon 1994; 51: 75–79.
Jones MG, Noble WC. A study of fatty acids as a taxonomic tool for dermatophyte fungi. J Appl Bacteriol 1981; 50: 577–583.
Leal JA, Gómez-Miranda B, Bernabé M, Cano J, Guarro J. The chemical composition of the wall of six species of Aphanoascus: the taxonomic significance of the presence of α-(1-2) (1-6) mannan and α-(1-4) glucan. Mycol Res 1992; 96: 363–368.
Guarro J, Cano J, Leal JA, Gómez-Miranda B, Bernabe M. Composition of the cell wall polysaccharides in some geophilic dermatophytes. Mycopathologia 1993; 122: 69–77.
Symoens F, Willenz P, Rouma V, Planard C, Nolard N. Isoelectric focusing applied to taxonomic differentiation of the Trichophyton mentagrophytes complex and the related Trichophyton interdigitale. Mycoses 1989; 32: 652–63.
Bruns TD, White TJ, Taylor JW. Fungal molecular systematics. Annu Rev Ecol Syst 1991; 22: 525–564.
Davidson FD, Mackenzie DWR. DNA homology in the taxonomy of dermatophytes. J Med Vet Mycol 1984; 22: 116–123.
Michelmore RW, Hullbert SH. Molecular markers for genetic analysis of phytopathological fungi. Ann Rev Phytopathol 1987; 25: 383–404.
Groosman LI, Hudspeth MES. Fungal mitochondrial genomes. In: Bennett JW, Lasure LL (eds.), Gene Manipulations in Fungi. Orlando: Academic Press, 1985: 65–103.
Kurtmanz CP. Molecular taxonomy of the fungi. In: Bennett JW, Lasure LL (eds.), Gene Manipulations in Fungi. Orlando: Academic Press, 1985: 35–63.
Taylor JW. Fungal evolutionary biology and mitochondrial DNA. Exp Mycol 1986; 10: 259–269.
Gardes M, Mueller GM, Fortin JA, Kropp BR. Mitochondrial DNA polymorphisms in Laccaria bicolor, L. laccata, L. proxima and L. amethystina. Mycol Res 1991; 95: 206–216.
Houben P, Clark-Walker GD. An approach to yeast classification by mapping mitochondrial DNA from Dekkera/Brettanomyces and Einiella genera. Curr Genet 1986; 8: 621–628.
Hintz WE, Mohan M, Anderson JB, Horgen PA. The mitochondrial DNA's of Agaricus: heterogeneity in A. bitorquis and homogeneity in A. brunnencens. Curr Genet 1985; 9: 127–132.
Weber CA, Hudspeth MES, Moore GP, Grossman LI. Analysis of the mitochondrial and nuclear genomes of two Basidiomycetes, Coprinus cinereus and Coprinus stercorarius. Curr Gen 1986; 10: 515–525.
Anderson JB, Petsche DM, Smith ML. Restriction fragment length polymorphisms in biological species of Armillaria mellea. Mycologia 1987; 79: 69–76.
Bruns TD, Palmer JD, Shumard DS, Hudspeth MES. Mitochondrial DNA of Suillus: Three fold size changes in molecules that share a common gene order. Exp Mycol 1988; 15: 316–325.
Guillamón J, Barrio E, Huerta T, Querol A. Rapid characterization of four species of the Saccharomyces sensu stricto complex according to mitochondrial DNA patterns. Int J Syst Bacteriol 1994; 44: 708–714.
Garber RC, Yoder OC. Mitochondrial DNA of the filamentous ascomycete Cochliobolus heterostrophus. Curr Genet 1985; 8: 621–628.
Hulbert SH, Michelmore RW. DNA restriction fragment length polymorphisms and somatic variation in the lettuce downy mildew fungus, Bremia lactucae. Mol Plant Microbe Interact 1988; 1: 17–24.
Forster H, Kinshcerf TG, Leong SA, Maxwell DP. Restriction fragment length polymorphisms of the mitochondrial DNA of Phytophthora megasperma isolated from soybean, alfalfa and fruit trees. Can J Bot 1989; 67: 529–537.
Querol A, Barrio E, Ramón D. A comparative study of different methods of yeast strain characterization. System Appl Microbiol 1992; 15: 439–446.
Bates MR, Buck KW, Brassier CM. Molecular relationships between Ophiostoma ulmi and the NAN and EAN races of O. novo-ulmi determined by restriction fragment length polymorphisms of nuclear DNA. Mycol Res 1993; 97: 449–455.
Thomas V, Rutherford MA, Bridge PD. Molecular differentiation of two races of Fusarium oxysporum special form cubense. Lett Appl Microbiol 1994; 18: 193–196.
Marriot AC, Archer SA, Buck KW. Mitochondrial DNA in Fusarium oxysporum is a 46.5 kilobase pair circular molecule. J Gen Microbiol 1984; 3001–3008.
Cooley RN. The use of RFLP analysis, electrophoretic karyotyping and PCR in studies of plant pathogenic fungi. In: de Stahl U, Tudzynski (eds.), Molecular Biology of Filamentous Fungi. Weinheim: VCH, 1992: 13–26.
Estruch JJ, Antuña C, Ferrer S, Ramón D. Aislamiento de DNA genómico de Trichophyton mentagrophytes. Rev Iber Micol 1989; 6: 62–66.
Sambrook J, Fritsch EF, Maniatis T. Molecular Cloning: A Laboratory Manual, 2nd ed. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press, 1989.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Cano, J., Guillamón, J.M., Vidal, P. et al. The utility of mitochondrial DNA restriction analysis in the classification of strains of Chrysosporium (hyphomycetes). Mycopathologia 134, 65–69 (1996). https://doi.org/10.1007/BF00436866
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00436866