Abstract
Purpose of Review
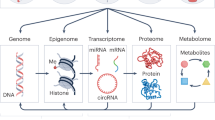
This systematic review evaluated existing evidence linking air pollution exposure in humans to major epigenetic mechanisms: DNA methylation, microRNAs, long noncoding RNAs, and chromatin regulation.
Recent Findings
Eighty-two manuscripts were eligible, most of which were observational (85%), conducted in adults (66%) and based on DNA methylation (79%).
Summary
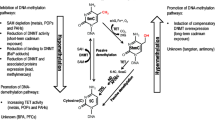
Most observational studies, except panel, demonstrated modest effects of air pollution on the methylome. Panel and experimental studies revealed a relatively large number of significant methylome alterations, though based on smaller sample sizes. Particulate matter levels were positively associated in several studies with global or LINE-1 hypomethylation, a hallmark of several diseases, and with decondensed chromatin structure. Several air pollution species altered the DNA methylation clock, inducing accelerated biological aging. The causal nature of identified associations is not clear, however, especially that most originate from countries with low air pollution levels. Existing evidence, gaps, and perspectives are highlighted herein.



Similar content being viewed by others
References
Papers of particular interest, published recently, have been highlighted as: •• Of major importance
Cohen AJ, Brauer M, Burnett R, Anderson HR, Frostad J, Estep K, et al. Estimates and 25-year trends of the global burden of disease attributable to ambient air pollution: an analysis of data from the Global Burden of Diseases Study 2015. Lancet. 2017;389(10082):1907–18. https://doi.org/10.1016/S0140-6736(17)30505-6.
WHO. WHO Global Urban Ambient Air Pollution 2016 Database http://www.who.int/phe/health_topics/outdoorair/databases/cities/en/. 2016.
Silva RA, West JJ, Lamarque J-F, Shindell DT, Collins WJ, Faluvegi G, et al. Future global mortality from changes in air pollution attributable to climate change. Nat Clim Chang. 2017;7(9):647–51. https://doi.org/10.1038/nclimate3354.
Forman HJ, Finch CE. A critical review of assays for hazardous components of air pollution. Free Radic Biol Med. 2018;117:202–17. https://doi.org/10.1016/j.freeradbiomed.2018.01.030.
Brook RD, Rajagopalan S, Pope CA 3rd, Brook JR, Bhatnagar A, Diez-Roux AV, et al. Particulate matter air pollution and cardiovascular disease: an update to the scientific statement from the American Heart Association. Circulation. 2010;121(21):2331–78. https://doi.org/10.1161/CIR.0b013e3181dbece1.
Nawrot TS, Perez L, Kunzli N, Munters E, Nemery B. Public health importance of triggers of myocardial infarction: a comparative risk assessment. Lancet. 2011;377(9767):732–40. https://doi.org/10.1016/S0140-6736(10)62296-9.
Fiorito G, Vlaanderen J, Polidoro S, Gulliver J, Galassi C, Ranzi A, et al. Oxidative stress and inflammation mediate the effect of air pollution on cardio- and cerebrovascular disease: a prospective study in nonsmokers. Environ Mol Mutagen. 2018;59(3):234–46. https://doi.org/10.1002/em.22153.
Hoek G, Krishnan RM, Beelen R, Peters A, Ostro B, Brunekreef B, et al. Long-term air pollution exposure and cardio- respiratory mortality: a review. Environ Health. 2013;12(1):43. https://doi.org/10.1186/1476-069X-12-43.
Hamra GB, Guha N, Cohen A, Laden F, Raaschou-Nielsen O, Samet JM, et al. Outdoor particulate matter exposure and lung cancer: a systematic review and meta-analysis. Environ Health Perspect. 2014;122(9):906–11. https://doi.org/10.1289/ehp.1408092.
Loomis D, Grosse Y, Lauby-Secretan B, El Ghissassi F, Bouvard V, Benbrahim-Tallaa L, et al. The carcinogenicity of outdoor air pollution. Lancet Oncol. 2013;14(13):1262–3.
Clifford A, Lang L, Chen R, Anstey KJ, Seaton A. Exposure to air pollution and cognitive functioning across the life course—a systematic literature review. Environ Res. 2016;147:383–98. https://doi.org/10.1016/j.envres.2016.01.018.
Saenen ND, Plusquin M, Bijnens E, Janssen BG, Gyselaers W, Cox B, et al. In utero fine particle air pollution and placental expression of genes in the brain-derived neurotrophic factor signaling pathway: an ENVIRONAGE birth cohort study. Environ Health Perspect. 2015;123(8):834–40. https://doi.org/10.1289/ehp.1408549.
Lanki T, Pekkanen J, Aalto P, Elosua R, Berglind N, D'Ippoliti D, et al. Associations of traffic related air pollutants with hospitalisation for first acute myocardial infarction: the HEAPSS study. Occup Environ Med. 2006;63(12):844–51. https://doi.org/10.1136/oem.2005.023911.
Harris MH, Gold DR, Rifas-Shiman SL, Melly SJ, Zanobetti A, Coull BA, et al. Prenatal and childhood traffic-related air pollution exposure and childhood executive function and behavior. Neurotoxicol Teratol. 2016;57:60–70. https://doi.org/10.1016/j.ntt.2016.06.008.
Saenen ND, Provost EB, Viaene MK, Vanpoucke C, Lefebvre W, Vrijens K, et al. Recent versus chronic exposure to particulate matter air pollution in association with neurobehavioral performance in a panel study of primary schoolchildren. Environ Int. 2016;95:112–9. https://doi.org/10.1016/j.envint.2016.07.014.
Goodson JM, Weldy CS, MacDonald JW, Liu Y, Bammler TK, Chien WM, et al. In utero exposure to diesel exhaust particulates is associated with an altered cardiac transcriptional response to transverse aortic constriction and altered DNA methylation. FASEB Journal. 2017;31(11):4935–45. https://doi.org/10.1096/fj.201700032R.
•• Plusquin M, Chadeau-Hyam M, Ghantous A, Alfano R, Bustamante M, Chatzi L, et al. DNA methylome marks of exposure to particulate matter at three time points in early life. Environ Sci Technol. 2018;52(9):5427–37. https://doi.org/10.1021/acs.est.7b06447 Using longitudinal measurements of blood DNA methylation from birth to adolescence provides evidence that residential PM10 exposure affects methylation of sites involved in neurological and cell division control mechanism.
Martens DS, Cox B, Janssen BG, Clemente DBP, Gasparrini A, Vanpoucke C, et al. Prenatal air pollution and newborns’ predisposition to accelerated biological aging. JAMA Pediatr. 2017;171(12):1160–7. https://doi.org/10.1001/jamapediatrics.2017.3024.
Waterland RA, Michels KB. Epigenetic epidemiology of the developmental origins hypothesis. Annu Rev Nutr. 2007;27:363–88. https://doi.org/10.1146/annurev.nutr.27.061406.093705.
Rakyan VK, Down TA, Balding DJ, Beck S. Epigenome-wide association studies for common human diseases. Nat Rev Genet. 2011;12(8):529–41. https://doi.org/10.1038/nrg3000.
Jones PA, Liang G. The human epigenome. In: Michels KB, editor. Epigentic epidemiology. 1 ed.: Springer, Dordrecht; 2012. p. 5–20.
Herceg Z, Ghantous A, Wild CP, Sklias A, Casati L, Duthie SJ, et al. Roadmap for investigating epigenome deregulation and environmental origins of cancer. Int J Cancer. 2018;142(5):874–82. https://doi.org/10.1002/ijc.31014.
von Elm E, Altman DG, Egger M, Pocock SJ, Gotzsche PC, Vandenbroucke JP. The Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement: guidelines for reporting observational studies. J Clin Epidemiol. 2008;61(4):344–9. https://doi.org/10.1016/j.jclinepi.2007.11.008.
Moher D, Liberati A, Tetzlaff J, Altman DG. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. J Clin Epidemiol. 2009;62(10):1006–12. https://doi.org/10.1016/j.jclinepi.2009.06.005.
Abraham E, Rousseaux S, Agier L, Giorgis-Allemand L, Tost J, Galineau J, et al. Pregnancy exposure to atmospheric pollution and meteorological conditions and placental DNA methylation. Environ Int. 2018;118:334–47. https://doi.org/10.1016/j.envint.2018.05.007.
Gruzieva O, Xu CJ, Breton CV, Annesi-Maesano I, Antó JM, Auffray C, et al. Epigenome-wide meta-analysis of methylation in children related to prenatal NO2 air pollution exposure. Environ Health Perspect. 2017;125(1):104–10. https://doi.org/10.1289/EHP36.
Breton CV, Gao L, Yao J, Siegmund KD, Lurmann F, Gilliland F. Particulate matter, the newborn methylome, and cardio-respiratory health outcomes in childhood. Environ Epigenetics. 2016;2(2):dvw005. https://doi.org/10.1093/eep/dvw005.
Goodrich JM, Reddy P, Naidoo RN, Asharam K, Batterman S, Dolinoy DC. Prenatal exposures and DNA methylation in newborns: a pilot study in Durban, South Africa. Environ Sci. 2016;18(7):908–17. https://doi.org/10.1039/c6em00074f.
Rossnerova A, Tulupova E, Tabashidze N, Schmuczerova J, Dostal M, Rossner Jr P et al. Factors affecting the 27K DNA methylation pattern in asthmatic and healthy children from locations with various environments. Mutation Research - Fundamental and Molecular Mechanisms of Mutagenesis. 2013;741–742:18–26. doi:https://doi.org/10.1016/j.mrfmmm.2013.02.003.
Maghbooli Z, Hossein-Nezhad A, Adabi E, Asadollah-Pour E, Sadeghi M, Mohammad-Nabi S, et al. Air pollution during pregnancy and placental adaptation in the levels of global DNA methylation. PLoS One. 2018;13(7):e0199772. https://doi.org/10.1371/journal.pone.0199772.
Janssen BG, Byun HM, Gyselaers W, Lefebvre W, Baccarelli AA, Nawrot TS. Placental mitochondrial methylation and exposure to airborne particulate matter in the early life environment: an ENVIRONAGE birth cohort study. Epigenetics. 2015;10(6):536–44. https://doi.org/10.1080/15592294.2015.1048412.
Herbstman JB, Tang D, Zhu D, Qu L, Sjodin A, Li Z, et al. Prenatal exposure to polycyclic aromatic hydrocarbons, benzo[a]pyrene-DNA adducts, and genomic DNA methylation in cord blood. Environ Health Perspect. 2012;120(5):733–8. https://doi.org/10.1289/ehp.1104056.
Li Y, Mu Z, Wang H, Liu J, Jiang F. The role of particulate matters on methylation of IFN-γ and IL-4 promoter genes in pediatric allergic rhinitis. Oncotarget. 2018;9(25):17406–19. https://doi.org/10.18632/oncotarget.24227.
Nawrot TS, Saenen ND, Schenk J, Janssen BG, Motta V, Tarantini L, et al. Placental circadian pathway methylation and in utero exposure to fine particle air pollution. Environ Int. 2018;114:231–41. https://doi.org/10.1016/j.envint.2018.02.034.
Neven KY, Saenen ND, Tarantini L, Janssen BG, Lefebvre W, Vanpoucke C, et al. Placental promoter methylation of DNA repair genes and prenatal exposure to particulate air pollution: an ENVIRONAGE cohort study. Lancet Planetary Health. 2018;2(4):e174–e83. https://doi.org/10.1016/s2542-5196(18)30049-4.
Prunicki M, Stell L, Dinakarpandian D, de Planell-Saguer M, Lucas RW, Hammond SK, et al. Exposure to NO2, CO, and PM2.5 is linked to regional DNA methylation differences in asthma. Clin Epigenetics. 2018;10:2. https://doi.org/10.1186/s13148-017-0433-4.
Alvarado-Cruz I, Sanchez-Guerra M, Hernandez-Cadena L, De Vizcaya-Ruiz A, Mugica V, Pelallo-Martinez NA, et al. Increased methylation of repetitive elements and DNA repair genes is associated with higher DNA oxidation in children in an urbanized, industrial environment. Mutat Res. 2017;813:27–36. https://doi.org/10.1016/j.mrgentox.2016.11.007.
Cai J, Zhao Y, Liu P, Xia B, Zhu Q, Wang X et al. Exposure to particulate air pollution during early pregnancy is associated with placental DNA methylation. Sci Total Environ. 2017;607–608:1103–8. doi:https://doi.org/10.1016/j.scitotenv.2017.07.029.
Lovinsky-Desir S, Jung KH, Jezioro JR, Torrone DZ, de Planell-Saguer M, Yan B, et al. Physical activity, black carbon exposure, and DNA methylation in the FOXP3 promoter. Clin Epigenetics. 2017;9(1):65. https://doi.org/10.1186/s13148-017-0364-0.
Saenen ND, Vrijens K, Janssen BG, Roels HA, Neven KY, Vanden Berghe W, et al. Lower placental leptin promoter methylation in association with fine particulate matter air pollution during pregnancy and placental nitrosative stress at birth in the ENVIRONAGE cohort. Environ Health Perspect. 2017;125(2):262–8. https://doi.org/10.1289/EHP38.
Breton CV, Yao J, Millstein J, Gao L, Siegmund KD, Mack W, et al. Prenatal air pollution exposures, DNA methyl transferase genotypes, and associations with newborn line1 and Alu methylation and childhood blood pressure and carotid intima-media thickness in the children’s health study. Environ Health Perspect. 2016;124(12):1905–12. https://doi.org/10.1289/EHP181.
Somineni HK, Zhang X, Biagini Myers JM, Kovacic MB, Ulm A, Jurcak N, et al. Ten-eleven translocation 1 (TET1) methylation is associated with childhood asthma and traffic-related air pollution. J Allergy Clin Immunol. 2016;137(3):797–805.e5. https://doi.org/10.1016/j.jaci.2015.10.021.
Hew KM, Walker AI, Kohli A, Garcia M, Syed A, McDonald-Hyman C, et al. Childhood exposure to ambient polycyclic aromatic hydrocarbons is linked to epigenetic modifications and impaired systemic immunity in T cells. Clin Exp Allergy. 2015;45(1):238–48. https://doi.org/10.1111/cea.12377.
Breton CV, Salam MT, Wang X, Byun HM, Siegmund KD, Gilliland FD. Particulate matter, DNA methylation in nitric oxide synthase, and childhood respiratory disease. Environ Health Perspect. 2012;120(9):1320–6. https://doi.org/10.1289/ehp.1104439.
Salam MT, Byun HM, Lurmann F, Breton CV, Wang X, Eckel SP, et al. Genetic and epigenetic variations in inducible nitric oxide synthase promoter, particulate pollution, and exhaled nitric oxide levels in children, J Allergy Clin Immunol. 2012;129(1):232–9.e7. https://doi.org/10.1016/j.jaci.2011.09.037.
Tang WY, Levin L, Talaska G, Cheung YY, Herbstman J, Tang D et al. Maternal exposure to polycyclic aromatic hydrocarbons and 5′-CpG methylation of interferon-γ in cord white blood cells. Environ Health Perspect 2012;120(8):1195–1200. doi:https://doi.org/10.1289/ehp.1103744.
Nadeau K, McDonald-Hyman C, Noth EM, Pratt B, Hammond SK, Balmes J, et al. Ambient air pollution impairs regulatory T-cell function in asthma. J Allergy Clin Immunol. 2010;126(4):845–52.e10. https://doi.org/10.1016/j.jaci.2010.08.008.
Perera F, Tang WY, Herbstman J, Tang D, Levin L, Miller R et al. Relation of DNA methylation of 5′-CpG island of ACSL3 to transplacental exposure to airborne polycyclic aromatic hydrocarbons and childhood asthma. PLoS One 2009;4(2):e4488. doi:https://doi.org/10.1371/journal.pone.0004488.
•• Mostafavi N, Vermeulen R, Ghantous A, Hoek G, Probst-Hensch N, Herceg Z, et al. Acute changes in DNA methylation in relation to 24h personal air pollution exposure measurements: a panel study in four European countries. Environ Int. 2018;120:11–21. https://doi.org/10.1016/j.envint.2018.07.026 Using panel design revealed an association between 24-hour personal exposure to air pollution and DNA-methylation both at single sites and regional clusters.
Lichtenfels AJFC, Van Der Plaat DA, De Jong K, Van Diemen CC, Postma DS, Nedeljkovic I, et al. Long-term air pollution exposure, genome-wide DNA methylation and lung function in the lifelines cohort study. Environ Health Perspect. 2018;126(2):027004. https://doi.org/10.1289/EHP2045.
Dai L, Mehta A, Mordukhovich I, Just AC, Shen J, Hou L, et al. Differential DNA methylation and PM2.5 species in a 450K epigenome-wide association study. Epigenetics. 2017;12(2):139–48. https://doi.org/10.1080/15592294.2016.1271853.
Nwanaji-Enwerem JC, Dai L, Colicino E, Oulhote Y, Di Q, Kloog I, et al. Associations between long-term exposure to PM2.5 component species and blood DNA methylation age in the elderly: the VA normative aging study. Environ Int. 2017;102:57–65. https://doi.org/10.1016/j.envint.2016.12.024.
•• Panni T, Mehta AJ, Schwartz JD, Baccarelli AA, Just AC, Wolf K, et al. Genome-wide analysis of DNA methylation and fine particulate matter air pollution in three study populations: KORA F3, KORA F4, and the Normative Aging Study. Environ Health Perspect. 2016;124(7):983–90. https://doi.org/10.1289/ehp.1509966 Combining evidence from three independent adult cohorts suggests novel plausible systemic pathways linking ambient PM exposure to adverse health effect through variations in DNA methylation.
Plusquin M, Guida F, Polidoro S, Vermeulen R, Raaschou-Nielsen O, Campanella G, et al. DNA methylation and exposure to ambient air pollution in two prospective cohorts. Environ Int. 2017;108:127–36. https://doi.org/10.1016/j.envint.2017.08.006.
Chi GC, Liu Y, MacDonald JW, Barr RG, Donohue KM, Hensley MD, et al. Long-term outdoor air pollution and DNA methylation in circulating monocytes: results from the multi-ethnic study of atherosclerosis (MESA). Environ Health. 2016;15(1):119. https://doi.org/10.1186/s12940-016-0202-4.
Nwanaji-Enwerem JC, Colicino E, Trevisi L, Kloog I, Just AC, Shen J et al. Long-term ambient particle exposures and blood DNA methylation age: findings from the VA normative aging study. Environ Epigenetics. 2016;2(2). doi:https://doi.org/10.1093/eep/dvw006.
Ward-Caviness CK, Nwanaji-Enwerem JC, Wolf K, Wahl S, Colicino E, Trevisi L, et al. Long-term exposure to air pollution is associated with biological aging. Oncotarget. 2016;7(46):74510–25. https://doi.org/10.18632/oncotarget.12903.
Carmona JJ, Sofer T, Hutchinson J, Cantone L, Coull B, Maity A, et al. Short-term airborne particulate matter exposure alters the epigenetic landscape of human genes associated with the mitogen-activated protein kinase network: a cross-sectional study. Environ Health. 2014;13(1):94. https://doi.org/10.1186/1476-069X-13-94.
De Nys S, Duca RC, Nawrot T, Hoet P, Van Meerbeek B, Van Landuyt KL, et al. Temporal variability of global DNA methylation and hydroxymethylation in buccal cells of healthy adults: association with air pollution. Environ Int. 2018;111:301–8. https://doi.org/10.1016/j.envint.2017.11.002.
Liu J, Xie K, Chen W, Zhu M, Shen W, Yuan J, et al. Genetic variants, PM2.5 exposure level and global DNA methylation level: a multi-center population-based study in Chinese. Toxicol Lett. 2017;269:77–82. https://doi.org/10.1016/j.toxlet.2017.02.003.
Sanchez-Guerra M, Zheng Y, Osorio-Yanez C, Zhong J, Chervona Y, Wang S, et al. Effects of particulate matter exposure on blood 5-hydroxymethylation: results from the Beijing truck driver air pollution study. Epigenetics. 2015;10(7):633–42. https://doi.org/10.1080/15592294.2015.1050174.
De Prins S, Koppen G, Jacobs G, Dons E, Van de Mieroop E, Nelen V, et al. Influence of ambient air pollution on global DNA methylation in healthy adults: a seasonal follow-up. Environ Int. 2013;59:418–24. https://doi.org/10.1016/j.envint.2013.07.007.
Wang C, Chen R, Shi M, Cai J, Shi J, Yang C, et al. Possible mediation by methylation in acute inflammation following personal exposure to fine particulate air pollution. Am J Epidemiol. 2018;187(3):484–93. https://doi.org/10.1093/aje/kwx277.
Wang C, Chen R, Cai J, Shi J, Yang C, Tse LA, et al. Personal exposure to fine particulate matter and blood pressure: a role of angiotensin converting enzyme and its DNA methylation. Environ Int. 2016;94:661–6. https://doi.org/10.1016/j.envint.2016.07.001.
Sofer T, Maity A, Coull B, Baccarelli A, Schwartz J, Lin X. Multivariate gene selection and testing in studying the exposure effects on a gene set. Stat Biosci. 2012;4(2):319–38. https://doi.org/10.1007/s12561-012-9072-7.
Peng C, Bind MAC, Colicino E, Kloog I, Byun HM, Cantone L, et al. Particulate air pollution and fasting blood glucose in nondiabetic individuals: associations and epigenetic mediation in the normative aging study, 2000-2011. Environ Health Perspect. 2016;124(11):1715–21. https://doi.org/10.1289/EHP183.
Ouyang B, Baxter CS, Lam HM, Yeramaneni S, Levin L, Haynes E, et al. Hypomethylation of dual specificity phosphatase 22 promoter correlates with duration of service in firefighters and is inducible by low-dose benzo[a]pyrene. J Occup Environ Med. 2012;54(7):774–80. https://doi.org/10.1097/JOM.0b013e31825296bc.
Madrigano J, Baccarelli A, Mittleman MA, Sparrow D, Spiro A, Vokonas PS, et al. Air pollution and DNA methylation: interaction by psychological factors in the VA normative aging study. Am J Epidemiol. 2012;176(3):224–32. https://doi.org/10.1093/aje/kwr523.
Madrigano J, Baccarelli A, Mittleman MA, Wright RO, Sparrow D, Vokonas PS, et al. Prolonged exposure to particulate pollution, genes associated with glutathione pathways, and DNA methylation in a cohort of older men. Environ Health Perspect. 2011;119(7):977–82. https://doi.org/10.1289/ehp.1002773.
Liu Y, Lan Q, Shen M, Jin J, Mumford J, Ren D, et al. Aberrant gene promoter methylation in sputum from individuals exposed to smoky coal emissions. Anticancer Res. 2008;28(4 B):2061–6.
Lepeule J, Bind MA, Baccarelli AA, Koutrakis P, Tarantini L, Litonjua A, et al. Epigenetic influences on associations between air pollutants and lung function in elderly men: the normative aging study. Environ Health Perspect. 2014;122(6):566–72. https://doi.org/10.1289/ehp.1206458.
Hou L, Zhang X, Zheng Y, Wang S, Dou C, Guo L, et al. Altered methylation in tandem repeat element and elemental component levels in inhalable air particles. Environ Mol Mutagen. 2014;55(3):256–65. https://doi.org/10.1002/em.21829.
Guo L, Byun HM, Zhong J, Motta V, Barupal J, Zheng Y, et al. Effects of short-term exposure to inhalable particulate matter on DNA methylation of tandem repeats. Environ Mol Mutagen. 2014;55(4):322–35. https://doi.org/10.1002/em.21838.
Chen R, Qiao L, Li H, Zhao Y, Zhang Y, Xu W, et al. Fine particulate matter constituents, nitric oxide synthase DNA methylation and exhaled nitric oxide. Environ Sci Technol. 2015;49(19):11859–65. https://doi.org/10.1021/acs.est.5b02527.
Cantone L, Iodice S, Tarantini L, Albetti B, Restelli I, Vigna L, et al. Particulate matter exposure is associated with inflammatory gene methylation in obese subjects. Environ Res. 2017;152:478–84. https://doi.org/10.1016/j.envres.2016.11.002.
Callahan CL, Bonner MR, Nie J, Han D, Wang Y, Tao MH, et al. Lifetime exposure to ambient air pollution and methylation of tumor suppressor genes in breast tumors. Environ Res. 2018;161:418–24. https://doi.org/10.1016/j.envres.2017.11.040.
Bind MA, Coull BA, Peters A, Baccarelli AA, Tarantini L, Cantone L, et al. Beyond the mean: quantile regression to explore the association of air pollution with gene-specific methylation in the Normative Aging Study. Environ Health Perspect. 2015;123(8):759–65. https://doi.org/10.1289/ehp.1307824.
Bind MA, Lepeule J, Zanobetti A, Gasparrini A, Baccarelli A, Coull BA, et al. Air pollution and gene-specific methylation in the Normative Aging Study: association, effect modification, and mediation analysis. Epigenetics. 2014;9(3):448–58. https://doi.org/10.4161/epi.27584.
Baccarelli A, Wright RO, Bollati V, Tarantini L, Litonjua AA, Suh HH, et al. Rapid DNA methylation changes after exposure to traffic particles. Am J Respir Crit Care Med. 2009;179(7):572–8. https://doi.org/10.1164/rccm.200807-1097OC.
Zhong J, Karlsson O, Wang G, Li J, Guo Y, Lin X, et al. B vitamins attenuate the epigenetic effects of ambient fine particles in a pilot human intervention trial. Proc Natl Acad Sci U S A. 2017;114(13):3503–8. https://doi.org/10.1073/pnas.1618545114.
Clifford RL, Jones MJ, MacIsaac JL, McEwen LM, Goodman SJ, Mostafavi S, et al. Inhalation of diesel exhaust and allergen alters human bronchial epithelium DNA methylation. J Allergy Clin Immunol. 2017;139(1):112–21. https://doi.org/10.1016/j.jaci.2016.03.046.
Jiang R, Jones MJ, Sava F, Kobor MS, Carlsten C. Short-term diesel exhaust inhalation in a controlled human crossover study is associated with changes in DNA methylation of circulating mononuclear cells in asthmatics. Particle Fibre Toxicol. 2014;11(1):71. https://doi.org/10.1186/s12989-014-0071-3.
Tobaldini E, Bollati V, Prado M, Fiorelli EM, Pecis M, Bissolotti G, et al. Acute particulate matter affects cardiovascular autonomic modulation and IFN-Γ methylation in healthy volunteers. Environ Res. 2018;161:97–103. https://doi.org/10.1016/j.envres.2017.10.036.
Chen R, Meng X, Zhao A, Wang C, Yang C, Li H, et al. DNA hypomethylation and its mediation in the effects of fine particulate air pollution on cardiovascular biomarkers: a randomized crossover trial. Environ Int. 2016;94:614–9. https://doi.org/10.1016/j.envint.2016.06.026.
Bellavia A, Urch B, Speck M, Brook RD, Scott JA, Albetti B, et al. DNA hypomethylation, ambient particulate matter, and increased blood pressure: findings from controlled human exposure experiments. J Am Heart Assoc. 2013;2(3):e000212. https://doi.org/10.1161/JAHA.113.000212.
•• Krauskopf J, Caiment F, van Veldhoven K, Chadeau-Hyam M, Sinharay R, Chung KF, et al. The human circulating miRNome reflects multiple organ disease risks in association with short-term exposure to traffic-related air pollution. Environ Int. 2018;113:26–34. https://doi.org/10.1016/j.envint.2018.01.014 Proposes cmiRNA signature comprised of organ-enriched miRNAs as a highly specific candidate for biomarker-based health risk assessment allowing the early detection and prevention of TRAP-induced health outcomes.
Liu PF, Yan P, Zhao DH, Shi WF, Meng S, Liu Y, et al. The effect of environmental factors on the differential expression of miRNAs in patients with chronic obstructive pulmonary disease: a pilot clinical study. Int J Chronic Obstructive Pulmonary Dis. 2018;13:741–51. https://doi.org/10.2147/copd.S156865.
Pergoli L, Cantone L, Favero C, Angelici L, Iodice S, Pinatel E, et al. Extracellular vesicle-packaged miRNA release after short-term exposure to particulate matter is associated with increased coagulation. Particle and fibre toxicology. 2017;14(1):32. https://doi.org/10.1186/s12989-017-0214-4.
Hou L, Barupal J, Zhang W, Zheng Y, Liu L, Zhang X, et al. Particulate air pollution exposure and expression of viral and human microRNAs in blood: the Beijing Truck Driver Air Pollution Study. Environ Health Perspect. 2016;124(3):344–50. https://doi.org/10.1289/ehp.1408519.
Motta V, Favero C, Dioni L, Iodice S, Battaglia C, Angelici L, et al. MicroRNAs are associated with blood-pressure effects of exposure to particulate matter: results from a mediated moderation analysis. Environ Res. 2016;146:274–81. https://doi.org/10.1016/j.envres.2016.01.010.
Rider CF, Yamamoto M, Gunther OP, Hirota JA, Singh A, Tebbutt SJ, et al. Controlled diesel exhaust and allergen coexposure modulates microRNA and gene expression in humans: effects on inflammatory lung markers. J Allergy Clin Immunol. 2016;138(6):1690–700. https://doi.org/10.1016/j.jaci.2016.02.038.
Rodosthenous RS, Coull BA, Lu Q, Vokonas PS, Schwartz JD, Baccarelli AA. Ambient particulate matter and microRNAs in extracellular vesicles: a pilot study of older individuals. Particle Fibre Toxicol. 2016;13:13. https://doi.org/10.1186/s12989-016-0121-0.
Fry RC, Rager JE, Bauer R, Sebastian E, Peden DB, Jaspers I, et al. Air toxics and epigenetic effects: ozone altered microRNAs in the sputum of human subjects. Am J Physiol Lung Cell Mol Physiol. 2014;306(12):L1129–L37. https://doi.org/10.1152/ajplung.00348.2013.
Yamamoto M, Singh A, Sava F, Pui M, Tebbutt SJ, Carlsten C. MicroRNA expression in response to controlled exposure to diesel exhaust: attenuation by the antioxidant N-acetylcysteine in a randomized crossover study. Environ Health Perspect. 2013;121(6):670–5. https://doi.org/10.1289/ehp.1205963.
Chen R, Li H, Cai J, Wang C, Lin Z, Liu C, et al. Fine particulate air pollution and the expression of microRNAs and circulating cytokines relevant to inflammation, coagulation, and vasoconstriction. Environ Health Perspect. 2018;126(1):017007. https://doi.org/10.1289/EHP1447.
Tsamou M, Vrijens K, Madhloum N, Lefebvre W, Vanpoucke C, Nawrot TS. Air pollution-induced placental epigenetic alterations in early life: a candidate miRNA approach. Epigenetics. 2018:1–12. https://doi.org/10.1080/15592294.2016.1155012.
Louwies T, Vuegen C, Panis LI, Cox B, Vrijens K, Nawrot TS, et al. miRNA expression profiles and retinal blood vessel calibers are associated with short-term particulate matter air pollution exposure. Environ Res. 2016;147:24–31. https://doi.org/10.1016/j.envres.2016.01.027.
Vriens A, Nawrot TS, Saenen ND, Provost EB, Kicinski M, Lefebvre W, et al. Recent exposure to ultrafine particles in school children alters miR-222 expression in the extracellular fraction of saliva. Environ Health. 2016;15(1):80. https://doi.org/10.1186/s12940-016-0162-8.
Fossati S, Baccarelli A, Zanobetti A, Hoxha M, Vokonas PS, Wright RO, et al. Ambient particulate air pollution and microRNAs in elderly men. Epidemiology (Cambridge, Mass). 2014;25(1):68–78. https://doi.org/10.1097/EDE.0000000000000026.
Wei MM, Zhou YC, Wen ZS, Zhou B, Huang YC, Wang GZ, et al. Long non-coding RNA stabilizes the Y-box-binding protein 1 and regulates the epidermal growth factor receptor to promote lung carcinogenesis. Oncotarget. 2016;7(37):59556–71. https://doi.org/10.18632/oncotarget.10006.
Liu C, Xu J, Chen Y, Guo X, Zheng Y, Wang Q, et al. Characterization of genome-wide H3K27ac profiles reveals a distinct PM2.5-associated histone modification signature. Environ Health. 2015;14(1):65. https://doi.org/10.1186/s12940-015-0052-5.
Zheng Y, Sanchez-Guerra M, Zhang Z, Joyce BT, Zhong J, Kresovich JK, et al. Traffic-derived particulate matter exposure and histone H3 modification: a repeated measures study. Environ Res. 2017;153:112–9. https://doi.org/10.1016/j.envres.2016.11.015.
Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013;14(10):R115. https://doi.org/10.1186/gb-2013-14-10-r115.
Hannum G, Guinney J, Zhao L, Zhang L, Hughes G, Sadda S, et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell. 2013;49(2):359–67. https://doi.org/10.1016/j.molcel.2012.10.016.
Prins GS, Calderon-Gierszal EL, Hu WY. Stem cells as hormone targets that lead to increased cancer susceptibility. Endocrinology. 2015;156(10):3451–7. https://doi.org/10.1210/en.2015-1357.
Vineis P, Chatziioannou A, Cunliffe VT, Flanagan JM, Hanson M, Kirsch-Volders M, et al. Epigenetic memory in response to environmental stressors. FASEB Journal. 2017;31(6):2241–51. https://doi.org/10.1096/fj.201601059RR.
Joubert BR, Felix JF, Yousefi P, Bakulski KM, Just AC, Breton C, et al. DNA methylation in newborns and maternal smoking in pregnancy: genome-wide consortium meta-analysis. Am J Hum Genet. 2016;98(4):680–96. https://doi.org/10.1016/j.ajhg.2016.02.019.
Yang AS, Estecio MR, Doshi K, Kondo Y, Tajara EH, Issa JP. A simple method for estimating global DNA methylation using bisulfite PCR of repetitive DNA elements. Nucleic Acids Res. 2004;32(3):e38. https://doi.org/10.1093/nar/gnh032.
Price EM, Cotton AM, Penaherrera MS, McFadden DE, Kobor MS, Robinson W. Different measures of “genome-wide” DNA methylation exhibit unique properties in placental and somatic tissues. Epigenetics. 2012;7(6):652–63. https://doi.org/10.4161/epi.20221.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74. https://doi.org/10.1016/j.cell.2011.02.013.
Felix JF, Joubert BR, Baccarelli AA, Sharp GC, Almqvist C, Annesi-Maesano I, et al. Cohort profile: pregnancy and childhood epigenetics (PACE) consortium. Int J Epidemiol. 2018;47(1):22–3u. https://doi.org/10.1093/ije/dyx190.
Vineis P, Chadeau-Hyam M, Gmuender H, Gulliver J, Herceg Z, Kleinjans J, et al. The exposome in practice: design of the EXPOsOMICS project. Int J Hyg Environ Health. 2017;220(2 Pt A):142–51. https://doi.org/10.1016/j.ijheh.2016.08.001.
Yang SJ, Yang SY, Wang DD, Chen X, Shen HY, Zhang XH, et al. The miR-30 family: versatile players in breast cancer. Tumour Biol. 2017;39(3):1010428317692204. https://doi.org/10.1177/1010428317692204.
Qu K, Lin T, Pang Q, Liu T, Wang Z, Tai M, et al. Extracellular miRNA-21 as a novel biomarker in glioma: evidence from meta-analysis, clinical validation and experimental investigations. Oncotarget. 2016;7(23):33994–4010. https://doi.org/10.18632/oncotarget.9188.
Sweeney CL, Teng R, Wang H, Merling RK, Lee J, Choi U, et al. Molecular analysis of neutrophil differentiation from human induced pluripotent stem cells delineates the kinetics of key regulators of hematopoiesis. Stem Cells. 2016;34(6):1513–26. https://doi.org/10.1002/stem.2332.
Paterson MR, Kriegel AJ. MiR-146a/b: a family with shared seeds and different roots. Physiol Genomics. 2017;49(4):243–52. https://doi.org/10.1152/physiolgenomics.00133.2016.
Ding S, Huang H, Xu Y, Zhu H, Zhong C. MiR-222 in cardiovascular diseases: physiology and pathology. Biomed Res Int. 2017;2017:4962426. https://doi.org/10.1155/2017/4962426.
Testa U, Pelosi E, Castelli G, Labbaye C. miR-146 and miR-155: two key modulators of immune response and tumor development. Noncoding RNA. 2017;3(3). doi:https://doi.org/10.3390/ncrna3030022.
Thum T, Gross C, Fiedler J, Fischer T, Kissler S, Bussen M, et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature. 2008;456(7224):980–4. https://doi.org/10.1038/nature07511.
Huarte M. The emerging role of lncRNAs in cancer. Nat Med. 2015;21(11):1253–61. https://doi.org/10.1038/nm.3981.
Bartonicek N, Maag JL, Dinger ME. Long noncoding RNAs in cancer: mechanisms of action and technological advancements. Mol Cancer. 2016;15(1):43. https://doi.org/10.1186/s12943-016-0530-6.
Jantsch MF, Quattrone A, O'Connell M, Helm M, Frye M, Macias-Gonzales M, et al. Positioning Europe for the EPITRANSCRIPTOMICS challenge. RNA Biol. 2018:1–3. https://doi.org/10.1080/15476286.2018.1460996.
Baccarelli A, Bollati V. Epigenetics and environmental chemicals. Curr Opin Pediatr. 2009;21(2):243–51. https://doi.org/10.1097/MOP.0b013e32832925cc.
Chervona Y, Costa M. The control of histone methylation and gene expression by oxidative stress, hypoxia, and metals. Free Radic Biol Med. 2012;53(5):1041–7. https://doi.org/10.1016/j.freeradbiomed.2012.07.020.
Bayarsaihan D. Epigenetic mechanisms in inflammation. J Dent Res. 2011;90(1):9–17. https://doi.org/10.1177/0022034510378683.
Byun HM, Colicino E, Trevisi L, Fan T, Christiani DC, Baccarelli AA. Effects of air pollution and blood mitochondrial DNA methylation on markers of heart rate variability. J Am Heart Assoc. 2016;5(4). https://doi.org/10.1161/JAHA.116.003218.
McCreanor J, Cullinan P, Nieuwenhuijsen MJ, Stewart-Evans J, Malliarou E, Jarup L, et al. Respiratory effects of exposure to diesel traffic in persons with asthma. N Engl J Med. 2007;357(23):2348–58. https://doi.org/10.1056/NEJMoa071535.
Houseman EA, Accomando WP, Koestler DC, Christensen BC, Marsit CJ, Nelson HH, et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics. 2012;13:86. https://doi.org/10.1186/1471-2105-13-86.
Bakulski KM, Feinberg JI, Andrews SV, Yang J, Brown S, McKenney SL, et al. DNA methylation of cord blood cell types: applications for mixed cell birth studies. Epigenetics. 2016;11(5):354–62. https://doi.org/10.1080/15592294.2016.1161875.
Houseman EA, Kile ML, Christiani DC, Ince TA, Kelsey KT, Marsit CJ. Reference-free deconvolution of DNA methylation data and mediation by cell composition effects. BMC Bioinformatics. 2016;17:259. https://doi.org/10.1186/s12859-016-1140-4.
Bauer M, Linsel G, Fink B, Offenberg K, Hahn AM, Sack U, et al. A varying T cell subtype explains apparent tobacco smoking induced single CpG hypomethylation in whole blood. Clin Epigenetics. 2015;7(1):81. https://doi.org/10.1186/s13148-015-0113-1.
Green BB, Karagas MR, Punshon T, Jackson BP, Robbins DJ, Houseman EA, et al. Epigenome-wide assessment of DNA methylation in the placenta and arsenic exposure in the New Hampshire Birth Cohort Study (USA). Environ Health Perspect. 2016;124(8):1253–60. https://doi.org/10.1289/ehp.1510437.
Langie SA, Szarc Vel Szic K, Declerck K, Traen S, Koppen G, Van Camp G, et al. Whole-genome saliva and blood DNA methylation profiling in individuals with a respiratory allergy. PLoS One. 2016;11(3):e0151109. https://doi.org/10.1371/journal.pone.0151109.
Funding
Support for this work was provided by the project EXPOSOMICS, grant agreement 308610-FP7 European Commission, and by the Plan Cancer-Eva-Inserm research grant.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
Rossella Alfano has a PhD fellowship from Bijzonder Onderzoeksfonds (BOF) Hasselt University.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Environmental Epigenetics
Electronic supplementary material
ESM 1
(DOCX 105 kb)
Rights and permissions
About this article
Cite this article
Alfano, R., Herceg, Z., Nawrot, T.S. et al. The Impact of Air Pollution on Our Epigenome: How Far Is the Evidence? (A Systematic Review). Curr Envir Health Rpt 5, 544–578 (2018). https://doi.org/10.1007/s40572-018-0218-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40572-018-0218-8