Abstract
Analysis of medical videos from the human gastrointestinal (GI) tract for detection and localization of abnormalities like lesions and diseases requires both high precision and recall. Additionally, it is important to support efficient, real-time processing for live feedback during (i) standard colonoscopies and (ii) scalability for massive population-based screening, which we conjecture can be done using a wireless video capsule endoscope (camera-pill). Existing related work in this field does neither provide the necessary combination of accuracy and performance for detecting multiple classes of abnormalities simultaneously nor for particular disease localization tasks. In this paper, a complete end-to-end multimedia system is presented where the aim is to tackle automatic analysis of GI tract videos. The system includes an entire pipeline ranging from data collection, processing and analysis, to visualization. The system combines deep learning neural networks, information retrieval, and analysis of global and local image features in order to implement multi-class classification, detection and localization. Furthermore, it is built in a modular way, so that it can be easily extended to deal with other types of abnormalities. Simultaneously, the system is developed for efficient processing in order to provide real-time feedback to the doctors and for scalability reasons when potentially applied for massive population-based algorithmic screenings in the future. Initial experiments show that our system has multi-class detection accuracy and polyp localization precision at least as good as state-of-the-art systems, and provides additional novelty in terms of real-time performance, low resource consumption and ability to extend with support for new classes of diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Rapid development of technologies in areas of sensors, imaging devices and diagnostic methods shifts the paradigm in medical diagnostic from manual analysis by trained doctors to wide usage of automated computer-assisted diagnostic systems. In our research, we are working at the intersection between computer science and pathological medicine, where we target a scalable, real-time, multi-disease detection system for the gastrointestinal (GI) tract. Our aim is to develop both a computer-aided, live analysis system of endoscopy videos and a scalable detection system for population-wide screening using a wireless video capsule endoscope (VCE). This small capsule with one or more image sensors is swallowed and captures videos while it traverses the entire GI tract.
In the context of object detection, localization and tracking in images and videos, a lot of research carried out. Particularly, current systems have been developed to detect general objects from the surrounding world, for example human faces, cars and logos. Our research targets a totally different domain, which is inside the body of a human being. Both the general objects and the GI tract irregularities can have different sizes, shapes, textures, colors and orientations, they can be located anywhere in the frame and also partially be hidden and covered by other objects and obstacle. However, GI tract findings can also have a color, texture and shape properties similar for the different diseases, as well as different for the similar diseases on the various developing stages. The GI findings can be covered by the biological substances, like for example seeds or stool, and lighted by direct and reflected light. Moreover, the images coming from the endoscopic equipment itself can be interleaved, noisy, blurry and over- or under-exposed, and it can contain borders, sub-images and a lot of specular reflections (flares) caused by endoscope’s light source. Therefore, detecting abnormalities and diseases in the GI tract is very different from detecting the objects from the surrounding world listed above. The GI tract can potentially be affected by a wide range of diseases with visible lesions (see Fig. 1d–e), but endoscopic findings may also include benign (normal) or man-made lesions. The most common diseases are gastric and colorectal cancer (CRC), which are both lethal when detected in a late stage. The 5-year survival rate of CRC ranges from 93% in stage I to 8% in stage IV [29]. Consequently, early detection is crucial. There are several ways of detecting pathology in the GI tract, but systematic population-wide screening is important. However, current methods have limitations regarding sensitivity, specificity, access to qualified medical staff and overall cost.
In this scenario, both high precision and recall are important, but so is the frequently ignored system performance in order to provide feedback in real-time. The most recent and most complete related work is the Polyp-Alert polyp detection system [52], which can provide near real-time feedback during colonoscopies. However, it is limited to polyp detection, it uses edges, colors and texture in the images, and, at the moment, it is not fast enough for live examinations.
To further aid and scale such examinations, we have earlier presented EIRFootnote 1 [32, 37], an efficient and scalable automatic analysis and feedback system for medical videos and images. The system is designed to support endoscopists in the detection and interpretation of diseases in the GI tract. EIR has initially been tested in video analysis of the lower portions (large bowel) of the GI tract. However, our main objective is to automatically detect abnormalities in the whole GI tract. Therefore, we are developing a complete system for detection and in-frame position localization of different endoscopic findings like polyps, tumors, diseases and landmark objects (like the Z-line and cecum). The aim is to use next-generation-EIR for both (i) a computer assisted diagnosis tool for live endoscopic examinations and (ii) a future fully automated and scalable screening system used together with VCEs. These goals impose strict requirements on the accuracy of the detection to maximize number of true positives and to avoid false negatives (overlooking a disease), as well as low computational resource consumption to provide massive population screening with VCEs. The live-assisted system also introduces a real-time processing requirement defined as being able to process at least 30 HD frames per second, i.e., a common frame rate and resolution in modern endoscopic devices.
Our first version [32, 37] was developed for detection of polyps, i.e., possible cancer precursors, and it was built on content-based information retrieval methodology using global image features for image content analysis. In this paper, the next generation of our system is presented, where we extend our system using out-of-the-box and improved deep learning neural network approaches and multi-class global-feature classification methods for detection and localization of endoscopic findings. We evaluate our prototype by training new and improved classifiers that are based on various image-recognition approaches. We compare the performance of feature-based analysis and neural network-based analysis in terms of accuracy and real-time processing, and thereby evaluate the different approaches for feasibility of multi-class detection and colonic polyp localization in real use-case scenarios.
The results from our experimental evaluation show that, (i) the detection and localization accuracy can reach the same performance or outperform other current state-of-the-art methods, (ii) the processing performance enables frame rates for real-time analysis at high definition resolutions, (iii) the localization-system performance can be improved further using a combination of our basic localization algorithms and neural network approaches, (iv) in our experiments, the global-feature multi-class detection approach slightly outperforms the deep learning neural network approach both in training speed and detection performance, and (v) the system proves to be easily extended by adding new types of abnormalities. Thereby, a system based on global features seems to be preferable and gives better performance in multi-class object detection than given existing deep learning network approaches. For the localization, additional research is needed to achieve better performance using a combination of local feature detection and deep learning neural networks.
The rest of the paper is organized as follows: First, in Section 2, we briefly introduce our medical case study. Next, we present related work in the field and compare it to the presented system in Section 3. This is followed by a presentation of the complete system in Section 4. We present an evaluation of the system in Section 5, and in Section 6, we discuss two cases where our system will be used in two medical examinations by medical experts. Finally, we conclude our results in Section 7.
2 Gastrointestinal endoscopy
The GI tract can potentially be affected by various abnormalities and diseases. Some examples of possible findings are shown in Fig. 1b–e. CRC is a major health issue world-wide, and early detection of CRC or polyps as predecessors of CRC is crucial for survival. Several studies demonstrate that a population-wide screening program improves the prognosis and can even reduce the incidences of CRC [17]. As a consequence, in the current European Union guidelines, screening for colorectal cancer is recommended for all people over 50 years old [50]. Colonoscopy, a common medical examination and the gold standard for visualizing the mucosa and the lumen of the entire colon, may be used either as a primary screening tool or in a second step after other positive screening tests [25]. However, traditional rectal endoscopic procedures are invasive and may lead to great discomfort for patients, and extensive training of physicians and nurses is required to perform the examination. They are performed in real-time, and, therefore, it is challenging to scale the number of examinations to a large population. Additionally, the classical endoscopic procedures are expensive. In the US, for example, colonoscopy is the most expensive cancer screening process, with an annual cost of 10 billion dollars (1,100$-6,000$/person) [47], and a time consumption of about one medical doctor-hour and two nurse-hours per examination.
In our research, we aim for an algorithmic system that detects multiple mucosal pathologies in videos of the GI tract. The idea is to assist endoscopists (physicians, who are highly trained in the procedure) during live examinations. Additionally, alternatives to traditional endoscopic examinations have recently emerged with the development of non-invasive VCEs. The GI tract is visualized using a pill-sized camera (available from vendors such as Medtronics/Given and Olympus) that is swallowed and then records a video of the entire GI tract. The challenge in this context is that medical experts still need to view the full-length video. Our system should provide a scalable tool that can be used in a first-order population screening system where the VCE-recorded video is used to determine whether an additional traditional endoscopic examination is needed or not. As a first step, we target the detection and the localization of colorectal polyps, which are known precursors of CRC (see for example Fig. 1d). The reason for starting with this scenario is that most colon cancers arise from benign, adenomatous polyps (around 20%) containing dysplastic cells, which may progress to cancer. Detection and removal of polyps prevent the development of cancer, and the risk of getting CRC in the following 60 months after a colonoscopy depends largely on the endoscopist’s ability to detect polyps [20]. Next, we extend our system to support detection of multiple abnormalities and diseases of the GI tract (see Fig. 1) by training the classifiers using multi-class datasets.
3 Related work
Detection of diseases in the GI tract has so far primarily focused on polyps. This is most probably due to the lack of alternative data in the medical field, but also that polyps are precursors of CRC. Several algorithms, methods and partial systems have, at first glance, achieved promising results [37] in their respective testing environment. However, none of the related works is able to perform real-time detection or support doctors by computer-aided diagnosis in real-time during colonoscopies. Furthermore, all of them are limited to a very specific use case, which in most cases is polyp detection for a specific type of camera [37]. Furthermore, in some cases, it is unclear how well the approach would perform as a real system used in hospitals. Most of the research conducted in this field uses rather small amounts of training and testing data, making it difficult to generalize the methods beyond the specific cleansed and prepared datasets and test scenarios. Therefore, overfitting for the specific datasets can be a problem and can lead to unreliable results.
The approach from Wang et al. [52] is the most recent and probably best-working system in the field of polyp detection. This system, called Polyp-Alert [52], is able to give near real-time feedback during colonoscopies. It uses an advanced edge-finding procedure to locate visual features and a rule-based classifier to detect an edge along the contour of a polyp. The system can recognize the same polyp across a sequence of video frames and can process up to 10 frames per second. The researchers report a performance of 97.7% correctly detected polyps with around 4.3% of frames incorrectly marked as containing polyps. Their results are based on a dataset that consists of 53 videos taken from different colonoscopes. Despite the promising polyp detection rate, the relatively high false positive rate makes the overall system detection performance not good enough for medical use cases. Unfortunately, the dataset used in this research is not publicly available, and therefore, a direct detection-performance comparison with our system is not possible. Moreover, most of the existing publications about polyp detection systems (see Tables 6 and 7 in Section 5) report detection accuracy on a per-polyp basis, counting the fact of successfully detected or missed polyp across the number of frames or even across the full video, which makes it difficult to perform a fair comparison. In our evaluation, we use a per-frame polyp detection and localization performance measurement. This gives a more realistic and better estimation of the performance of the developed method in the medical domain.
Other promising polyp detection approaches utilize quite old, but recently reborn neural networks and their advanced implementation called deep learning neural networks. Neural networks are conceptually easy to understand, and large amounts of research has been done in this direction in the last years. Results recently reported on, for example, the ImageNet dataset, look promising [13] in the areas of indexing, retrieving, organizing and annotating multimedia data. Despite the fact that the neural network model training process is very complicated and time-consuming [12], their ability to detect and localize various objects can potentially help us to improve our system. However, such an improvement is possible only after careful investigation, to ensure that our system will still run in real-time and be able to deal with the required amount of lesion categories. This is important since we deal with patient health, and the outcome can make the difference between life and death.
Most modern deep learning frameworks state that they can be used out-of-the-box for different types of input data. This statement sounds promising, but most state-of-the-art neural networks in multimedia research are designed to process images from everyday life, like cats, dogs, bicycles, cars, pedestrians, etc. It needs to be proven that they can be used in medical domains, because it is difficult to evaluate their performance and robustness properly [28] due to the lack of relevant training and test data. In fact, obtaining such datasets is one of the biggest challenges related to deep learning approaches in connection with the medical field, due to a lack of medical experts needed to annotate data, and legal and ethical issues. Some common conditions, like colon polyps, may already have the number of collected images and videos required to perform training of a neural network, while other endoscopic findings, like tattoos from previous endoscopic procedures (black-colored parts of the mucosa), are not that well documented, but still interesting to detect [40]. Recent research [8] on the topic of transfer learning promises a solution for the problem of insufficient amounts of available training data. Transferring the knowledge learned by the deep network on a large dataset, e.g. ImageNet, to train a specialized network on a small medically oriented dataset, together with a saliency prediction used to emphasize key image points, can result in better performance of the endoscopic finding detection and localization. Thus, in this research, we perform some preliminary experiments to see how neural networks can deal with small training datasets.
In summary, related work primarily targets specialized problems or elements of the more general, holistic medical problem we are attempting to solve. Existing systems are either (i) too narrow for a flexible, multi-disease detection system; (ii) have been tested on limited datasets too small to show whether the method would work in a real scenario, or; (iii) provide a processing performance too low for a real-time system or ignore the system performance entirely. Last, but not least, we are targeting a holistic end-to-end system where a VCE that traverses the entire tract with its video signals is algorithmically analyzed. To solve the fundamental systems problems, we are targeting and developing a close to fully automated, accurate, low false positive, scalable, privacy-preserving and low-cost screening system that will, if we may say so, have significant potential impact on the society.
4 The EIR system
Our objective is to develop a system that supports doctors in multi-disease detection in the GI tract. The system must (i) be easy to use and less invasive for the patients than existing methods, (ii) support multiple classes of detected GI objects, (iii) be easy to extend to new different diseases and findings, (iv) handle multimedia content in-real time (30 frames per second or more for Full HD videos), (v) be usable for real-time computer-aided diagnosis, (vi) achieve high classification performance with minimal false-negative classification results and (vii) have a low computational resource consumption. These properties potentially provide a scalable system with regard to reduced number of specialists required for a larger population, and dramatically increased number of users potentially willing to be screened. Therefore, EIR consists of three parts: the annotation subsystem [2], the detection and automatic analysis subsystem and the visualization and computer-aided diagnosis subsystem [35].
The subsystems for algorithmic analysis are designed in a modular way, so that they can be extended to different diseases or subcategories of diseases, as well as other tasks like size determination, etc. Currently, we have implemented two types of analysis subsystems: the detection subsystem that detects different irregularities in video frames and images, and the localization subsystem that localizes the exact position of the disease (only polyp localization is supported at the moment) in the frame. The detection subsystem is not designed to determine the location of the detected irregularity. The exact lesion position finding is done by the localization subsystem, so that we can use the same localization subsystem for different detection subsystems. The localization subsystem uses the output of the detection system as input and processes only frames marked as containing a localizable disease.
4.1 Detection subsystem
The detection subsystem performs lesion recognition and classification. It is intended for abnormality-presence detection without searching for the precise position of the lesion. The detection is performed using various visual similarity finding techniques. For each lesion that has to be detected, we use a set of reference frames that contains examples of this lesion occurring in different parts of the GI tract. This set can be seen as the model of the specific disease. We also use sets of frames containing examples of all kinds of healthy tissue, normal findings like stool, food, liquids, etc. The final goals of the detection subsystem is to decide if this particular frame analyzed contain any lesion or not, and to detect the exact type of the lesion. The detection system is designed in a modular way and can easily be extended with new diseases. This would, for example, allow not only to detect a polyp, but to distinguish between a polyp with low or high risk for developing CRC by using the NICE classification.Footnote 2
4.1.1 Basic EIR system
In our previous work, we presented our basic EIR system [32, 36, 37] that implements a single-class global-feature-based detector able to recognize the abnormalities in a given video frame. Global image features were chosen, because they are easy and fast to calculate, and the exact lesion’s position is not needed for detection, i.e., identifying frames that contain a disease. We showed that the global features we chose, Tamura feature [45] and Joint Composite Descriptor (JCD) [53], which is a combination of Fuzzy Color and Texture Histogram (FCTH) [10] and Color and Edge Directivity Descriptor (CEDD) [9], can indeed outperform or at least reach the same results as local features.
The basic algorithm is based on an improved version of a search-based method for image classification. The overall structure and the data flow in the basic EIR system is depicted in Fig. 2. First, we create the index containing the visual features extracted from the training images and videos, which can be seen as a model of the diseases and normal tissue. The index also contains information about the presence and type of the disease in the particular frame. The resulting size of the index is determined by the feature vector sizes and the number of required training samples, which is rather low compared to other methods. Thus, the size of the index is relatively small compared to the size of the training data, and it can be easily fit into main memory on a modern computer. Next, during the classification stage, a classifier performs a search of the index for the frames that are visually most similar to a given input frame (see Section 4.1.3 for a detailed description of the method). The whole basic detector is implemented as two separate tools, an indexer and a classifier. We have released the indexer and the classifier as an open-source project called OpenSea Footnote 3
[37].
The indexer is implemented as a batch-processing tool. Creating the models for the classifier does not influence the real-time capability of the system and can be done off-line, because it is only done once when the training data is first inserted into the system. Visual features to calculate and store in the indexes are chosen based on the type of the disease because different sets of features or combinations of features are suitable for different types of diseases. For example, bleeding is easier to detect using color features, whereas polyps require shape and texture information.
The classifier can be used to classify video frames from an input video into as many classes as the detection subsystem model consists of. The classifier uses indexes generated by the indexer. In contrast to other classifiers that are commonly used, this classifier is not trained in a separate learning step. Instead, the classifier searches previously generated indexes, which can be seen as the model, for similar visual features. The output is weighted based on the ranked list of the search results. Based on this, a decision is made. The classifier is parallelized and can utilize multiple CPU cores for the extraction of features and the searching in indexes. To increase performance even more, we implemented the most compute intensive parts of the system with GPU computation support.
4.1.2 Deep-EIR
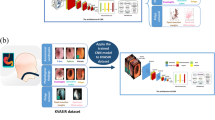
The neural network version of EIR called Deep-EIR is based on a pre-trained convolutional neural network architecture and transfer learning [8]. We trained a model based on the Inception v3 architecture [43] using the ImageNet dataset [13] and then re-trained and fine-tuned the last layers. We did not perform complex data augmentation at this point and only relied on transfer learning. We are currently in the process of data collection, and for future work, we will also look into data augmentation and training a network from scratch using the newly collected data, which might lead to better results than transfer learning. Figure 3 gives a detailed overview of the complete pipeline for the neural network-based implementation of the detection.
Inception v3 achieves good results regarding single-frame classification and has reasonable computational resource consumption. The top one result error is 21.2%, and the top five error is 5.6% with less than 25 million parameters. The training of the Inception v3 network is performed from scratch using Google Tensorflow v1.2rc [1]. The training takes several weeks on a single modern computer with GPU support. Tensorflow is an open source framework that allows all kinds of numerical computations using graphs. Nodes within the flow graphs represent mathematical operations, and the edges represent data arrays (called tensors in Tensorflow). It is especially built to support scalable machine learning, which includes neural network-based architectures [1].
The trained Inception v3 model is then used in a retraining step. For this step, we follow the approach presented in [14]. Basically, we froze all the basic convolutional layers of the network and only retrained the two top fully connected (FC) layers. The FC layers were retrained using the RMSprop [48] optimizer that allows an adaptive learning rate during the training process. After 1,000 epochs, we stopped the retraining of the FC layers and started fine-tuning the convolutional layers. For that step, we did the analysis of the Inception v3 model layer structure and decided to apply fine-tuning on the top two convolutional layers. This step finalizes the transfer-learning scenario and performs an additional tuning of all the NNs layers according to our dataset. For this training step, we used a stochastic gradient descent method with a low learning rate of 10−4 to achieve the best effect in terms of speed and accuracy [27]. This comes with the advantage that little training data is needed to train the network, which is an advantage for our medical use case. Additionally, it is fast, requiring just about one day to retrain the model. The re-trainer is based on an open source implementation of Tensorflow.Footnote 4 To increase the number of training samples, we also performed distortion operations on the images. Specifically, we performed random cropping, random rescaling and random change of brightness. The grade of distortion was set to 25% per image. After the model has been retrained, we use it for a multi-class classifier that provides the top five classes based on probability for each class.
4.1.3 Multi-class global-feature-based EIR
The new multi-class global-feature-based version of EIR is based on the initial version of EIR with some extensions. The basic search-based classification part of EIR is used to create a classifier for each disease that we want to classify. Figure 2 gives a detailed overview of the classifier’s pipeline for the global-feature-based implementation of the detection. The difference to the basic EIR version is that the ranked lists of each search-based classifier are then used in an additional classification step to determine the final class.
For features extraction in the detection step and for the training procedure, the indexing is performed using the basic EIR indexer implementation [32, 37]. The same set of two global features, namely Tamura and JCD, is used. These features were selected by a simple features efficiency estimation by testing different combinations of features on smaller reference datasets to find the best combinations in terms of processing speed and classification accuracy. The selected features can be combined in two different ways. The first is called feature values fusion or early fusion, and it basically combines the feature value vectors of the different features into a single representation before they are used in a decision-making step. The second one is called decision fusion or late fusion where the features are combined after a decision-making step. Our multi-class global-feature-based approach implements feature combination using the late fusion.
During the detection step, a term-based query from the hashed feature values of the query image is created for each image, and a comparison with all images in the index is performed, resulting in a ranked list of similar images. The ranked list is sorted by a distance or dissimilarity function associated with the low-level features. This is done by computing the distance between the query image and all images in the index. The distance function for our ranking is the Tanimoto distance [46]. A smaller distance between an image in the index and the query image means a better rank [46]. The final ranked list is used in the classification step, which implements a simple k-nearest neighbors algorithm [4]. This algorithm can be used for supervised and unsupervised learning, two or multi-class classification and different types of input data ranging from features extracted from images to videos to meta-data. Its main advantages are its simplicity, that it achieves state-of-the-art classification results and that it is very fast in terms of processing time.
For the final classification, we use the random forest classifier [6], an ensemble learning method for classification that operates by constructing a multitude of decision trees at training time and outputting the class that is the mode of the classes of the individual trees. A decision tree can be seen as a classifier, which basically performs decision-based classification on the given data. To get the final class, the classifier combines decision trees into a final decision implementing a late fusion for the multi-class classification. The advantage of the random forest algorithm is that the training of the classifier is very fast because the classification steps can be parallelized since each tree is processed separately. Additionally, it is shown that the random forest is very efficient for large datasets due to the ability to find distinctive classes in the dataset and also to detect the correlation between these classes. The disadvantage is that the training time increases linearly with the number of trees, which means a longer training time when many trees are used at the same time. However, this is not a problem for our use-case since the training is done offline, where time is less critical. Our implementation of the random forest classifier uses the version provided by the Weka machine learning libraryFootnote 5 [16], which is a collection of algorithms for machine learning and data mining. We chose the random forest approach, because it is fast and achieves good results [49]. It is important to point out that for this step, another classification algorithm can also be used.
4.2 Localization subsystem
The localization subsystem is intended for finding the exact positioning of a lesion, which is used to show markers on the frame containing the disease. This information is then used by the visualization subsystem. All images that we process during the localization step come from the positive frames list generated by the detection subsystem. Processing of the images is implemented as a sequence of intra-frame pre- and main-filters. Pre-filtering is needed because we use local image features to find the exact position of objects in the frames. Lesion objects or areas can have different shapes, textures, colors and orientations. They can be located anywhere in the frame and also partially be hidden and covered by biological substances, like seeds or stool, and lighted by direct light. Moreover, the image itself can be interlaced, noisy, blurry and over- or under-exposed, and it can contain borders and sub-images. Apart from that, images can have various resolutions depending on the type of endoscopy equipment used. Endoscopic images usually have a lot of flares and flashes caused by a light source located close to the camera. All these nuances affect the local feature-based detection methods negatively and have to be specially treated to reduce localization precision impact. In our case, several sequentially applied filters are used to prepare raw input images for the following analysis. These filters are border and sub-image removal, flare masking and low-pass filtering. After pre-filtering, the images are ready to be used for further analysis.
4.2.1 Basic EIR system
Previously, we have implemented the localization of colon polyps using our hand-crafted approach based on local image features [35]. The main idea of the localization algorithm is to use the polyp’s physical shape to find the exact position in the frame. In most cases, the polyps have the shape of a hill located on a relatively flat underlying surface or the shape of a more or less round rock connected to an underlying surface with stalks of varying thickness. These polyps can be approximated with an elliptically shaped region consisting of local features that differ from the surrounding tissue with high probability. To detect these types of objects, we process the frames marked by the detection subsystem as containing polyps by a sequence of various image processing procedures, resulting in a set of possible abnormality coordinates within each frame. Figure 4 gives a detailed overview of a localization pipeline for the basic EIR algorithm implementation. The pipeline consists of the following steps: non-local means de-noising [7]; 2D Gaussian blur and 2D image gradient vector extraction; border extraction by gradient vector threshold binarization; border line isolated binary noise removal; estimation of ellipses locations; ellipse size estimation by analyzing border pixel distribution; ellipse fitting to extracted border pixels; selection of a predefined number of non-overlapping local peaks and outputting their coordinates as possible polyp locations. For the possible locations of ellipses, we use the coordinates of local maxima in the insensitivity image, created by additive drawing of straight lines starting at each border pixel in the direction of its gradient vector. Ellipse fitting is then performed using an ellipse fitting function [15]. This version of the subsystem is implemented in C++, and it uses the OpenCVFootnote 6 open source library for routine image content manipulation and the CUDAFootnote 7 toolkit for GPU computation support.
4.2.2 Deep-EIR
The existing localization scheme can be extended to support different diseases by implementation of lesion-specific shape, color and texture detection, but such an extension requires experimental studies for each new type of abnormality. In order to reduce the system improvement costs, we performed an evaluation of two universal object localization frameworks, based on deep learning neural network approaches. First is TensorBoxFootnote 8 [41], which extends Google’s reference implementation of the machine-learning framework called Tensorflow [1]. Second approach is based on the Darknet [33] open-source deep learning neural network implementation called YOLOFootnote 9 [34]. Both of these frameworks are designed to provide not only object detection, but also object localization inside frames. They implement GPU-accelerated deep learning algorithms that can work with near to real-time performance and provide the capability of locating various objects out-of-the-box.
The TensorBox approach introduces an end-to-end algorithm for detecting objects in images. As input, it accepts images and directly generates a set of object bounding boxes as output. The main advantage of the algorithm is the capability of avoiding multiple detections of the same object by using a recurrent neural network (RNN) with long short-term memory (LSTM) units together with fine-tuned image features from the implementation of a convolutional neural network (CNN) for visual objects classification and detection called GoogLeNet [42].
The Darknet-YOLO approach introduces a custom CNN, designed to simultaneously predict multiple bounding boxes and class probabilities for these boxes within each input frame. The main advantage of the algorithm is that the CNN sees the entire image during the training process, so it implicitly encodes contextual information about classes as well as their appearance, resulting in a better generalization of objects’ representation. The custom CNN in this approach is also inspired by the GoogLeNet [42] model.
As initial models for both approaches, we used database models pre-trained on ImageNet [19] . Our custom training and testing data for the algorithms consists of frames and corresponding text files describing ground truth data with defined rectangular areas around objects: a JSON file for TensorBox and one text file per frame for Darknet-YOLO. Ground truth data was generated using a binary-masked frame set (example shown in Fig. 5) by the localization validation software used in our experimental studies. Both frameworks were trained using the same training dataset, where all frames contained one or more visible polyps. No special filtering or data preprocessing was used, thus the training dataset contained high quality and clearly visible polyp areas as well as blurry, noisy, over-exposed frames and partially visible polyps. The models were trained from scratch using corresponding default-model training settings [34, 41]. After the training, the test dataset was processed by both neural networks in testing mode. As a result, the frameworks output JSON (TensorBox) and plain-text (Darknet-YOLO) files containing sets of rectangles, one set per frame, marking possible polyp locations with corresponding location confidence values. These results have been processed using our localization algorithms.
4.3 Visualization and computer aided diagnosis subsystem
The visualization subsystem is developed as a flexible multi-purpose tool. First, it should help in evaluating the performance of the system and get insights into why things work well or not. Second, it can be used as a computer-aided diagnostic system for medical experts. Third, it should help us in the creation of new datasets, allow us to extend the number of detected diseases and help doctors to create annotations in a time-saving manner. Previously, we have developed the TagAndTrack subsystem [2] that can be used for visualization and computer-aided diagnosis. We developed a web-based visualization toolkit that can be used to support medical experts while being very easy to use and distribute. This tool takes the output of the detection and localization subsystems and creates a web-based representation of the detection and localization results. The web-based visualization is then combined with a video sharing and annotating platform where doctors are able to watch, archive, annotate and share information. To break through low availability of high quality training and testing datasets for different GI track diseases, we developed a new ClusterTag application for the visualization subsystem. The main purpose of ClusterTag is to provide an easy-to-use and convenient user interface to huge image and video frame collections captured during endoscopic procedures, including conventional colonoscopies and VCEs.
Figure 6 illustrates our ClusterTag application while processing a dataset containing 36,476 images with the exact lesion areas marked. The application implements image and ground truth loading and analyzing, image tagging, creation and editing of ground truth data, global feature extraction and semi-automatic dataset clustering using our previously developed algorithms [38]. With the main focus on the interactive visual representation of huge image collections, the visualization module helps users create and interact with the new or already defined clusters. We use the Weka library to help the user in building clusters. For the image attribute extraction required for machine-learning-based classification we use global image features, which are extracted using the image retrieval framework called LIRE.Footnote 10 In our approach, we use global features describing the image in terms of different visual attributes, such as sharpness, color distribution and histogram of brightness. A detailed description of the used global features, the corresponding clustering algorithm and the clustering performance metrics can be found in [38]. Both the WEKA and LIRE libraries can be easily replaced by other machine learning or feature extraction libraries if desired.
Applying unsupervised clustering on huge unsorted and unannotated datasets significantly reduces the amount of work required from skilled doctors during image labeling and grouping. Together with unsupervised clustering, our application provides the users with the ability of tagging and analyzing multiple single images at once and putting them into appropriate groups together. The ClusterTag application is released as open-source softwareFootnote 11 and might help other research groups in the creation and analysis of new datasets.
5 Evaluation
For our experimental evaluation, we use two different use-cases. First, we evaluated the performance of our multi-class classification and detection algorithms in automated colonoscopy video processing. Here, we tested our system using six different classes of endoscopic findings that can be found in the colon (shown in Fig. 1). The classes to be detected are (a) frames with normal colon mucosa (healthy colon wall), (b) frames of the cecum area which is an intraperitoneal pouch that is considered to be the beginning of the colon (an anatomic landmark helping doctors and VCE video analysis algorithms to orientate in the colon), (c) frames displaying the Z-line which is the gastroesophageal junction that joins the esophagus to the stomach (an anatomic landmark), (d) frames containing one or more polyps, (e) frames with visible tumor areas, and (f) useless blurry frames without any visible and recognizable objects. Thus, the developed multi-class classification and detection system should split all the video frames into six classes that can be observed in the human GI tract. The developed method allows us to implement a new generation of endoscopy video processing systems able to efficiently detect various lesions of the GI tract.
Second, we evaluated the performance of the state-of-the-art object localization approaches based on deep learning algorithms, and then we compared it with our basic polyp localization algorithm. In this use-case, we compared the ability of different methods to find the location of polyps inside a frame. The main goal of this evaluation is to decide if we can improve the polyp localization performance of our system using a combination of different algorithms.
During the evaluation, wherever it was possible, we compared the performance of our method with the best state-of-the-art competitors. Nevertheless, a direct comparison is hard as different datasets and detection measures are used in state-of-the-art system evaluations. Thus, we compared the metrics we found in the relevant publications.
For all of the subsequent measurements, we used the same computer. It has an Intel Core i7-6700K CPU running at 4.00GHz, 16 GB of RAM, a GeForce GTX TITAN X GPU, and it runs a 64-bit Ubuntu Linux v16.04.
5.1 Multi-class classification
In the multi-class classification experiments, we used cross-validation because of the relatively small number of images in the annotated dataset. For the performance measurement, we used the standard tool from WEKA for evaluating multi-class classifiers. This tool uses the ground truth to compute a confusion matrix and the common standard metrics: recall (sensitivity), precision, specificity, accuracy and F1 score. We created a new dataset from colonoscopy images that we got from Vestre Viken Hospital, Norway. From the whole unannotated dataset, we manually selected 50 different frames of 6 different classes (described in Section 2): blurry frames, cecum, normal colon mucosa, polyps, tumor, and Z-line. The selected frames were used to create 10 separate datasets, each containing training and test subsets with equal numbers of images. Training and test subsets were created by equally splitting random-ordered frame sets for each of the 6 classes. The total number of frames used in this evaluation is 300: 150 in the training subsets and 150 in the test subsets. Each training and test subset contains 25 images per class. Multi-class classification is then performed on all 10 splits and then combined and averaged. Following this strategy, an accurate enough estimation about the performance can be made even with a smaller number of images.
5.1.1 Deep-EIR
First, we performed an evaluation of Deep-EIR that implements the deep learning neural network multi-class detection approach. Table 1 shows the resulting confusion matrix. The detailed performance metrics presented in Table 2 and the results can be considered as good, they confirm that Deep-EIR performs well. All blurry and Z-line frames were classified correctly. Cecum and normal colon mucosa were often cross-mis-classified, which is a normal behavior, because from a medical point of view, normal colon mucosa is part of the cecum, and under real-world circumstances, this would not be a relevant mistake. Interesting polyps and tumors were detected correctly in most cases, as well as the Z-line landmark, which is important for our medical use case.
5.1.2 Multi-class global-feature-based EIR
Second, we performed an evaluation of the multi-class global-feature-based EIR, which implements a global-feature multi-class detection approach. The multi-class global-feature-based EIR classifier allows us to use a number of different global image features for the classification. The more image features we use, the more precise the classification becomes. We generated indexes containing all possible image features for all frames of all different classes of findings from our training and test dataset. These indexes were used for multi-class classification, different performance measurements and also for leave-one-out cross-validation. Using our detection system, the built-in metrics functionality can provide information on the different performance metrics for benchmarking. Further, it provides us with the late fusion of all the selected image features and performs the selection of the exact class for each frame in test dataset. All used features are described in detail in [24].
Table 3 shows the resulting confusion matrix, which shows, like the Deep-EIR results, that the global feature-based detection approach performs well, too. Again, all blurry and Z-line frames were classified correctly. Cecum and normal colon mucosa were sometimes cross-misclassified. Polyps and tumors were detected correctly in most cases. The detailed performance metrics are presented in Table 4 and can also be considered as good.
5.1.3 Deep-EIR vs multi-class global-feature-based EIR
The comparison of these two approaches shows that both approaches have equal excellent overall F1 score of 100% in Z-line detection. The global-feature approach with the 100% F1 score outperforms the neural network approach by a small margin in blurry frame detection. The neural network F1 score detection for tumors is 98.2%, which is 1% better than the global-feature approach. Detection of other classes is better for the global-feature approach, giving the F1 scores of 79.2% and 74.3% for cecum and normal mucosa. Most importantly for our case study, polyp detection performed much better using the global-feature approach, giving the 92.8% F1 score (13.1% better than the neural network approach).
The performance evaluation of the cross-validation for both multi-class classification approaches (see Table 5) confirms the high stability of the models used for the classification.
The processing performance of both Deep-EIR and global-feature-based EIR in terms of processing speed meets real-time demands with a good margin for the real-time medical use case. Both can process Full HD images at a frame rate of 30 frames per second.
Our experimental comparison of the Deep-EIR and the global-feature-based EIR of the detection system shows clearly that the global-feature approach outperforms the deep learning neural network approach and gives better accuracy for almost all target detection classes (except several cases of misclassification of tumors) in conjunction with high 92.8% and 97.2% F1 scores for the most important findings: polyps and tumors. Moreover, when a sufficiently large training dataset covering all possible detectable lesions of the GI tract is used, the proposed global-feature approach for multi-class detection requires relatively little time for training [35] compared to days and weeks for the deep learning neural network approach.
A comparison of Deep-EIR and global-feature-based EIR with existing competitive approaches is shown in Table 6. The basic-, Deep- and multi-class global feature-based EIR detector versions are depicted in the last table’s rows. As one can see, the global feature-based EIR approach gives the best performance in terms of precision (90.6%), specificity (98.1%) and accuracy (96.9%), and comparable recall/sensitivity (90.6%). In other words, the results indicate that we can detect different classes of GI tract findings with a precision of almost 91%. If we compare this to the best performing system in Table 6, we see that Polyp-Alert reaches slightly higher detection accuracy on a different dataset. However, our system is faster and can detect colonoscopic findings in real-time, and furthermore, it is not designed and restricted to detect only polyps, it can detect multiple classes of diseases, and EIR can further be expanded to any additional diseases if we have the correct training data.
The performance comparison of different multi-class detection and classification approaches in terms of frame processing speed is depicted in Fig. 7. Deep-EIR, multi-class global feature-based EIR and basic EIR perform better in terms of speed than competitors. The single-class basic EIR detector can process up to 300 Full HD frames per second (for a GPU-accelerated implementation) [35]. Deep- and global feature-based EIR classifiers showed 30 frames per second, which fits our medical use case. For further processing speed improvements, we plan to implement additional GPU acceleration for a random-trees classifier and feature index search, as we have for our initial polyp detection version of EIR [32].
5.2 Polyp localization
The multi-class dataset from Vestre Viken Hospital does not contain the ground truth for the localization of the findings. Therefore, in this experiment, we used the available ASU-Mayo Clinic polyp database.Footnote 12 It consists of training and test sets of images and videos with corresponding ground truth showing the exact polyp location areas. This was the biggest publicly available dataset (until recently, when the owners decided to withdrawn it from the public), consisting of 20 videos from standard colonoscopies with a total of 18,781 frames and different resolutions up to full HD [44]. For this particular evaluation, we selected only frames containing polyps, which gave us 8,169 frames in total: 3,856 in the training subset and 4,313 in the test subset. The frames with polyps contain various polyp types, fully visible and particularly hidden, clearly visible and blurry, clean and covered by stool. Figure 8 depicts variations in polyp sizes (in terms of number of pixels showing polyp bodies within images) across the datasets. As one can see, there are huge variations in polyp sizes in terms of video-frame pixels from very small up to one third of the full video frame size. This reflects real colonoscopy video-capturing scenarios and introduces a big challenge for object localization algorithms.
For the localization-performance measurement, we used the common metrics: recall (sensitivity), precision, specificity, accuracy and F1 score. To count the corresponding localization events correctly, we took into account that polyps can have different shapes, they are often not located in compact pixel space areas (in contrast to, e.g., people faces). The shape of the polyps is marked in the ground truth data by binary masks. Before computing the localization subsystem performance, we need to figure out how to convert output of different localization algorithms into performance metrics. Our initial assumption (from practical experience) was to count each of the neural networks’ location rectangles as a true positive localization event if and only if it covers at least 10% of the corresponding ground truth area. Otherwise, we count it as a false positive. In our use case, multiple detection of the same polyp does not improve medical outcome. Therefore, we count multiple true positives on the same polyp ground truth area as one true positive. Polyp misses are counted if, after processing all resulting rectangles for a particular frame, we still have one or more ground truth areas without corresponding true positives. We count such misses as false negatives. Thus, there is a possibility of multiple false negatives per one frame, in case we have multiple lesions in the same frame. In this experiment, we process only frames that contain one or more polyps. This means that we do not have true negatives. Therefore, specificity of the algorithms can be assumed as 100%. To check our assumptions about minimal coverage areas, we performed an initial performance evaluation and built a graph showing unfiltered output from neural networks. In our EIR system, the base localization algorithm outputs points instead of rectangular areas. Thus, we count a true positive if a point is located inside of a polyp ground truth area, keeping other rules the same. An example of a polyp localization algorithm output is depicted in Fig. 9. The polyp-location ground truth marked by light green ellipses is computed based on the ground truth binary masks (see Fig. 5) using the closest elliptical region approximation. Due to the limitations of the current version of the localization algorithm, it produces four possible polyp locations per frame without any location ranking. In this evaluation, we consider all four points as equal and always use all of them for calculating the performance metrics. These points are marked by the green and red crosses. The green crosses correspond to the true positive events, and the red crosses show the false positive events.
The deep learning neural network frameworks tested in this experiment require training before they are able to perform polyp localization. Thus, both networks were trained using their default model training parameters. For TensorBox, the neural network model training took 6.5 days, and for Darknet-YOLO, we needed 5.1 days. After the training, we performed model validation using the corresponding frameworks’ routines, and the training dataset as input. The validation confirmed the correctness of the trained models for both TensorBox and Darknet-YOLO. The deep learning approaches are capable of correctly localizing polyps that were previously detected by the detection subsystem within the training dataset with 98% accuracy for the TensorBox model and 95% accuracy for the Darknet-YOLO model.
Next, we performed a main localization run of both frameworks on the test dataset and validation using the corresponding ground truth. Both TensorBox and Darknet-YOLO can be finely tuned by setting confidence threshold values, which limits the number of returned location rectangles to only highly confident ones. In order to investigate how the output of both can be affected by a confidence threshold value, it was set to zero during the first test run, which should give us the full unfiltered localization output. The reason for studying this dependency is that it is the only network tuning parameter in the unseen data process mode, which can help us to maximize their localization accuracy. Figure 10 shows a histogram of true polyps’ area coverage by location boxes found by TensorBox. We counted only location boxes that cover at least one pixel of a true polyp area. As one can see, the histogram has clearly visible maximum around 16% coverage rate, followed by an exponential decrease to almost constant level. A comparable analysis with the same type of histogram for the Darknet-YOLO output is depicted in Fig. 11. We observe a similar distribution for coverage rate (higher than 10%). A much higher number of location rectangles with zero coverage rate indicates that TensorBox implements additional localization result filtering. Thus, the effect of the confidence threshold level adjustment cannot be as significant as for Darknet-YOLO, which has the expected output with a high number of location boxes covering small parts of true polyp areas. Therefore, Darknet-YOLO should show a strong response to confidence threshold level. For the following validation and performance evaluation of both frameworks, we used 10% as the threshold value for the minimal required polyp ground truth coverage for true positive events, i.e., 10% must be covered for the event to be counted. Figures 12 and 13 confirm our assumption about output result filtering in TensorBox. Its output contains a relatively small number of found locations with high number of highly-confident locations compared to Darknet-YOLO, which has a large number of low-confident locations, exactly as expected with the choice of a zero-confidence threshold.
The histogram shows polyps area coverage by location boxes found by the TensorBox localization algorithm with the maximum around 16% coverage rate with following exponential decrease to the almost constant level. The low number of found location rectangles around zero coverage rate is an evidence of some output results pre-filtering
The histogram shows polyps area coverage by location boxes found by the Darknet-YOLO localization algorithm with near to exponential distribution for coverage rate higher than 10%. The higher number of found location rectangles around zero coverage rate gives clear indications that algorithm output unfiltered results
The performance results depending on the confidence threshold value are depicted in Fig. 14 for TensorBox and Fig. 15 for Darknet-YOLO. As one can see, TensorBox localization performance does not depend on the confidence threshold value in any significant way. The best performance in terms of minimizing the number of false negative events with an acceptable number of false positive events can be achieved by maximizing the algorithm’s accuracy metrics. For TensorBox, the maximum accuracy reaches a level of 31.6% for a confidence threshold value of zero with a corresponding polyp miss rate of 66.2%. For TensorBox, this is the best value, and it cannot be improved by adjusting the confidence threshold value. For Darknet-YOLO, maximum accuracy is reached at a 42.2% confidence threshold. The accuracy is 8% with a corresponding polyp miss rate of 47.9%. Darknet-YOLO showed more flexibility and a good response to the confidence threshold value. For Darknet-YOLO, the polyp miss rate can be significantly reduced by decreasing the confidence threshold value, but this gives a significant increase in the number of false positives, making the whole system too noisy. Nevertheless, combining Darknet-YOLO and the basic EIR localizer approaches can potentially give better overall system performance and better polyp miss rate.
Performing a comparison with well-known existing approaches in polyp localization is difficult due to lack of publicly available information (see Table 7) about other researchers’ algorithms’ performance and evaluation methods, and due to prevalent non-disclosure restrictions that prevent sharing of datasets in the research community. The available data shows, that our EIR basic localization approach has good performance with an F1 score of 41.6%.
The performance of the TensorBox approach (see Table 7) is too low for our real-time use-case. But, as depicted in Table 7, Darknet-YOLO performs well in terms of processing speed and can run at 45 frames per second. Our basic approach runs at 120 frames per second, thus a combination of both approaches can give us better localization performance while staying within the required real-time frame rate limits.
6 Real-world use cases
In this section, we describe two real-world use cases where the presented system can be used. The first one is a live system that will assist medical doctors during endoscopies. Currently, we are deploying a proof-of-applicabilty prototype in one of our partner hospitals. The second is a system that will automatically analyze videos captured by VCEs. Several hospitals are involved in this more concrete and applied research, and currently we are setting up the data-sharing agreements and collect the data for a new multi-disease dataset that will be released open-source. The first use case requires fast and reliable processing, and the second requires a system that is able to process a large amount of data in a reliable and scalable way.
6.1 Live system
The aim of the live system is to provide live feedback to the doctors, i.e., a computer-aided diagnosis in real-time. While the endoscopist performs the colonoscopy, the system analyzes the video frames that are captured by the colonoscope. To provide helpful information for the operating doctor, we combine the visual information from the endoscope with our marks. For the detection, we alter the frame borders and show the name of the detected finding in the auxiliary area of the endoscope device monitor. For the implemented localization classes, we put a cross on top of the localized findings (polyps in this system version). At the moment, we have implemented a demo version of the live system [39]. The live demo supports detection and localization of polyps. It is able to process a FullHD video stream with 30 FPS in real-time. An example of the graphical output of the live system is depicted in Fig. 16.
A screenshot of the live system showing the combination of the visual information from the endoscope with feedback information from the detection and localization system. The pink fame surrounding background shows a positive detection. The name of the detected finding is shown in the frame auxiliary screen area, and the cross shows the location of the polyp
In addition to supporting the medical expert during the colonoscopy, we are working on an extension of the system, where the system is used to document the examination procedure. We will implement the generation of a document with an overview of the colonoscopic procedure. The doctors will be able to make changes or corrections, and add additional information to that document. The document will be stored or used as an appendix to the written endoscopy report.
6.2 Wireless video capsule endoscope
The current existing VCEs have a resolution of around 256 × 256, frame rates of 3-35 frames per second (adaptive frame rate with a feedback loop from the receiver to the transmitter). They do not have optimum lighting, making it more difficult to detect endoscopic findings in the captured images than in images from traditional endoscopes. Also, during VCE procedures, the intestine is not expanded, unlike in traditional endoscopy, where the expansion allows for clear and non-obfuscated pictures of the intestine walls. Nevertheless, ongoing research aims at improving the VCEs’ hardware capabilities and at improving the methods and algorithms developed for colonoscopies to work also for VCEs [22]. The multi-sensor VCE is swallowed in order to visualize the GI tract for subsequent diagnosis and detection of GI diseases. Thus, people may in the future be able to buy VCEs at the pharmacy, and deliver the video stream from the GI tract to the phone over a wireless connection. In the best case, the first screening results are available within eight hours after swallowing the VCE, which is the time the camera typically spends traversing the GI tract. Thus, the ability to implement and perform mass-screening of the GI tract highly depends on two main research areas. First, it requires the development of a new generation of VCEs with better image quality and the ability to communicate with widely used mobile phones. Second, mass-screening requires a new generation of lesion detection algorithms able to process the captured GI tract multimedia data and video footage fully automatically in the mobile phone with public cloud computing support. Here, a preliminary analysis and task-oriented compression of a captured video footage before uploading into the cloud is important due to huge amounts of video data generates by VCEs. In our future research for this use case, we will work on the adaptation of the detection algorithms to the common mobile platforms. We will create a new mobile application to demonstrate the ability of our system to perform on hardware with the limited resources available.
7 Conclusion
In this paper, a complex automated diagnosis system built for different GI tract disease detection scenarios, colonic polyp localization and big dataset visualization has been presented. We briefly described the whole system from data collection for medical knowledge transfer and system learning, evaluation of the experimental results to visualization of the findings. A detailed evaluation of detection of multiple endoscopic findings, polyp-localization accuracy and system performance has been performed. We introduced two new multi-class classification methods, one based on a deep learning neural network approach and another new multi-class classification algorithm based on global image features. For the localization, we evaluated existing localization approaches based on deep learning neural networks and compared the results to our initial localization method.
The novelty of the research includes an end-to-end implementation of the whole EIR system pipeline, from frame capture, annotation and analysis to user (doctor) feedback, as a combination of many out-of-the-box and modified existing components, as well as several new ones. The experiments showed that the proposed system (i.e., both the global feature-based and the neural network-based implementations) can achieve equal results to state-of-the-art methods in terms of detection performance for state-of-the-art endoscopic data, and a comparable localization performance. Further, we showed that the new EIR system outperforms state-of-the-art systems in terms of system performance, that it scales in terms of data throughput and that it can be used in a real-time scenario. We concluded, based on our initial experiments, that the global features multi-class detection approach slightly outperforms the tested neural network approaches, and that the localization algorithm can benefit from combining local features and neural network approaches. We also presented automatic analysis of VCE videos and live support of colonoscopies as two real-world use cases that can potentially benefit from the proposed system where clinical tests are currently being planned in our partner hospitals. The experimental evaluation of the system as well as dataset creation are performed in collaboration with the Cancer Registry of Norway, and in the near future, the system will be tested in a real-world environment, i.e., it will have a real societal impact.
For future work, we plan to further improve the multi-class detection and localization accuracy of the system and support detection and localization of more abnormalities. In this respect, we are currently working with medical experts to collect more training data, annotate them and create new, larger training and testing datasets [30, 31]. Finally, to further improve the performance of the system, we work on a universal system extension that will allow the system to utilize the computing power of one or more GPUs on single or multiple nodes. Implementing such an extension will allow parallelization of the detection and localization workloads [32], which is important in our multi-disease analysis system of GI tract [32, 35, 37,38,39].
Notes
In Scandinavian mythology, EIR is a goddess with medical skill.
References
Abadi M, Barham P, Chen J, Chen Z, Davis A, Dean J, Devin M, Ghemawat S, Irving G, Isard M, etal (2016) Tensorflow: a system for large-scale machine learning. In: Proceedings of OSDI
Albisser Z, Riegler M, Halvorsen P, Zhou J, Griwodz C, Balasingham I, Gurrin C (2015) Expert driven semi-supervised elucidation tool for medical endoscopic videos. In: Proceedings of MMSys, pp 73–76
Alexandre LA, Casteleiro J, Nobreinst N (2007) Polyp detection in endoscopic video using svms. In: Proceedings of PKDD, pp 358–365
Altman NS (1992) An introduction to kernel and nearest-neighbor nonparametric regression. Am Stat 46(3):175–185
Ameling S, Wirth S, Paulus D, Lacey G, Vilarino F (2009) Texture-based polyp detection in colonoscopy. In: Proceedings of bfm, pp 346–350
Breiman L (2001) Random forests. Mach Learn 45(1):5–32
Buades A, Coll B, Morel JM (2011) Non-local means denoising. Image Processing On Line 1:208–212
Chaabouni S, Benois-Pineau J, Amar CB (2016) Transfer learning with deep networks for saliency prediction in natural video. In: Proceedings of ICIP, pp 1604–1608
Chatzichristofis S, Boutalis Y (2008) Cedd: color and edge directivity descriptor: a compact descriptor for image indexing and retrieval. Comput Vis Syst 312–322
Chatzichristofis SA, Boutalis YS (2008) Fcth: Fuzzy color and texture histogram-a low level feature for accurate image retrieval. In: 9th international workshop on image analysis for multimedia interactive services, 2008. WIAMIS’08. IEEE, pp 191–196
Cheng DC, Ting WC, Chen YF, Pu Q, Jiang X (2008) Colorectal polyps detection using texture features and support vector machine. In: Proceedigns of MDAISM, pp 62–72
Chin C, Brown DE (2000) Learning in science: a comparison of deep and surface approaches. J Res Sci Teach 37(2):109–138
Deng J, Dong W, Socher R, Li LJ, Li K, Fei-Fei L (2009) Imagenet: a large-scale hierarchical image database. In: Proceedings of CVPR, pp 248–255
Donahue J, Jia Y, Vinyals O, Hoffman J, Zhang N, Tzeng E, Darrell T (2014) Decaf: a deep convolutional activation feature for generic visual recognition. In: Proceedings of ICML, pp 647–655
Fitzgibbon AW, Fisher RB et al (1996) A buyer’s xguide to conic fitting. DAI Research paper
Hall M, Frank E, Holmes G, Pfahringer B, Reutemann P, Witten IH (2009) The weka data mining software: an update. ACM SIGKDD Explor Newslet 11 (1):10–18
Holme Ø., Bretthauer M, Fretheim A, Odgaard-Jensen J, Hoff G (2013) Flexible sigmoidoscopy versus faecal occult blood testing for colorectal cancer screening in asymptomatic individuals. The Cochrane Library
Hwang S, Oh J, Tavanapong W, Wong J, de Groen P (2007) Polyp detection in colonoscopy video using elliptical shape feature. In: Proceedings of ICIP, pp 465–468
Imagenet ImageNet Challenge Datasets. http://www.image-net.org/. [last visited, March 06, 2016]
Kaminski MF, Regula J, Kraszewska E, Polkowski M, Wojciechowska U, Didkowska J, Zwierko M, Rupinski M, Nowacki MP, Butruk E (2010) Quality indicators for colonoscopy and the risk of interval cancer. N Engl J Med 362 (19):1795–1803
Kang J, Doraiswami R (2003) Real-time image processing system for endoscopic applications. In: Proceedings of CCECE, vol 3, pp 1469–1472
Khaleghi A, Balasingham I (2015) Wireless communication link for capsule endoscope at 600 mhz. In: Proceedings of EMBC, pp 4081–4084
Li B, Meng MH (2012) Tumor recognition in wireless capsule endoscopy images using textural features and svm-based feature selection. IEEE Trans Inf Technol Biomed 16(3):323–329
Lux M, Marques O (2013) Visual information retrieval using java and lire. Synt Lect Inform Conc Retri Serv 5(1):1–112
Mallery S, Van Dam J (2000) Advances in diagnostic and therapeutic endoscopy. Med Clin N Am 84(5):1059–1083
Mamonov A, Figueiredo I, Figueiredo P, Tsai YH (2014) Automated polyp detection in colon capsule endoscopy. IEEE Trans Med Imaging 33(7):1488–1502
Ngiam J, Coates A, Lahiri A, Prochnow B, Le QV, Ng AY (2011) On optimization methods for deep learning. In: Proceedings of ICML, pp 265–272
Nguyen A, Yosinski J, Clune J (2014) Deep neural networks are easily fooled: High confidence predictions for unrecognizable images. arXiv:1412.1897
O’Connell JB, Maggard MA, Ko CY (2004) Colon cancer survival rates with the new american joint committee on cancer sixth edition staging. J Natl Cancer Inst 96(19):1420–1425
Pogorelov K, Randel KR, Griwodz C, Eskeland SL, de Lange T, Johansen D, Spampinato C, Dang-Nguyen DT, Lux M, Schmidt PT, Riegler M, Halvorsen P (2017) Kvasir: a multi-class image dataset for computer aided gastrointestinal disease detection. In: Proceedings of MMSYS, pp 164–169
Pogorelov K, Randel KR, de Lange T, Eskeland SL, Griwodz C, Johansen D, Spampinato C, Taschwer M, Lux M, Schmidt PT, Riegler M, Halvorsen P (2017) Nerthus: a bowel preparation quality video dataset. In: Proceedings of MMSYS, pp 170–174
Pogorelov K, Riegler M, Halvorsen P, Schmidt PT, Griwodz C, Johansen D, Eskeland SL, de Lange T (2016) GPU-Accelerated real-time gastrointestinal diseases detection. In: Proceedings of CBMS, pp 185–190
Redmon J Darknet: Open source neural networks in C. http://pjreddie.com/darknet/. [last visited, March 06, 2016]
Redmon J, Divvala S, Girshick R, Farhadi A (2015) You only look once: Unified, real-time object detection. arXiv:1506.02640
Riegler M, Griwodz C, Spampinato C, de Lange T, Eskeland SL, Pogorelov K, Tavanapong W, Schmidt PT, Gurrin C, Johansen D, Johansen H, Halvorsen P (2016) Multimedia and medicine: Teammates for better disease detection and survival. In: Proceedings of ACM MM, pp 968–977
Riegler M, Pogorelov K, Eskeland SL, Thelin Schmidt P, Albisser Z, Johansen D, Griwodz C, Halvorsen P, de Lange T (2017) From annotation to computer aided diagnosis: Detailed evaluation of a medical multimedia system. ACM Trans Multimed Comput Commun Appl 9(4)
Riegler M, Pogorelov K, Halvorsen P, de Lange T, Griwodz C, Johansen D, Schmidt PT, Eskeland SL (2016) Eir - efficient computer aided diagnosis framework for gastrointestinal endoscopies. In: Proceedings of CBMI, pp 1–6
Riegler M, Pogorelov K, Lux M, Halvorsen P, Griwodz C, de Lange T, Eskeland SL (2016) Explorative hyperbolic-tree-based clustering tool for unsupervised knowledge discovery. In: Proceedings of CBMI, pp 1–4
Riegler M, Pogorelov K, Markussen J, Lux M, Stensland HK, de Lange T, Griwodz C, Halvorsen P, Johansen D, Schmidt PT, Eskeland SL (2016) Computer aided disease detection system for gastrointestinal examinations. In: Proceedings of MMSys, p 29
Schmidhuber J (2015) Deep learning in neural networks: an overview. Neural Netw 61:85–117
Stewart R, Andriluka M (2015) End-to-end people detection in crowded scenes. arXiv
Szegedy C, Liu W, Jia Y, Sermanet P, Reed SE, Anguelov D, Erhan D, Vanhoucke V, Rabinovich A (2014) Going deeper with convolutions. CoRR 1409.4842
Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z (2015) Rethinking the inception architecture for computer vision. arXiv:1512.00567
Tajbakhsh N, Gurudu SR, Liang J (2016) Automated polyp detection in colonoscopy videos using shape and context information. IEEE Trans Med Imaging 35 (2):630–644
Tamura H, Mori S, Yamawaki T (1978) Textural features corresponding to visual perception. IEEE Trans Syst Man Cybern 8(6):460–473
Tanimoto TT (1958) Elementary mathematical theory of classification and prediction
The New York Times: The 2.7 Trillion Medical Bill. http://goo.gl/CuFyFJ. [last visited, Nov. 29, 2015]
Tieleman T, Hinton G (2012) Lecture 6.5-rmsprop: Divide the gradient by a running average of its recent magnitude COURSERA. Neural Networks for Machine Learning 4(2)
Van Essen B, Macaraeg C, Gokhale M, Prenger R (2012) Accelerating a random forest classifier: Multi-core, GP-GPU, or FPGA?. In: Proceedings of FCCM, pp 232–239
von Karsa L, Patnick J, Segnan N (2012) European guidelines for quality assurance in colorectal cancer screening and diagnosis. first edition–executive summary. Endoscopy 44(S 03):SE1–SE8
Wang Y, Tavanapong W, Wong J, Oh J, de Groen PC (2014) Part-based multiderivative edge cross-sectional profiles for polyp detection in colonoscopy. In: Proceedings of BHI, vol 18, pp 1379–1389
Wang Y, Tavanapong W, Wong J, Oh JH, De Groen PC (2015) Polyp-alert: Near real-time feedback during colonoscopy. Comput Meth Programs Biomed 120 (3):164–179
Zagoris K, Chatzichristofis SA, Papamarkos N, Boutalis YS (2010) Automatic image annotation and retrieval using the joint composite descriptor. In: 14th panhellenic conference on informatics (PCI), 2010. IEEE, pp 143–147
Zhou M, Bao G, Geng Y, Alkandari B, Li X (2014) Polyp detection and radius measurement in small intestine using video capsule endoscopy. In: Proceedings of BMEI, pp 237–241
Acknowledgements
This work is founded by the FRINATEK project “EONS” #231687.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Pogorelov, K., Riegler, M., Eskeland, S.L. et al. Efficient disease detection in gastrointestinal videos – global features versus neural networks. Multimed Tools Appl 76, 22493–22525 (2017). https://doi.org/10.1007/s11042-017-4989-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11042-017-4989-y