Abstract
Rho GTPases are a class of G-proteins involved in several aspects of cellular biology, including the regulation of actin cytoskeleton. The most studied members of this family are RHOA and RAC1 that act in concert to regulate actin dynamics. Recently, Rho GTPases gained much attention as synaptic regulators in the mammalian central nervous system (CNS). In this context, ARHGAP22 protein has been previously shown to specifically inhibit RAC1 activity thus standing as critical cytoskeleton regulator in cancer cell models; however, whether this function is maintained in neurons in the CNS is unknown. Here, we generated a knockout animal model for arhgap22 and provided evidence of its role in the hippocampus. Specifically, we found that ARHGAP22 absence leads to RAC1 hyperactivity and to an increase in dendritic spine density with defects in synaptic structure, molecular composition, and plasticity. Furthermore, arhgap22 silencing causes impairment in cognition and a reduction in anxiety-like behavior in mice. We also found that inhibiting RAC1 restored synaptic plasticity in ARHGAP22 KO mice. All together, these results shed light on the specific role of ARHGAP22 in hippocampal excitatory synapse formation and function as well as in learning and memory behaviors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Rho family of small GTPases (Rho GTPases) is a class of signaling G-proteins known to orchestrate several cellular processes, such as cell migration, vesicle trafficking, and differentiation [1]. Moreover, it has been shown that Rho GTPases control actin cytoskeleton dynamics [2] and therefore regulates CNS synapses [3].
The most extensively studied members of this family in the nervous system are RHOA, RAC1, and CDC42 [4]. While the former is involved in the formation of actin-myosin contractile fibers and focal adhesion, RAC1 and CDC42 play a key role in regulating actin polymerization through the activation of several downstream proteins, such as actin-related protein 2 (ARP2), serine/threonine-protein kinase 1 (PAK1), and Wiskott-Aldrich syndrome protein family member verprolin-homologous protein (WAVE). In addition, these proteins act in concert to exert their functions through reciprocal regulation [5]. In particular, PAK1 is able to directly bind active RAC1 triggering a signal cascade that involves different effectors modulating actin polymerization [6, 7]. Moreover, WAVE is recruited by RAC1-GTP at the plasma membrane and in turn it activate ARP2, a part of the ARP2/3 complex which works as a nucleator platform for the formation of new actin filaments [8, 9].
Rho GTPases act as molecular switches cycling between active and inactive states, when bound to GTP or GDP, respectively [10, 11]. Two classes of proteins are known to regulate Rho GTPases activity: (1) guanine nucleotide exchange factors (GEFs) that promote GDP exchange with GTP and are considered positive modulators and (2) GTPase activating proteins (GAPs) that enhance the intrinsic activity of Rho GTPases thereby inactivating them [12]. Misregulated RHOA, RAC1/RAC3, and CDC42 activity have been associated with cognitive disorders [13,14,15,16,17].
The arhgap22 gene, located on the chromosome 10 in humans and 14 in mice, encodes for ARHGAP22 protein (also called RhoGAP2 and RhoGAP22), a member of the ARHGAP family. This protein is structurally composed of N-terminal pleckstrin-homology (PH) domain, followed by RhoGAP and C-terminal coiled-coil (CC) domains. ARHGAP22 is ubiquitously expressed in mammals, with a prevalence in highly vascularized tissues [18], and has been demonstrated to specifically inhibit RAC1 through its GAP activity [18, 19], thus standing as a critical cytoskeleton regulator. Previous reports have linked ARHGAP22 activity to cell movement and morphology as well as to actin dynamics in cancer, through its RAC1 regulating activity [19, 20]. In addition, ARHGAP22 has also been proposed as cellular transcription regulator [18]. Previously, we found that ARHGAP22 is localized at post-synaptic site of the excitatory synapses and interacts with interleukin-1 receptor accessory protein-like 1 (IL1RAPL1) protein in order to induce dendritic spines formation [21].
In this study, we showed that ARHGAP22 is expressed in the CNS peaking during synaptogenesis. The absence of arhgap22 in mouse brain causes alterations in excitatory synaptic structure, function, and in mice behavior. We also observed that arhgap22 loss of function induces RAC1 hyperactivation and the treatment with NSC23766, an inhibitor of RAC1 activity, rescues some of the defects.
These findings establish the important role of ARHGAP22 protein in excitatory synapse formation with key repercussions on cognitive functions.
Material and Methods
Generation of arhgap22 KO Mice
Arhgap22 KO mice were generated using gene-trapping technique. Mice (strain C57BL/6) were generated from an embryonic stem (ES) cell line (IST13119C7 clone, Texas Institute for Genomic Medicine, TIGM). The retroviral (Omni- Bank Vector 76) cassette contained a splice acceptor sequence (SA) followed by a 5′ selectable marker –geo (a functional fusion between the beta galactosidase and neomycin resistance genes) for identification of successful gene trap events, followed by a poly-adenylation signal (pA). Insertion of the retroviral vector into the arhgap22 gene (intron 3) led to the splicing of the endogenous first 3 exons and the cassette to produce a truncated transcript of arhgap22 gene. 3′ RACE was used to verify the insertion of the cassette into the correct genomic location. The ES cell clone, containing the retroviral cassette in the Arhgap22 gene, was microinjected into C57BL/6 host blastocysts to generate chimeras using standard procedures. Chimeric males were bred to C57BL/6 wild type females for germ line transmission of the knock-out Arhgap22 allele.
Real-Time PCR
mRNA was extracted from murine tissues using Nucleozol Reagent (Macherey Nagel) following manufacturer’s instructions. A total of 1.5 µg of extracted mRNA was used to synthetize cDNA using SuperScript™ VILO™ cDNA Synthesis Kit (Thermo Fisher). Following, arhgap22 specific sequence was amplified from 60 ng of cDNA in the presence of SYBR Green PCR Master Mix (Applied Biosystems) using an Applied Biosystems 7000 Real-Time thermocycler. Parallel PCR reaction was performed with α-actin-specific primers as housekeeping control gene.
The sequences of primers (Sigma Aldrich) were the following: arhgap22 Fw97 (TTCGGCCACAGATAGAGGAT), arhgap22 Rev97 (GTCATCAGATGCTGAACCAGAG), α-actin Fw (AGATGACCCAGATCATGTTTGAGA), α-actin Rev (CCTCGTAGATGGGCACAGTGT).
Each sample was run in triplicate, and the results were analyzed by ABI PRISM 7000 software using the ΔΔCT method to allow the normalization of each sample to the internal standard and comparison with the calibrator.
Rac1-GTP Pulldown Assay
pGex-PakCRIB construct (kind gift from G. Pietrini) was used to produce GST-CRIB fusion protein. GST fusion protein was prepared in BL21 Escherichia coli and purified using standard procedures [22]. To evaluate the levels of active RAC1, we performed the Rac1-GTP pulldown assay. Briefly, hippocampal and cortical samples from adult mice were collected and homogenized in RAC Buffer (10% glycerol, 100 mM NaCl, 10 mM MgCl2, 50 mM Tris HCl pH 7.4, 10% triton, protease inhibitor). Lysates were then centrifuged at 12000 g for 10 min at 4 °C. Supernatants were incubated with 40 μl of S-transferase PAK glutathione-coupled Sepharose 4B beads (GE Healthcare) for 30 min at 4 °C. After 3 washes with RAC buffer, beads were resuspended in sample buffer (4% SDS, 10% 2-mercaptoethanol, 20% glycerol, 0.004% bromophenol blue, 0.125 M Tris–HCl). Bound protein as well as inputs were loaded on polyacrylamide gels and analyzed by Western blotting.
Crude Synaptosomes Preparation
Cortices and hippocampi were dissected from arhgap22 WT and KO adult animals as previously reported [23] and homogenized with a glass-teflon homogenizer in cold HEPES/sucrose buffer (4 mM HEPES [pH 7.4], 0.32 M sucrose, protease inhibitor). After centrifugation at 1000 g at 4 °C for 10 min, Supernatant (S1) were centrifuged at 10000 g at 4 °C for 15 min. Resulting supernatant contained mainly cytosolic components while the pellet was composed of crude synaptosomes (P2). After resuspension in 10 volumes of HEPES/sucrose buffer, pellet was centrifuged at 10000 g for 15 min at 4 °C to purify the preparation. P2 fractions were resuspended in modified RIPA buffer (50 mM TRIS–HCl, 200 mM NaCl, 1 mM EDTA, 1% NP40, 1% Triton X-100, pH 7.4, protease inhibitor cocktail) and then quantified with bicinchoninic acid assay (BCA) (Euroclone) prior to SDS-PAGE and Western blotting.
F/G Actin Ratio
To study F/G ratio, we followed a previously described protocol [24]. Briefly, hippocampal and cortical samples from adult mice [23] were resuspended in actin lysis buffer (10 mM K2HPO4, 100 mM NaF, 50 mM KCl, 2 mM MgCl2, 1 mM EGTA, 0.2 mM DTT, 0.5% Triton, 1 mM sucrose [pH 7]) and centrifuged at 15000 g for 30 min at 4 °C. Supernatants containing G-actin were collected while the pellet was resuspended in 1 volume of actin lysis buffer/1 volume of Buffer II (1.5 mM guanidine hydrochloride, 1 mM sodium acetate, 1 mM CaCl2, 1 mM ATP, 20 mM Tris HCl [pH 7.5]). After 1-h incubation on ice, samples were centrifuged at 15000 g for 30 min at 4 °C. Supernatant containing F-actin was collected. G-actin and F-actin samples were then resuspended in sample buffer prior to Western blotting analyses.
SDS-PAGE and Western Blot
Samples were loaded on polyacrylamide gels at different concentrations (7.5–15%) and then transferred onto nitrocellulose membrane (0.22 μm, GE Healthcare). Membranes were incubated, at room temperature for 2–3 h or overnight at 4 °C in Blocking buffer (non-fat dry milk 5% and TBS Tween-20), with the primary antibodies against the following protein categories: (1) RAC1 and downstream effectors (α-RAC1 (Abcam, 1:500), α-WAVE (SantaCruz Biotechnology,1:500); α-ACTIN (Sigma Aldrich,1:4,000); α-PAK1 and pPAK1 (Cell Signaling, 1:1000), α-ARP2 (SantaCruz Biotechnology,1:500)); (2) α-TUBULIN (Sigma Aldrich,1:40,000) and α-GAPDH (Santa Cruz Biotechnology, 1:2000) as loading controls; (3) subunits of glutamatergic receptors (α-NR1 (Gift from C. Gotti, 1:500), α-NR2A (Sigma Aldrich, 1:1500), α-NR2B (Chemicon, 1:1000); α-GLUA1 (Chemicon, 1:1000); α-GLUA2/3 (Gift from C. Gotti, 1:2500); α-GLUK2 (Sigma Aldrich 1:1000)); (4) post-synaptic proteins (α-CASK1 (Neuromab 1:2500); α-PSD-95 (Neuromab, 1:2500), α-PICK1 (Neuromab; 1:1000), α-GRIP1 (BD transduction laboratories, 1:2000)); and (5) pre-synaptic markers (α-VGAT (SySy, 1:1000), α-synaptophysin (SySy, 1:5000)). After washing, the blots were incubated at room temperature for 1 h with α-rabbit or α-mouse IR Dyes- (LI-COR) conjugated antibodies (1:7500) in Blocking buffer accordingly to the host species of primary antibody. Immunoreactive bands were visualized by Odyssey Infrared Imager (LI-COR).
Golgi and DiI Staining
We performed Golgi staining using FD Rapid GolgiStainTM Kit (FD Neurotechnologies Inc.) according to manufacturer’s instructions. Adult arhgap22 WT and KO male mice (at least 3 per genotype) were deeply anesthetized with isoflurane and transcardially perfused with saline solution (0.9%). Brains were then immersed in impregnation solution for 14 days in the dark at room temperature and subsequently in solution C for 3 days at room temperature. The brains were cut in 100-μm coronal sections with a vibratome (Leica VT1000S). Slices were mounted on gelatin-coated glasses and maintained at room temperature in the dark, allowing them to dry naturally. Sections were then washed in MilliQ water and placed in solution D for 10 min. Subsequently, slices were washed and dehydrated with increasing concentrations of ethanol (50, 70, 95, and 100%) for 4 min each and, as last step, with xylene (3 times, 4 min each). At the end of the procedure, slices were mounted with a coverslip using Permount mounting medium (Fisher Scientific). Acquisition of the stained neurons from the hippocampus was performed using a Leica DMI6000 spinning disk microscope (Leica). Stacks were collected every 0.5 μm using × 63 objective. Analysis of the dendritic spine density was calculated on 100 μm dendrites/neuron using NeuroStudio software.
For DiI analyses, arhgap22 WT and KO mice (at least 3 per genotype) were deeply anesthetized with isoflurane and transcardially perfused with 1.5% paraformaldehyde solution (PFA). Brains were incubated in 1.5% PFA ON at 4 °C and then cut in 3-mm coronal sections with a vibratome (Leica VT1000S). Dil 1,1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine perchlorate (“DiI”; DiIC18(3)) crystals were placed on top of each slice immersed in PBS overnight. A total of 100-μm coronal sections were cut with a vibratome (Leica VT1000S) and mounted with a coverslip using Fluoromount mounting medium (Sigma Aldrich). Acquisition of the stained neurons from the hippocampus was performed using a Zeiss 510 confocal microscope (Carl Zeiss). Stacks were collected every 0.5 μm using × 63 objective. Analysis of the dendritic spine density and spine morphology were performed on 100 μm dendrites/neuron using NeuroStudio software using parameters to define mushroom, stubby, and thin spines as in [25].
Electron Microscopy
Adult arhgap22 WT and KO male mice (3 per genotype) were deeply anesthetized with isoflurane before transcardial perfusion with EM buffer (2.5% glutaraldehyde and 2% paraformaldehyde in 0.15 M sodium cacodylate buffer, pH 7.4). The brains were further fixed in EM buffer for 24 h at 4 °C. A total of 100 μm slices were obtained with a vibratome (Leica VT1000S) and hippocampi manually dissected. Samples were then post-fixed with 2% osmium tetroxide, rinsed, en bloc stained with 1% uranyl acetate, dehydrated, and embedded in epoxy resin (Electron Microscopy Science, Hatfield, PA, USA). Thin Sects. (70–90 nm) were obtained with EM UC6 ultramicrotome (Leica Microsystems, Austria), stained with a saturated solution of uranyl acetate in ethanol 20%, and observed under a Philips CM10 transmission electron microscope (TEM) (FEI, Eindhoven, Netherlands). Analyzed excitatory synapses (apical dendrite layer of the hippocampal CA1 region) were identified on the base of three conditions: (1) the presence in their postsynaptic terminal of the electron-dense post-synaptic density (PSD); (2) the presence of at least 3 synaptic vesicles in the pre-synaptic compartment; (3) the presence of a defined synaptic cleft. Images were acquired at a final magnification of × 25–34,000 using a Morada CCD camera (Olympus, Munster, Germany) and analyzed with ImageJ software (NIH Image).
Electrophysiology
Adult arhgap22 WT and KO male mice (at least 3 per genotype) were used for electrophysiological recordings. Coronal hippocampal slices (thickness, 300/400 μm) were prepared as previously described [26] using a vibratome VT1000S (Leica). After the cutting procedure, brain slices were incubated in aCSF (125 mM NaCl, 2.5 mM KCl, 1.25 mM NaH2PO4, 1 mM MgCl2, 2 mM CaCl2, 25 mM glucose, and 26 mM NaHCO3; pH 7.3, equilibrated with 95% O2 and 5% CO2) for 40 min at 35 °C for miniature excitatory and inhibitory post-synaptic currents (mEPSCs/mIPSCs) or at room temperature for field excitatory post-synaptic potentials (fEPSPs) recordings. mEPSCs and mIPSCs were recorded from CA1 pyramidal neurons in voltage-clamp mode in aCSF supplemented with lidocaine (500 µM) to block voltage-gated sodium channels, and using an internal solution containing (in mM) 130 CsGluconate, 8 CsCl, 2 NaCl, 10 HEPES, 4 EGTA, 4 MgATP, and 0.3 Tris-GTP (pH 7.4). The excitation/inhibition (E/I) balance was calculated through the ratio between mEPSCs and mIPSCs amplitudes recorded from the same neuron [27].
fEPSPs were evoked through Schaffer collateral (SC) stimulation (0.05 Hz of frequency) and recorded from the stratum radiatum of the hippocampal CA1 using aCSF-filled capillaries.
Input–output (I–O) relationship was obtained measuring the slope of the fEPSPs evoked in response to increased stimulation intensity (0–1.0 mA). Stimulus strength was adjusted to give 50% maximal response and long-term potentiation (LTP) was induced by high-frequency stimulation (HFS) (100 stimuli at 100 Hz).
For paired pulse ratio (PPR) experiments, pairs of stimuli were delivered at 50-ms intervals every 20 s (0.05 Hz) and the ratio was calculated by dividing the amplitude of the second response by the first one [28]. Currents and fEPSPs were filtered at 2 kHz and digitized at 20 kHz using Clampex 10.1 software through the patch-clamp amplifier. All the analyses were performed offline with Clampfit 10.1 software.
Microelectrode Array
Adult mice were anesthetized with isoflurane prior to decapitation. Coronal hippocampal slices (400 μm thick) were cut using a vibratome (Leica VT1000S) in an ice-cold high-sucrose protective solution containing (in mM) 87 NaCl, 25 NaHCO3, 2.5 KCl, 0.5 CaCl2, 7 MgCl2, 25 glucose, 75 sucrose, and saturated with 95% O2 and 5% CO2. Before recordings, slices were left to recover for at least 1 h at room temperature in aCSF. APS-MEA has been previously described [29, 30]. Recordings were performed at room temperature and slices were continuously perfused at a rate of 4–5 ml/min with aCSF. After a slice stabilization period (at least 30 min), 4-amynopyridine (4-AP; 100 μm, Sigma-Aldrich) was applied for at least 30 min before activity recordings (5 min per session). All the analyses were conducted off-line using BrainWave software (3Brain Gmbh, Switzerland). Hippocampal activation clusters were defined manually according to mouse brain atlas. A band-pass filter (from 100 to 250 Hz) was applied to the raw data to extract extracellular local field potentials (LFPs). Spikes and LFPs were detected and analyzed through hard threshold (HT) algorithm (spikes, threshold − 100 μV; refractory period, 1 ms; LFPs, high threshold 100 μV; low threshold, − 100 μV; with maximum LFP event duration < 1 s).
Behavioral Tests
All the behavioral tests were conducted on adult arhgap22 WT and KO male mice (6 to 10 per genotype), and depending on the type of test, we replicated the experiments 2/3 times. Animals were kept in 12-h light/dark cycle with food and water ad libitum.
General Health Assessment
Animals were evaluated once a week to check their general health status by measuring their body weight and food intake.
Spontaneous Motor Activity
Mice locomotion integrity was evaluated by recording animals in automated activity cage (43 × 43 × 32 cm) (Ugo Basile, Varese, Italy). Cumulative horizontal and vertical beam breaks were counted for 180 min [31].
Balance Beam
Balance beam test was used to assess mice balance and coordination. Mice were trained and tested to cross beams of different widths (6 and 12 mm). Latency to traverse each beam was recorded [32].
Pole Test
In the vertical pole task, the mouse was placed on a vertical wire-mesh pole with its head facing upwards. Mice were habituated to the task in 2 trials per day for 2 days. On test day (third day), mice were subjected to 5 trials: the total time taken to turn the body and to descend was recorded according to the literature [33]. A cut-off of 60 s was given. Data were shown as mean of 5 trials evaluated during the test day.
Wire Hanging
Muscle strength was evaluated by positioning the mouse on the top of a wire cage lid. After shaking the lid to give the mouse the possibility to grip to the wire, the operator turned upside down the lid. Latency to fall off the wire was recorded [34].
Novel Object Recognition
Novel object recognition test was divided into two distinct phases: the familiarization phase and the object recognition phase. After 10 min of habituation in a cage, two different objects were placed at the center of the arena and mice were free to explore the objects for 20 min. After the familiarization phase, animals were returned to the home cage. The experiment was repeated at different time points (5 min, 120 min, and 24 h later) but every time, one of the familiar object was replaced by a novel one. Object recognition was achieved when mice stayed within 0.5 cm from the novel object with the nose toward the object [35].
Spatial Object Recognition
Spatial object recognition test was divided into two distinct phases: the familiarization phase and the spatial object recognition phase. After 10 min of habituation in an open-field arena, two different objects were placed at different corners of the arena. Mice were able to explore the objects for 20 min. After the familiarization phase, animals were returned to the home cage. After 5 and 120 min, animals were put again in the arena but every time, one of the two object was relocated [36]. The discrimination index was calculated as described in [37].
T-maze
T-maze test was performed to evaluate hippocampal-dependent spatial working memory in rodents. Before the experiment, mice experienced food deprivation until they had reached 85–90% of their free-feeding body weight. Then, animals were placed in the start arm of a maze with a T shape. After leaving the start arm, mice could choose between entering either the left or the right arm to explore the maze. In the acquisition phase, one arm (left or right) was used as the reinforcing area as it contained food. The percentage of animals and number of days to reach the criterion (80% of correct choices for 3 days) was calculated. Each mouse that met the criterion for acquisition was then tested using a reversal phase in which the reinforcer arm was changed [31].
Morris Water Maze
Water maze paradigm was performed as described in [38]. The maze consisted in a tank (diameter 1.5 m) containing water and a hidden platform placed about 1 cm below the water surface. The pool was divided into four quadrants: NW, NE, SW, and SE as reference. Several posters and furniture that provided visual cues were mounted near the maze. During the acquisition phase, mice began the experiment from different points of the pool, while the platform position was maintained (4 trials/day for 4 days). Twenty-four hours after the last trial, the platform was removed from the pool and a probe test was performed. Escape latency during acquisition phase and the time spent in the target zone of the maze during probe phase were recorded.
Elevated Plus Maze
Elevated plus maze was used to evaluate anxiety-related behavior. It is based on an apparatus with a cross shape: two opposite open arms and two opposite enclosed arms separated by a central platform. The apparatus was 50 cm far from the floor and it was placed in a quiet room. After 20 min adaptation to the new environment, mice were placed individually in the center of the apparatus facing an open arm. The number of open- and total-arm entries and the time spent in open arms were recorded for 5 min [39].
Marble Burying
Marble burying test was used to evaluate anxiety-like behavior in mice. Briefly, animals were placed in a standard cage filled with 5-cm depth of bedding and 12 marbles regularly interspersed. The number of marble buried with bedding and the latency to bury were measured [40].
Mice Pharmacological Treatment
Adult arhgap22 WT and KO male mice (at least 3 per group), housed in groups of maximum 4 per home cage and maintained on a 12-h light/dark cycle with food and water ad libitum, received intraperitoneal (IP) injection once a day for 5 days (at 8:00–9:00 AM). The arhgap22 WT and KO mice weight were daily checked before the treatment. Arhgap22 WT mice were treated with saline as a control, while KO mice were treated with saline or the RAC1 inhibitor NSC23766 (Tocris Bioscience; 2,5 mg/kg) [41]. Arhgap22 KO mice were randomly assigned to the saline or the NSC23766 group. Mice received the last IP injection 2 h before the experiments.
Statistical Analyses
All the experiments were performed using at least 3 animals per genotype. Data were expressed as mean ± standard error of the mean (SEM). Statistical analyses were performed using Prism 6 software (GraphPad, San Diego, CA). Statistical significance was determined either by Student t test or ANOVA followed by Bonferroni post hoc tests. A p-value lower than 0.05 was considered statistically significant (*p < 0.05, **p < 0.01, ***p < 0.001).
Animal Use Guidelines
All the experiments followed the guidelines established by the Italian Council on Animal Care and were approved by the Italian Government decree No. 747/2015-PR. All efforts were made to minimize the number of subjects used and their suffering.
Results
Arhgap22 Transcript Is Present in Mouse Brain
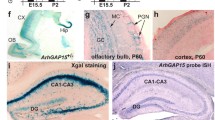
Since little is known about ARHGAP22 expression in the murine organism, we decided to evaluate arhgap22 transcript levels by RT-PCR. First, we analyzed arhgap22 transcript in different murine organs, showing that arhgap22 is expressed in all the analyzed samples including the brain (Fig. 1A). To assess whether the transcript was present at similar levels in different brain areas, we performed RT-PCR on cortical, hippocampal, and cerebellar lysates showing that ARHGAP22 is expressed in all the regions checked with predominance in cortex and hippocampus (Fig. 1B).
ARHGAP22 expression in mouse brain and generation of arhgap22 KO mouse. A Arhgap22 mRNA relative expression in different murine organ. ARHGAP22 is particularly expressed in the kidney, liver, and brain. Values are normalized on α-actin. Error bars indicate ± s.e.m.; B Arhgap22 mRNA expression in different brain areas of adult mice. Arhgap22 transcript is expressed in the cortex and hippocampus while it is present at lower levels in the cerebellum. Values are normalized on α-actin. Error bars indicate ± s.e.m.; C Arhgap22 mRNA expression at different time-points: embryonal day 15 (E15), post-natal day 0 (P0), 7 (P7), 14 (P14), 40 (P40). The highest degree of expression is present at P14 and P40. Values are normalized on α-actin. Error bars indicate ± s.e.m. D Schematic representation of the vector used to randomly insert the gene-trap cassette into arhgap22 wild type locus (intron 3). The gene-trap cassette includes the following elements: 5′ and 3′ flanking long terminal repeats (LTR), splicing acceptor (SA), βGeo marker (βGal and Neo fusion), and a polyadenylation site. E Characteristic genotyping PCR bands of the resulting phenotypes. Arhgap22Fw/Arhgap22Rev genotyping primer pairs hybridize on intron 4 at either side of the insertion point resulting in amplification only from the wild-type allele, whereas theArhgap22/V76R pairs result in amplification from the gene-trap cassette. F Arhgap22 mRNA expression in WT and KO mice brain. KO mice show the almost complete absence of arhgap22 transcript. Values are normalized on α-actin. Error bars indicate ± s.e.m. Arhgap22 WT and KO mice have been tested to assess their general health state. G KO mice present decrease in body weight and food intake. Arhgap22 WT e KO mice have been tested to evaluate also general motor functions. No differences were detected in spontaneous motor activity (H), balance beam (I), pole test (L), and wire hanging (M)
We therefore used cortical and hippocampal samples to study temporal distribution of arhgap22 transcript and we found an increase in the expression during development, reaching the highest expression levels at P14 which are then maintained stable in adulthood (Fig. 1C).
Importantly, the increase in Arhgap22 transcript levels observed at P14 coincides with the temporal window in which synaptogenesis is most prominent [42], thus suggesting that ARHGAP22 might play a role in synapse formation.
Arhgap22 KO Mice Present Hyper-Activation of the RAC1 Pathway in Cortex and Hippocampus
To test this hypothesis and gain additional insights on Arhgap22 function in the CNS, we used a KO mouse for arhgap22 generated by using gene-trapping strategy (Fig. 1D). To confirm the disruption of the functional arhgap22 gene in KO mice, mutant and normal alleles were amplified by PCR. The specific arhgap22 normal allele (400 bp band) was detected in WT and heterozygous animals, while mutated product (200 bp band) was amplified in heterozygous and KO mice (Fig. 1E). We also evaluated arhgap22 transcript levels in cortical and hippocampal lysates from WT and KO mice by RT-PCR. Quantification demonstrated that arhgap22 mRNA was almost completely absent in KO mice (Fig. 1F).
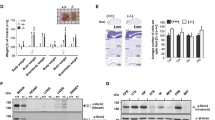
Since this animal model was never characterized before, we evaluated arhgap22 KO mice general health. Although adult KO mice presented a reduction in food intake (area under the curve (AUC)l WT 15.980 ± 0.284 vs KO 14.660 ± 0.402; nmice = 4–5, Student t test * p < 0.05) that led to a slight reduction in body weight (area under the curve (AUC); WT 591.600 ± 7.419 vs KO 552.700 ± 10.730; nmice = 4–5, Student t test * p < 0.05) (Fig. 1G), they did not show alterations in spontaneous motor activity (Fig. 1H), balance beam (Fig. 1I), or in pole and wire hanging test (Fig. 1L, M). Taken together, these results suggest that Arhgap22 KO mice have a good general health, comparable to WT animals.
Considering that ARHGAP22 was shown to selectively inhibit RAC1 in other cellular systems [19], we wondered if the protein had the same function in the brain. Therefore, we performed a GST-pulldown experiment to evaluate the levels of activated RAC1 (RAC1-GTP) in arhgap22-deficient mice compared to WT. Briefly, we incubated cortical and hippocampal lysates with the GST-tagged CRIB domain of PAK1 protein, known to selectively bind active RAC1-GTP [43, 44]. Western blot analysis showed that arhgap22 KO mice presented higher levels of RAC1-GTP compared to WT mice (RAC1GTP/Total RAC1; WT 0.460 ± 0.033 vs KO 0.754 ± 0.041 nmice = 3, Student t test ** p < 0.01) (Fig. 2A).
Arhgap22 KO mice present hyper-active RAC1 pathway in the brain. A Activated RAC1 GST-pulldown on hippocampal and cortical lysates. Red ponceau show the amount of protein in the lysates and GST-CRIB beads used for the experiment. Western blot of total RAC1 (total Rac1) and active RAC1 (Rac1-GTP) are shown. Tubulin was used as internal control (A, top). (A, right histogram) Quantification of the level of active Rac1 normalized on total Rac1 protein. KO animals present elevated level of Rac1-GTP. B Western blot analyses on hippocampal and cortical lysates of WT and KO mice. Levels of RAC1 downstream effectors ARP2/3, WAVE, PAK1, and pPAK1 have been quantified and normalized on GAPDH (left panel). ARP2/3 and WAVE protein levels and the ratio between the active (phosphorylated) and total PAK1 are increased in KO mice compared to WT (right panel). Error bars indicate ± s.e.m. C Analyses of actin levels in cortical and hippocampal lysates from Arhgap22 KO and WT mice. Representative Western blot image (left) and quantification (right) indicate no changes between genotypes. Error bars indicate ± s.e.m. D Analyses of filamentous (F) and globular actin (G) ratio in cortical and hippocampal lysates from Arhgap22 KO and WT mice. Representative Western blot image (left) and quantification (right) indicate that KO animals presented higher F-actin/G-actin ratio compared to WT. Error bars indicate ± s.e.m
The strong increase in RAC1 activity prompted us to study the downstream RAC1 pathway analyzing the levels of different proteins positively regulated by RAC1 activity (ARP2, PAK1, and WAVE) in cortical and hippocampal synaptosomes from WT and KO animals. These data demonstrated that KO mice presented an increase in the total levels of WAVE (WT 1.000 ± 0.061 vs KO 1.24 ± 0.065 nmice = 5, Student t test * p < 0.05) and ARP2 (WT 1.000 ± 0.054 vs KO 1.271 ± 0.089 n = 4mice, Student t test * p < 0.05) proteins and in the levels of active PAK1 (phosphorylated form) (ratio pPAK/PAK; WT 1.000 ± 0.072 vs KO 1.321 ± 0.085 nmice = 5, Student t test * p < 0.05) (Fig. 2B).
Lastly, since the aforementioned proteins are involved in the polymerization of actin in neurons [45, 46], we first evaluated the total level of actin in cortical and hippocampal samples from WT and KO mice and found no statistical difference (Fig. 2C ); and then, we analyzed the ratio between the polymeric (F) and the monomeric (G) form of actin (F/G ratio) in cortex and hippocampus of WT and KO mice. This revealed a strong increase in the F/G ratio (F/G ratio; WT 1.000 ± 0.160 vs KO 1.730 ± 0.090; nmice = 3, Student t test * p < 0.05) (Fig. 2D) in KO animals, suggesting an increased RAC1-mediated actin polymerization.
These results suggest that arhgap22 KO mice present hyper-activated RAC1 which, in turn, might induce actin cytoskeleton defects.
Arhgap22 KO Mice Present Increased Spine Density in Hippocampus
The key role of RAC1 and actin cytoskeleton in the formation and maturation of dendritic spine is well established [47, 48]. Therefore, we evaluated the number of dendritic spines in hippocampal excitatory neurons in fixed brain slices of arhgap22 KO mice by Golgi staining technique. Knockout animals presented a huge increase in the number of dendritic spines (WT 1.000 ± 0.065 vs KO 1.402 ± 0.18; nneurons = 15–16, Student t test * p < 0.05,) (Fig. 3A). However, using this technique, it was not possible to evaluate the morphology of dendritic spines due to the limited resolution of light microscopy [49]. Consequently, in order to perform a morphological analysis of dendritic spines, we used the lipophilic fluorescent molecule DiI (1,1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine perchlorate (“DiI”; DiIC18(3))) which, for its lateral diffusion properties, is able to stain neurons in fixed brain [50, 51]. We, then, performed DiI staining on brain slices of arhgap22 WT and KO mice and analyzed hippocampal excitatory neurons at a confocal microscope. However, after confocal microscopy analysis, we did not find any alteration in spine morphology (Fig. 3B).
Arhgap22 KO mice present increased spine density and altered synaptic molecular composition in hippocampus. A Representative Golgi staining images of hippocampal pyramidal neurons and high magnification of dendrites segments from WT (left) and KO (right) mice. Dendritic spine density measurement indicates increased number of dendritic spines in KO mice compared to WT mice. B Representative images of dendrites segments from WT (left) and KO (right) hippocampal neurons after DiI staining. Scale bar, 5 μm. No differences in spine morphology between genotypes are detectable. C Electron micrographs of asymmetrical synapses of the hippocampal CA1 regions of WT (left) and KO (right) mice. Scale bar, 100 nm. Analyses confirm increased spine density in KO mice and reveal alterations in PSD length and thickness. No pre-synaptic ultrastructural defects are detectable. The total surface area analyzed with stereology was 350 µm2 for each animal. D Representative Western blot images (top) and histograms (bottom) showing the quantification of synaptic markers on crude hippocampal and cortical synaptosomes from adult arhgap22 WT and KO mice. Densitometry analyses performed with Li-Cor technology show that GLUA1 and GLUA2/3 subunits are significantly reduced in arhgap22 KO mice compared to WT
Afterwards, we studied the ultrastructure of hippocampal excitatory synapses through electron microscopy. Firstly, thanks to stereological analyses [28], we confirmed the increased dendritic spine density in KO mice (WT 0.967 ± 0.057 vs KO 1.430 ± 0.053, Student t test ** p < 0.01, with n = 3) (Fig. 3C). Moreover, we found that arhgap22 KO post-synaptic density (PSD) was longer and thinner than WT animals (PSD length (nm), WT 216.25 ± 4.17 vs KO 251.55 ± 6.79; nPSD = 180 nmice = 3, Student t test *** p < 0.001; PSD thickness (nm), WT 38.97 ± 0.54 vs KO 23.03 ± 0.41; nPSD = 180 nmice = 3, Student t test *** p < 0.001) (Fig. 3C). On the pre-synaptic side, we did not find differences in vesicle density between the two genotypes (Fig. 3C).
In conclusion, these data suggest that the hyperactivation of RAC1 and the consequent actin cytoskeleton alterations might be responsible for PSD ultrastructural defects and increased dendritic spines density in arhgap22 KO mice.
Arhgap22-Deficient Synapses Present Altered Molecular Composition
Given the increased dendritic spine density and altered PSD, we asked whether ARHGAP22 absence could alter the molecular composition of synapses. To explore this, we analyzed crude synaptosomes from WT and KO mice by Western blot. We evaluated several synaptic markers including the following: glutamate receptors, PSD scaffold,proteins and pre-synaptic markers. We found reduced levels of GLUA1 (integrated intensity, WT 1.00 ± 0.121 vs KO 0.640 ± 0.041; nmice = 4, Student t test * p < 0.05) and GLUA2/3 (integrated intensity, WT 1.000 ± 0.061 vs KO 0.800 ± 0.043; nmice = 4, Student t test * p < 0.05) AMPA receptors subunits in arhgap22 KO mice compared to WT littermates. No statistical differences for all the other proteins analyzed were detected, even if a tendency toward the decrease for NMDA receptor subunit NR1, GluK2 subunit of kainate receptor, CASK1, and PSD95 was present (Fig. 3D). All together, these data demonstrate altered dendritic spine molecular composition, suggesting an immature profile of arhgap22 KO mice synapses.
Arhgap22 KO Mice Display Reduced Neuronal Excitability
Considering the modifications in spine density and in GLUA1 and GLUA2/3 AMPAR subunits expression, we investigated whether ARHGAP22 absence could induce alterations in Schaffer collateral (SC)-CA1 synaptic functions. Therefore, we recorded mEPSCs and mIPSCs from CA1 hippocampal pyramidal neurons in acute brain slices from arhgap22 WT and KO animals. Interestingly, we found a significant decrease in mEPSC frequency and amplitude in KO mice (frequency (Hz), WT 1.333 ± 0.287 vs KO 0.678 ± 0.287; nneurons = 17–18 nmice = 3, Student t test *p < 0.05; amplitude (pA), WT 14.16 ± 1.21 vs KO 10.21 ± 0.63; nneurons = 17–18 nmice = 3, Student t test *p < 0.05) (Fig. 4A). On the other hand, we did not observe modifications in mIPSCs currents (Fig. 4A), suggesting that ARHGAP22 absence affects specifically the excitatory synapse.
Arhgap22 KO animals present altered excitation/inhibition balance, impaired LTP, and network activity. A Representative traces and quantification of mIPSCs (up) and mEPSCs (down) recorded from CA1 hippocampal pyramidal neurons from arhgap22 WT and KO mice. Quantification (right panel) shows reduced frequency and amplitude of mEPSCs in KO animals while no alterations have been found for mIPSCs. These data reflect the impaired E/I balance in arhgap22 KO mouse. B Representative traces of fEPSPs recorded from hippocampal CA1 of arhgap22 WT and KO mice and quantification of the input/output relationship. C Representative traces and quantification of paired pulse ratio experiments showing no differences in the glutamate release probability between genotypes. D fEPSPs slope quantification before and after HFS shows impairment in LTP at Schaffer’s collaterals-CA1 synapses in arhgap22 KO mice compared to WT. E Time laps representations of propagating events from representative WT (top) and Arhgap22 KO mice (bottom) slices upon 4AP (100 µM) chemical manipulation. Signals from 64 × 64-electrode array are represented in a false color map where each pixel shows the maximal signal variation of each microelectrode (µV). F 5 min raster plots of network hippocampal activity recorded by 200 channels from WT and KO mice. Each dot represents a detected spike, and each line is an electrode. Interictal-like activity is highlighted in black rectangles. G Quantification of mean firing rate (MFR), mean bursting rate (MBR), mean burst duration (MBD), LFP’s events, and Global Synchrony index for Arhgap22 WT, and KO mice are shown
A balanced excitatory and inhibitory (E/I) transmission is fundamental in the CNS, and our data suggested a disruption of this equilibrium in KO mice. As expected, arhgap22 KO mice showed a decrease in the E/I ratio, calculated dividing the mEPSCs and mIPSCs amplitudes for every recorded neuron (E/I ratio, WT 0.6971 ± 0.0561 vs KO 0.4977 ± 0.0402 nneurons = 16 nmice = 3, Student t test ** p < 0.01) (Fig. 4A), suggesting a neurotransmission unbalance toward the inhibition.
Afterwards, we studied if the aforementioned alterations could affect synaptic plasticity in KO mice. To do this, we recorded field excitatory post-synaptic potentials (fEPSPs) at the SC-CA1 synapse in WT and KO mice, stimulating the SC and recording fEPSP from the stratum radiatum of the CA1 hippocampal region. Firstly, we generated an input/output (I/O) curve stimulating the SC with increasing intensity (0–1 mA) and analyzing the fEPSP slope in the CA1 as percentage of the maximal response. We did not found alteration for the I/O curve between genotypes (Fig. 4B). Since we found a decrease in mEPSCs frequency that could suggest an altered glutamate release probability, we performed paired-pulse ratio (PPR) experiments recording two fEPSP responses separated by 50 ms in WT and KO mice. The results demonstrated no changes for glutamate release probability in KO mice, confirming that ARHGAP22 acts mainly at the post-synapse (Fig. 4C).
Next, we investigated if arhgap22 loss-of-function impaired also long-term potentiation (LTP) at SC-CA1 synapse. LTP was induced using a typical NMDA-dependent protocol (100 stimuli at 100 Hz) and fEPSP responses were recorded before (10 min baseline at half-maximal response) and after SC high-frequency stimulation. The analysis of fEPSP slope showed a significant LTP impairment in arhgap22 KO mice compared to WT animals (last 10 min LTP (%), WT 172.6 ± 10.7 vs KO 137.4 ± 11.5 nslices = 7–9 nmice = 3, Student t test *p < 0.05) (Fig. 4D).
All together, these results suggested that ARHGAP22 absence causes significant defects in hippocampal excitatory synaptic function and plasticity, sparing the inhibitory transmission.
Finally, we investigated the hippocampal network activity from adult arhgap22 KO and WT mice using a high-resolution microelectrode array (MEA) [30].
Since we did not observe any spontaneous activity under control conditions (aCSF, not shown), we induced spontaneous epileptiform discharges superfusing slices with aCSF containing 4-aminopyridine (4AP, 100 μM), a classical K+ channel blocker [52]. 4-AP is reported to enhance both inhibitory and excitatory synaptic transmission [53], representing a useful tool to uncover changes in excitability. Thus, arhgap22 WT and KO mice hippocampal slices global activity was recorded onto MEA chips for 5 min (Fig. 4E). Visual inspection of the raster plots indicated that 4-AP elicited isolated spikes, single-channel bursts, and synchronous interictal (I-IC)-like discharges in both genotypes (Fig. 4F) [54]. In particular, I-IC-like discharges are considered to represent local field potentials (LFPs), and involve large neuronal populations in the hippocampus [55].
Specifically, we observed a significant decrease of mean firing rate (MFR (Hz), WT 1.19 ± 0.16 vs KO 0.79 ± 0.08; nslices = 7–9 nmice = 3; Student t test *p < 0.05) and mean bursting rate (MBR (burst/min), WT 4.73 ± 0.95 vs KO 2.69 ± 0.36 burst/min; nslices = 7–9 nmice = 3, Student t test *p < 0.05) in arhgap22 KO mice compared to WT, while mean burst duration (MBD) was unaltered (Fig. 4G, first three left histograms). We also found that LFP events were decreased in KO mice compared to WT mice (LFP/min, WT 11.35 ± 1.29 vs KO 7.27 ± 1.14; nslices = 7–9 nmice = 3, Student t test *p < 0.05) (Fig. 4G, fourth histogram from the left). Furthermore, no difference in global synchronization of the hippocampal networks was observed between KO and WT mice (Fig. 4G, last histogram from the left). These results suggest that the hippocampal network of arhgap22 KO mice is less excitable than WT mice.
Arhgap22 KO Mice Present Learning/Memory Defects and Reduced Anxiety-Like Behavior
Since we found different alterations at morphological, biochemical, and electrophysiological level, we hypothesized that arhgap22 KO animals could present behavioral defects. In particular, considering the impairment in LTP, one of the fundamental mechanisms at the basis of learning and memory [56], we subjected arhgap22 WT and KO mice to cognitive tests.
First, we tested arhgap22 WT and KO littermates in novel object (NOR) and spatial object recognition tests (SOR). In NOR, KO mice presented a significant reduction of the discrimination index between the old and the new object compared to WT at all the time-points analyzed (index 5 min (N-F/N + F), WT 0.2167 ± 0.0606 vs KO − 0.2025 ± 0.0741; index 30 min (N-F/N + F), WT 0.2320 ± 0.0330 vs KO − 0.0900 ± 0.0987; index 120 min (N-F/N + F), WT 0.2790 ± 0.0638 vs KO, − 0.0540 ± 0.0840; nmice = 10 per group, two-way ANOVA **p < 0.01) (Fig. 5A). In particular, the discrimination index can vary between + 1 and − 1. Positive scores indicate that the animals spent more time with the novel object while negative scores indicate preference for the familiar ones and values approaching zero indicate a null preference [57]. The preference for the novel object demonstrate that the familiar object is present in mice memory [58]; on the contrary, a null preference indicates that mice interact by chance with the two objects, suggesting that they forgot the previously encountered object, and thus reflecting a memory impairment. Accordingly, our analysis demonstrated that KO mice discrimination index values, even if negative, are not statistical different respect to zero, suggesting that KO mice interact randomly with the two objects. On the contrary, WT mice showed positive values statistically significant at all the time points tested, indicating a preference for the new object. In SOR, KO animals failed to discriminate between two objects based on spatial cues both at 5 and 120 min after familiarization (index 5 min (N-F/N + F), WT 0.2975 ± 0.0536 vs KO − 0.2100 ± 0.1460; index 120 min (N-F/N + F), WT 0.1938 ± 0.0454 vs KO − 0.0900 ± 0.06071; nmice = 9 per group, two-way ANOVA *p < 0.05, **p < 0.01) (Fig. 5B). These results indicate that alteration in hippocampal functions in arhgap22 KO mice are reflected by defects in recognition and spatial learning and/or memory.
Arhgap22 KO mice present learning/memory defects and reduced anxiety-like behaviors. A, B Representative scheme of the NOR test (A, top) and SOR test (B, top) are shown. Arhgap22 KO mice present altered capability for episodic (novel object) and spatial memory (spatial object recognition). C Schematic representation of T-Maze test is shown. As indicated by quantification, arhgap22 KO mice present a significant impairment during T-Maze acquisition phase but not in the reversal phase. D Scheme of Morris Water maze test during trial and probe phase. Arhgap22 KO mice spent more time trying to reach the target zone during the trial phase. E Representative scheme of elevated plus maze test (left). Time spent in the open arm and number of entries in the open arm indicate that KO mice are less anxious than WT mice as shown by quantification. F Representative scheme of marble-burying test (left). Marble-burying test presents reduced number of marbles buried and an increase latency to the first burial
Spatial learning and memory were additionally analyzed through T-maze test. Also, in this case, we found differences between the two genotypes; indeed, arhgap22 KO mice showed an increased latency to reach the criterion in the acquisition phase compared to WT (days, WT 4.00 ± 0.03 vs KO 6.60 ± 1.03; nmice = 6 per group, two-way ANOVA with Bonferroni post hoc test *p < 0.05) (Fig. 5C). In contrast, no differences were found during the reversal phase although a trend was detectable.
We therefore subjected mice to the water maze test, a hippocampus-dependent paradigm [59]. Knock-out mice showed an upward trend in the latency to find the platform during the trial phase. During the probe test, arhgap22 KO mice spent less time in the target quadrant compared to WT mice (time in the target (%), WT 51.875 ± 0.048331 vs KO 14.82 ± 6.2982; nmice = 6 per group, Student t test **p < 0.01) (Fig. 5D).
Since the hippocampus has been reported to influence the innate anxiety behavior [60], we analyzed also this aspect in our mice. We therefore subjected the mice to two commonly used tests: elevated plus maze (EPM) and marble-burying tests. In EPM, KO mice entered more times (open arm entries (N), WT 39.330 ± 3.199 vs KO 66.180 ± 5.117; nmice = 6 per group, Student t test **p < 0.01) and spent more time in the open arm compared to WT (open arm time (%), 18.600 ± 2.185 vs KO, 62.270 ± 9.474; nmice = 6 per group, Student t test ***p < 0.001) (Fig. 5E). No differences in the total number of entries were detected. Lastly, in marble-burying test, KO animals buried fewer marble stones (buried marbles (N), WT 10.000 ± 1.633 vs KO 3.714 ± 0.8081 nmice = 7 per group, Student t test **p < 0.01) and presented an increased latency to bury compared to WT littermates (latency (s), WT 186.500 ± 77.690 vs KO 479.700 ± 98.520 nmice = 6 per group, Student t test *p < 0.05) (Fig. 5F).
These data demonstrate that arhgap22 KO mice present impairment in learning and memory formation and reduced anxiety-like behavior.
Treatment with Rac1 Inhibitor NSC23766 Rescued Synaptic Alterations in arhgap22 KO Mice
Next, we investigated whether treatment with an inhibitor of RAC1 could rescue the defects observed in arhgap22 KO mice For this purpose, we evaluated synaptic functions after treatment with NSC23766, a compound previously reported to inhibit RAC1 activity [61]. The chronic treatment with NSC23766 (2 mg/kg of NSC23766 or vehicle once a day for 5 days) induced a normalization of RAC1 activity in KO mice (Fig. 6A, B). Noteworthy, vehicle-treated KO mice showed increased level of Rac1-GTP, supporting the specificity of our result. The NSC23766 treatment completely restored the normal protein expression level of GluA1 subunit of AMPA receptor as well as WAVE and ARP2, but not of GluA2/3 AMPA receptor subunit (Fig. 6C and data not shown). Moreover, the drug administration induced full recovery of LTP in KO mice (Fig. 6D).
RAC1 inhibition by NSC23766 ameliorates the synaptic phenotype in Arhgap22 KO mice. A Representative scheme of pharmacologic treatment. B Western blot (left) and quantification (right) showing the normalization of RAC1 activity after NSC treatment in KO mice. C Western blot (left) and quantification (right) showing the normalization of GluA1, WAVE, and ARP2 protein expression after NSC treatment in KO mice. D Quantification of LTP induction upon RAC1 activity normalization showing the full recovery of synaptic plasticity in KO mice
All together, these results demonstrated that RAC1 activity normalization was able to restore some of the defects showed by KO mice.
Discussion
This work stems from two main previous observations: (1) ARHGAP22 regulates RAC1 activity in cancer cell models; and (2) RAC1 is a key player in controlling actin cytoskeleton dynamics in neurons. These evidences prompted us to investigate whether ARHGAP22-mediated regulation of RAC1 in neurons could coordinate synapse formation and plasticity, given the importance of actin cytoskeleton dynamics in these processes.
We observed that ARHGAP22 is present in several mouse tissues [18] including the brain, where is predominantly expressed in cortex and hippocampus. This is not surprising, since ARHGAP22 protein was previously detected in the post-synaptic compartment of cultured rat hippocampal neurons [21]. Interestingly, ARHGAP22 reaches the maximum expression level at P14, when synaptogenesis peaks [42], similarly to some RAC1 GEF and GAP proteins [62, 63]. Moreover, in arhgap22 KO mice, we found a hyper-activity of RAC1 and an upregulation of ARP2, PAK1, and WAVE, proteins known to be involved in neuronal actin dynamics [45, 46]. Coherently, in absence of ARHGAP22, we found an increased F/G actin ratio and, as expected, an increased hippocampal dendritic spine density.
Generally, an increased number of dendritic spines positively correlates with an increase in AMPAR content. Surprisingly, in KO mice, we found a decrease in AMPAR subunits expression. On the other hand, several studies have demonstrated that actin dynamic is critical for AMPAR trafficking and function [64]. Therefore, we can speculate that the reduction in Arhgap22 KO mice AMPAR subunits expression could be the effect of an impaired exocytosis, endocytosis, and/or endosomal recycling that makes dendritic spines less functionally active.
Structurally, we found that KO mice present a roughly 40% reduction in the PSD thickness. This parameter has been suggested to depend on protein translocation from cytoplasm to PSD [65] and on AMPA receptors content, with smaller PSD containing less AMPA receptors and vice versa [66,67,68,69]. This result might support the hypothesis of impairment in AMPAR trafficking. It is widely accepted that neuronal post-synaptic function depends on post-synaptic receptor density and/or efficacy. Coherently, in arhgap22 KO mice, we found decreased glutamatergic currents, while the inhibitory transmission was unchanged, suggesting that ARHGAP22 absence affects specifically the excitatory synapse. In accordance, we found an unbalance toward inhibition in arhgap22 KO mice. Interestingly, arhgap22 KO mice showed decreased LTP at the SC-CA1 synapse. A similar effect was reported in mice after the hippocampal expression of a constitutively active form of RAC1 [70]. This result might be explained by two main hypotheses: (1) the reduction in AMPAR subunits impairs the removal of the Mg2+ block on NMDA receptor, which activation is crucial for LTP induction [26]; and/or (2) a defect in the dendritic spines structural plasticity [71], driven by the increased RAC1-mediated actin polymerization. Since a proper actin dynamic is essential for spinogenesis [48, 72], synaptic plasticity [71], and AMPAR trafficking [64], it is likely that in arghap22 KO mice, dendritic spines, even if increased in number, are in a structural and molecular “crystallized” conformation due to the strong increase in actin polymerization. This phenomenon would make dendritic spines less dynamic and less responsive to stimuli, both during formation and plasticity-induced reorganization.
As discussed above, patch-clamp data suggested an altered E/I balance in arhgap22 KO mice. However, despite being the gold standard for studying neuronal function, patch-clamp gives information about a limited number of neurons and offers poor spatial resolution. To integrate patch-clamp results in a more circuit-based view, we performed MEA experiments using high electrode density probes (4096 electrodes). We found a decreased activity in terms of mean firing and bursting rate in arhgap22 KO mice hippocampus, suggesting a hypo-functionality of the circuit and supporting the neuronal E/I unbalance. Additionally, we also found that arhgap22 KO mice present a decrease in the frequency of LFP insurgence. Interestingly, blocking ionotropic glutamatergic receptors decreases amplitude and frequency of LFPs [73]; we can therefore speculate that the reduced LFP frequency in our model might be the result of the diminished AMPA-dependent glutamatergic activity. Furthermore, being the LFP, a summation of excitatory and inhibitory signals of a large neuronal population, it is also reasonable to assume that an impaired E/I balance could induce alteration in LFP.
Alterations in hippocampal synaptic structure and function are often reported in animal models of intellectual disability that, in turn, present impairments in learning and memory behaviors [26, 74, 75]. Concordantly, when arhgap22 KO and WT mice were subjected to a battery of behavioral test, we found that the former presented strong impairments in learning and memory formation. This is not surprising, since several proteins with RHOGAP activity, such as OLIGOPHRENIN1, ARHGAP15, and ARHGAP33 for instance, have been linked to higher cognitive functions [76,77,78]. In addition, an upregulation of RAC1 activity has also been found in fragile X syndrome, which is characterized by cognitive dysfunctions, aberrant plasticity, and immature dendritic spines [79], partially mimicking what we found in arhgap22 KO.
Moreover, we also observed a significant decrease in anxiety-like behaviors in arhgap22 KO mice. This is in accordance with evidence demonstrating that a disequilibrium between excitation and inhibition might be responsible for anxiety [80]. In particular, it has been demonstrated that GABAA receptors antagonism increases anxiety-like behaviors while blocking AMPA receptors induces an opposite effect [80], resembling what we found in arhgap22 KO mice.
We hypothesized RAC1 hyper-activity as leading cause of the KO mice defects, then we tried to pharmacologically inhibit RAC1 activity in our model. Interestingly, treating KO mice with NSC23766 drug, previously reported to inhibit RAC1 activity (56), we were able to rescue some of the KO mice defects. In particular, we found that NSC23766 treatment of KO mice re-established RAC1 activity, and restored the expression of GluA1 AMPAR subunit as well as WAVE and ARP2. Importantly, RAC1 inhibition fully recovered LTP.
In conclusion, our results shed light on ARHGAP22 protein as an important player in excitatory synapse formation and function as well as in cognition. These activities are likely to be orchestrated through the regulation of RAC1-mediated actin dynamics since we demonstrated that normalizing RAC1 activity rescued several defects in KO mice, supporting RAC1 activity modulation as a promising approach for cognitive dysfunctions.
Data Availability
The authors confirm that the data supporting the findings of this study are available within the article and are available on request from the corresponding authors, LM and MP.
References
Zamboni V, Jones R, Umbach A, et al. (2018) Rho gtpases in intellectual disability: from genetics to therapeutic opportunities. Int J Mol Sci. https://doi.org/10.3390/ijms19061821
Sit S-T, Manser E (2011) Rho GTPases and their role in organizing the actin cytoskeleton. J Cell Sci 124:679–683. https://doi.org/10.1242/jcs.064964
Ba W, Nadif Kasri N (2017) RhoGTPases at the synapse: an embarrassment of choice. Small GTPases 8:106–113. https://doi.org/10.1080/21541248.2016.1206352
Burridge K, Wennerberg K (2004) Rho and Rac take center stage. Cell 116:167–179. https://doi.org/10.1016/s0092-8674(04)00003-0
Guilluy C, Garcia-Mata R, Burridge K (2011) Rho protein crosstalk: another social network? Trends Cell Biol 21:718–726. https://doi.org/10.1016/j.tcb.2011.08.002
Sells MA, Knaus UG, Bagrodia S et al (1997) Human p21-activated kinase (Pak1) regulates actin organization in mammalian cells. Curr Biol 7:202–210. https://doi.org/10.1016/s0960-9822(97)70091-5
Mierke CT, Puder S, Aermes C et al (2020) Effect of PAK inhibition on cell mechanics depends on rac1. Front Cell Dev Biol 8:13. https://doi.org/10.3389/fcell.2020.00013
Chen B, Chou H-T, Brautigam CA, et al. (2017) Rac1 GTPase activates the WAVE regulatory complex through two distinct binding sites. Elife. https://doi.org/10.7554/eLife.29795
Tahirovic S, Hellal F, Neukirchen D et al (2010) Rac1 regulates neuronal polarization through the WAVE complex. J Neurosci 30:6930–6943. https://doi.org/10.1523/JNEUROSCI.5395-09.2010
Hodge RG, Ridley AJ (2016) Regulating Rho GTPases and their regulators. Nat Rev Mol Cell Biol 17:496–510. https://doi.org/10.1038/nrm.2016.67
Jaffe AB, Hall A (2005) Rho GTPases: biochemistry and biology. Annu Rev Cell Dev Biol 21:247–269. https://doi.org/10.1146/annurev.cellbio.21.020604.150721
Bos JL, Rehmann H, Wittinghofer A (2007) GEFs and GAPs: critical elements in the control of small G proteins. Cell 129:865–877. https://doi.org/10.1016/j.cell.2007.05.018
Kim IH, Wang H, Soderling SH, Yasuda R (2014) Loss of Cdc42 leads to defects in synaptic plasticity and remote memory recall. Elife. https://doi.org/10.7554/eLife.02839
Richter M, Murtaza N, Scharrenberg R et al (2019) Altered TAOK2 activity causes autism-related neurodevelopmental and cognitive abnormalities through RhoA signaling. Mol Psychiatry 24:1329–1350. https://doi.org/10.1038/s41380-018-0025-5
Wang X, Liu D, Wei F et al (2020) Stress-sensitive protein Rac1 and its involvement in neurodevelopmental disorders. Neural Plast 2020:8894372. https://doi.org/10.1155/2020/8894372
Reijnders MRF, Ansor NM, Kousi M et al (2017) RAC1 missense mutations in developmental disorders with diverse phenotypes. Am J Hum Genet 101:466–477. https://doi.org/10.1016/j.ajhg.2017.08.007
Ramakers GJA, Wolfer D, Rosenberger G et al (2012) Dysregulation of Rho GTPases in the αPix/Arhgef6 mouse model of X-linked intellectual disability is paralleled by impaired structural and synaptic plasticity and cognitive deficits. Hum Mol Genet 21:268–286. https://doi.org/10.1093/hmg/ddr457
Aitsebaomo J, Wennerberg K, Der CJ et al (2004) p68RacGAP is a novel GTPase-activating protein that interacts with vascular endothelial zinc finger-1 and modulates endothelial cell capillary formation. J Biol Chem 279:17963–17972. https://doi.org/10.1074/jbc.M311721200
Sanz-Moreno V, Gadea G, Ahn J et al (2008) Rac activation and inactivation control plasticity of tumor cell movement. Cell 135:510–523. https://doi.org/10.1016/j.cell.2008.09.043
Mori M, Saito K, Ohta Y (2014) ARHGAP22 localizes at endosomes and regulates actin cytoskeleton. PLoS ONE 9:e100271. https://doi.org/10.1371/journal.pone.0100271
Valnegri P, Montrasio C, Brambilla D et al (2011) The X-linked intellectual disability protein IL1RAPL1 regulates excitatory synapse formation by binding PTPδ and RhoGAP2. Hum Mol Genet 20:4797–4809. https://doi.org/10.1093/hmg/ddr418
Sambrook J, Russell DW (2006) Detection of protein-protein interactions using the GST fusion protein pulldown technique. CSH Protoc. https://doi.org/10.1101/pdb.prot3757
Spijker S (2011) Dissection of Rodent Brain Regions. In: Li KW (ed) Neuroproteomics. Humana Press, Totowa, NJ, pp 13–26
Pyronneau A, He Q, Hwang J-Y et al (2017) Aberrant Rac1-cofilin signaling mediates defects in dendritic spines, synaptic function, and sensory perception in fragile X syndrome. Sci Signal. https://doi.org/10.1126/scisignal.aan0852
Moretto E, Longatti A, Murru L et al (2019) TSPAN5 enriched microdomains provide a platform for dendritic spine maturation through neuroligin-1 clustering. Cell Rep 29:1130-1146.e8. https://doi.org/10.1016/j.celrep.2019.09.051
Murru L, Vezzoli E, Longatti A et al (2017) Pharmacological modulation of AMPAR rescues intellectual disability-like phenotype in Tm4sf2-/y mice. Cereb Cortex 27:5369–5384. https://doi.org/10.1093/cercor/bhx221
Sakimoto Y, Mizuno J, Kida H et al (2019) Learning promotes subfield-specific synaptic diversity in hippocampal CA1 neurons. Cereb Cortex 29:2183–2195. https://doi.org/10.1093/cercor/bhz022
Folci A, Murru L, Vezzoli E et al (2016) Myosin ixa binds AMPAR and regulates synaptic structure, LTP, and cognitive function. Front Mol Neurosci 9:1. https://doi.org/10.3389/fnmol.2016.00001
Imfeld K, Neukom S, Maccione A et al (2008) Large-scale, high-resolution data acquisition system for extracellular recording of electrophysiological activity. IEEE Trans Biomed Eng 55:2064–2073. https://doi.org/10.1109/TBME.2008.919139
Ferrea E, Maccione A, Medrihan L et al (2012) Large-scale, high-resolution electrophysiological imaging of field potentials in brain slices with microelectronic multielectrode arrays. Front Neural Circuits 6:80. https://doi.org/10.3389/fncir.2012.00080
Sala M, Braida D, Lentini D et al (2011) Pharmacologic rescue of impaired cognitive flexibility, social deficits, increased aggression, and seizure susceptibility in oxytocin receptor null mice: a neurobehavioral model of autism. Biol Psychiatry 69:875–882. https://doi.org/10.1016/j.biopsych.2010.12.022
Luong TN, Carlisle HJ, Southwell A, Patterson PH (2011) Assessment of motor balance and coordination in mice using the balance beam. J Vis Exp. https://doi.org/10.3791/2376
Hickey MA, Kosmalska A, Enayati J et al (2008) Extensive early motor and non-motor behavioral deficits are followed by striatal neuronal loss in knock-in Huntington’s disease mice. Neuroscience 157:280–295. https://doi.org/10.1016/j.neuroscience.2008.08.041
Crawley JN (1999) Behavioral phenotyping of transgenic and knockout mice: experimental design and evaluation of general health, sensory functions, motor abilities, and specific behavioral tests. Brain Res 835:18–26. https://doi.org/10.1016/s0006-8993(98)01258-x
Pan D, Sciascia A, Vorhees CV, Williams MT (2008) Progression of multiple behavioral deficits with various ages of onset in a murine model of Hurler syndrome. Brain Res 1188:241–253. https://doi.org/10.1016/j.brainres.2007.10.036
Kenney JW, Adoff MD, Wilkinson DS, Gould TJ (2011) The effects of acute, chronic, and withdrawal from chronic nicotine on novel and spatial object recognition in male C57BL/6J mice. Psychopharmacology 217:353–365. https://doi.org/10.1007/s00213-011-2283-7
Pitsikas N, Rigamonti AE, Cella SG et al (2001) Effects of molsidomine on scopolamine-induced amnesia and hypermotility in the rat. Eur J Pharmacol 426:193–200. https://doi.org/10.1016/s0014-2999(01)01164-5
Morris R (1984) Developments of a water-maze procedure for studying spatial learning in the rat. J Neurosci Methods 11:47–60. https://doi.org/10.1016/0165-0270(84)90007-4
Hogg S (1996) A review of the validity and variability of the elevated plus-maze as an animal model of anxiety. Pharmacol Biochem Behav 54:21–30. https://doi.org/10.1016/0091-3057(95)02126-4
Silverman JL, Pride MC, Hayes JE et al (2015) GABAB receptor agonist R-baclofen reverses social deficits and reduces repetitive behavior in two mouse models of autism. Neuropsychopharmacology 40:2228–2239. https://doi.org/10.1038/npp.2015.66
Veluthakal R, Sidarala V, Kowluru A (2016) NSC23766, a known inhibitor of Tiam1-Rac1 signaling module, prevents the onset of type 1 diabetes in the NOD mouse model. Cell Physiol Biochem 39:760–767. https://doi.org/10.1159/000445666
Moyer CE, Zuo Y (2018) Cortical dendritic spine development and plasticity: insights from in vivo imaging. Curr Opin Neurobiol 53:76–82. https://doi.org/10.1016/j.conb.2018.06.002
Manser E, Leung T, Salihuddin H et al (1994) A brain serine/threonine protein kinase activated by Cdc42 and Rac1. Nature 367:40–46. https://doi.org/10.1038/367040a0
Massari S, Perego C, Padovano V et al (2009) LIN7 mediates the recruitment of IRSp53 to tight junctions. Traffic 10:246–257. https://doi.org/10.1111/j.1600-0854.2008.00854.x
Hotulainen P, Hoogenraad CC (2010) Actin in dendritic spines: connecting dynamics to function. J Cell Biol 189:619–629. https://doi.org/10.1083/jcb.201003008
Costa JF, Dines M, Lamprecht R (2020) The role of rac gtpase in dendritic spine morphogenesis and memory. Front Synaptic Neurosci 12:12. https://doi.org/10.3389/fnsyn.2020.00012
Govek E-E, Newey SE, Van Aelst L (2005) The role of the Rho GTPases in neuronal development. Genes Dev 19:1–49. https://doi.org/10.1101/gad.1256405
Basu S, Lamprecht R (2018) The role of actin cytoskeleton in dendritic spines in the maintenance of long-term memory. Front Mol Neurosci 11:143. https://doi.org/10.3389/fnmol.2018.00143
Kang HW, Kim HK, Moon BH, et al. (2017) Comprehensive review of golgi staining methods for nervous tissue. AM 47:63–69. doi: https://doi.org/10.9729/AM.2017.47.2.63
Kim BG, Dai H-N, McAtee M et al (2007) Labeling of dendritic spines with the carbocyanine dye DiI for confocal microscopic imaging in lightly fixed cortical slices. J Neurosci Methods 162:237–243. https://doi.org/10.1016/j.jneumeth.2007.01.016
Mahmmoud RR, Sase S, Aher YD et al (2015) Spatial and working memory is linked to spine density and mushroom spines. PLoS ONE 10:e0139739. https://doi.org/10.1371/journal.pone.0139739
Avoli M, de Curtis M (2011) GABAergic synchronization in the limbic system and its role in the generation of epileptiform activity. Prog Neurobiol 95:104–132. https://doi.org/10.1016/j.pneurobio.2011.07.003
Rutecki PA, Lebeda FJ, Johnston D (1987) 4-Aminopyridine produces epileptiform activity in hippocampus and enhances synaptic excitation and inhibition. J Neurophysiol 57:1911–1924
D’Antuono M, Köhling R, Ricalzone S et al (2010) Antiepileptic drugs abolish ictal but not interictal epileptiform discharges in vitro. Epilepsia 51:423–431. https://doi.org/10.1111/j.1528-1167.2009.02273.x
Queiroz CM, Gorter JA, Lopes da Silva FH, Wadman WJ (2009) Dynamics of evoked local field potentials in the hippocampus of epileptic rats with spontaneous seizures. J Neurophysiol 101:1588–1597. https://doi.org/10.1152/jn.90770.2008
Neves G, Cooke SF, Bliss TVP (2008) Synaptic plasticity, memory and the hippocampus: a neural network approach to causality. Nat Rev Neurosci 9:65–75. https://doi.org/10.1038/nrn2303
Antunes M, Biala G (2012) The novel object recognition memory: neurobiology, test procedure, and its modifications. Cogn Process 13:93–110. https://doi.org/10.1007/s10339-011-0430-z
Burke SN, Wallace JL, Nematollahi S et al (2010) Pattern separation deficits may contribute to age-associated recognition impairments. Behav Neurosci 124:559–573. https://doi.org/10.1037/a0020893
Morris RG, Garrud P, Rawlins JN, O’Keefe J (1982) Place navigation impaired in rats with hippocampal lesions. Nature 297:681–683. https://doi.org/10.1038/297681a0
Jimenez JC, Su K, Goldberg AR et al (2018) Anxiety cells in a hippocampal-hypothalamic circuit. Neuron 97:670-683.e6. https://doi.org/10.1016/j.neuron.2018.01.016
Tejada-Simon MV (2015) Modulation of actin dynamics by Rac1 to target cognitive function. J Neurochem 133:767–779. https://doi.org/10.1111/jnc.13100
Tcherkezian J, Lamarche-Vane N (2007) Current knowledge of the large RhoGAP family of proteins. Biol Cell 99:67–86. https://doi.org/10.1042/BC20060086
Ma X-M, Huang J, Wang Y et al (2003) Kalirin, a multifunctional Rho guanine nucleotide exchange factor, is necessary for maintenance of hippocampal pyramidal neuron dendrites and dendritic spines. J Neurosci 23:10593–10603
Hanley JG (2014) Actin-dependent mechanisms in AMPA receptor trafficking. Front Cell Neurosci 8:381. https://doi.org/10.3389/fncel.2014.00381
Tao-Cheng J-H (2019) Stimulation induces gradual increases in the thickness and curvature of postsynaptic density of hippocampal CA1 neurons in slice cultures. Mol Brain 12:44. https://doi.org/10.1186/s13041-019-0468-x
Takumi Y, Ramírez-León V, Laake P et al (1999) Different modes of expression of AMPA and NMDA receptors in hippocampal synapses. Nat Neurosci 2:618–624. https://doi.org/10.1038/10172
Shinohara Y, Hirase H, Watanabe M et al (2008) Left-right asymmetry of the hippocampal synapses with differential subunit allocation of glutamate receptors. Proc Natl Acad Sci USA 105:19498–19503. https://doi.org/10.1073/pnas.0807461105
Fukazawa Y, Shigemoto R (2012) Intra-synapse-type and inter-synapse-type relationships between synaptic size and AMPAR expression. Curr Opin Neurobiol 22:446–452. https://doi.org/10.1016/j.conb.2012.01.006
Chen X, Levy JM, Hou A et al (2015) PSD-95 family MAGUKs are essential for anchoring AMPA and NMDA receptor complexes at the postsynaptic density. Proc Natl Acad Sci USA 112:E6983–E6992. https://doi.org/10.1073/pnas.1517045112
Liu Y, Du S, Lv L et al (2016) Hippocampal activation of rac1 regulates the forgetting of object recognition memory. Curr Biol 26:2351–2357. https://doi.org/10.1016/j.cub.2016.06.056
Borovac J, Bosch M, Okamoto K (2018) Regulation of actin dynamics during structural plasticity of dendritic spines: Signaling messengers and actin-binding proteins. Mol Cell Neurosci 91:122–130. https://doi.org/10.1016/j.mcn.2018.07.001
Sekino Y, Kojima N, Shirao T (2007) Role of actin cytoskeleton in dendritic spine morphogenesis. Neurochem Int 51:92–104. https://doi.org/10.1016/j.neuint.2007.04.029
Gonzalez-Sulser A, Wang J, Queenan BN et al (2012) Hippocampal neuron firing and local field potentials in the in vitro 4-aminopyridine epilepsy model. J Neurophysiol 108:2568–2580. https://doi.org/10.1152/jn.00363.2012
Zapata J, Moretto E, Hannan S et al (2017) Epilepsy and intellectual disability linked protein Shrm4 interaction with GABABRs shapes inhibitory neurotransmission. Nat Commun 8:14536. https://doi.org/10.1038/ncomms14536
Belichenko NP, Belichenko PV, Kleschevnikov AM et al (2009) The “Down syndrome critical region” is sufficient in the mouse model to confer behavioral, neurophysiological, and synaptic phenotypes characteristic of Down syndrome. J Neurosci 29:5938–5948. https://doi.org/10.1523/JNEUROSCI.1547-09.2009
Billuart P, Bienvenu T, Ronce N et al (1998) Oligophrenin-1 encodes a rhoGAP protein involved in X-linked mental retardation. Nature 392:923–926. https://doi.org/10.1038/31940
Zamboni V, Armentano M, Sarò G et al (2016) Disruption of ArhGAP15 results in hyperactive Rac1, affects the architecture and function of hippocampal inhibitory neurons and causes cognitive deficits. Sci Rep 6:34877. https://doi.org/10.1038/srep34877
Schuster S, Rivalan M, Strauss U et al (2015) NOMA-GAP/ARHGAP33 regulates synapse development and autistic-like behavior in the mouse. Mol Psychiatry 20:1120–1131. https://doi.org/10.1038/mp.2015.42
Bongmba OYN, Martinez LA, Elhardt ME et al (2011) Modulation of dendritic spines and synaptic function by Rac1: a possible link to Fragile X syndrome pathology. Brain Res 1399:79–95. https://doi.org/10.1016/j.brainres.2011.05.020
Berg L, Eckardt J, Masseck OA (2019) Enhanced activity of pyramidal neurons in the infralimbic cortex drives anxiety behavior. PLoS ONE 14:e0210949. https://doi.org/10.1371/journal.pone.0210949
Acknowledgements
We thank all the authors for feedback on the manuscript and fruitful discussions.
Funding
This work was supported by Telethon Italy (Grant number GGP17283) to MP, Ministero della Salute (Grant number GR‐2016‐02361366) to LM, and Fondazione Zardi Gori (for providing fellowship to LP) and Regione Lombardia for providing fellowship to AL (Grant iPS Light ID 227333).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by A. Longatti, L. Ponzoni, E. Moretto, G. Giansante, N. Lattuada, MN. Colombo, and L. Murru. The first draft of the manuscript was written by A. Longatti, L. Murru, and M. Passafaro and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics Approval
Italian Government decree No. 747/2015-PR.
Consent to Participate
N/A
Consent for Publication
All the authors consent to publish this manuscript in Molecular Neurobiology.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Luca Murru and Maria Passafaro share senior authorship
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Longatti, A., Ponzoni, L., Moretto, E. et al. Arhgap22 Disruption Leads to RAC1 Hyperactivity Affecting Hippocampal Glutamatergic Synapses and Cognition in Mice. Mol Neurobiol 58, 6092–6110 (2021). https://doi.org/10.1007/s12035-021-02502-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-021-02502-x