Abstract
Phylogeographic studies of highly mobile large carnivores suggest that intra-specific genetic differentiation of modern species might be the consequence of the most recent Pleistocene glaciation. However, the relative influence of biogeographical processes and subsequent human-induced population fragmentation requires a better understanding. Poland represents the western edge of relatively continuous distributions of many wide-ranging species, e.g. lynx (Lynx lynx), wolves (Canis lupus), moose (Alces alces) and, therefore, a key area for understanding historic and contemporary patterns of gene flow in central Europe. We examined wolf genetic structure in Poland and in a recently recolonized area in eastern Germany using microsatellite profiles (n = 457) and mitochondrial DNA sequencing (mtDNA, n = 333) from faecal samples. We found significant genetic structure and high levels of differentiation between wolves in the Carpathian Mountains and the Polish lowlands. Our findings are consistent with previously reported mtDNA subdivision between northern lowlands and southern mountains, and add new and concordant findings based on autosomal marker variation. Wolves in western Poland and eastern Germany showed limited differentiation from northeastern Poland. Although the presence of private alleles suggests immigration also from areas not sampled in this study, most individuals seem to be immigrants from northeastern Poland or their descendants. We observed moderate genetic differentiation between certain northeastern lowland regions separated by less than 50 km. Moreover, mtDNA results indicated a southeastern subpopulation near the border with Ukraine. The observed structure might reflect landscape fragmentation and/or ecological differences resulting in natal habitat-biased dispersal.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Pleistocene glaciations have had profound implications for biogeographical processes such as distributions, population fragmentation and gene flow in wild species (e.g. Taberlet et al. 1998; Hewitt 2000). Data from highly mobile large carnivores indicate weak phylogeographic structuring prior to the last glacial maximum, suggesting that the current distributions of distinct genetic lineages arose in consequence of the most recent glaciation (Hofreiter et al. 2004). However, ensuing human activity has resulted in further and more recent fragmentation in populations of species such as European brown bears (Ursus arctos) (Zedrosser et al. 2001), Eurasian lynx (Lynx lynx) (Schmidt et al. 2011) and Canadian wolves (Canis lupus) (Stronen et al. 2012). The extent to which historical biogeographical processes and recent human-induced fragmentation have contributed to shape the extant population genetic subdivisions in vagile taxa requires further understanding.
An abrupt ecological transition between the Great European Plains and the Carpathian Mountains occurs in southern Poland. These areas represent potential contact zones between genetically divergent lineages of organisms that recolonized central Europe during the Holocene from different glacial refugia in the Balkans, the Carpathians, and in other parts of eastern Europe (Bhagwat and Willis 2008 and references therein) In several species, contact zones between different mitochondrial DNA (mtDNA) lineages have been detected in Poland. Examples include the Norway spruce Picea abies (Tollefsrud et al. 2008), the bank vole Myodes glareolus (Wójcik et al. 2010), and the red deer Cervus elaphus (Niedziałkowska et al. 2011). Poland represents the western edge of relatively continuous distributions of wide-ranging Palaearctic species, including the lynx (Schmidt et al. 2009), moose (Alces alces) (Schmölcke and Zachos 2005), and wolf (Jędrzejewski et al. 2010). Some of these populations are now expanding westward, recolonizing parts of central Europe [e.g., the moose (Schmölcke and Zachos 2005, and references therein); the wolf (Nowak et al. 2011)]. Other populations are geographically and perhaps genetically isolated. Lynx inhabiting northeastern Poland were reported to be genetically distinct from their conspecifics in Latvia and Estonia and are presumably isolated by unsuitable dispersal habitats (Schmidt et al. 2009). Moose appear to be expanding from their western range edge in Poland into the Czech Republic and Germany (reviewed in Schmölcke and Zachos 2005). Although the population structure of red deer in Poland and neighbouring countries has been mainly shaped by human translocation, some differentiation between populations in Carpathian Mountains and northern Poland is still detectable (Niedziałkowska et al. 2011, 2012 and references therein).
Wolves in eastern Europe show high levels of genetic diversity and distinct mtDNA haplogroups (Pilot et al. 2010). Two mtDNA haplogroups, which might have diverged about 200,000 years ago, converge in the Carpathian Mountains (Pilot et al. 2010). However, earlier results from microsatellite markers could not identify distinct populations in NE Poland and the Carpathian Mountain (Pilot et al. 2006), underlining the need to clarify patterns of gene flow and the origin of expanding populations.
Distinct genetic structuring of the Polish wolf population was suggested by an earlier large-scale study of central and eastern European wolves (Pilot et al. 2006). Although Pilot et al. (2006) sampled only a portion of the Polish wolf range, they detected 3–4 subpopulations delimited by mtDNA and 2 subpopulations based on microsatellite loci. Western Poland has abundant wolf habitat, and the country could support two to three times its current population of approximately 650 wolves (as estimated by Jędrzejewski et al. 2008, 2010). Wolves have recently recolonized parts of western Poland and eastern Germany, probably from northeastern Poland (Jędrzejewski et al. 2004, 2005a; Ansorge et al. 2006; Nowak et al. 2011). Consequently, Poland represents a key area for understanding historic and contemporary patterns of wolf gene flow in central Europe.
In this study we sampled the entire wolf distribution range in Poland and a portion of eastern Germany that now represents a natural extension of the population in western Poland. The wolf is a protected species, and we consequently based our investigations mainly on non-invasive molecular techniques using DNA extracted from faecal samples. The aim of our study was to answer the following questions: (1) Are wolf populations genetically structured across Poland? (2) Do mitochondrial and nuclear genetic markers show concordant results? Because the central European wolf range has expanded over the past decades, we also asked: (3) Were western Poland and eastern Germany recolonized by wolves from northeastern or southern Poland?
Materials and methods
Study area
We collected wolf scat and tissue samples throughout Poland (311,904 km2, 49°00′–54°50′N, 14°08′–24°09′E). This territory extends throughout various geographical regions, with lowlands (<300 m a.s.l.) dominating in the northern and central part and uplands (301–500 m a.s.l.) and mountains (501–2,499 m a.s.l.) in the south. The climate of Poland is temperate with transitional character (oceanic in the north and west, continental in the east). The mean annual temperature decreases from south-west (8.5 °C) to north-east (6 °C). Annual precipitation is 500–650 mm in the lowlands and 1,200–1,500 mm in the mountains. The snow cover persists from an average of 40 days in south-western Poland to 100 days in the north-eastern part of the country and 200 days in the mountains (Concise Statistical Yearbook of Poland 2011). The mean human population density is 122 inhabitants/km2 and ranges from 20 in north-western and north-eastern Poland to 500 inhabitants/km2 in Upper Silesia in south-western Poland (Demographic Yearbook of Poland 2011).
Forests cover 29 % of the country; the rest is primarily farmlands (60 %) with predominance of arable land. Most forests are coniferous stands (51 %) dominated by Scots pine (Pinus sylvestris, 60 % of the forest area) and Norway spruce (Picea abies, 6 %). Deciduous and mixed forest with oak (Quercus robur and Q. petraea), ash (Fraxinus excelsior), maple (Acer platanoides and A. pseudoplatanus), beech (Fagus sylvatica), hornbeam (Carpinus betulus), birch (Betula pubescens and B. pendula) and alder (Alnus glutinosa) constitute about one-fourth of all forest land. About 98 % of woodlands are commercial stands and only 2 % are national parks and reserves (Forestry 2011).
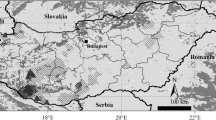
Almost all of Poland is inhabited by three native species of ungulates: red deer, wild boar (Sus scrofa), and roe deer (Capreolus capreolus). In north-eastern and central parts of the country, moose and three populations of European bison (Bison bonasus) occur. The Carpathian Mountains harbour isolated populations of chamois (Rupicapra rupicapra) and European bison. Populations of fallow deer (Dama dama), sika deer (Cervus nippon) and mouflon (Ovis musimon) have been introduced to certain forests (Wawrzyniak et al. 2010). Large predators—wolf and lynx—occur mainly in the north-eastern, eastern, and south-eastern parts of the country (Jędrzejewski et al. 2004, 2008; Niedziałkowska et al. 2006). However, wolves have recently been recolonizing the large woodlands of western Poland (Nowak et al. 2011) and the current range of wolves in Poland is shown in Fig. 1.
Wolf (Canis lupus) genotypes analysed in this study shown on the background of wolf range and forest cover in Poland. See Table 1 for region names (1–12)
Sample collection
Wolf scat and tissue samples were collected during 2001–2009. Foresters, national park rangers, students, volunteers, and personnel of the Mammal Research Institute of the Polish Academy of Sciences (hereafter MRI PAS), the Association for Nature “Wolf”, and the Institute of Nature Conservation PAS surveyed areas within the presently known wolf range and gathered wolf scats throughout the year. Small fragments of fresh wolf faeces were either stored in plastic tubes (5–30 ml) filled with 96 % alcohol or ALS buffer (QIAGEN), or kept frozen at −20 °C. In total, 1696 scat samples were collected in Poland (Table 1). In addition, 19 scat samples from eastern Germany were added to supplement data from recently colonized areas near the Polish-German border (Lower Silesia Forest). Information on dead wolves (mainly poached or killed in vehicle collisions) was also recorded and tissue or skin samples (n = 38) were collected. We henceforth refer to geographic sampling areas as sampling regions (n = 12, Fig. 1) to distinguish these units from clustering results obtained from genetic analyses. Finally, twenty-five dog samples (blood and tissue), to be used as a reference group for detecting possible wolf-dog hybrids, were collected in Poland from private owners and from individuals killed by vehicles.
Laboratory methods
DNA from faecal samples was isolated using the QIAamp DNA Stool Mini Kit (QIAGEN) and DNA from tissue samples was extracted with the DNAeasy tissue kit (QIAGEN).
For a portion of the samples collected in southern Poland (n = 185), DNA was isolated using a phenol–chloroform method (Sambrook and Russel 2001) with two additional phenol extractions at the beginning of the procedure, at the Institute of Nature Conservation PAS in Cracow, Poland. In order to reduce contamination risk, scat extraction was carried out in a room dedicated to non-invasive samples. Negative controls were included in each extraction set to monitor for contamination.
Amplification of a 230 bp mtDNA fragment of the HV1 region was performed using primers described by Savolainen et al. (1997). Amplifications were carried out in 10 μl reaction volumes containing 1U Taq polymerase, 200 μM dNTP, 2.0 μl 10× concentrated PCR buffer, 1.5 mM MgCl2, 0.1 mM of primers, 0.2 μl of BSA (Fermentas) plus 0.2 μl PCR Anti-inhibitor (DNA Gdańsk, Poland) and 2 or 3.6 μl of DNA extract, respectively, for tissue or scat samples. The reaction conditions were as follows: 2 min at 94 °C of initial denaturation, 30 cycles (tissue) or 36 cycles (scats) of 20 s at 94 °C, 30 s at 69 °C, 40 s at 72 °C, and the final extension step for 10 min at 72 °C. We subsequently amplified mtDNA sequences for each region using a portion of the samples identified as having distinct microsatellite profiles. PCR products were purified using Clean Up (A&A Biotechnology, Gdańsk, Poland). Sequencing reactions were carried out in 10 μl volumes using the Big Dye ver. 3.1 sequencing kit (Applied Biosystems) with the forward primer. Products were purified with the Exterminator kit (A&A Biotechnology) and separated on an ABI PRISM 3100 Genetic Analyser (Applied Biosystems). Sequencing results were analysed with ABI PRISM DNA Sequencing Analysis software and aligned by eye in BioEdit ver. 7.0.9. (Hall 1999). Haplotypes were compared to previously recorded sequences from Pilot et al. (2006) using Collapse ver. 1.1 (D. Posada, http://darwin.uvigo.es/software/collapse.html).
Microsatellite genotyping was performed using 11 polymorphic loci: FH2001, FH2010, FH2017, FH2054, FH2079, FH 2088, FH2096, FH2137, and FH2140 (Francisco et al. 1996), C213 (Ostrander et al. 1993) and VWF (Shibuya et al. 1994). Amplifications were carried out in four multiplex reactions using Multiplex PCR Kit (QIAGEN) and PCR conditions described in the manufacturer’s instructions with modifications by adding 0.1 μl BSA (Fermentas) and 0.1 μl PCR Anti-inhibitor (DNA Gdańsk). Cycling was performed on a DNA Engine Dyad Peltier Thermal Cycler (BIO RAD) with the following profile: 16 cycles of 94 °C for 30 s, 58 °C for 90 s, 72 °C for 60 s, followed by 10 cycles of 94 °C for 30 s, 57 °C for 90 s, 72 °C for 60 s and 10 cycles of 94 °C for 30 s, 56 °C for 90 s, 72 °C for 60 s with a final extension of 60 °C for 30 min, after an initial denaturation step of 95 °C for 15 min. Fragments were separated using genetic analyzer ABI3100 and allele lengths were determined with GENEMAPPER 3.5 (Applied Biosystems) and GENEMARKER 1.51 (SoftGenetics LLC). Each scat sample was amplified at least three independent times through a multiple-tube approach (Taberlet et al. 1996). We accepted alleles confirmed by a minimum of two independent PCR amplifications. Only individuals for which six or more loci had been successfully amplified were included in subsequent analyses.
Additionally, 66 samples were analysed at nine single nucleotide polymorphism (SNP) loci: 372M9, BLA22, BLB52, 168J14, 1C06, 38K22, 182B11, 218J14, 309N24, using the TaqMan Assay protocol described in Fabbri et al. (2012) at the Laboratory of Genetics at Istituto Superiore per la Protezione e la Ricerca Ambientale (ISPRA), Italy, to verify individual identification, with focus on samples genotyped at ≤9 loci. If not stated otherwise, all analyses were done at MRI PAS, Poland.
Probability of identity and test for the presence of wolf-dog hybrids
We used GIMLET 1.3.3 (Valière 2002) to identify false homozygotes (drop-out alleles) and false alleles by comparing repeated genotypes, construct a consensus genotype from the sets of PCR repetitions for each sample of faeces, and identify multiple samples from the same individual. Multilocus genotypes were analysed with 25 reference dog genotypes to detect any dogs or wolf-dog hybrids using the Bayesian clustering procedure implemented in STRUCTURE (Pritchard et al. 2000) as outlined in previous analyses of wolves from Italy (Verardi et al. 2006; Randi 2008). We ran STRUCTURE using 105 iterations, following a burn-in period of 104 iterations, to assign individuals to clusters. Five independent runs were completed for each K value (1–10). We verified that likelihood values converged for each run of K. All samples with membership q i < 0.90 in the wolf cluster (in total 17 dogs and possible wolf-dog hybrids, not concentrated in any particular region) were excluded, and we continued analyses only with samples considered to be wolves.
Probability of identity (PI) for an increasing number of loci, i.e. the probability that different individuals share an identical multilocus genotype by chance, and the PI between sibs (PIsib) (Waits et al. 2001) were calculated in GenAlEx ver. 6.1 (Peakall and Smouse 2006). Additionally, the number of matches between wolf genotypes was calculated to determine the minimum number of loci needed for individual identification.
Statistical analyses
Genetic diversity statistics for mtDNA sequences including the number of haplotypes and polymorphic sites, haplotype and nucleotide diversity were estimated using DnaSP ver. 5 (Librado and Rozas 2009). To infer population genetic structure based on mtDNA haplotype frequency we used the spatial analysis of molecular variance implemented in the SAMOVA software (Dupanloup et al. 2002). This approach defines groups of populations that are geographically homogenous and maximally differentiated from each other. The method is based on a simulated annealing procedure that aims to maximize the proportion of total genetic variance due to differences between groups of populations. This approach requires the a priori definition of the number (K) of groups. We ran SAMOVA with K ranging from 2 to 6. Each analysis was performed twice to check for consistency of results. In each run, 100 simulated annealing processes were performed. The recognition of the most probable number of groups was based on the pattern of changes in values of Φ-statistic parameters with K.
Microsatellite variability statistics, including the number of alleles per locus and the number of private alleles, observed (H O) and expected (H E) heterozygosity, and tests for departures from Hardy–Weinberg equilibrium were calculated in GenAlEx. Allelic richness, which corrects the observed number of alleles for differences in sample size, and the inbreeding estimator Wright’s F IS were computed with FSTAT ver. 2.9.3 (Goudet 1995). To test for genetic differentiation among sampling regions, pairwise F ST values (Weir and Cockerham 1984) were calculated in Arlequin ver. 3.11 (Excoffier et al. 2005). Genetic differentiation between populations was characterized by exact tests (Raymond and Rousset 1995), using GENEPOP ver. 3.1. The levels of significance for multiple tests were adjusted by the sequential Bonferroni method (Rice 1989).
Two complementary Bayesian clustering algorithms, STRUCTURE ver. 2.3 (Pritchard et al. 2000) and GENELAND ver. 3.3 (Guillot et al. 2005), were applied to infer population structure and assign individuals to subpopulations (clusters) based on individual multilocus genotypes and—in GENELAND—their spatial location. The posterior probability of the data [LnP(D)] was estimated from four replicate runs for a number of groups K = 1 to 10, each with a burn-in of 104 followed by 105 iterations, using the admixture model and correlated allele frequencies (Falush et al. 2003). Inferences in GENELAND were done in a single step as recommended by the authors (Guillot 2008). This approach makes inferences faster and avoids the issue of ghost populations. Thus, we ran GENELAND 50 times allowing K to vary from 1 to 15, with the following parameters: 205 MCMC iterations, maximum rate of Poisson process fixed to 100, uncertainty attached to spatial coordinates fixed to 0.5°, and the maximum number of nuclei in the Poisson–Voronoi tessellation fixed to 300. We computed the posterior probability of subpopulation membership for each pixel of the spatial domain and the modal subpopulation for each individual in all 50 runs. Finally, we examined all runs for consistency.
Subsequently, we evaluated population genetic structure performing principal component analyses (PCA) on individual genotypes from 4 subpopulations defined by STRUCTURE (with region 4 and 5 separated from regions 8–12 because of geographic distance and the GENELAND results) using the adegenet-package (Jombart 2008) in R 2.15.0 (R development Core Team 2012). The PCA does not assume Hardy–Weinberg equilibrium (HWE) and is highly efficient in revealing genetic structure in the form of clines, which is more difficult to detect than clusters (Jombart et al. 2009). We calculated pairwise genetic distances among wolves in the 12 regions based on mitochondrial and nuclear markers, and compared the results using the partial Mantel test in R with the vegan package (Oksanen et al. 2011).
Results
Genetic variability and structuring based on mtDNA
We found six mtDNA haplotypes (H1, H2, H3, H6, H8, H14—nomenclature consistent with Fig. 2 in Pilot et al. 2006) among 333 analysed samples. They are all known from previous studies (Vila et al. 1999; Randi et al. 2000; Jedrzejewski et al. 2005a; Pilot et al. 2006). H1, H2, H3 and H8 belong to haplogroup 1, which is widespread in north-eastern and central Europe and the Iberian Peninsula, whereas H6 and H14 belong to haplogroup 2 that dominates in south-eastern Europe and Italy (Pilot et al. 2010). The number of mtDNA haplotypes per region ranged from one to four (Table 2). We observed no relationship between the number of samples and the number of haplotypes per region (r 2 = 0.004, F 1,10 = 0.038, P > 0.8). Notably, we found the highest number of haplotypes (4) in regions 4 (n = 25 individuals analysed) and 5 (n = 10 inds). The latter region also showed the highest number of polymorphic sites (Table 2). Conversely, samples from regions 2 (n = 28 inds) and 7 (n = 27 inds) revealed only one haplotype, which were H6 and H2, respectively.
Four subpopulations (S1–S4) of wolves in Poland, based on mtDNA, delimited by SAMOVA (left panel) and haplotype frequencies for subpopulations (right). Sampling information is provided in Table 1
Results of STRUCTURE analysis for Polish wolves, based on 11 microsatellite loci, assuming K = 2 to 5 subpopulations of wolves. Black lines separate wolves from different sampling regions. Sampling information is provided in Table 1
Structuring of the Polish wolf population inferred by GENELAND in 50 runs. Individuals were assigned to subpopulation G1 in 43 of 50 runs and to population G2 in 40 of 50 runs. Other individuals were inconsistently clustered. Sampling information is provided in Table 1
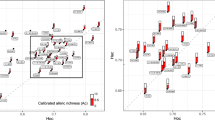
Principal Component Analysis (PCA) of Polish wolves representing 5 subpopulations suggested by STRUCTURE and GENELAND. The plot shows individual wolves organized by sampling regions (detailed in Table 1). Black ovals are 95 % inertia ellipses. Thirteen outliers (from seven different regions) were excluded to improve resolution of the figure. None of the 13 outliers appeared to have dog ancestry, and nine (for which mtDNA was analysed) had common wolf haplotypes
Relationship between pairwise genetic distances for Polish wolves from the 12 sampling regions based on microsatellite (F ST) and mtDNA (Φ ST) markers. Sampling information is provided in Table 1
Pairwise Φ ST values between the 12 geographically predefined regions were generally very high (>0.25) and statistically significant, which indicated isolation among many regions according to the mtDNA (Table 3). Only seven of the 66 pairwise distance values were low (<0.05). These low values occurred among the regions located in the same part of the country within the lowlands and the Carpathian Mountains.
The results of SAMOVA indicated significant population genetic structure for each assumed number of groups. The highest increase in Φ CT value occurred between K = 3 and K = 4 and all parameters of the Φ-statistics stabilized from K = 4 (Appendix: Fig. 7). Thus, we assumed four subpopulations as the most parsimonious clustering configuration that maximized variation among groups. Wolves from the regions 1, 2, and 3 in the Carpathians formed subpopulation S1 (Fig. 2). Subpopulation S2 was comprised exclusively of individuals from region 4 in south-eastern Poland. Wolves from regions 5, 8, 10, 11, and 12 formed the geographically disjunct subpopulation S3. Finally, wolves from regions 6, 7, and 9 formed subpopulation S4. Subpopulation S1 in the Carpathian Mountains was strongly dominated by haplogroup 2 (97 % of the wolves), and four haplotypes from haplogroup 1 were present there at low frequencies. In contrast, subpopulations S2, S3 and S4 were primarily comprised of individuals carrying haplotypes from the previously identified haplogroup 1 (S2—88 %, S3—96 %, and S4—99 %) and included haplogroup 2 at very low frequencies (Fig. 2). Pairwise Φ ST values among four subpopulations (S1–S4) were all very high (>0.25), which indicates little or no mtDNA gene flow among them (Appendix: Table 5).
Changes in Φ-statistics for K = 2 to 6 subpopulations of wolves in Poland, on the basis of mtDNA and inferred from SAMOVA. Φ SC—proportion of the variance among local populations within groups. Φ ST—proportion of the variance among local populations within the total population. Φ CT—proportion of the total variance explained by the grouping
Genetic differentiation and structuring based on microsatellites
We identified a total of 457 wolf genotypes. The allelic drop-out rate ranged from 0.079 to 0.360 among regions. The highest values were observed for the loci FH2010, FH2017 and FH2079 (Appendix: Table 6), which had fragment length >200 bp and thus higher probability of drop-out for low quality-samples (Broquet et al. 2007). Based on all 11 loci, the cumulative PI of all genotypes was 1.35 × 10−11 and the probability of identity between sibs (PIsib) equaled 5.0 × 10−5 (detailed data available upon request). Data from seven microsatellites were sufficient to distinguish all 457 genotypes with PI = 3.13 × 10−7. We used three samples genotyped at 6 loci. They were assigned to the region of origin, and so their inclusion is unlikely to have biased our results. SNP data for the 66 samples with ≤9 microsatellite loci amplified confirmed individual identifications with cumulative PI = 0.002 and PIsibs = 0.042. Microsatellite data for these 66 samples were therefore included in subsequent analyses.
The 11 loci were polymorphic in all regions and the average number of microsatellite alleles per locus varied between 4.27 and 6.64 (Table 2). Observed heterozygosity (H O) values (range 0.45–0.58) were lower than expected (H E, range 0.63–0.69). In the total population of Polish wolves we found significant deviation from HWE at all 11 loci (Appendix: Table 6). We also tested for HWE in subpopulations defined by STRUCTURE (Fig. 3). Four to seven of 11 loci (depending on subpopulation) showed significant deviation from HWE (after correcting for multiple tests), except for subpopulation 5 (regions 8–12) where 10 loci showed significant deviation. Only locus FH2137 displayed consistent deviation from HWE across all subpopulations, the results for other loci varied among subpopulations. F IS values ranged from 0.122 to 0.310 and were not correlated to the number of wolf genotypes per group (r 2 = 0.15, F 1,10 = 1.779, P > 0.2). We found no relationship between the number of mtDNA haplotypes and the number of microsatellite alleles per locus in the 12 regions (Pearson r = 0, F 1,10 = 0.002, P > 0.9). Individuals from regions 5, 9, and 10 did not show private alleles (Table 2). The highest number of private alleles was found in region 12 (western Poland and eastern Germany), which comprised 57 individuals distributed across one-third of the sampling area (see Fig. 1).
Pairwise F ST values between the 12 geographically predefined regions ranged from 0.011 to 0.212 (Table 3) and 59 of 66 pairwise comparisons remained significant after Bonferroni correction. F ST values generally indicated high (>0.15; Balloux and Lugon-Moulin 2002) or moderate (0.05–0.15) genetic differentiation among wolves from the Carpathian Mountains (regions 1, 2, 3) and other sampling regions. Within northern Poland, we observed high differentiation only between regions 6 and 7. The lowest differentiation (F ST < 0.05) was observed among regions 10, 11 and 12 (Table 3).
Microsatellite results from STRUCTURE suggested division of Polish wolves into two subpopulations: the Carpathian Mountains (regions 1–3) and the lowlands (regions 4–12) (Fig. 3). Although LnP(D) continued to increase when augmenting K in the STRUCTURE analyses, ΔK (Evanno et al. 2005) showed highest support for K = 2 followed by K = 5 groups (Appendix: Table 4). However, the pattern of individual assignment into five subpopulations only followed a visible spatial structure for four of the clusters: regions 1–3, region 4, region 6, and region 7 (Fig. 3). Removal of locus FH2137 did not change the genetic clusters identified in STRUCTURE with ΔK showing highest support for K = 2 followed by K = 4 groups.
Inferences made in GENELAND supported clear separation of the Carpathian Mountains (regions 1–3) and south-eastern areas comprising wolves from regions 4 and 5 (Fig. 4). Twenty-six of 50 runs gave a modal value at five whereas 24 gave a mode at six for the posterior distribution of K. None of the 50 runs indicated ghost populations. The location of the two southern clusters was constant among all 50 runs, and 147 wolves from regions 1, 2 and 3 were consistently assigned to the Carpathian subpopulation in 43 of 50 runs. Similarly, 35 individuals from region 4 were assigned to the southeastern subpopulation in 40 of 50 runs. The remaining 275 wolves from regions 6–12 were randomly and inconsistently divided into three or four groups.
Finally, the first two PCA components clearly divided wolves into 2 main groups. PC-1 differentiated the Carpathian wolves (region 1–3) from lowland individuals, and PC-2 separated regions 6 and 7, which both overlapped with regions 8–12 (Fig. 5). Wolves from regions 4–5 were placed on the PCA plot between the Carpathian and lowland individuals, which accords with the STRUCTURE and GENELAND results.
Genetic structuring of wolves based on microsatellite markers and inferred by three analytical tools (STRUCTURE, GENELAND, PCA) and results from mtDNA (SAMOVA) consistently divided the Polish wolf population into a Carpathian and a lowland subpopulation. Within the lowlands, only differentiation of region 4 (Roztocze) by mtDNA was supported by the microsatellite results (GENELAND, STRUCTURE with K = 5). We found concordance between pairwise genetic distances among wolves in 12 regions based on mitochondrial and nuclear markers (Fig. 6). A partial Mantel test (Smouse et al. 1986) where the geographic distance matrix was held constant, while the relationship of F ST and Φ ST was determined, showed a significant positive correlation of the two measures of genetic distance (Mantel r = 0.521, P = 0.001, test with 999 permutations).
Discussion
Are wolf populations structured across Poland?
We found significant genetic structure between wolf populations in the Carpathians and the Polish lowlands. Our mtDNA results seem to correspond with those reported from earlier research encompassing a larger area of eastern Europe. A Carpathian subpopulation indicated in Pilot et al. (2006) corresponds with subpopulation S1 identified in our study. Similarly, S2 corresponds with a subpopulation spanning from southeastern Poland through southern Belarus and northern Ukraine into Russia. S3 and S4 appear to be part of a subpopulation extending from northeastern Poland and eastward into Russia (Pilot et al. 2006). A subsequent study examined the historical distribution of haplogroups in (primarily) eastern Europe (Pilot et al. 2010). These findings suggest that most haplotypes from S1 belong to haplogroup 2 from southern Europe, whereas S2, S3, and S4 are principally composed of wolves from haplogroup 1. The present contact zone between the two lineages in central Europe appears to be a result of one haplogroup (1) partially replacing a more ancient haplogroup (2) that had been predominant over the last several millennia and remains widely distributed in southern Europe (Leonard et al. 2007; Pilot et al. 2010). The presence of mainly one haplotype (H6) from haplogroup 2 in the Polish Carpathians indicates low genetic diversity at its current northern range. Consequently, Poland can be seen as a meeting zone between two lineages of wolves. Despite the geographical proximity well within wolf dispersal distance (e.g. Wabakken et al. 2007), our results suggest very limited gene flow between the two areas and, hence, a restricted contact zone between the north and the south. Pilot et al. (2006) observed a single population cluster encompassing northern and southern Polish wolves. Our microsatellite results showed strong genetic structure between the Carpathians and the Polish lowlands, and thus demonstrate additional substructuring within Poland when compared with previous findings. The observed deviation from HWE may be due several reasons: the presence of null alleles, high allelic dropout rate in non-invasive studies, the existence of local genetic structure (Wahlund’s effect), lack of random mating, or the presence of closely related individuals (members of the family groups) in a sample. Although genotyping error and null alleles may have contributed to the excess of homozygotes, these factors are unlikely to explain the observed mountain-lowland structure in Polish wolves. Other studies on wolf population structure (that used tissue samples) also showed significant heterozygote deficit and positive values of F IS, which was explained by moderate inbreeding, the presence of closely related individuals, or the presence of additional undetected structure (Lucchini et al. 2004, Pilot et al. 2006, Jansson et al. 2012).
In addition to the north–south structure, our results indicated clustering within the lowland area. F ST values between small groups may reflect social structure and can show relatively high values between local family groups. For example, Thiessen (2007) reported between-pack differentiation of F ST = 0.179 based on n = 36 wolf packs in western Canada. Wolves typically live in social and territorial groups of 2–11 individuals (Fuller et al. 2003; Jędrzejewski et al. 2010). Although wolf social structure may have contributed to the high F ST value (0.156) observed between region 6 (n = 55) and region 7 (n = 45), the samples from these regions represent members of multiple packs (Jędrzejewski et al. 2004). Consequently, wolf pack structure alone is not expected to produce such high F ST values. The moderate F ST values (0.05–0.15) observed between a number of lowland regions require further investigation, as well as the high F ST value (0.156) seen between wolves in regions 6 and 7 that are separated by <50 km. Assessment of samples from contiguous regions, landscape features, and additional genetic markers could improve understanding of the extent to which these differences might be explained by landscape fragmentation or ecological differences resulting in natal habitat-biased dispersal (Geffen et al. 2004; Sacks et al. 2004; Pilot et al. 2006).
Factors that could maintain divisions between genetically distinct wolf populations in Poland
MtDNA and microsatellite results consistently showed differentiation between wolves in the Carpathian Mountains and the Polish lowlands. We examined only non-coding fragments of DNA, and future analyses of genes under selection (using e.g. SNP mapping within exons or regulatory sequences of functional genes) could help clarify whether adaptive genetic differences might play a role in the observed structure. Associations between environmental factors (habitat type, climate, prey abundance) and genetic variants might have resulted in the development of wolf ecotypes adapted to different habitats (e.g. Carmichael et al. 2001; Musiani et al. 2007; Muñoz-Fuentes et al. 2009). This is consistent with the findings that (1) ecological factors (habitat, prey, climate) appear to explain much of the spatial variation in wolf genetic diversity in east-central Europe (Pilot et al. 2006), and that (2) wolves from three genetically distinct populations in Poland show significant differences in prey composition and prey preferences (Jedrzejewski et al. 2012). In northeastern Poland, wolves prey on four ungulate species (moose, red and roe deer, wild boar), although only red deer was killed in a higher proportion than that expected from availability. In eastern Poland (regions 4 and 5) wolves preferred roe deer, whereas in the south-east (regions 1–3) they hunted mostly red deer and only a small proportion of roe deer and wild boar occurred in their diet (Jedrzejewski et al. 2012).
The genetic structure of wolves in Poland shows strong and abrupt spatial division, and the primary separation occurs between lowland and Carpathian Mountain wolves. Within the lowlands, both mtDNA and microsatellite results indicate a further division between region 4 (Roztocze) in the southeast and northern Poland. This appears to concur with previous mtDNA and microsatellite results that supported the presence of a subpopulation extending from southeastern Poland and eastward into Russia (Pilot et al. 2006). Interestingly, separation of Carpathian and lowland wolves by a ‘wolf-free belt’ has also been suggested to occur in Ukraine (Gursky 1985). Wolves are highly mobile (mean daily movement distance in Polish wolves: 23 km, max 64 km, Jędrzejewski et al. 2001). Moreover, long-distance dispersers can travel several hundred kilometers (Wabakken et al. 2001, 2007; Schede et al. 2010), so individuals from different populations should be able to meet and interbreed. The observed genetic isolation of Carpathian wolves is thus surprising given the species’ remarkable dispersal ability. Several factors could nevertheless contribute to the observed divisions. First, analyses of habitat structure in and around wolf ranges conducted in southern and northern Poland (Jędrzejewski et al. 2004, 2005a) showed very rapid deterioration of habitat connectivity immediately north of the Carpathians. In contrast, northeastern Poland provides better wolf habitat because the human density and network of transportation infrastructure is lower here than in southern Poland (Jędrzejewski et al. 2004, 2005a). In southern Poland, wolves occurred only in a narrow belt of <100 km, whereas in northern Poland stable wolf populations persist more than 200 km from the continuous wolf range (Jędrzejewski et al. 2004, 2005a).
Huck et al. (2010, 2011) analysed dispersal costs among patches of wolf habitat and modeled dispersal corridors in Poland, and found that dispersal from the Carpathians to any other patch would be much more costly than dispersal among other regions of the country. Densely populated and urbanized areas in southern Poland along the Carpathians may act as a serious barrier to wolf movement and limit wolf dispersal (Huck et al. 2011). Moreover, modeling of suitable wolf habitat showed that the eastern portion of Poland was already ‘filled’ by wolves (Jędrzejewski et al. 2008). Western Poland still has much suitable wolf habitat not occupied at present, which could support a large population of wolves (Jędrzejewski et al. 2008). Hence, dispersers would likely prefer to settle in western Poland than the more saturated east.
The current landscape structure and wolf distribution in Poland are important factors that are likely to limit gene flow between Carpathian and lowland wolves. However, wolves can move through highly heterogeneous and human dominated landscapes (Blanco and Cortes 2007) and cross a range of natural and anthropogenic barriers (Blanco et al. 2005; Wabakken et al. 2007; Ciucci et al. 2009). Other factors might therefore contribute to the observed structure. Natal-habitat biased dispersal seems a possible explanation in the case of Carpathian wolves, and is consistent with findings from Europe and North America (Geffen et al. 2004; Sacks et al. 2004; Pilot et al. 2006). Population genetic structure consistent with the presence of highland and lowland habitats has been reported in coyotes (Sacks et al. 2004, 2005) and merits further attention in wolf populations from the Carpathians Mountains and surrounding lowland areas. Differences in the legal status and protection of the species in the Carpathians might also influence genetic structure, as only wolves in the Polish and Czech parts of the north-western edge of the Carpathian Mountains are protected. In Slovakia and Ukraine, wolves are regularly hunted or persecuted. This probably causes a source-sink effect and thus dispersal southward from the Polish to the Slovakian and Ukrainian parts of the Carpathian Mountains, where dispersers repopulate vacant territories (Nowak et al. 2008).
Recolonization of western Poland and eastern Germany
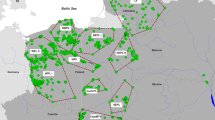
Our results suggest that wolves colonizing western Poland and eastern Germany primarily originate from northeastern Poland. In particular, it appears that westward dispersal from regions 10 and 11 has been relatively frequent. The location of these sampling regions on the western border of the established wolf range in northeastern Poland, and the relatively contiguous forest habitat in this area (Huck et al. 2011) suggest that regions 10 and 11 represent a natural starting point for westward expansion (Jędrzejewski et al. 2008; Huck et al. 2010, 2011). Wolves in western Poland and eastern Germany appear to represent the expanding western edge of a vast, northeastern European wolf population that primarily inhabits boreal and temperate forests and extends through the Baltic States, northern Belarus, and northwestern Russia (Pilot et al. 2006, 2010).
Importantly, our study detected private alleles in region 12 (western Poland and eastern Germany). Although these alleles might be present in unsampled northeastern Polish wolves, the most likely explanation is immigration from areas not covered by our investigation. East-European countries (Belarus, Latvia) harbour the region’s largest wolf populations, although human harvest is high and potentially unsustainable in some areas (Jędrzejewski et al. 2010), which might affect source-sink dynamics (Jedrzejewski et al. 2005b). Recently documented movements of a radio-collared wolf between Germany and Belarus further support such dispersal (Schede et al. 2010). The apparent recolonization of western Poland and eastern Germany from various source populations should help ensure high levels of genetic variation and subsequent potential for adaptation to new and altered environments.
References
Ansorge H, Kluth G, Hahne S (2006) Feeding ecology of wolves Canis lupus returning to Germany. Acta Theriol 51:99–106
Balloux F, Lugon-Moulin N (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165
Bhagwat SA, Willis KJ (2008) Species persistence in northerly glacial refugia of Europe: a matter of chance or biogeographical traits? J Biogeogr 35:464–482
Blanco JC, Cortes Y (2007) Dispersal patterns, social structure and mortality of wolves living in agricultural habitats in Spain. J Zool 273:114–124
Blanco JC, Cortes Y, Virgos E (2005) Wolf response to two kinds of barriers in an agricultural habitats in Spain. Can J Zool 83:312–323
Broquet T, Menard N, Petit E (2007) Nonivasive population genetics: a review of sample source, diet, fragment length and microsatellite motif effects on amplification success and genotyping error rate. Conserv Genet 8:249–260
Carmichael LE, Nagy JA, Larter NC, Strobeck C (2001) Prey specialization may influence patterns of gene flow in wolves of the Canadian Northwest. Mol Ecol 10:2787–2798
Ciucci P, Reggioni W, Maiorano L, Boitani L (2009) Long-distance dispersal of a rescued wolf from the northern Apennines to the western Alps. J Wildl Manage 73:1300–1306
Concise Statistical Yearbook of Poland (2011) Central Statistical Office, Warszawa
Demographic Yearbook of Poland (2011) Central Statistical Office, Warszawa
Dupanloup I, Schneider S, Excoffier L (2002) A simulated annealing approach to define the genetic structure of populations. Mol Ecol 11:2571–2581
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinformatics Online 1:47–50
Fabbri E, Caniglia R, Mucci N, Thomsen HP, Krag K, Pertoldi C, Loeschcke V, Randi E (2012) Comparison of single nucleotide polymorphisms and microsatellites in non-invasive genetic monitoring of a wolf population. Arch Biol Sci Belgrade 64:321–335
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Forestry (2011) Central Statistical Office, Warszawa
Francisco LV, Langston AA, Mellersh CS, Neal CL, Ostrander EA (1996) A class of highly polymorphic tetranucleotide repeats for canine genetic mapping. Mamm Genome 7:359–362
Fuller TK, Mech LD, Cochrane JF (2003) Wolf population dynamics. In: Mech LD, Boitani L (eds) Wolves: behaviour, ecology and conservation. University of Chicago Press, Chicago, pp 161–191
Geffen E, Anderson MJ, Wayne RK (2004) Climate and habitat barriers to dispersal in the highly mobile gray wolf. Mol Ecol 13:2481–2490
Goudet J (1995) FSTAT (version 1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Guillot G (2008) Inference of structure in subdivided populations at low levels of genetic differentiation. The correlated allele frequencies model revisited. Bioinformatics 24:2222–2228
Guillot G, Mortier F, Estoup A (2005) Geneland: a program for landscape genetics. Mol Ecol Notes 5:712–715
Gursky IG (1985) Ukraine and Moldavia. In: Bibikov DI (ed) The wolf. History, systematics, morphology, ecology. Nauka Publishers, Moskva, pp 487–493 (In Russian)
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hewitt GM (2000) The genetic legacy of the Quaternary ice ages. Nature 405:907–913
Hofreiter M, Serre D, Rohland N, Rabeder G, Nagel D, Conard N, Münzel S, Pääbo S (2004) Lack of phylogeography in European mammals before the last glaciations. PNAS 101:12963–12968
Huck M, Jędrzejewski W, Borowik T, Miłosz-Cielma M, Schmidt K, Jędrzejewska B, Nowak S, Mysłajek RW (2010) Habitat suitability, corridors and dispersal barriers for large carnivores in Poland. Acta Theriol 55:177–192
Huck M, Jędrzejewski W, Borowik T, Jędrzejewska B, Nowak S, Mysłajek RW (2011) Analyses of least cost paths for determining effects of habitat types on landscape permeability: wolves in Poland. Acta Theriol 56:91–101
Jansson E, Ruokonen M, Kojola I, Aspi J (2012) Rise and fall of wolf population: genetic diversity and structure during recovery, rapid expansion and drastic decline. Mol Ecol 21:5178–5193
Jędrzejewski W, Schmidt K, Theuerkauf J, Jędrzejewska B, Okarma H (2001) Daily movements and territory use by radio-collared wolves (Canis lupus) in Białowieża Primeval Forest in Poland. Can J Zool 79:1993–2004
Jędrzejewski W, Niedziałkowska M, Mysłajek RW, Nowak S, Jędrzejewska B (2004) Habitat variables associated with wolf (Canis lupus) distribution and abundance in northern Poland. Divers Distrib 10:225–233
Jędrzejewski W, Niedziałkowska M, Mysłajek RW, Nowak S, Jędrzejewska B (2005a) Habitat selection by wolves Canis lupus in the uplands and mountains of southern Poland. Acta Theriol 50:417–428
Jędrzejewski W, Branicki W, Veit C, Medugorac I, Pilot M, Bunevich AN, Jędrzejewska B, Schmidt K, Theuerkauf J, Okarma H, Gula R, Szymura L, Förster M (2005b) Genetic diversity and relatedness within packs in an intensely hunted population of wolves Canis lupus. Acta Theriol 50:1–22
Jędrzejewski W, Jędrzejewska B, Zawadzka B, Borowik T, Nowak S, Mysłajek RW (2008) Habitat suitability model for Polish wolves based on long-term national census. Anim Conserv 11:377–390
Jędrzejewski W, Jędrzejewska B, Andersone-Lilley Ž, Balčiauskas L, Männil P, Ozoliņš J, Sidorovich VE, Bagrade G, Kübarsepp M, Ornicāns A, Nowak S, Pupila A, Žunna A (2010) Synthesizing wolf ecology and management in eastern Europe: similarities and contrasts with North America. In: Musiani M, Boitani L, Paquet PC (eds) The world of wolves: new perspectives on ecology, behaviour and management. University of Calgary Press, Calgary, pp 207–233
Jędrzejewski W, Niedziałkowska M, Hayward MW, Goszczyński J, Jędrzejewska B, Borowik T, Bartoń KA, Nowak S, Harmuszkiewicz J, Juszczyk A, Kałamarz T, Kloch A, Koniuch J, Kotiuk K, Mysłajek RW, Nędzyńska M, Olczyk A, Teleon M, Wojtulewicz M (2012) Prey choice and diet in wolves related to differentiation of ungulate communities and corresponding wolf subpopulations in Poland. J Mamm 33:1480–1492
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Biochem Genet 10:149–163
Jombart T, Pontier D, Durfour A-B (2009) Genetic markers in the playground of multivariate analysis. Heredity 102:330–341
Leonard JA, Vila C, Fox-Dobbs K, Koch PL, Wayne RK, Van Valkenburgh B (2007) Megafaunal extinctions and the disappearance of a specialized wolf ecomorph. Curr Biol 17:1146–1150
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Muñoz-Fuentes V, Darimont CT, Wayne RK, Paquet PC, Leonard JA (2009) Ecological factors drive differentiation in wolves from British Columbia. J Biogeogr 36:1516–1531
Musiani M, Leonard JA, Cluff HD, Gates CC, Mariani S, Paquet PC, Vilà C, Wayne RK (2007) Differentiation of tundra/taiga and boreal coniferous forest wolves: genetics, coat colour and association with migratory caribou. Mol Ecol 16:4149–4170
Niedziałkowska M, Jędrzejewski W, Mysłajek RW, Nowak S, Jędrzejewska B, Schmidt K (2006) Environmental correlates of Eurasian lynx occurrence in Poland—large scale census and GIS mapping. Biol Conserv 133:63–69
Niedziałkowska M, Jędrzejewska B, Honnen A-C, Thurid O, Sidorovich VE, Perzanowski K, Skog A, Hartl GB, Borowik T, Bunevich AN, Lang J, Zachos FE (2011) Molecular biogeography of red deer Cervus elaphus from eastern Europe: insights from mitochondrial DNA sequences. Acta Theriol 56:1–12
Niedziałkowska M, Jędrzejewska B, Wójcik JM, Goodman SJ (2012) Genetic structure of red deer population in northeastern Poland in relation to the history of human interventions. J Wildl Manage 76:1264–1276
Nowak S, Mysłajek RW, Jędrzejewska B (2008) Density and demography of wolf Canis lupus population in the western-most part of the Polish Carpathian Mountains, 1996–2003. Folia Zool 57:392–402
Nowak S, Mysłajek RW, Kłosińska A, Gabryś G (2011) Diet and prey selection of wolves Canis lupus recolonising Western and Central Poland. Mamm Biol 76:709–715
Oksanen JF, Blanchet G, Kindt R, Legendre P, Minchin PR, O’Hara R et al. (2011) vegan: Community Ecology Package. R package version 2. 0–2
Ostrander EA, Sprague GF Jr, Rine J (1993) Identification and characterization of dinucleotide repeat (CA)n markers for genetic mapping in the dog. Genomics 16:207–332
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pilot M, Jędrzejewski W, Branicki W, Sidorovich VE, Jędrzejewska B, Stachura K, Funk SM (2006) Ecological factors influence population genetic structure of European grey wolves. Mol Ecol 15:4533–4553
Pilot M, Branicki W, Jędrzejewski W, Goszczyński J, Jędrzejewska B, Dykyy I, Shkvyrya M, Tsingarska E (2010) Phylogeographic history of grey wolves in Europe. BMC Evol Biol 10:104
Pritchard J, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Randi E (2008) Detecting hybridization between wild species and their domesticated relatives. Mol Ecol 17:285–293
Randi E, Lucchini V, Christensen MF, Mucci N, Funk SM, Dolf G, Loeschcke V (2000) Mitochondrial DNA variability in Italian and East European wolves: detecting the consequences of small population size and hybridization. Conserv Biol 14:464–473
Raymond M, Rousset F (1995) Genepop (version 3.1): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
R Development Core Team (2012) R: A language and environment for statistical computing. R Foundation for Statistical Computing,Vienna, Austria. ISBN 3-900051-07-0, URL http://www.R-project.org/.
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Sacks BN, Brown SK, Ernest HB (2004) Population structure of California coyotes corresponds to habitat-specific breaks and illuminate species history. Mol Ecol 13:1265–1275
Sacks BN, Mitchell BR, Williams CL, Ernest HB (2005) Coyote movements and social structure along a cryptic population genetic subdivision. Mol Ecol 14:1241–1249
Sambrook J, Russel DW (2001) Purification of nucleic acids. In: Molecular cloning: a laboratory manual, 3rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp A8.9–A8.10
Savolainen P, Rosen B, Holmberg A, Leitner T, Uhlen M, Lundeberg J (1997) Sequence analysis of domestic dog mitochondrial DNA for forensic use. J Forensic Sci 42:593–600
Schede J-U, Schumann G, Wersin-Sielaff A (2010) Wölfe in Brandenburg—eine spurensuche im märkischen Sand. Ministerium für Umwelt, Gesundheit und Verbraucherschutz des Landes Brandenburg, Potsdam
Schmidt K, Kowalczyk R, Ozolins J, Männil P, Fickel J (2009) Genetic structure of the Eurasian lynx population in north-eastern Poland and the Baltic states. Conserv Gen 10:497–501
Schmidt K, Ratkiewicz M, Konopinski M (2011) The importance of genetic variability and population differentiation in the Eurasian lynx Lynx lynx for conservation, in the context of habitat and climate change. Mamm Rev 41:112–124
Schmölcke U, Zachos FE (2005) Holocene distribution and extinction of the moose (Alces alces, Cervidae) in Central Europe. Mamm Biol 70:329–344
Shibuya H, Collins BK, Huang TH-M, Johnson GS (1994) A polymorphic (AGGAAT)n tandem repeat in an intron of the canine von Willebrand factor gene. Anim Gen 25:112
Smouse PE, Long JC, Sokal RR (1986) Multiple regression and correlation extension of the Mantel test of matrix correspondence. Syst Zool 35:627–632
Stronen AV, Forbes GJ, Paquet PC, Goulet G, Sallows T, Musiani M (2012) Dispersal in a plain landscape: short-distance genetic differentiation in southwestern Manitoba wolves, Canada. Conserv Gen 13:359–371
Taberlet P, Griffin S, Goossens B, Questiau S, Manceau V, Escaravage N, Waits LP, Bouvet J (1996) Reliable genotyping of samples with very low DNA quantities using PCR. Nucleic Acid Res 24:3189–3194
Taberlet P, Fumagalli L, Wust-Saucy AG, Cosson JF (1998) Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol 7:453–464
Thiessen CD (2007) Population structure and dispersal of wolves in the Canadian Rocky Mountains. MSc thesis, Department of Biology, University of Alberta, Edmonton, Canada
Tollefsrud MM, Kissling R, Gugerli F, Johnsen Ø, Skrøppa T, Rachid C, Cheddadi R, van der Knaap WO, Latałowa M, Terhurne-Berson R, Litt T, Geburek T, Brochman C, Sperisen C (2008) Genetic consequences of glacial survival and postglacial colonization in Norway spruce: combined analysis of mitochondrial DNA and fossil pollen. Mol Ecol 17:4134–4150
Valière N (2002) GIMLET: a computer program for analysing genetic individual identification data. Mol Ecol Notes 2:377–379
Verardi A, Lucchini V, Randi E (2006) Detecting introgressive hybridization between free-ranging domestic dogs and wild wolves (Canis lupus) by admixture linkage disequilibrium analysis. Mol Ecol 15:2845–2855
Vila C, Amorim IR, Leonard JA, Posada D, Castroviejo J, Petrucci-Fonseca F, Crandall KA, Ellegren H, Wayne RK (1999) Mitochondrial DNA phylogeography and population history of the grey wolf Canis lupus. Mol Ecol 8:2089–2103
Wabakken P, Sand H, Liberg O, Bjärvall A (2001) The recovery, distribution and population dynamics of wolves on the Scandinavian Peninsula, 1978–1998. Can J Zool 79:710–725
Wabakken P, Sand H, Kojola I, Zimmermann B, Arnemo JM, Pedersen HC, Liberg O (2007) Multistage, long-range natal dispersal by a global positioning system-collared Scandinavian wolf. J Wildl Manage 71:1631–1634
Waits LP, Luikart G, Taberlet P (2001) Estimating the probability of identity among genotypes in natural populations: cautions and guidelines. Mol Ecol 10:249–256
Wawrzyniak P, Jędrzejewski W, Jędrzejewska B, Borowik T (2010) Ungulates and their management in Poland. In: Apollonio M, Andersen R, Putman R (eds) European ungulates and their management in the 21st century. Cambridge University Press, Cambridge, pp 223–242
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wójcik JM, Kawałko A, Marková S, Searle JB, Kotlík P (2010) Phylogeographic signatures of northward post-glacial colonization from high-latitude refugia: a case study of bank voles using museum specimens. J Zool 281:249–262
Zedrosser A, Dahle B, Swenson JE, Gerstl N (2001) Status and management of the brown bear in Europe. Ursus 12:9–20
Acknowledgments
We thank the General Directorate of the State Forests and the Department of Nature Conservation of the Ministry of Environment for their cooperation during the National Wolf Census. We are grateful to numerous volunteers, students, personnel of Polish State Forests and national parks for their help in collecting wolf scats and to Barbara Marczuk for help with laboratory analyses. German samples were provided by Ilka Reinhard and Gesa Kluth. The study was financed by the Mammal Research Institute, Polish Academy of Sciences, the European Nature Heritage Fund EURONATUR (Germany), and by the grants 2 P04F 019 27 and 2 P06L 006 29 from the Ministry of Science and Higher Education, Poland. AVS was supported by the project Research Potential in Conservation and Sustainable Management of Biodiversity—BIOCONSUS in the 7th Framework Programme (contract no. 245737). CP was supported by a Marie Curie Transfer of Knowledge Fellowship (project BIORESC in the 6th FP, contract no. MTKD-CT-2005-029957). We thank the Danish Natural Science Research Council for financial support to CP (grant numbers: 11-103926, 09-065999, 95095995) and the Carlsberg Foundation (grant number 2011-01-0059). SN and RWM were supported by EURONATUR (Germany), International Fund for Animal Welfare (USA), and Wolves and Humans Foundation (Great Britain). We thank C. Vilà and three anonymous reviewers for their helpful comments on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
See Tables 4, 5, 6 and Fig. 7.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Czarnomska, S.D., Jędrzejewska, B., Borowik, T. et al. Concordant mitochondrial and microsatellite DNA structuring between Polish lowland and Carpathian Mountain wolves. Conserv Genet 14, 573–588 (2013). https://doi.org/10.1007/s10592-013-0446-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-013-0446-2