Summary
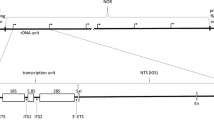
1. Total Neurospora crassa DNA was restricted with endonucleases and fragments carrying rRNA coding sequences were identified by hybridization with Xenopus laevis ribosomal DNA probes. 2. The repeating unit of the rRNA gene cluster was found to be 8.6 kbp, arranged in a head-to-tail fashion. 3. Digestion with Hind III yielded fragments of 3.4 kbp and 5.2 kbp and both were cloned. 4. Digestion with Eco RI yieled fragments of 2.2 kbp, 3.0 kbp and 3.4 kbp; the 3.0 kbp fragment was cloned. 5. Sequences coding for RNA (S-rRNA)1 of the smaller subribosomal particle were found (at least 90%) in the 2.2 kbp EcoRI subfragment of the 5.2 kbp Hind III fragment. 6. The coding sequences for the major RNA species (L-rRNA) of the larger subribosomal particle were located mainly (at least 95%) in the 3.4kbp Hind III fragment. 7. For comparison, a Hind III digest of total yeast DNA was cloned and recombinants containing a 6.4 kbp rDNA fragment were isolated.
Similar content being viewed by others
References
Bell, G.I., De Gennaro, L.J., Gelfand, D.H., Bishop, R.J., Valenzuela, P., Rutter, W.J.: Ribosomal RNA genes of Saccharomyces cerevisiae. J. Biol. Chem. 252, 8118–8125 (1977)
Benton, W.D., Davis, R.W.: Screening λgt recombinant clones by hybridisation to single plaques in situ. Science 196, 180–182 (1977)
Birnstiel, M.L., Grunstein, M.: The ribosomal cistrons of eukaryotes — a model system for the study of evolution of serially repeated genes. FEBS Symp. 23, 349–365 (1972)
Brooks, R.R., Huang, P.C.: Redundant DNA of Neurospora crassa. Biochem. Genet. 6, 41–49 (1972)
Cameron, J.R., Philippsen, P., Davis, R.W.: Analysis of chromosomal integration and deletions of yeast plasmids. Nucleic Acids Res. 4, 1429–1448 (1977)
Chattopadhyay, S.K., Kohne, D.E., Dutti, S.K.: Ribosomal RNA genes of Neurospora: isolation and characterision. Proc. Natl. Acad. Sci. U.S.A. 69, 3256–3259 (1972)
Clewell, D.D., Helinski, D.R.: Supercoiled circular DNA-protein complex in Escherichia coli: purification and conversation to an open circular DNA form. Proc. Natl. Acad. Sci. U.S.A. 62, 1159–1166 (1969)
Cox, R.A.: Ribonucleic acid from ribonucleoprotein particles. Biochem. Prep. 11, 104–109 (1966)
Cox, R.A.: Structure and function of prokaryotic and eukaryotic ribosomes. Prog. Biophys. Mol. Biol. 32, 193–231 (1977)
Cox, R.A., Hirst, W.: A study of the influence of magnesium ions on the confirmation of ribosomal ribonucleic acid and on the stability of the larger subribosomal particle of rabbit reticulocytes. Biochem. J. 160, 505–519 (1976)
Cox, R.A., Thompson, R.: A novel recombinant of phage λ and a 3 kbp Hind III/Bam HI fragment of Xenopus laevis deoxyribonucleic acid that encompasses the conserved sequences of the largest ribonucleic acid component of cytoplasmic ribosomes. Biochem. Soc. Trans. 6, 1232–1233 (1978)
Cramer, J.H., Farrelly, F.W., Barnitz, J.T., Rownd, R.H.: Construction and restriction endonuclease mapping of hybrid plasmids containing Saccharomyces cerevisiae ribosomal DNA. Mol. Gen. Genet. 151, 229–244 (1977)
Denhardt, D.T.: A membrane-filter technique for the detection of complementary DNA. Biochem. Biophys. Res. Commun. 23, 641–646 (1966)
Engberg, J., Anderson, P., Leick, V., Collins, J.: Free ribosomal DNA molecules from Tetrahymena pyriformis GL are giant palindromes. J. Mol. Biol. 104, 455–470 (1976)
Ferguson, J., Davis, R.W.: A new electron microscopic technique for establishing the positions of genes: an analysis of the yeast ribosomal RNA coding region. J. Mol. Biol. 123, 417–430 (1978)
Gerbi, S.: Fine structure of ribosomal RNA I. Conservation of homologous regions within ribosomal RNA of eukaryotes. J. Mol. Biol. 106, 791–816 (1976)
Gillespie, D., Spiegelman, S.: A quantitative assay of DNA-RNA hybrids with DNA immobilized on a membrane. J. Mol. Biol. 12, 829–842 (1965)
Glover, D.M., Hogness, D.S.: A novel arrangement of the 15S and 25S sequences in a repeating unit of Drosophila melanogaster rDNA. Cell 10, 167–176 (1977)
Glover, D.M., White, R.L., Finnegan, D.J., Hogness, D.S.: Characterization of six cloned DNAs from Drosophila melanogaster, including one that contains the genes for rRNA. Cell, 5, 149–157 (1975)
Hagenbuchle, O., Santer, M., Steitz, J.: Conservation of the primary structure at the 3′-end of 18 S-rRNA from eukaryotic cells. Cell 13, 551–563 (1978)
Hohn, B., Murray, K.: Packaging recombinant DNA molecules into bacteriophage particles in vitro. Proc. Natl. Acad. Sci. U.S.A. 74, 3259–3263 (1977)
Jeffreys, A.J., Flavell, R.A.: A physical map of the DNA regions flanking the rabbit β-globin gene. Cell 12, 429–439 (1977)
Karrer, K.M., Gall, J.G.: The macronuclear ribosomal DNA of Tetrahymena pyriformis is a palindrome. J. Mol. Biol. 104, 421–453 (1976)
Kramer, R.A., Cameron, J.R., Davis, R.W.: Isolation of bacteriophage containing yeast ribosomal RNA genes: screening by in situ RNA hybridization to plaques. Cell 8, 227–232 (1976)
Kramer, R.A., Philippsen, P., Davis, R.W.: Divergent transcription in the yeast ribosomal RNA coding region as shown by hybridization to separated stands and sequence analysis of cloned DNA. J. Mol. Biol. 123, 405 (1978)
Laskey, R.A., Mills, A.D.: Enhanced autoradiographic detection of 32P and 125I using intensifying screens and hypersensitized film. FEBS Lett. 82, 314–316 (1977)
Maniatis, T., Jeffrey, A., Kleid, D.G.: Nucleotide sequence of the rightward operator in phage λ. Proc. Natl. Acad. Sci. U.S.A. 72, 1184–1188 (1975)
Martin, T.E., Bicknell, J.N., Kumar, A.: Hybrid 80S monomers formed from subunits of ribosomes protozoa, fungi, plants and mammals. Biochem. Genet. 4, 603–615 (1970)
Molgaard, H.V., Matthews, H.R., Bradbury, E.M.: Organisation of genes for ribosomal RNA in Physarum polycephalum. Eur. J. Biochem. 68, 541–549 (1976)
Murray, N.E., Brammer, W.J., Murray, K.: Lambdoid phages that simplify the recovery of in vitro recombinants. Mol. Gen. Genet. 150, 53–61 (1977)
Nath, K., Bollon, A.P.: Restriction analysis of tandemly repeated yeast ribosomal RNA genes. Mol. Gen. Genet. 160, 235–245 (1978)
Perkins, D.D., Barry, E.G.: The cytogenetics of Neurospora. Adv. Genet. 19, 133–285 (1977)
Philippsen, P., Thomas, M., Kramer, R.A., Davis, R.W.: Unique arrangement of coding sequences for 5S, 5.8S, 18S and 25S. J. Mol. Biol. 123, 387 (1978)
Sinclair, J.H., Brown, D.D.: Retention of common nucleotide sequences in the ribosomal deoxyribonucleic acid of eukaryotes and some of their physical characteristics. Biochemistry 10, 2761–2769 (1971)
Southern, E.M.: Detection of specific sequences among DNA fragments separated by gel electrophoresis. J. Mol. Biol. 98, 503–517 (1975)
Valenzuela, P., Bell, G.I., Venegas, A., Sewell, E.T., Marsiaz, F.R., De Gennero, L.J., Weinberg, F., Rutter, W.J.: Ribosomal RNA genes of Saccharomyces cerevisiae II Physical map and nucleotide sequence of the 5S ribosomal RNA gene and adjacent intergenic regions. J. Biol. Chem. 252, 8126–8135 (1977)
Vogt, V.M., Braun, R.: Structure of ribosomal DNA in Physarum polycephalum. J. Mol. Biol. 106, 567–587 (1976)
Wellauer, P.K., David, I.B.: Organisation of members within the repeating families of genes coding for ribosomal RNA in Xenopus laevis and Drosophila melanogaster. In: Recombinant molecules: Impact on science and society (Beers, R.F., Jr. and Bassett, E.G., eds.), pp. 379–398. New York: Raven Press, 1977
Wood, D.D., Luck, D.J.L.: Hybridization of mitochondrial ribosomal RNA. J. Mol. Biol. 41, 211–224 (1969)
Author information
Authors and Affiliations
Additional information
Communicated by Ch. Auerbach
Containment Facilities. The isolation of recombinant plasmids and phage were carried out under Category 2 conditions as described in the Williams report. Recombinant DNA was handled under Category 1 conditions. These procedures were approved by GMAG and the Departmental Safety Committee.
Rights and permissions
About this article
Cite this article
Cox, R.A., Peden, K. A study of the organisation of the ribosomal ribonucleic acid gene cluster of Neurospora crassa by means of restriction endonuclease analysis and cloning in bacteriophage lambda. Molec. Gen. Genet. 174, 17–24 (1979). https://doi.org/10.1007/BF00433300
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00433300