Abstract
The fungicide tolprocarb (TPC) is a melanin biosynthesis inhibitor, but it may also have another mode of action. Here in tests of TPC for inducing plant systemic acquired resistance (SAR), TPC induced promoter activity of the tobacco pathogenesis-related gene PR-1a in Arabidopsis thaliana and genes for PBZ1, β-1,3-glucanase, and chitinase 1 in the defense-related salicylic acid (SA) signaling pathway in rice, but not genes for the jasmonate signaling pathway. Probenazole (PBZ), a commercially used plant defense activator, induced genes in both signaling pathways. The antibacterial activity of TPC was equivalent to that of PBZ. Irrigation with 200 μM TPC prevented growth by Pseudomonas syringae pv. maculicola in A. thaliana, and 30 μM TPC inhibited Xanthomonas oryzae pv. oryzae growth in rice. The results of this study suggest that TPC functions not only as a melanin biosynthesis inhibitor but also as an SAR inducer and is applicable as a novel bacterial control agent that induces SAR activity in both A. thaliana and rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tolprocarb (TPC) is a fungicide that is used to control rice blast, which is caused by the fungal pathogen Magnaporthe grisea, and is recognized as a member of the group of the polyketide synthase inhibitors of melanin biosynthesis (melanin biosynthesis polyketide synthase inhibitor, MBI-P; Banba et al. 2017; Hamada et al. 2014). TPC controls M. grisea by inhibiting melanization of appressoria, a process that is essential to increase turgor pressure (8 MPa) and facilitate physical penetration of host cell walls (Chumley and Valent 1990; Howard et al. 1991). TPC also inhibits the dispersal of conidia from conidiophores, but it does not affect other fungal stages, such as conidia formation and germination, appressorial formation or mycelial growth (Araki et al. 2015). In practice, TPC is potent when applied to M. grisea, including those strains that are resistant to scytalone dehydratase (SH) inhibitors of melanin biosynthesis (melanin biosynthesis dehydratase inhibitors, MBI-Ds) and quinone outside inhibitors (Schindler et al. 2019).
Previous studies have shown that carpropamid, which is an MBI-D, was reported to activate systemic acquired resistance (SAR) in plants by inducing the production of some pathogenesis-related proteins and diterpene phytoalexins (Araki and Kurahashi 1999; Kuchii et al. 2002; Thieron et al. 1988). However, its control of infection of any plant pathogens has not been attributed to the induction of SAR activity; thus, whether the infection control offered by carpropamid is in fact due to inducing systemic immunity is still unclear.
SAR is a nonspecific systemic immunity system in plants (Horvath and Chua 1994) and induced by the signaling compound salicylic acid (SA) (Malamy et al. 1990; Mertraux et al. 1990). Two genes involved with the SA-signaling pathway, NPR1 and WRKY45, regulate various pathogenesis-related genes, such as PR-1, PR-2, PR-5, and PBZ1, which encode pathogenesis-related proteins such as glucanase and chitinase. NPR1 and WRKY45 also regulate the phytoalexin synthetic pathways (Lattanzio et al. 2006; Van Loon and Van Strien 1999). Similarly, jasmonic acid (JA) is a key compound of the JA-signaling pathway, which regulates a group of defense-related genes such as PDF1.2 and VSP1 (Tamaoki et al. 2013). Pathogenesis-related proteins induced by JA defend plants against pathogenic fungi and herbivores (Stintzi et al. 2001; Vijayan et al. 1998; Wang and Wu 2013). The SA- and JA-signaling pathways interact in a coordinated manner during plant defense responses. Pathogenesis-related genes are not regulated independently from both signaling pathways. For example, PR-1b, commonly used as a marker of defense resistance, is induced by both SA and JA treatments (Agrawel et al. 2000).
Bacterial diseases pose major challenges to crop production globally, due to the limited availability of control strategies. Although some antibiotic compounds control bacteria on crops, the emergence of drug-resistant bacterial isolates is a particular problem. However, plant defense activators can control bacterial diseases without triggering the development of resistant strains. For example, acibenzolar-S-methyl, or probenazole (PBZ), is reported to control bacterial and viral activity through the induction of pathogenesis-related genes (Hong et al 2011; Iwai et al. 2007; Midoh and Iwata 1997; Soylu et al. 2003; Takeshita et al. 2013; Yoshioka et al. 2001). Numerous substances that enhance SAR have been reported, including synthetic chemicals, natural products such as rare sugars, and microorganisms such as avirulent Pseudomonas syringae (Cameron et al. 1994; Kano et al. 2010).
In this study, we investigated the TPC-responsive expression of defense-related genes and bacterial disease resistance in Arabidopsis thaliana and Oryza sativa to determine the SAR-inducing activity of TPC.
Materials and methods
Solutions of TPC and PBZ
TPC was manufactured and supplied by Mitsui Chemicals Agro (Tokyo, Japan); PBZ was purchased from FUJIFILM Wako Pure Chemical Corporation (Osaka, Japan). For the trials, except for the gene expression analysis in rice, the compounds were dissolved in dimethyl sulfoxide (DMSO) at 25 g/l. For gene expression analyses in rice, all compounds were dissolved in acetone at 25 g/l to avoid any potential phytotoxicity associated with DMSO.
Monitoring PR-1a gene expression in A. thaliana
Trials to monitor the promoter activity of PR-1a were done as described previously (Ono et al. 2011). In brief, transgenic A. thaliana seeds harboring the tobacco PR-1a promoter-Fluc (firefly luciferase) reporter gene fragment (PR-1a::Fluc) were germinated in water in 96-well microplates. TPC or PBZ dissolved in DMSO was added to the water with the new seedlings, and the final concentrations were adjusted to 30 µM. The seedlings were incubated in a growth chamber (constant light at 20 °C). To monitor PR-1a::Fluc activity, we added d-luciferin before observing the activation of bioluminescence by the Fluc reporter. Bioluminescence images were obtained with a VIM camera (Hamamatsu Photonics, Hamamatsu, Japan) using the photon counting method and normalized to respective luminescence at 0 h (relative activity). PR-1a::Fluc activity was monitored at 0, 24, 48, 72, 96, 120, 144, 168, 192, 216, and 240 h after treatment. Means (±SE) were calculated from six replicates.
Alternatively, A. thaliana seedlings were grown on 1/2 Murashige and Skoog (MS) medium containing sucrose (10 g/l) for 3 weeks in a growth chamber (constant light at 20 °C). Adult plants were then transplanted into water in 12-well microplates and incubated for 48 h in a growth chamber. After incubation, 30 µM of TPC and PBZ were applied to the water. PR-1a::Fluc activity was observed 96 and 120 h after treatment. Means were calculated from three replicates with standard errors, and statistical differences among respective data were determined by Tukey’s test.
Control activity of TPC and PBZ against Pseudomonas syringae pv. maculicola in A. thaliana
Resistance of adult A. thaliana to Pseudomonas syringae pv. maculicola (MAFF 730010; Psm) was evaluated on 5 ml 1/2 MS agar media containing sucrose (10 g/l) in 6-well microplates. Three-week-old plants were incubated in a growth chamber (constant light at 20 °C). TPC and PBZ were dissolved in DMSO at concentrations of 25 g/l and diluted in distilled water to give final concentrations of 200 µM each. TPC or PBZ (0.5 ml/well) were applied to agar 10 days before inoculation with bacteria. Psm cells were prepared by washing the bacterial cells from a Petri dish culture with distilled water and adjusting the concentration to 1 × 105 colony forming units (CFU)/ml. Three leaves on each 31-day-old plant were infiltrated with the inoculum using a needleless syringe. The population of Psm in leaves on day 3 after inoculation was quantified using DNA-based real-time PCR and normalized against the plant DNA as described previously (Ross and Somssich 2016). Three leaves collected from a single plant were combined to form a replicate, with three replicates used to calculate the mean and standard error. Significant differences among treatments were determined using Tukey’s test.
Monitoring the expression of pathogenesis-related genes in rice
The expression of pathogenesis-related genes in rice, Oryza sativa L. cv. Nipponbare (japonica group), was analyzed by quantitative-reverse transcription (qRT)-PCR. Rice plants were grown to the three-leaf stage in Kimura B broth in growth chambers (16 h day at 26 °C, 8 h night at 22 °C). TPC and PBZ were dissolved in acetone at concentrations of 25 g/l. Acetone solutions were added to the broth to give final concentrations of 30 μM TPC and 30 or 100 μM PBZ. The third leaves of six rice seedlings were collected at 24 and 72 h after treatment, and total RNA was extracted with an RNeasy Plant Mini Kit (Qiagen, Hilden, Germany). cDNAs were synthesized immediately using PrimeScript RT reagent Kit (Takara Bio, Otsu, Japan). The qRT-PCR reactions were carried out with a LightCycler 480 (Roche, Basel, Switzerland) using TB Green Premix Ex Taq II (Takara Bio). Gene expression analyses were conducted for 10 pathogenesis-related genes, including PBZ1 (AK071613), PR1b (AK107926), POX22.3 (AK073202), β-1,3-glucanase (AK104862), chitinase 1 (AK059767), KSL7 (AK068310), CYP99A2 (AK071546), RBBI2-3 (AK064050), RCI-1 (AK072241), and NOMT (AB692949). Actin (AK06893) gene expression was used to normalize the expression levels of target genes. The qRT-PCR primers used for the genes have previously been reported (Table 1) (Cho et al. 2004; Hasegawa et al. 2014; Kano et al. 2010; Miyamoto et al. 2014a). The ΔCt (dCt) values were calculated, and the means and standard errors were obtained from six replicates; − ΔΔCt (− ddCt) values were calculated based on dCt values of acetone sections. Statistical differences among respective dCt data were determined using Tukey’s test.
Control activity of TPC and PBZ against Xanthomonas oryzae pv. oryzae in rice
Rice (Oryza sativa L. cv. Nipponbare) seedlings were grown in a greenhouse to the five-leaf stage in 45-mm-diameter plastic pots with autoclaved sterilized soil collected from Agro Research Center, Mitsui Chemicals Agro, Ibaraki, Japan. Solutions of TPC or PBZ in DMSO (25 g/l) were diluted in water to give final concentrations of 30 μM each. A volume of 2000 μl of the solution was applied to the soil for each plant, and the seedlings continued to grow in the greenhouse. The plants were inoculated with bacterial culture 10 days after the chemical treatments. Xanthomonas oryzae pv. oryzae (MAFF 210749) (Xoo) cells grown on Luria–Bertani (LB) medium were collected with distilled water and adjusted to 1 × 105 CFU/ml for inoculation. Fifth leaves of rice seedlings were inoculated using a clipping technique (Kauffman et al. 1973), then the seedlings were incubated in a growth chamber (16 h day at 25 °C, 8 h night at 20 °C). After 10 days, the length of lesions on inoculated leaf blades was measured. The means and standard errors of 16 replicates for each treatment were calculated, and significant differences among treatments were determined using Tukey’s test.
Results
Monitoring PR-1a gene expression in A. thaliana
Tobacco PR-1a promoter gene expression in A. thaliana plants treated with 30 μM TPC or PBZ was evaluated using PR-1a::Fluc activity. In seedlings, PBZ accelerated PR-1a gene expression from 120 h after treatment, whereas TPC accelerated PR-1a gene expression from 96 h after treatment. The acceleration and peak period of TPC activity were observed earlier than those of PBZ (Fig. 1a). In the second trial using adult A. thaliana plants, PR-1a::Fluc bioluminescence activity was determined at 96 and 120 h after treatment in water. The results clearly indicated that the PR-1a promoter was induced in response to treatments with TPC and PBZ (Fig. 1b, c).
Monitoring tobacco PR-1a::Fluc reporter activity in transgenic Arabidopsis thaliana plants treated with TPC. Relative activity, luminescence level at each time/luminescence level at zero time. Error bars indicate standard error. aPR-1a::Fluc reporter activity in A. thaliana seedlings. Seedlings were dipped into liquid medium containing TPC or PBZ. Relative activity was assessed at 0, 24, 48, 72, 96, 120, 144, 168, 192, 216, and 240 h after treatment. bPR-1a::Fluc reporter activity in 3-week-old A. thaliana leaves. Compounds were added to agar around the roots. Relative activity was assessed at 96 or 120 h after treatment. c Bright-field and Fluc luminescence images from 3-week-old A. thaliana plants treated with TPC. Fluc luminescence was measured using the photon counting method. Bright-field images 96 h after treatments and Fluc luminescence images 96 and 120 h after treatments with 30 µM TPC or PBZ are indicated. Significant difference between treatment and DMSO in Tukey’s test: *P < 0.05, **P < 0. DMSO dimethyl sulfoxide, TPC tolprocarb, PBZ probenazole
TPC and PBZ control of Psm in A. thaliana
When the DNA of Psm in A. thaliana was quantified after the TPC and PBZ treatments, bacterial population had been reduced compared to the DMSO section (Fig. 2a). However, due to large standard errors, the difference between the PBZ and the DMSO (control) treatments was not significant (Fig. 2).
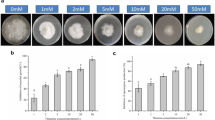
TPC control of Pseudomonas syringae pv. maculicola in Arabidopsis thaliana 3 days after inoculation. Compounds (200 µM TPC or PBZ) were added to medium 10 days before inoculation. aPsm populations in A. thaliana leaves were quantified using DNA-based qPCR and normalized against the plant DNA (− ΔCt). ΔCt was calculated by subtracting the Ct of A. thaliana DNA (At4g26410) from the Ct of Psm DNA (NC_004578.1). Error bars indicate standard error. Significant difference between treatment and DMSO in Tukey’s test: *P < 0.05. bPsm lesions in A. thaliana at 3 days after inoculation. Three replicates of three inoculated leaves are shown. DMSO dimethyl sulfoxide, TPC tolprocarb, PBZ probenazole, PsmPseudomonas syringae pv. maculicola
Monitoring pathogenesis-related gene expression in rice
After treatment with TPC or PBZ, the qRT-PCR showed that two genes, PBZ1 and β-1,3-glucanase, were significantly upregulated by 24 h after treatment with 30 μM TPC, and chitinase 1 was induced by 72 h. PBZ at 100 μM upregulated the expression of PBZ1 and chitinase 1 by 72 h after treatment. Interestingly, PBZ upregulated RCI-1 at 72 h after treatment, but the gene was not upregulated after the TPC treatment. In fact, TPC significantly suppressed the expression of RCI-1 at 24 h after treatment. TPC and PBZ each reduced the expression of KSL7, CYP99A2 and RBBI2-3 by 24 after treatment, but no significant differences were found between the TPC treatment sections and the controls.
Control of Xoo by TPC and PBZ in rice
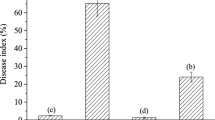
When control activity of TPC and PBZ against Xoo was estimated at 10 days after inoculation by measuring lesion lengths, lesions on TPC- and PBZ-treated leaves were significantly shorter than those on DMSO-treated rice leaves (Fig. 3). Therefore, TPC and PBZ provided significant control of Xoo.
Effects of TPC treatment with on rice bacterial blight 10 days after inoculation with Xanthomonas oryzae pv. oryzae (Xoo). a Mean (±SE) length of lesions on fifth leaves of rice seedlings (n = 16). Compounds (30 µM) were applied 10 days before inoculation. **Significant difference between treatment and DMSO in Tukey’s test: P < 0.01. b Bacterial blight on rice leaves. DMSO dimethyl sulfoxide, TPC tolprocarb, PBZ probenazole
Discussion
The commercial potential of numerous compounds that induce SAR for the control of bacterial diseases has been explored in past studies. However, few compounds have proven effective; therefore, novel control agents are required. PBZ is recognized as a plant defense activator. PBZ and the metabolite 1,2-benzisothiazol-3(2H)-one 1,1-dioxide (BIT) induce SAR in A. thaliana, rice, and tobacco (Midoh and Iwata 1997; Nakashita et al. 2002; Yoshioka et al. 2001).
In A. thaliana, TPC and PBZ similarly induced promoter activity of the tobacco PR-1a gene (Fig. 1) in adult plants. However, in new seedlings, TPC induced the activity faster than PBZ did. The reason for this is yet to be determined.
Our data revealed that TPC induces pathogenesis-related genes in rice. The SA- and JA-signaling pathways have important roles in rice defense against pathogens (Yamada et al. 2012; Yuan et al. 2007). In these defense-related pathways, SA regulates the NPR1 and WRKY45 pathways independently; NPR1 regulates pathogenesis-related proteins, whereas WRKY45 regulates the diterpene phytoalexin biosynthetic pathway (Miyamoto et al. 2014a; Nakayama et al. 2013; Shimono et al. 2007). In addition, lipoxygenase, which is regulated by JA, is also upregulated after the acceleration of the SA-signaling pathway by a plant defense activator such as PBZ (Shimono et al. 2007). PBZ acts upstream of the SA-mediated signaling pathway in rice and A. thaliana and induces pathogenesis-related genes downstream of NPR1 and WRKY45, along with JA-related genes (Nakashita et al. 2002; Yoshioka et al. 2001), although the detailed mechanisms remain unclear.
In the present study, the expression of seven of the 10 pathogenesis-related genes tested changed after treatment with the tested compounds: PBZ1, β-1,3-glucanase, chitinase 1, KSL7, and CYP99A2 related to the SA-signaling pathway and RBBI2-3 and RCI-1 related to the JA-signaling pathway (Cho et al. 2004; Hwang et al. 2008, 2016; Liu et al. 2005; Marla and Singh 2012; Miyamoto et al. 2014b; Rakwal and Komatsu 2000; Ryan 1990; Van Loon and Van Strien 1999). KSL7 and CYP99A2 are associated with the diterpene phytoalexin pathway, KSL7 participates in the phytocassane synthesis pathway, and CYP99A2 participates in the momilactone synthesis pathway (Cho et al. 2004; Miyamoto et al. 2014b). RCI-1 is a lipoxygenase gene that participates in the oxylipin biosynthesis process and is induced by plant defense activators such as PBZ and JA (Schaffrath et al. 2000).
qRT-PCR analyses of the third leaves of rice seedlings revealed that TPC induced SA-related genes such as PBZ1, β-1,3-glucanase, and chitinase 1. In particular, PBZ1 and β-1,3-glucanase were induced just 24 h after treatment (Fig. 4). However, no significant induction of JA-related genes was observed. After PBZ treatment, two SA-related genes, PBZ1 and chitinase 1, and one JA-related gene, RCI-1, were significantly induced, which is consistent with previous findings that PBZ induces both SA-related and JA-related genes (Iwai et al. 2007; Liu et al. 2005; Midoh and Iwata 1997). The results suggest that SAR activity could be a second mode of action for TPC. Further investigation is thus needed to clarify the molecular mechanisms of TPC on the activation of SA-related gene expression.
Relative expression levels of 10 pathogenesis-related genes following treatments with TPC and PBZ. The − ddCt for six replicates and their standard errors are indicated. Rice seedlings at the 3-leaf stage were cultured in Kimura B broth with each compound for 24 or 72 h. Total RNA was extracted and used for qRT-PCR analyses. The − ddCt for each gene was calculated on the basis of the value for acetone sections. Data were normalized against actin (AK06893). Accessions used: PBZ1 (AK071613), PR1b (AK107926), POX22.3 (AK073202), β-1,3-glucanase (AK104862), chitinase 1 (AK059767), KSL7 (AK068310), CYP99A2 (AK071546), RBBI2-3 (AK064050), RCI-1 (AK072241), and NOMT (AB692949). On the basis of Tukey’s test comparing treatments against acetone controls at P < 0.05, asterisk (*) indicates upregulation, pound (#) indicates downregulation. Key to compound and concentration: gray, tolprocarb 30 µM; white, probenazole 30 µM; vertical lines, probenazole 100 µM. The x-axis labels show in order the sampling period (24, 24 h after treatment; 72, 72 h after treatment), compound (T tolprocarb, P probenazole) and its concentration (0, 30 µM; 100, 100 µM), upper and lower of whiskers, highest or lowest observations; upper or bottom line of boxes, upper or lower quartiles; middle line of boxes, medians; X, averages; circles, outliers
Some genes were downregulated at 24 h after treatment with TPC or PBZ. These included RCI-1 after treatment with 30 μM TPC; KSL7, CYP99A2, and RBBI2-3 after treatment with 30 μM PBZ; and CYP99A2 after treatment with 100 μM PBZ. However, the levels of expression had reverted to control levels by 72 h after treatment. Hence, the downregulation of these genes is apparently transient.
This study demonstrates that TPC has two modes of action, i.e., MBI-P activity and the induction of SAR. Other fungicides have been reported to exhibit SAR activity as a second mode of action (Araki and Kurahashi 1999; Ishikawa et al. 2005; Skandalis et al. 2016; Thieron et al. 1988). Because of their dual actions, these fungicides, including TPC, could be important models for the production of novel fungicides.
TPC did not inhibit the growth of Psm or Xoo on LB agar at 145 μM (data not shown), suggesting that the compound has no anti-bacterial activity and that the efficacy against the pathogenic bacteria can be attributed to its SAR-inducing activity. Efficacy against bacterial pathogens is a critical factor in determining whether plant defense activators should be applied commercially. In the present study, the treatment of A. thaliana with TPC inhibited Psm considerably. Infection pressure after infiltration was severe compared to natural infection, but regardless, TPC showed significant efficacy. On the other hand, in TPC-treated rice Xoo was substantially inhibited. Xoo infected from wound of leaves, so the present study of Xoo infection was similar to natural infection. The efficacy of the control was equivalent to that after treatment with PBZ, which has been used commercially as a plant defense activator to treat rice, some Brassicaceae, and other crops.
In a previous study, the response to PBZ was related to the developmental stage of rice (Iwai et al. 2007). According to the study, more pathogenesis-related proteins accumulate in response to PBZ in older rice leaves, such as leaf eight of 8-leaf-stage rice plants, than in younger leaves, such as leaf four of a 4-leaf-stage rice seedlings. The present study demonstrated that TPC treatment suppressed bacterial blight disease caused by Xoo at the 5-leaf stage in rice. Although the efficacy of control was not tested at a younger leaf stage, the expression of pathogenesis-related genes was induced even at the 3-leaf stage. We also demonstrated that 30 μM TPC strongly induced some genes in the SA-related pathway more than 30 or 100 μM PBZ did, which suggests that TPC may also be potent in younger rice leaves.
Some plant defense activators are known to cause growth defects (Fukumoto et al. 2013; Kano et al. 2010); however, in this study, no growth defects were observed in TPC- or PBZ-treated A. thaliana and rice plants. TPC was effective in controlling disease control without causing phytotoxicity under our conditions.
In summary, TPC has commercial potential not only as a fungicide but also as a bactericide.
References
Agrawel GK, Rakwel R, Jwa N (2000) Rice (Oryza sativa L.) OsPR1b gene is phytohormonally regulated in close interaction with light signals. Biochem Biophys Res Commun 278:290–298
Araki Y, Kurahashi Y (1999) Enhancement of phytoalexin synthesis during rice blast infection of leaves by pre-treatment with carpropamid. J Pestic Sci 24:369–374
Araki N, Ezaki R, Akase T, Tomura N (2015) Studies on a novel fungicide tolprocarb (4): effects of tolprocarb against the life cycle of Magnaporthe oryzae (abstract in Japanese). Jpn J Phytopathol 81:252–253
Banba S, Hamada T, Araki N, Ebihara K (2017) Synthesis and activities of tolprocarb derivatives against Pyricularia oryzae: relationships among the activities for polyketide synthase, melanin biosynthesis, and rice blast. J Pestic Sci 42:25–31
Cameron RK, Dixon RA, Lamb CJ (1994) Biologically induced systemic acquired resistance in Arabidopsis thaliana. Plant J 5:715–725
Cho E, Okada A, Kenmoku H, Otomo K, Toyomasu T, Mitsuhashi W, Sassa T, Yajima A, Yabuta G, Mori K, Oikawa H, Toshima H, Shibuya N, Nojiri H, Omori T, Michiyama M, Yamane H (2004) Molecular cloning and characterization of a cDNA encoding ent-cassa-12,15-diene synthase, a putative diterpenoid phytoalexin biosynthetic enzyme, from suspension-cultured rice cells treated with a chitin elicitor. Plant J 37:1–8
Chumley GF, Valent B (1990) Genetic analysis of melanin-deficient, nonpathogenic mutants of Magnaporthe grisea. Mol Plant Microbe Interact 3:135–143
Fukumoto T, Kano A, Ohtani K, Inoue M, Yoshihara A, Izumori K, Tajima S, Shigematsu Y, Tanaka K, Ohkouch T, Ishida Y, Nishizawa Y, Tada Y, Ichimura K, Gomi K, Yoo SD, Sheen J, Akimitsu K (2013) Phosphorylation of d-allose by hexokinase involved in regulation of OsABF1 expression for growth inhibition in Oryza sativa L. Planta 237:1379–1391
Hamada T, Asanagi M, Satozawa T, Araki N, Banba S, Higashimura N, Akase T, Hirase K (2014) Action mechanism of the novel rice blast fungicide tolprocarb distinct from that of conventional melanin biosynthesis inhibitors. J Pestic Sci 39:152–158
Hasegawa M, Mitsuhara I, Seo S, Okada K, Yamane H, Iwai T, Ohashi Y (2014) Analysis on blast fungus-responsive characters of a flavonoid phytoalexin sakuranetin; accumulation in infected rice leaves, antifungal activity and detoxification by fungus. Molecules 19:11404–11418
Hong JC, Momol MT, Ji P, Olson SM, Colee J, Jones JB (2011) Management of bacterial wilt in tomatoes with thymol and acibenzolar-S-methyl. Crop Prot 30:1340–1345
Horvath D, Chua N (1994) The role of salicylic acid in systemic acquired resistance. Curr Opin Biotechnol 5:131–136
Howard RJ, Ferrari MA, Roach DH, Money NP (1991) Penetration of hard substrates by a fungus employing enormous turgor pressures. Proc Natl Acad Sci USA 88:11281–11284
Hwang SH, Lee IH, Yie SW, Hwang DJ (2008) Identification of an OsPR10a promoter region responsive to salicylic acid. Planta 227:1141–1150
Hwang SH, Kwon SI, Jang JY, Fuang IL, Lee H, Choi C, Park S, Ahn I, Bae SC, Hwang DJ (2016) OsWRKY51, a rice transcription factor, functions as a positive regulator in defense response against Xanthomonas oryzae pv. oryzae. Plant Cell Res 35:1975–1985
Ishikawa R, Shirouzu K, Nakashita H, Lee H, Motoyama T, Yamaguchi I, Teraoka T, Aie T (2005) Foliar spray of validamycin A or validoxylamine A controls tomato Fusarium wilt. Phytopathology 95:1209–1216
Iwai T, Seo S, Mitsuhara I, Ohashi Y (2007) Probenazole-induced accumulation of salicylic acid confers resistance to Magnaporthe grisea in adult rice plants. Plant Cell Physiol 48:915–924
Kano A, Gomi K, Yamasaki-Kokudo Y, Satoh M, Fukumoto T, Ohtani K, Tajima S, Izumori K, Tanaka K, Ishida Y, Toda Y, Nishizawa Y, Akimitsu K (2010) A rare sugar, d-allose, confers resistance to rice bacterial blight with upregulation of defense-related genes in Oryza sativa. Phytopathology 100:85–90
Kauffman HE, Reddy APK, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Rep 57:537–541
Kuchii Y, Araki Y, Sawada H (2002) Effect of carpropamid on enzymatic activities of rice plant that are related to the resistant reactions (abstract in Japanese). In: Abst 27th Ann Meet J Pesticide Sci, p 81
Lattanzio V, Lattanzino VMT, Cardinali A (2006) Role of phenolics in the resistance mechanisms of plants against fungal pathogens and insects. In: Imperato F (ed) Phytochemistry: advances in research. Research Signpost, Trivandrum, pp 23–67
Liu XQ, Bai XQ, Qian Q, Wang XJ, Chen MS, Chu CC (2005) OsWRKY03, a rice transcriptional activator that functions in defense signaling pathway upstream of OsNPR1. Cell Res 15:593–603
Malamy JE, Carr JP, Klessing DF, Raskin I (1990) Salicylic acid: a likely endogenous signal in the resistance response of tobacco to viral infection. Science 250:1002–1004
Marla SS, Singh VK (2012) LOX genes in blast fungus (Magnaporthe grisea) resistance in rice. Funct Integr Genom 12:265–275
Mertraux J, Signer H, Ryals JA, Inverardi B (1990) Increase in salicylic acid at the onset of systemic acquired resistance in cucumber. Science 250:1004–1006
Midoh N, Iwata M (1997) Expression of defense-related genes by probenazole or 1,2-benzisothiazole-3(2H)-one 1,1-dioxide. J Pestic Sci 22:45–47
Miyamoto K, Matsumoto T, Okada A, Komiyama K, Chujo T, Yoshikawa H, Nojiri H, Yamane H, Okada K (2014a) Identification of target genes of the bZIP transcription factor OsTGAP1, whose overexpression causes elicitor-induced hyperaccumulation of diterpenoid phytoalexins in rice cells. PLoS ONE 9:e105823
Miyamoto K, Shimizu T, Okada K (2014b) Transcriptional regulation of the biosynthesis of phytoalexin: a lesson from specialized metabolites in rice. Plant Biotechnol 31:377–388
Nakashita H, Yoshioka K, Yasuda M, Nitta T, Arai Y, Yoshida S, Yamaguchi I (2002) Probenazole induces systemic acquired resistance in tobacco through salicylic acid accumulation. Physiol Mol Plant Pathol 61:197–203
Nakayama A, Fukushima S, Goto S, Matsushita A, Shimono M, Sugano S, Jiang C, Akagi A, Yamazaki M, Inoue H, Takatsuji H (2013) Genome-wide identification of WRKY45-regulated genes that mediate benzothiadiazole-induced defense responses in rice. BMC Plant Biol 13:150
Ono S, Kusama M, Ogura R, Hiratsuka K (2011) Evaluation of the use of the tobacco PR-1a promoter to monitor defense gene expression by the luciferase bioluminescence reporter system. Biosci Biotechnol Biochem 75:1796–1800
Rakwal R, Komatsu S (2000) Role of jasmonate in the rice (Oryza sativa L.) self-defense mechanism using proteome analysis. Electrophoresis 21:2492–2500
Ross A, Somssich IE (2016) A DNA-based real-time PCR assay for robust growth quantification of the bacterial pathogen Pseudomonas syringae on Arabidopsis thaliana. Plant Methods 12:48. https://doi.org/10.1186/s13007-016-0149-z
Ryan CA (1990) Protease inhibitors in plants: genes for improving defenses against insects and pathogens. Annu Rev Phytopathol 28:425–449
Schaffrath U, Zabbai F, Dudler R (2000) Characterization of RCI-1, a chloroplastic rice lipoxygenase whose synthesis is induced by chemical plant resistance activators. Eur J Biochem 267:5935–5942
Schindler M, Sawada H, Tietjen K, Hamada T, Hagiwara H, Banba S (2019) Melanin synthesis in the cell wall. In: Jeschke P, Witschel M, Krämer W, Schirmer U (eds) Modern crop protection compounds, 3rd edn. Wiley, Weinheim, pp 879–909
Shimono M, Sugano S, Nakayama A, Jiang C, Ono K, Yoki S, Takatsuji H (2007) Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 19:2064–2076
Skandalis N, Dimopoulou A, Beri D, Tzima A, Malandraki I, Theologidis I, Bitivanos S, Varveri C, Klitsinaris T, Vassilakos N (2016) Effect of pyraclostrobin application on viral and bacterial diseases of tomato. Plant Dis 100:1321–1330
Soylu S, Baysal O, Soylu E (2003) Induction of disease resistance by the plant activator, acibenzolar-S-methyl (ASM), against bacterial canker (Clavibacter michiganensis subsp. michiganensis) in tomato seedlings. Plant Sci 165:1069–1075
Stintzi A, Weber H, Reymond P, Browse J, Farmer EE (2001) Plant defense in the absence of jasmonic acid: the role of cyclopentenones. Proc Natl Acad Sci USA 98:12837–12842
Takeshita M, Okuda M, Hyodo A, Hamano K, Furuya N, Tsuchiya K (2013) Induction of antiviral responses by acibenzolar-S-methyl against cucurbit chlorotic yellows virus in melon. Phytopathology 103:960–965
Tamaoki D, Seo S, Yamada S, Kano A, Miyamoto A, Shishido H, Miyoshi S, Taniguchi S, Gomi K (2013) Jasmonic acid and salicylic acid activate a common defense system in rice. Plant Signal Behav 8:e24260. https://doi.org/10.4161/psb.24260
Thieron M, Pontzen R, Kurahashi Y (1988) Carpropamid: a rice fungicide with two modes of action. Pflanzenschutz-Nachrichten Bayer 51:257–278
Van Loon LC, Van Strien EA (1999) The families of pathogenesis-related proteins, their activities, and comparative analysis of PR-1 type proteins. Physiol Mol Plant Pathol 55:85–97
Vijayan P, Shockey J, Lévesque CA, Cook RJ, Browse J (1998) A role for jasmonate in pathogen defense of Arabidopsis. Proc Natl Acad Sci USA 95:7209–7214
Wang L, Wu J (2013) The essential role of jasmonic acid in plant–herbivore interactions — using the wild tobacco Nicotiana attenuata as a model. J Genet Genom 40:597–606
Yamada S, Kano A, Tamaoki D, Miyamoto A, Shishido H, Miyoshi S, Taniguchi S, Akimitsu K, Gomi K (2012) Involvement of OsJAZ8 in jasmonate-induced resistance to bacterial blight in rice. Plant Cell Physiol 53:2060–2072
Yoshioka K, Nakashita H, Klessing DF, Yamaguchi I (2001) Probenazole induces systemic acquired resistance in Arabidopsis with a novel type of action. Plant J 25:149–157
Yuan Y, Zhong S, Li Q, Zhu Z, Lou Y, Wang L, Wang J, Wang M, Li Q, Yang D, He Z (2007) Functional analysis of rice NPR1‐like genes reveals that OsNPR1/NH1 is the rice orthologue conferring disease resistance with enhanced herbivore susceptibility. Plant Biotechnol J 5:313–324
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Hagiwara, H., Ogura, R., Fukumoto, T. et al. Novel bacterial control agent tolprocarb enhances systemic acquired resistance in Arabidopsis and rice as a second mode of action. J Gen Plant Pathol 86, 39–47 (2020). https://doi.org/10.1007/s10327-019-00891-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10327-019-00891-5