Abstract
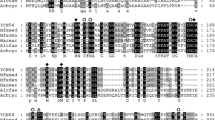
The structural gene, nirK, for the respiratory Cu-containing nitrite reductase from denitrifying Pseudomonas aureofaciens was isolated and sequenced. It encodes a polypeptide of 363 amino acids including a signal peptide of 24 amino acids for protein export. The sequence showed 63.8% positional identity with the amino acid sequence of “Achromobacter cycloclastes” nitrite reductase. Ligands for the blue, type I Cu-binding site and for a putative type-II site were identified. The nirK gene was transferred to the mutant MK202 of P. stutzeri which lacks cytochrome cd 1 nitrite reductase due to a transposon Tn5 insertion in its structural gene, nirS. The heterologous enzyme was active in vitro and in vivo in this background and restored the mutationally interrupted denitrification pathway. Transfer of nirK to Escherichia coli resulted in an active nitrite reductase in vitro. Expression of the nirS gene from P. stutzeri in P. aureofaciens and E. coli led to nonfunctional gene products. Nitrite reductase activity of cell extract from either bacterium could be reconstituted by addition of heme d 1, indicating that both heterologous hosts synthesized a cytochrome cd 1 without the d 1-group.
Similar content being viewed by others
Abbreviations
- Cu-NIR:
-
Cu-containing nitrite reductase
- DDC:
-
diethyldithiocarbamate
- EPR:
-
electron paramagnetic resonance
- IPTG:
-
isopropyl-β-D-galactoside
- SDS:
-
sodium dodecyl sulfate
- LB medium:
-
Luria-Bertani medium
References
Abraham Z, Lowe D, Smith BE (1992) Trimeric nitrite reductase of Achromobacter xylosoxidans has two types of copper centre. J Inorg Chem 47: 45, P01
Boyer HW, Roulland-Dussoix D (1969) A complementation analysis of the restriction and modification of DNA in Escherichia coli. J Mol Biol 41: 459–472
Chang CK, Timkovich R, Wu W (1986) Evidence that heme d 1 is a 1,3-porphyrindione. Biochemistry 25: 8447–8453
Cole JA (1988) Assimilatory and dissimilatory reduction of nitrate to ammonia. In: Cole JA, Ferguson SJ (eds) The nitrogen and sulphur cycles. Cambridge University Press, Cambridge, pp 281–329
Coyle CL, Zumft WG, Kroneck PMH, Körner H, Jakob W (1985) Nitrous oxide reductase from denitrifying Pseudomonas perfectomarina, purification and properties of a novel multicopper enzyme. Eur J Biochem 153: 459–467
Coyne MS, Arunakumari A, Averill BA, Tiedje JM (1989) Immunological identification and distribution of dissimilatory heme cd 1 and nonheme copper nitrite reductases in denitrifying bacteria. Appl Environ Microbiol 55: 2924–2931
Coyne MS, Arunakumari A, Pankratz HS, Tiedje JM (1990) Localization of the cytochrome cd 1 and copper nitrite reductases in denitrifying bacteria. J Bacteriol 172: 2558–2562
Fenderson FF, Kumar S, Adman ET, Liu MY, Payne WJ, LeGall J (1991) Amino acid sequence of nitrite reductase: a copper protein from Achromobacter cycloclastes. Biochemistry 30: 7180–7185
Frunzke K, Zumft WG (1984) Rapid, single sample analysis of H2, O2, NO, CO, N2O and CO2 by isothermal gas chromatography: applications to the study of bacterial denitrification. J Chromatogr 299: 477–483
Glockner AB, Zumft WG (1992) Primary structure of the copper-containing respiratory nitrite reductase of Pseudomonas aureofaciens and functional expression in Escherichia coli. BioEngineering 8: 76, P424
Godden JW, Turley S, Teller DC, Adman ET, Liu MY, Payne WJ, LeGall J (1991) The 2.3 Angstrom X-ray structure of nitrite reductase from Achromobacter cycloclastes. Science 253: 438–442
Grisshammer R, Oeckl C, Michel H (1991) Expression in Escherichia coli of c-type cytochrome genes from Rhodopseudomonas viridis. Biochim Biophys Acta 1088: 183–190
Grossberger D (1987) Minipreps of DNA from bacteriophage lambda. Nucleic Acids Res 15: 6737
Heijne G von (1988) Transcending the impenetrable, how proteins come to terms with membranes. Biochim Biophys Acta 947: 307–333
Hill KE, Wharton DC (1978) Reconstitution of the apoenzyme of cytochrome oxidase from Pseudomonas aeruginosa with heme d 1 and other heme groups. J Biol Chem 253: 489–495
Holmes DS, Quigley M (1981) A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem 114: 193–197
Hulse CL, Tiedje JM, Averill BA (1988) A spectrophotometric assay for dissimilatory nitrite reductases. Anal Biochem 172: 420–426
Hulse CL, Averill BA, Tiedje JM (1989) Evidence for a coppernitrosyl intermediate in denitrification by the copper-containing nitrite reductase of Achromobacter cycloclastes. J Am Chem Soc 111: 2322–2323
Huynh TV, Young RA, Davis RW (1985) Constructing and screening cDNA libraries in lambda gt10 and gt11. In: Glover DM (ed) DNA cloning, vol. 1. IRL Press Oxford, pp 49–78
Iwasaki H, Matsubara T (1972) A nitrite reductase from Achromobacter cycloclastes. J Biochem 71: 645–652
Iwasaki H, Shidara S, Suzuki H, Mori T (1963) Studies on denitrification. VII. Further purification and properties of denitrification enzyme. J Biochem 53: 299–303
Jackson RH, Cornish-Bowden A, Cole JA (1981) Prosthetic groups of the NADH-dependent nirite reductase from Escherichia coli K12. Biochem J 193: 861–867
Jüngst A, Braun C, Zumft WG (1991a) Close linkage in Pseudomonas stutzeri of the structural genes for respiratory nitrite reductase and nitrous oxide reductase, and other essential genes for denitrification. Mol Gen Gent 225: 241–248
Jüngst A, Zumft WG (1992) Heterologous expression of respiratory nitrite reductase (cytochrome cd 1) of Pseudomonas stutzeri in Escherichia coli and Pseudomonas aureofaciens. BioEngineering 8: 76, P423
Jüngst A, Wakabayashi S, Matsubara H, Zumft WG (1991b) The nirSTBM region coding for cytochrome cd 1-dependent nitrite respiration of Pseudomonas stutzeri consists of a cluster of mono-, di-, and tetraheme proteins. FEBS Lett 279: 205–209
Kajie S, Anraku Y (1986) Purification of a hexaheme cytochrome c 552 from Escherichia coli K12 and its properties as a nitrite reductase. Eur J Biochem 154: 457–463
Kakutani T, Watanabe H, Arima K, Beppu T (1981) A blue protein as an inactivating factor for nitrite reductase from Alcaligenes faecalis strain S-6. J Biochem 89: 463–472
Körner H, Mayer F (1992) Periplasmic location of nitrous oxide reductase and its apoform in denitrifying Pseudomonas stutzeri. Arch Microbiol 157: 218–222
Körner H, Frunzke K, Döhler K, Zumft WG (1987) Immunochemical patterns of distribution of nitrous oxide reductase and nitrite reductase (cytochrome cd 1) among denitrifying pseudomonads. Arch Microbiol 148: 20–24
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature (London) 227: 680–685
Laurell CB, McKay EJ (1981) Electroimmunoassay. Meth Enzymol 73: 339–369
Libby E, Averill BA (1992) Evidence that the type 2 copper centers are the site of nitrite reduction by Achromobacter cycloclastes nitrite reductase. Biochem Biophys Res Commun 187: 1529–1535
Liu MC, Payne WJ, Peck HD, Jr, LeGall J (1983) Comparison of cytochromes from anaerobically and aerobically grown cells of Pseudomonas perfectomarinus. J Bacteriol 154: 278–286
Liu MY, Liu MC, Payne WJ, LeGall J (1986) Properties and electron transfer specifity of copper proteins from the denitrifier Achromobacter cycloclastes. J Bacteriol 166: 604–608
Masuko M, Iwasaki H, Sakurai T, Suzuki S, Nakahara A (1984) Characterization of the nitrite reductase from a denitrifier, Alcaligenes Sp. NCIB 11015. A novel copper protein. J Biochem 96: 447–454
Matsubara T, Zumft WG (1982) Identification of a copper protein as part of the nitrous oxide-reducing system in nitrite-respiring (denitrifying) pseudomonads. Arch Microbiol 132: 322–328
Messing J, Gronenborn B, Müller-Hill B, Hofschneider PH (1977) Filamentous coliphage M13 as a cloning vehicle: insertion of a HindII fragment of the lac regulatory region in M13 replicative form in vitro. Proc Natl Acad Sci USA 74: 3642–3646
Page MD, Ferguson SJ (1989) A bacterial c-type cytochrome can be translocated to the periplasm as an apoform: the biosynthesis of cytochrome cd 1 (nitrite reductase) from Paracoccus denitrificans. Mol Microbiol 3: 653–661
Priefer UB, Simon R, Pühler A (1985) Extension of the host range of Escherichia coli vectors by incorporation of RSF1010 replication and mobilization functions. J Bacteriol 163: 324–330
Ryden L, Lundgren JO (1976) Homology relationships among the small blue proteins. Nature 261: 344–346
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Shapleigh JP, Payne WJ (1985) Differentation of c, d 1 cytochrome and copper nitrite reductase production in denitrifiers. FEMS Microbiol Lett 26: 275–279
Silvestrini MC, Cutruzzola F, D'Alessandro R, Brunori M, Fochesato N, Zennaro E (1992) Expression of Pseudomonas aeruginosa nitrite reductase in Pseudomonas putida and characterization of the recombinant protein. Biochem J 285: 661–666
Simon R, Priefer U, Pühler A (1983) A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in Gram-negative bacteria. Bio/Technology 1: 784–791
Studier FW, Moffatt BA (1986) Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol 189: 113–130
Suzuki S, Yoshimura T, Kohzuma T, Shidara S, Masuko M, Sakurai T, Iwasaki H (1989) Spectroscopic evidence for a coppernitrosyl intermediate in nitrite reduction by blue copper-containing nitrite reductase. Biochem Biophys Res Commun 104: 1366–1372
Thomas PE, Ryan D, Levin W (1976) An improved staining procedure for the detection of peroxidase activity of cytochrome P-450 on sodium dodecyl sulfate polyacrylamide gels. Anal Biochem 75: 168–176
Timkovich R, Cork MS, Gennis RB, Johnson PY (1985) Proposed structure of heme d, a prosthetic group of bacterial terminal oxidases. J Am Chem Soc 107: 6096–6075
Wachenfeldt C von, Hederstedt L (1990) Bacillus subtilis holocytochrome c-550 can be synthesized in aerobic Escherichia coli. FEBS Lett 270: 147–151
Yamanaka T, Kijimoto S, Okunuki K (1963) Biological significance of Pseudomonas cytochrome oxidase in Pseudomonas aeruginosa. J Biochem 53: 416–421
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33: 103–119
Young RA, Davis RW (1983a) Yeast RNA polymerase II genes: isolation with antibody probes. Science 222: 778–782
Young RA, Davis RW (1983b) Efficient isolation of genes by using antibody probes. Proc Natl Acad Sci USA 89: 1194–1198
Young RA, Bloom BR, Crossinsky CM, Ivanyi J, Thomas D, Davis RW (1985) Dissection of Mycobacterium tuberculosis antigens using recombinant DNA. Proc Natl Acad Sci USA 82: 2583–2587
Zumft WG (1992) The denitrifying prokaryotes. In: Balows A, Trüper HG, Dworkin M, Harder W, Schleifer KH (eds) The prokaryotes. A handbook on the biology of bacteria: ecophysiology, isolation, identification, applications, 2nd ed, vol. 1. Springer Berlin, Heidelberg, New York, pp 554–582
Zumft WG, Döhler K, Körner H (1985) Isolation and characterization of transposon Tn5-induced mutants of Pseudomonas perfectomarina defective in nitrous oxide respiration. J Bacteriol 163: 918–924
Zumft WG, Gotzman DJ, Kroneck PMH (1987) Type 1, blue copper proteins constitute a respiratory nitrite-reducing system in Pseudomonas aureofaciens. Eur J Biochem 168: 301–307
Zumft WG, Döhler K, Körner H, Löchelt S, Viebrock A, and Frunzke K (1988) Defects in cytochrome cd 1-dependent nitrite respiration of transposon Tn5-induced mutants from Pseudomonas stutzeri. Arch Microbiol 149: 492–498
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Glockner, A.B., Jüngst, A. & Zumft, W.G. Copper-containing nitrite reductase from Pseudomonas aureofaciens is functional in a mutationally cytochrome cd 1-free background (NirS−) of Pseudomonas stutzeri . Arch. Microbiol. 160, 18–26 (1993). https://doi.org/10.1007/BF00258141
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00258141