Abstract
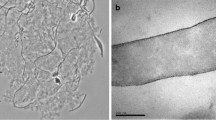
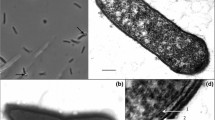
An agar-fermenting bacterium (strain 16AV) was isolated from the sediment of an abattoir effluent waste pond. The cells were curved rods, non-motile, gramnegative, and did not form endospores. Old cultures produced spherical bodies that were not refractile and did not survive pasteurization at 80°C for 5 min. Spherical cells were viable after at least 10 months storage at room temperature. The bacterium was metabolically restricted and grew only on agar, agarose, galactose, and cellobiose. No inorganic electron acceptors were reduced. Acetate and ethanol were produced as end products. Strain 16AV grew between 20 and 37°C, with optimum growth occurring at 28°C. Growth occurred in medium containing NaCl from 0 to 15 g l-1, with an optimum between 2.5 and 5 g l-1. The DNA base composition was 39.5 mol% G+C. 16S rDNA sequence analysis showed that strain 16AV falls within the radiation of members of the genus Clostridium and related taxa. This is the first description of an obligate anaerobe capable of degrading agar.
Similar content being viewed by others
References
Aeckersberg F, Bak F, Widdel F (1991) Anaerobic oxidation of saturated hydrocarbons to CO2 by a new type of sulfate-reducing bacterium. Arch Microbiol 156:5–14
Agbo JAC, Moss MO (1979) The isolation and characterization of agarolytic bacteria from a lowland river. J Gen Microbiol 115: 355–368
American Public Health Association (1980) Standard methods for the examination of water and wastewater. New York, pp 802–804
Atlas R (1993) Handbook of microbiological media. CRC Press, Boca Raton Ann Arbor London
Chapman VJ, Chapman DJ (1980) Seaweeds and their uses, 3rd edn. Chapman and Hall, London New York
Doetsch RN (1981) Determinative methods of light microscopy. In: Gerhardt P, Murray RGE, Costilow RN, Nester EW, Wood WA, Kreig NR, Phillips GB (eds) Manual of methods for general bacteriology. American Society for Microbiology, Washington DC, pp 21–33
Duckworth M, Yaphe W (1971) The structure of agar. 1. Fractionation of a complex mixture of polysaccharides. Carbohyd Res 16:189–197
Gregersen T (1978) Rapid method for distinction of gram-negative from gram-positive bacteria. Eur J Appl Microbiol Biotechnol 5:123–127
Hunger W, Claus D (1978) Reisolation and growth conditions of Bacillus agar-exedens. Antonie Van Leeuwenhoek 44:105–113
Imhoff-Stuckle D, Pfennig N (1983) Isolation and characterization of a nicotinic acid-degrading sulfate-reducing bacterium, Desulfococcus niacini sp. nov. Arch Microbiol 136:194–198
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro NH (ed) Mammalian protein metabolism. Academic Press, New York London, pp 21–132
Meulen HJ van der, Harder W, Veldkamp H (1974) Isolation and characterization of Cytophaga flevensis sp. nov., a new agarolytic flexibacterium. Antonic Van Leeuwenhoek 40:329–346
Mountfort DO, Asher RA (1986) Isolation from a methanogenic ferulate-degrading consortium of an anaerobe that converts methoxy groups of aromatic acids to volatile fatty acids. Arch Microbiol 144:55–61
Owen RJ, Lapage SP (1976) The thermal denaturation of partly purified bacterial deoxyribonucleic acid and its taxonomic applications. J Appl Bacteriol 41:335–340
Patel BKC, Morgan HW, Daniel RM (1985a) A simple and efficient method for preparing and dispensing anaerobic media. Biotechnol Lett 7:277–278
Patel BKC, Morgan HW, Daniel RM (1985b) Fervidobacterium nodosum gen. nov. and spec. nov., a new chemoorganotrophic, caldoactive, anaerobic bacterium. Arch Microbiol 141:63–69
Patel BKC, Monk C, Littleworth H, Morgan HW, Daniel RM (1987) Clostridium fervidus sp. nov., a new chemoorganotrophic, acetogenic thermophile. Int J Syst Bacteriol 37:123–126
Pfennig N, Trüper HG (1981) Isolation of members of the families Chromatiaceae and Chlorobiaceae. In: Starr MP, Stolp H, Trüper HG, Balows A, Schlegel HG (eds) The prokaryotes, vol 1. Springer, Berlin Heidelberg New York, pp 279–289
Rainey FA, Stackebrandt E (1993) 16S rDNA analysis reveals phylogenetic diversity among the polysaccharolytic clostridia. FEMS Microbiol Lett 113:125–128
Rainey FA, Dorsch M, Morgan HW, Stackebrandt E (1992) 16S rDNA analysis of Spirochaeta thermophila: its phylogenetic position and implications for the systematics of the order Spirochaetales. Syst Appl Microbiol 15:197–202
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schink B, Pfennig N (1982) Fermentation of trihydroxybenzenes by Pelobacter acidigallici gen. nov. sp. nov., a new strictly anaerobic, non-spore-forming bacterium. Arch Microbiol 133: 195–201
Smibert RM, Kreig NR (1981) General characterization. In: Gerhardt P, Murray RGE, Costilow RN, Nester EW, Wood WA, Kreig NR, Phillips GB (eds) Manual of methods for general bacteriology. American Society for Microbiology, Washington DC, pp 409–443
Stanier RY (1941) Studies on marine agar-digesting bacteria. J Bacteriol 42:527–588
Stanier RY (1942) Agar-decomposing strains of the actinomyces coelicolor species-group. J Bacteriol 44:555–570
Stouthamer AH (1979) The search for correlation between theoretical and experimental growth yields. In: Quayle JR (ed) International review of biochemistry, vol. 21. Microbial biochemistry. University Park Press, Baltimore, pp 1–47
Veldkamp H (1961) A study of two marine agar-decomposing, facultatively anaerobic myxobacteria. J Gen Microbiol 26:331–342
Widdel F, Kohring G-W, Mayer F (1983) Studies on dissimilatory sulfate-reducing bacteria that decompose fatty acids. III. Characterization of the filamentous gliding Desulfonema limicola gen. nov. sp. nov., and Desulfonema magnum sp. nov. Arch Microbiol 134:286–294
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rees, G.N., Rainey, F.A. & Harfoot, C.G. Characterization of a novel obligate anaerobe that ferments agar. Arch. Microbiol. 162, 395–400 (1994). https://doi.org/10.1007/BF00282103
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00282103