Summary
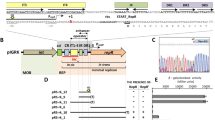
pTB19, a 27 kb plasmid originating from a thermophilic Bacillus species, contains integrated copies of two rolling-circle type plasmids on a 10.6 kb DNA fragment. In the present study we analysed the part of pTB19 that contains the rolling-circle plasmid pTB913 and the region in between the two rolling-circle plasmids. We show that, in the integrated state, pTB913 was flanked by a 55 by direct repeat that duplicated part of the replication initiation gene repB. Since repB was interrupted, the integrated pTB913 could not initiate rolling-circle replication. Autonomously replicating pTB913 was produced from pTB19, probably through recombination between the 55 by direct repeats; this was a rare event. Since the second integrated rolling-circle type plasmid also contained a non-functional replication initiation gene, replication of pT1319 must be controlled by the RepA determinant. Theta-type replication, controlled by RepA is likely to account for the high stability of pTB19. In between the two integrated rolling-circle plasmids was present an open reading frame (447 codons) which could encode a protein of unknown function.
Similar content being viewed by others
References
Bron S (1990) Plasmids. In: Harwood CR, Cutting SM (eds) Molecular biology methods for Bacillus. John Wiley & Sons, Chichester UK, pp 75–174
Bron S, Holsappel S, Venema G, Peeters BPH (1991) Plasmid deletion formation between short direct repeats in Bacillus subtilis is stimulated by single-stranded rolling-circle replication intermediates. Mol Gen Genet 226:88–96
Bron S, Luxen E (1985) Segregational instability of pUB110-derived recombinant plasmids in Bacillus subtilis. Plasmid 14:235–244
Bron S, Luxen E, Swart P (1988) Instability of recombinant pUB110 plasmids in Bacillus subtilis: plasmid-encoded stability function and effects of DNA inserts. Plasmid 19:231–241
Bron S, Venema G (1972) Ultraviolet inactivation and excision repair in Bacillus subtilis. I. Construction and characterization of a transformable eightfold auxotrophic strain and two ultraviolet-sensitive derivatives. Mutat Res 15:1–10
Bruand C, Ehrlich SD, Janniere L (1990) A method for detecting unidirectional theta replication in Bacillus subtilis plasmids. In: Ganeson AT, Hoch JA, Barnitz JT, Zukowski MM (eds) Genetics and biotechnology of bacilli. Academic Press, New York, pp 123–129
Byrne ME, Gillespie MT, Skurray RA (1991) 4′,4″adenyltransfer-ase activity on conjugative plasmids isolated from Staphylococcus aureus is encoded on an integrated copy of pUB110. Plasmid 25:70–75
Ehrlich SD (1989) Illegitimate recombination in bacteria. In: Berg D, Howe M (eds) Mobile DNA. American Society for Microbiology, Washington DC, pp 797–830
Ehrlich SD, Noirot P, Petit MA, Jannière L, Michel B, te Riele H (1986) Structural instability in Bacillus subtilis plasmids. In: Setlow JK, Hollaender A (eds) Genetic engineering, vol 8. Plenum Publishing, New York, pp 71–83
Gennaro ML, Kornblum J, Novick RP (1987) A site-specific recombination function in Staphylococcus aureus plasmids. J Bacteriol 169:2601–2610
Gruss A, Ehrlich SD (1989) The family of highly interrelated ssDNA plasmids. Microbiol Rev 53:231–241
Imanaka T, Fujii M, Aiba S (1981) Isolation and characterization of antibiotic resistance plasmids from thermophilic Bacilli and construction of deletion plasmids. J Bacteriol 146:1091–1097
Imanaka T, Ano T, Fujii M, Aiba S (1984) Two replication determinants of an antibiotic-resistance plasmid, pTB19, from a thermophilic Bacillus. J Gen Microbiol 130:1399–1408
Imanaka T, Ishikawa H, Aiba S (1986) Complete nucleotide sequence of the low copy number plasmid pRAT11 and replication control by the RepA protein in Bacillus subtilis. Mol Gen Genet 205:90–96
Jannière L, Ehrlich SD (1987) Recombination between short repeated sequences is more frequent in plasmids than in the chromosome. Mol Gen Genet 210:116–121
Jannière L, Bruand C, Ehrlich SD (1990) Structurally stable Bacillus subtilis cloning vectors. Gene 87:53–61
Koehler TM, Thorne CB (1987) Bacillus subtilis (natto) plasmid pLS20 mediates interspecies plasmid transfer. J Bacteriol 169:5271–5278
Maciag IE, Viret J-F, Alonso JC (1988) Replication and incompatibility properties of plasmid pUB110 in Bacillus subtilis. Mol Gen Genet 212:232–240
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Matsumura M, Katakura Y, Imanaka T, Aiba S (1984) Enzymatic and nucleotide sequence studies of a kanamycin-inactivating enzyme encoded by a plasmid from thermophilic Bacilli in comparison with that encoded by plasmid pUB110. J Bacteriol 160:413–420
Messing JW (1979) A multipurpose cloning system based on the single-stranded bacteriophage M13. Recomb DNA Tech Bull 2:43–48
Michel B, Ehrlich SD (1986) Illegitimate recombination occurs between the replication origin of plasmid pC194 and a progressing replication fork. EMBO J 5:3691–3696
Muller RE, Ano T, Imanaka T, Aiba S (1986) Complete nucleotide sequences of Bacillus plasmids pUB110dB, pRBH1 and its copy mutants. Mol Gen Genet 202:169–171
Oskam L, Hillenga DJ, Venema G, Bron S (1991) The large Bacillus plasmid pTB19 contains two integrated rolling-circle plasmids carrying mobilization functions. Plasmid 26:30–39
Pearson WR, Lipman DJ (1988) Improved tools for biological sequence comparison. Proc Natl Acad Sci USA 85:2444–2448
Peeters BPH, de Boer JH, Bron S, Venema G (1988) Structural plasmid instability in Bacillus subtilis: effect of direct and inverted repeats. Mol Gen Genet 212:450–458
Perkins JB, Youngman P (1983) Streptococcus plasmid pAMα1 is a composite of two separable replicons, one of which is closely related to Bacillus plasmid pBC16. J Bacteriol 155:607–615
Queen C, Korn LJ (1984) A comprehensive sequence analysis program for the IBM personal computer. Nucleic Acids Res 12:581–599
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Selinger LB, McGregor NF, Khachatourians GG, Hynes MF (1990) Mobilization of closely related plasmids pUB110 and pBC16 by Bacillus plasmid pXO503 requires trans-acting ORFβ. J Bacteriol 172:3290–3297
Tautz D, Renz M (1983) An optimized freeze-squeeze method for the recovery of DNA fragments from agarose gels. Anal Biochem 132:14–19
van der Lelie D, Bron S, Venema G, Oskam L (1989) Similarity of minus origins and flanking open reading frames of plasmids pUB110, pTB913 and pMV158. Nucleic Acids Res 17:7283–7294
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of M13mp18 and pUC19 vectors. Gene 33:103–119
Author information
Authors and Affiliations
Additional information
Communicated by W. Goebel
Rights and permissions
About this article
Cite this article
Oskam, L., Hillenga, D.J., Venema, G. et al. The integrated state of the rolling-circle plasmid pTB913 in the composite Bacillus plasmid pTB19. Molec. Gen. Genet. 233, 462–468 (1992). https://doi.org/10.1007/BF00265444
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00265444