Abstract
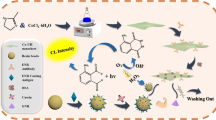
Detection of enzyme biomarkers originating from either bio-fluids or contaminating microorganisms is of utmost importance in clinical diagnostics and food safety. Herein, we present a simple, low-cost and easy-to-use sensing approach based on the switchable peroxidase-mimicking activity of plasmonic gold nanoparticles (AuNPs) that can catalyse for the oxidation of 3,3’,5’5-tetramethylbenzidine (TMB) for the determination of protease enzyme. The AuNP surface is modified with casein, showing dual functionalities. The first function of the coating molecule is to suppress the intrinsic peroxidase-mimicking activity of AuNPs by up to 77.1%, due to surface shielding effects. Secondly, casein also functions as recognition sites for the enzyme biomarker. In the presence of protease, the enzyme binds to and catalyses the degradation of the coating layer on the AuNP surface, resulting in the recovery of peroxidase-mimicking activity. This is shown visually in the development of a blue colored product (oxidised TMB) or spectroscopically as an increase in absorbance at 370 and 650 nm. This mechanism allows for the detection of protease at 44 ng·mL−1 in 90 min. The nanosensor circumvents issues associated with current methods of detection in terms of ease of use, compatibility with point-of-care testing, low-cost production and short analysis time. The sensing approach has also been applied for the detection of protease spiked in ultra-heat treated (UHT) milk and synthetic human urine samples at a limit of detection of 490 and 176 ng·mL−1, respectively, showing great potential in clinical diagnostics, food safety and quality control.

Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Meisner, M. Biomarkers of sepsis: Clinically useful? Curr. Opin. Crit. Care 2005, 11, 473–480.
Eckersall, P. D.; Bell, R. Acute phase proteins: Biomarkers of infection and inflammation in veterinary medicine. Vet. J. 2010, 185, 23–27.
Fadda, S.; López, C.; Vignolo, G. Role of lactic acid bacteria during meat conditioning and fermentation: Peptides generated as sensorial and hygienic biomarkers. Meat Sci. 2010, 86, 66–79.
Ossovskaya, V. S.; Bunnett, N. W. Protease-activated receptors: Contribution to physiology and disease. Physiol. Rev. 2004, 84, 579–621.
Otlewski, J.; Jelen, F.; Zakrzewska, M.; Oleksy, A. The many faces of protease–protein inhibitor interaction. EMBO J. 2005, 24, 1303–1310.
Zhang, S. G.; Janciauskiene, S. Multi-functional capability of proteins: a1- antichymotrypsin and the correlation with Alzheimer’s disease. J. Alzheimers Dis. 2002, 4, 115–122.
Chen, J.; Li, L.; Chin, L. S. Parkinson disease protein DJ-1 converts from a zymogen to a protease by carboxyl-terminal cleavage. Hum. Mol. Genet. 2010, 19, 2395–2408.
Travis, J.; Pike, R.; Imamura, T.; Potempa, J. The role of proteolytic enzymes in the development of pulmonary emphysema and periodontal disease. Am. J. Respir. Crit. Care Med. 1994, 150, S143–S146.
Singh, R. B.; Dandekar, S. P.; Elimban, V.; Gupta, S. K.; Dhalla, N. S. Role of proteases in the pathophysiology of cardiac disease. Mol. Cell. Biochem. 2004, 263, 241–256.
Medcalf, R. L. Fibrinolysis, inflammation, and regulation of the plasminogen activating system. J. Thromb. Haemost. 2007, 5, 132–142.
Arora, P.; Ricks, T. K.; Trejo, J. Protease-activated receptor signalling, endocytic sorting and dysregulation in cancer. J. Cell Sci. 2007, 120, 921–928.
Quirós, P. M.; Langer, T.; López-Otín, C. New roles for mitochondrial proteases in health, ageing and disease. Nat. Rev. Mol. Cell Biol. 2005, 16, 345–359.
Musante, L.; Tataruch, D.; Gu, D. F.; Liu, X. Y.; Forsblom, C.; Groop, P. H.; Holthofer, H. Proteases and protease inhibitors of urinary extracellular vesicles in diabetic nephropathy. J. Diabetes Res. 2015, 2015, 289734.
Edgington-Mitchell, L. E. Pathophysiological roles of proteases in gastrointestinal disease. Am. J. Physiol. Gastrointest. Liver Physiol. 2016, 310, G234–G239.
McCarty, S. M.; Cochrane, C. A.; Clegg, P. D.; Percival, S. L. The role of endogenous and exogenous enzymes in chronic wounds: A focus on the implications of aberrant levels of both host and bacterial proteases in wound healing. Wound Repair Regen. 2012, 20, 125–136.
Koop, G.; van Werven, T.; Roffel, S.; Hogeveen, H.; Nazmi, K.; Bikker, F. J. Protease activity measurement in milk as a diagnostic test for clinical mastitis in dairy cows. J. Dairy Sci. 2015, 98, 4613–4618.
Huis in’t Veld, J. H. J. Microbial and biochemical spoilage of foods: An overview. Int. J. Food Microbiol. 1996, 33, 1–18.
Shen, J. J.; Person, M. D.; Zhu, J. J.; Abbruzzese, J. L.; Li, D. H. Protein expression profiles in pancreatic adenocarcinoma compared with normal pancreatic tissue and tissue affected by pancreatitis as detected by two-dimensional gel electrophoresis and mass spectrometry. Cancer Res. 2004, 64, 9018–9026.
Wang, G. K.; Zhu, J. Q.; Zhang, J. T.; Li, Q.; Li, Y.; He, J.; Qin, Y. W.; Jing, Q. Circulating microRNA: A novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur. Heart J. 2010, 31, 659–666.
Rissin, D. M.; Kan, C. W.; Campbell, T. G.; Howes, S. C.; Fournier, D. R.; Song, L. N.; Piech, T.; Patel, P. P.; Chang, L.; Rivnak, A. J. et al. Singlemolecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nat. Biotechnol. 2010, 28, 595–599.
Liao, H.; Wu, J.; Kuhn, E.; Chin, W.; Chang, B.; Jones, M. D.; O'Neil, S.; Clauser, K. R.; Karl, J.; Hasler, F. et al. Use of mass spectrometry to identify protein biomarkers of disease severity in the synovial fluid and serum of patients with rheumatoid arthritis. Arthritis Rheum. 2004, 50, 3792–3803.
Lee, G.; Eom, K.; Park, J.; Yang, J.; Haam, S.; Huh, Y. M.; Ryu, J. K.; Kim, N. H.; Yook, J. I.; Lee, S. W. et al. Real-time quantitative monitoring of specific peptide cleavage by a proteinase for cancer diagnosis. Angew. Chem. 2012, 124, 5939–5943.
Shi, L. F.; De Paoli, V.; Rosenzweig, N.; Rosenzweig, Z. Synthesis and application of quantum dots FRET-based protease sensors. J. Am. Chem. Soc. 2006, 128, 10378–10379.
Chen, G. C.; Xie, Y. S.; Zhang, H. T.; Wang, P.; Cheung, H. Y.; Yang, M. S.; Sun, H. Y. A general colorimetric method for detecting protease activity based on peptide-induced gold nanoparticle aggregation. RSC Adv. 2014, 4, 6560–6563.
Park, S. Y.; Lee, S. M.; Kim, G. B.; Kim, Y. P. Gold nanoparticle-based fluorescence quenching via metal coordination for assaying protease activity. Gold Bull. 2012, 45, 213–219.
Ingram, A.; Byers, L.; Faulds, K.; Moore, B. D.; Graham, D. SERRS-based enzymatic probes for the detection of protease activity. J. Am. Chem. Soc. 2008, 130, 11846–11847.
Tseng, C. W.; Chang, H. Y.; Chang, J. Y.; Huang, C. C. Detection of mercury ions based on mercury-induced switching of enzyme-like activity of platinum/ gold nanoparticles. Nanoscale 2012, 4, 6823–6830.
Kim, S. H.; Park, M. K.; Kim, J. Y.; Chuong, P. D.; Lee, Y. S.; Yoon, B. S.; Hwang, K. K.; Lim, Y. K. Development of a sandwich ELISA for the detection of Listeria spp. using specific flagella antibodies. J. Vet. Sci. 2005, 6, 41–46.
Hvolbæk, B.; Janssens, T. V. W.; Clausen, B. S.; Falsig, H.; Christensen, C. H.; Nørskov, J. K. Catalytic activity of Au nanoparticles. Nano Today 2007, 2, 14–18.
Deng, H. H.; Weng, S. H.; Huang, S. L.; Zhang, L. N.; Liu, A. L.; Lin, X. H.; Chen, W. Colorimetric detection of sulfide based on target-induced shielding against the peroxidase-like activity of gold nanoparticles. Anal. Chim. Acta 2014, 852, 218–222.
Zhao, D.; Chen, C. X.; Lu, L. X.; Yang, F.; Yang, X. R. A label-free colorimetric sensor for sulfate based on the inhibition of peroxidase-like activity of cysteamine-modified gold nanoparticles. Sens. Actuators B Chem. 2015, 215, 437–444.
Hizir, M. S.; Top, M.; Balcioglu, M.; Rana, M.; Robertson, N. M.; Shen, F. F.; Sheng, J.; Yigit, M. V. Multiplexed activity of perAuxidase: DNA-capped AuNPs act as adjustable peroxidase. Anal. Chem. 2016, 88, 600–605.
Shah, J.; Purohit, R.; Singh, R.; Karakoti, A. S.; Singh, S. ATP-enhanced peroxidase-like activity of gold nanoparticles. J. Colloid Interface Sci. 2015, 456, 100–107.
Turkevich, J.; Stevenson, P. C.; Hillier, J. A study of the nucleation and growth processes in the synthesis of colloidal gold. Discuss. Faraday Soc. 1951, 11, 55–75.
Haddadi, K.; Moussaoui, F.; Hebia, I.; Laurent, F.; Le Roux, Y. E. coli proteolytic activity in milk and casein breakdown. Reprod. Nutr. Dev. 2005, 45, 485–496.
Cupp-Enyard, C. Use of the protease fluorescent detection kit to determine protease activity. J. Vis. Exp. 2009, 30, e1514.
Bellino, M. G.; Calvo, E. J.; Gordillo, G. Adsorption kinetics of charged thiols on gold nanoparticles. Phys. Chem. Chem. Phys. 2004, 6, 424–428.
Liu, Y.; Liu, L. L.; Yuan, M.; Guo, R. Preparation and characterization of casein-stabilized gold nanoparticles for catalytic applications. Colloids Surf. A Physicochem. Eng. Aspects 2013, 417, 18–25.
Liu, Y.; Guo, R. pH-dependent structures and properties of casein micelles. Biophys. Chem. 2008, 136, 67–73.
Wang, S.; Chen, W.; Liu, A. L.; Hong, L.; Deng, H. H.; Lin, X. H. Comparison of the peroxidase-like activity of unmodified, amino-modified, and citrate-capped gold nanoparticles. ChemPhysChem 2012, 13, 1199–1204.
Arnao, M. B.; Acosta, M.; Del Rio, J. A.; García-Canovás, F. Inactivation of peroxidase by hydrogen peroxide and its protection by a reductant agent. Biochim. Biophys. Acta 1990, 1038, 85–89.
Drozd, M.; Pietrzak, M.; Parzuchowski, P. G.; Malinowska, E. Pitfalls and capabilities of various hydrogen donors in evaluation of peroxidase-like activity of gold nanoparticles. Anal. Bioanal. Chem. 2016, 408, 8505–8513.
Chuang, Y. C.; Li, J. C.; Chen, S. H.; Liu, T. Y.; Kuo, C. H.; Huang, W. T.; Lin, C. S. An optical biosensing platform for proteinase activity using gold nanoparticles. Biomaterials 2010, 31, 6087–6095.
Laromaine, A.; Koh, L.; Murugesan, M.; Ulijn, R. V.; Stevens, M. M. Protease-triggered dispersion of nanoparticle assemblies. J. Am. Chem. Soc. 2007, 129, 4156–4157.
Guarise, C.; Pasquato, L.; De Filippis, V.; Scrimin, P. Gold nanoparticlesbased protease assay. Proc. Natl. Acad. Sci. USA 2006, 103, 3978–3982.
Richardson, B. C.; Pearce, K. N. The determination of plasmin in dairy products. N. Z. J. Dairy Sci. Technol. 1981, 16, 209–220.
Politis, I.; Hang, K. F. N. K.; Giroux, R. N. Environmental factors affecting plasmin activity in milk. J. Dairy Sc. 1989, 72, 1713–1718.
Buhl, K. B.; Oxlund, C. S.; Friis, U. G.; Svenningsen, P.; Bistrup, C.; Jacobsen, I. A.; Jensen, B. L. Plasmin in urine from patients with type 2 diabetes and treatment-resistant hypertension activates ENaC in vitro. J. Hypertens. 2014, 32, 1672–1677.
Acknowledgements
The author C. M. and N. L. thank the PhD studentship support from the Department of Employment and Learning for Northern Ireland (DEL); C. C. thanks the strong support from the Central Research Support Funds of Queen’s University Belfast via a start-up grant, the support from the Agri-Food Quest Competence Centre R&D funding programme sponsored by Invest Northern Ireland Agency (Invest NI), and the support from the Queen’s University of Belfast AMR Network (QUBAN) sponsored by the UK’s Engineering and Physical Sciences Research Council (EPSRC).
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at link.springer.com
Electronic supplementary material
Rights and permissions
Open Access: This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
McVey, C., Logan, N., Thanh, N.T.K. et al. Unusual switchable peroxidase-mimicking nanozyme for the determination of proteolytic biomarker. Nano Res. 12, 509–516 (2019). https://doi.org/10.1007/s12274-018-2241-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12274-018-2241-3