Summary
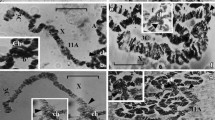
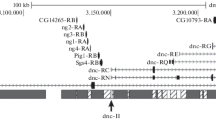
Four dominant suppressor and one enhancer of variegation loci were mapped in the polytene chromosome region extending from section 86C to section 88B of the Drosophila melanogaster third chromosome using a set of deficiencies. The suppressor locus Su-var(3) 14 maps in 86CD, Su-var(3) 13 in 86F4-7, Su-var(3)6 in 87B4-7 and Su-var(3)7 in 87E4-5. The enhancer locus E-var(3)3 maps in 87E12-F11. Su-var(3)13, Su-var(3)6 and Su-var(3)7 are also defined by point mutant alleles originally identified by other criteria (Reuter et al. 1986). Duplications covering the suppressor loci Su-var(3)14, Su-var(3)13, Su-var(3)6 and Su-var(3)7 were found to reduce considerably the haplo-abnormal effect of heterozygous point mutants of the corresponding loci. One suppressor locus, Su-var(3)7, maps within a region which has previously been cloned. The positions of deficiency breakpoints delimiting the suppressor locus indicate that all the necessary sequences for its function are located within 10 kb of cloned DNA.
Similar content being viewed by others
References
Ashburner M, Angel P, Detwiler C, Faithfull J, Gubb D, Harrington G, Littlewood T, Tsubota S, Velissariou V, Walker V (1981) New mutants report. Drosophila Inf Serv 56:186–191
Baker WK (1968) Position-effect variegation. Adv Genet 14:133–169
Bender W, Spierer P, Hogness DS (1983) Chromosome walking and jumping to isolate DNA from the Ace and rosy loci and the bithorax complex of Drosophila melanogaster. J Mol Biol 168:17–33
Bossy B, Hall LMC, Spierer P (1984) Genetic activity along 315 kb of the Drosophila chromosome. EMBO J 3:2537–2541
Bridges PN (1941) A revision of the salivary gland 3R- chromosome map of Drosophila melanogaster. J Hered 32:299–300
Dorn R, Heimann S, Lindigkeit R, Reuter G (1986) Suppressor mutation of position-effect variegation in Drosophila melanogaster affecting chromatin properties. Chromosoma 93:398–403
Gausz J, Bencze G, Gyurkovics H, Ashburner M, Ish-Horowicz D, Holden JJ (1979) Genetic characterization of the 87C region of the third chromosome of Drosophila melanogaster. Genetics 93:917–934
Gausz J, Awad AAM, Gyurkovics H (1980) New deficiencies for the kar locus of Drosophila melanogaster. Drosophila Inf Serv 55:45–46
Gausz J, Gyurkovics H, Bencze G, Awad AAM, Holden JJ, Ish-Horowicz D (1981) Genetic characterization of the region between 86F1,2 and 87B15 on the chromosome 3 of Drosophila melanogaster. Genetics 98:775–789
Gausz J, Hall LMC, Spierer A, Spierer P (1986) Molecular genetics of the rosy-Ace region of Drosophila melanogaster. Genetics 112:65–78
Hall LMC, Spierer P (1986) The Ace locus of Drosophila melanogaster: structural gene for acetylcholinesterase with an unusual 5′ leader. EMBO J 5:2949–2954
Hall LMC, Mason PJ, Spierer P (1983) Transcripts, genes and bands in 315,000 base-pairs of Drosophila DNA. J Mol Biol 169:83–96
Henikoff S (1979) Position effects and variegation enhancers in an autosomal region of Drosophila melanogaster. Genetics 93:105–115
Hilliker AJ, Clark SH, Chovnick A, Gelbart WM (1980) Cytogenetic analysis of the chromosomal region immediately adjacent to the rosy locus in Drosophila melanogaster. Genetics 95:95–110
Ish-Horowicz D, Holden JJ, Gehring WJ (1977) Deletions of two heat-activated loci in Drosophila melanogaster and their effects on heat-induced protein synthesis. Cell 12:643–652
Ising G, Block K (1981) Derivation-dependent distribution of insertion sites for a Drosophila transposon. Cold Spring Harbor Symp Quant Biol 45:527–544
Lewis EB (1950) The phenomenon of position effect. Adv Genet 8:73–115
Lindsley DL, Grell EH (1968) Genetic variations of Drosophila melanogaster. Carnegie Inst Wash Publ No 627
Reuter G, Szidonya J (1983) Cytogenetic analysis of variegation suppressors and a dominant temperature-sensitive lethal in region 23–26 of chromosome 2L in Drosophila melanogaster. Chromosoma 88:277–285
Reuter G, Wolff I (1981) Isolation of dominant suppressor mutations for position-effect variegation in Drosophila melanogaster. Mol Gen Genet 182:516–519
Reuter G, Dorn R, Hoffmann HJ (1982) Butyrate sensitive suppressor of position-effect variegation mutations in Drosophila melanogaster. Mol Gen Genet 188:480–485
Reuter G, Dorn R, Wustmann G, Friede B, Rauh G (1986) Third chromosome suppressor of position-effect variegation loci in Drosophila melanogaster. Mol Gen Genet 202:481–487
Rubin GM, Spradling AC (1982) Genetic transformation of Drosophila with transposable element vectors. Science 218:348–353
Scalenghe F, Ritossa F (1976) Controllo dell'attivita genica in Drosophila. Il puff at locus ebony e la glutamina sintetasi I. Atti della Academia Nazionale dei Lincei Ser 8:441–528
Sinclair DAR, Mottus RC, Grigliatti TA (1983) Genes which suppress position-effect variegation in Drosophila melanogaster are clustered. Mol Gen Genet 191:326–333
Spierer P, Spierer A, Bender W, Hogness DS (1983) Molecular mapping of genetic and chromomeric units in Drosophila melanogaster. J Mol Biol 168:35–50
Spofford JB (1976) Position-effect variegation in Drosophila. In: Ashburner M, Novitski E (eds) Genetics and biology of Drosophila, vol 1C. Academic Press New York, pp 955–1018
Tartof KD, Hobbs C, Jones M (1984) A structural basis for variegating position effects. Cell 37:869–878
Author information
Authors and Affiliations
Additional information
Communicated by J.A. Campos-Ortega
Rights and permissions
About this article
Cite this article
Reuter, G., Gausz, J., Gyurkovics, H. et al. Modifiers of position-effect variegation in the region from 86C to 88B of the Drosophila melanogaster third chromosome. Mol Gen Genet 210, 429–436 (1987). https://doi.org/10.1007/BF00327193
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00327193