Abstract
Introduction
The contribution of CDH1 germline variants to gastric cancer burden among young adults is unknown in Brazil. We aimed to evaluate the frequency of CDH1 germline variants and the diet/lifestyle habits in early age onset gastric cancer (EOGC, ≤ 55 years old) patients.
Methodology
From 2013 to 2015, a total of 88 unrelated and consecutive patients diagnosed with EOGC were enrolled. All CDH1 exons and intronic boundaries were sequenced, and large genomic rearrangements were screened by MLPA. CDH1 transcription analysis was performed for variants that could potentially induce an effect on splicing. The diet and lifestyle habits of EOGC patients were compared to Brazilian population diet and lifestyle, obtained from governmental databases.
Results
Of 88 patients, the mean age at EOGC diagnosis was 39 years and 55% fulfilled the criteria for hereditary diffuse gastric cancer. The majority of the tumors were diffuse (74%) and poorly differentiated (80%). In total, 4 novel missense variants of uncertain significance (VUS) were identified: c.313T>A, c.387G>T, c.1676G>A, and c.1806C>A. The MLPA results revealed no rearrangements and CDH1 transcription analysis for variants of interest were inconclusive. EOGC patients had a higher red (OR:2.6, 95%CI:1.4–4.9) and processed (OR:3.1, 95%CI:1.6–6.0) meat intake and higher fruit consumption (OR:0.4, 95%IC:0.3–0.7) compared to eating habits of the Brazilian population.
Conclusions
No unequivocal pathogenic germline CDH1 variants were identified in Brazilian EOGC patients. Dietary habits may be associated with the EOGC development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gastric cancer is diagnosed in approximately 1 million people globally, is responsible for 780.000 deaths each year, and is currently the third leading cause of cancer death worldwide [1]. Its incidence shows remarkable geographical variation. More than 70% of the cases occur in the developing countries, with the highest incidence rates observed in East Asia, Latin America, Central and Eastern Europe; and the lowest incidence in Africa and Northern America [2]. Brazil is considered a middle-/high-incidence country, with 21.290 new cases expected in 2018 [3].
Although environmental and lifestyle factors—such as Helicobacter pylori infection, obesity, tobacco, alcoholic drinks, and foods preserved by salting and processed meat—are major contributors to the etiology of this disease, familial aggregation is observed in approximately 10% of gastric cancer cases which are thought to be hereditary. Overall, only 1–3% arise as a result of inherited cancer predisposition syndromes [4]. Among the hereditary forms, the most important genetic mechanism is associated with germline mutations in the CDH1 gene (E-cadherin gene type 1, epithelial cadherin, and OMIM #192,090), which encodes the protein called E-cadherin that is a transmembrane calcium-dependent cell-adhesion molecule involved in cell-junction formation and the maintenance of epithelial integrity [5]. CDH1 germline pathogenic mutations cause hereditary diffuse gastric cancer syndrome (HDGC) [6, 7].
Even though gastric cancer remains a major public health issue in South America, where countries in the region have some of the highest mortality rates worldwide, there is a lack of research focusing on risk factors influencing gastric cancer burden, specially those involving genetic inheritance. To date, only five germline variants of CDH1 were described in gastric cancer patients in South America: 4 in Brazil [8,9,10] (c.185G>T, c.1018A>G, c.1763_1764delTG, c.1023T>G), 1 in Argentina [11] (c.1913G>A), and 1 in Colombia [6] (c.2245C>T). In general, germline CDH1 mutations were identified in subjects with a strong family history of diffuse gastric cancer or lobular breast cancer. However, a combined analysis of 264 sporadic early age onset gastric cancer (EOGC) cases from low-incidence countries found that 2.3% of the subjects had a CDH1 germline pathogenic mutation [12], highlighting the importance to investigate hereditary cancer in this subpopulation.
Given the fact that inherited risk factors involved in the development of gastric cancer in Brazil are largely unexplored, we investigated the incidence and mutational spectrum of germline CDH1 variants as well as environmental and lifestyle risk factors in Brazilian early onset gastric cancer patients.
Materials and methods
Study population
Consecutive and unrelated patients diagnosed up to 55 years old with gastric carcinoma, except those with neuroendocrine carcinoma, were invited to participate in the IRB approved study at Instituto do Câncer do Estado de São Paulo-Hospital das Clínicas da Faculdade de Medicina da Universidade de São Paulo (ICESP-HCFMUSP), Brazil. All patients who agreed to participate signed an informed consent, underwent genetic counseling, and donated blood for CDH1 complete sequencing. Personal and familial histories of cancer from EOCG patients were collected through a structured questionnaire. In addition, medical records were reviewed for all participants. Using the personal and family history data, probands were categorized based on whether they met International Gastric Cancer Linkage Consortium (IGCLC) criteria [13,14,15,16].
DNA extraction from blood
DNA was extracted from 8 mL of whole blood using the Biopur Kit Mini Spin Plus (Mobius Life Science, Pinhais, PR, Brazil) and Illustra Blood GenomicPrep Mini Spin Kit (GE Healthcare Bio-Sciences, Pittsburgh, PA, USA/28-9042-64), following the instructions of the manufacturer.
Polymerase chain reaction (PCR) amplification, Sanger sequencing, and multiplex ligation-dependent probe amplification of CDH1 gene
Briefly, all exons and intron boundaries of CDH1 gene were amplified and sequenced in both forward and reverse directions. Primers and conditions are described in Supplementary Table 1. Sequences obtained were visualized by Chromas (v2.33; Technelysium Pty, Ltd., Eden Prairie, MN, USA) and by Mutation Surveyor software (v3.20, SoftGenetics LLC, State College, PA, USA). All patients’ samples were submitted to Multiplex Ligation-Dependent Probe Amplification - MLPA technique (SALSA® MLPA® P083-050R probemix; MRC-Holland, Amsterdam, The Netherlands), to investigate the presence of large rearrangements, as described in the Supplementary Materials.
CDH1 sequencing analysis and reporting criteria
All variants were named according to CDH1 sequence available at GenBank (NM_004360.4) using the nomenclature reported by the Human Genome Variation Society, HGVS (http://www.hgvs.org). The variants were searched for their classification in two publicly accessible databases: Leiden Open Variation Database (LOVD v3.0 build 13) and CLINVAR [17], freeze January 2018.
Allele frequencies of CDH1 variants
The difference of the prevalence of identified germline variants was evaluated in publicly available population datasets (ExAC—https://doi.org/10.1038/nature19057) and in 609 Brazilian controls [18].
In silico analysis
Missense variants were analyzed in the following in silico prediction models: Polymorphism Phenotyping (PolyPhen; v2.2.2) [19], Sorting Intolerant From Tolerant (SIFT; v1.0.3) [20], Align-GV/GD [21], MutationTaster2 [22], and Protein Variation Effect Analyzer (Provean; v1.1) [23]. To check for intronic and exonic variants leading to potential splicing defects, the following prediction tools were used: Human Splicing Finder [24], Neural Network (NNS, v0.9) [25], MaxEntScan (MES) [26], and NetGene2 (NG2, v2.42) [27].
Variant classification
The variants were classified according to recommendations of the American College of Medical Genetics and Genomics in: pathogenic, likely pathogenic, benign, likely benign, and variant of uncertain significance (VUS) [28].
RNA extraction from paraffin-embedded samples and characterization of the impact on splicing for CDH1 variants
Briefly, samples harboring CDH1 variants of interest were chosen for RNA extraction, further cloning, and sequencing, as shown in the Supplementary material.
Diet and lifestyle habits analysis
Diet and lifestyle information from EOCG patients was collected through a structured questionnaire. The exposure to smoking and alcohol intake was assessed through categories (never, former, and present use). The food intake of fruits, vegetables, leaves, red meat, processed meat, and salty food was assessed by intake frequency categories (less than once a week, once to twice a week, three-to-five times a week, and six-to-seven times a week). To estimate the association of these factors and gastric cancer, information from Brazilian population diet and lifestyle databases was used as a control group. This information was retrieved from the following online public databases: Instituto Brasileiro de Geografia e Estatística (IBGE) [29], Sistema de Vigilância Alimentar e Nutricional (SISVAN) [30], and the Instituto Nacional de Ciência e Tecnologia para Políticas Públicas do Álcool e outras drogas (INPAD) [31]. These governmental data were obtained through population surveys from 2008 to 2015, and is representative of the Brazilian population. These surveys provided information regarding smoking habits, alcohol consumption, and food-intake frequency through similar categories as used in our patients´ questionnaires. The associations between exposures to diet and lifestyle factors and gastric cancer were estimated using logistic regression and calculating the odds ratios (ORs) and 95% confidence intervals (CIs). For this, SPSS version 20 was used. P < 0.05 was considered as statistically significant.
Results
Population characteristics
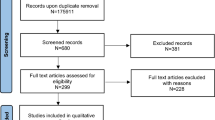
From October 2013 to August 2015, 93 consecutive and unrelated patients diagnosed with gastric cancer ≤ 55 years were enrolled. However, two patients were not successful in collecting blood and three patients were excluded, because the diagnosis changed after pathology review by a gastrointestinal pathologist at ICESP-HCFMUSP (two patients were diagnosed with neuroendocrine tumors, and in one patient, the malignancy was not confirmed in the histological study review).
The characteristics of the remaining 88 EOGC patients are shown in Table 1. The mean age at diagnosis was 39 years. Patients were born in all regions of Brazil; most of them were originally from Southeast (50%) and Northeast (38.6%) regions (Supplementary Fig. 1). There was no difference between sexes. The majority of the tumors were diffuse (74%), poorly differentiated (80%), and located in the middle and distal-third of the stomach (67%). Most patients were diagnosed with locally advanced disease (27.3%) or metastatic (39.8%) disease. More than half underwent gastrectomy (58%) and about 28% initially treated with curative intent, had tumor recurrence. The H. pylori infection was confirmed in 6 out of 32 cases (infection status was unknown in 56 cases).
The cancer family history of the study participants is described in Table 1. In total, 23% and 8% of the probands had 1st or 2nd degree relatives diagnosed with gastric and/or breast cancer, respectively. Nevertheless, the probands were not able to confirm neither inform the histologic subtype (e.g., diffuse or lobular) of those cancers diagnosed in their relatives. Because of that, none met the classical criteria postulated in 1999; however, the 2010 and 2015 IGCLC criteria were fulfilled by 55% of the patients; 47% met the criteria due to the diagnosis of diffuse gastric cancer before 40 years of age.
Complete CDH1 sequencing
Among these patients, 24 distinctive germline variants were identified (Table 2), including 5 (20.8%) missense, 6 (25%) synonymous, and 13 (54.2%) intronic variants. No large rearrangements were detected through MLPA.
Of the 24 variants, 9 were novel (c.49-61T>G, c.163+57G>A, c.163+59G>C, c.313T>A, c.387G>T, c.1676G>A, c.1806C>A, c.2164+16insA, and c.2439+56T>G) and 8 were classified as polymorphisms, because the population frequencies were greater than 1% in the ExAC databank (c.48+6C>T, c.531+10G>C, c.1896C>T, c.1937-13T>C, c.2076T>C, c.2164+16insA, c.2253C>T, and c.2634C>T).
Excluding the eight variants classified as polymorphisms, 32 patients presented CDH1 germline variants (regardless of their pathogenicity), corresponding to 36.4% of the cases (Supplementary Table 2).
In silico analyses of the missense variants
In total, five missense variants were identified; four of them were never previously reported: c.313T>A, c.387G>T, c.1676G>A, and c.1806C>A. In silico analyses of missense substitutions using five different prediction tools have shown conflicting results. All variants were considered benign by SIFT, PROVEAN, and PolyPhen-2 algorithms. On the other hand, Mutation Taster considered all the variants as potential disease causing. No variant has reached the maximum score of pathogenicity by the Align-GV/GD software, but the c.313T>A, c.387G>T e c.1676G>A achieved high scores (Table 3).
The missense mutation c.1849G>A has been previously reported. It was identified in four patients in our study: four women with diffuse gastric cancer diagnosed at 31, 35, 43, and 48 years. This variant was first described as a pathogenic somatic mutation in an endometrial [32] tumor and as a pathogenic germline mutation in a diffuse gastric cancer patient [33]. This variant is localized in the extracellular portion of E-cadherin, affecting a conserved sequence encoding one of the calcium-binding motifs. These calcium-binding motifs are functionally important, because the presence of calcium ions stabilizes the active conformation of the protein. Due to its position, it has been suggested that this mutation could lead to an unstable intercellular protein complex. In 2003, Suriano et al. identified this germline mutation in two African–American female patients diagnosed with diffuse gastric cancer at 43 years [34]. In this study, functional in vitro analysis of the c.1849G>A mutation in a cell model resulted only in minor functional changes. A recent study identified the same germline variant in 6% (10/165) of African–American patients diagnosed with ductal or mixed carcinoma of the breast [35]. This frequency was similar to the allele frequency identified in the African population in EXAC (0.04622-481/10406, with 15 appearances in homozygous; http://exac.broadinstitute.org/variant/16-68856041-G-A). In the Brazilian population controls, this mutation had allelic frequency of 0.006658. Therefore, due to the mild functional consequences observed in vitro assays and its high allele frequency, especially in the African-descendent population, this variant was classified as likely benign. This classification is in concordance with the majority of the CLINVAR submission in which 14 of the 17 submission classified this variant as benign or likely benign (https://www.ncbi.nlm.nih.gov/clinvar/variation/12232/, accessed November 2018).
In silico prediction of splice-affecting CDH1 germline variants
The results of in silico tools for the prediction of splicing defect are described in Table 4. Five variants have been previously described as benign and in silico analyses really indicated their low pathogenicity: c.324A>G, c.345G>A, c.532-18C>T, c.833-16C>G, and c.2439+10C>T. Only one novel variant showed potential to affect the splicing process by three prediction tools: c.387G>T. This missense variant is located in the last nucleotide of the exon 3, leading to an amino acid substitution (glutamine-to-histidine). This variant may cause the alteration of the donor site and has the potential to affect splicing.
CDH1 germline variants of unknown significance and patients’ characteristics
Based on allele frequency, literature/databases searches, and in silico analysis, four variants were classified as VUS according to the recommendations of the American College of Medical Genetics and Genomics [22] (Table 5). These variants were identified in patients diagnosed with diffuse gastric adenocarcinoma; however, E-cadherin immunoexpression was present in these tumor samples and these patients did not report a gastric cancer family history. These variants had never been described previously. It is noteworthy that VUS carriers were also exposed to environmental risk factors such as H. pylori infection, alcoholism, obesity, smoking, and red/processed meat consumption.
Evaluation of CDH1 mRNA from tumor samples
Nine FFPE tumor samples, presenting seven different variants, were further studied to evaluate mRNA splicing effects.
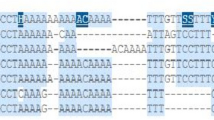
After mRNA extraction and cDNA synthesis, electrophoresis in agarose gel revealed no PCR product for six samples, probably due to mRNA degradation in the FFPE fragment (n=5) or insufficient material (n = 1). However, PCR products were detected for three tumor samples (Fig. 1): an amplicon of 178 bp, representing exons 3–4, on sample GH68 and an amplicon of 271 bp, representing exons 15–16, on sample GH80. There was an unexpected amplicon of 238 bp on sample GH12. These products were cloned for cDNA sequencing
PCR products were also submitted to electrophoresis in polyacrylamide gel: for sample GH68, besides the fragment of 178 bp, another one of approximately 300 bp was detected, and for sample GH80, besides the amplicon of 271 bp, an additional product of 100 bp was detected. Sequencing the products of samples GH12, GH68 (amplicon 300 bp) and GH80 (amplicon 100 bp) revealed only the universal primer M13 sequence. For sample GH68, sequencing of the 178 amplicon showed the wild-type exons 3–4 sequence, without the germline variant c.387G>T, which is located in a canonical splice region, indicating that only the mRNA from the normal allele was recovered (Supplementary Figure 2). In addition, sequencing of the 271 bp amplicon from sample GH80 disclosed the wild-type sequence of exons 15–16.
Electrophoresis of amplicons in agarose gel. RNA was extracted from FFPE tumor samples, and after RT-PCR, amplicons detected in agarose gel were cloned in bacteria and sequenced. NLC normal breast tissue (FFPE), NC negative control, MCF7 breast cancer cell line, GH gastric cancer FFPE samples, Primer pair 2CDH1 Exons 3–4, Primer pair 5CDH1 Exons 15–16, Primer pair 4CDH1 Exons 10–12
Diet and lifestyle habits of EOGC patients
The diet/lifestyle habits of EOGC patients were compared to Brazilian population diet and lifestyle, according to public governmental databases. EOGC patients had a higher red (OR 2.6, 95% CI 1.4–4.9) and processed (OR 3.1, 95% CI 1.6–6.0) meat intake as well as increased fruit consumption compared to eating habits of the Brazilian population (OR 0.4, 95% IC 0.3–0.7). Moreover, a trend to increased consumption of food preserved in salt was observed (OR 1.7, 95% CI 1.0–3.0; p = 0.051). However, there were no differences in tobacco and alcohol exposure in the Brazilian population and EOGC patients (Table 6).
Discussion
Gastric cancer remains one of the leading causes of cancer mortality in developing countries; however, studies evaluating the influence of hereditary factors on gastric cancer burden in these countries are scarce. In a cohort of Brazilian EOGC patients screened for CDH1 mutations, a total of 24 germline variants were identified, including nine variants never previously described in the literature. Although no definitive pathogenic mutations have been found, four novel missense VUS were detected. The analysis of the socio-environmental risk factors, such as diet and lifestyle habits, revealed that patients with EOGC reported a significantly higher consumption of processed meat and red meat. To our knowledge, the present study represents the largest series analyzing the incidence and spectrum of CDH1 germline mutations in consecutive and unrelated EOGC patients in Latin America.
HDGC is an autosomal dominant neoplastic syndrome described in 1998 [36, 37]. Male and female CDH1 mutation carriers have a 70% (95% CI 59–80%) and 56% (95% CI 44–69%) cumulative lifetime risk of developing diffuse gastric cancer, respectively. In addition, the lifetime risk of lobular breast cancer for female carriers is 42% (95% CI 23–68%) [38]. Over the past 2 decades, approximately 160 CDH1 germline variants have been published; the majority were described in probands with strong family history of cancer and from countries with a low-incidence rate of stomach cancer.
Our study population included predominantly patients diagnosed with diffuse gastric cancer under 40 years old and without family history of cancer. It is important to highlight that probands were originally from all the regions of Brazil. Approximately 50% of the participants were born in Brazilian states outside the Southeast region, with 38.6% coming from Northeast. Therefore, despite the fact that the recruitment was carried out at a single academic center in São Paulo city, the study population was not limited to the inhabitants of this part of Brazil.
Unequivocal pathogenic germline CDH1 variants were not identified in 88 EOCG patients in Brazil. In a systematic review, that compiled published series usually from regions of low incidence of gastric cancer, only 2.3% of the cases diagnosed with gastric cancer under 35 years carried pathogenic CDH1 variants [12]. In high-incidence areas, Corso et al. reported germline variants less frequently, of which 68.8% were missense mutations [39]. Therefore, an absence or a low frequency of definitely pathogenic mutations in our study was already expected, mainly because it was held in a middle/high-incidence country for gastric cancer, where exposure to external risk factors might predominate and, thereby, increase the risk of sporadic cases.
Hansford et al. have recently cataloged all CDH1 variants identified so far [38]. Among the 155 mutations described, 126 were pathogenic and 29 were VUS. Among the 126 pathogenic mutations, only 16% were missense. On the other hand, among the 29 VUS, 86% were missense. In our study, among the 24 variants identified, 33.3% (8/24) were classified as benign-polymorphisms (4 intronic and 4 synonymous), 20.8% (5/24) as benign (4 intronic and 1 synonyms), 29.2% (7/24) as probably benign (5 intronic, 1 synonymous and 1 missense), and 16.7% (4/24) as VUS (4 missense). Despite the fact that 16.7% of mutations found in CDH1 are novel and have low allele frequency, the ideal approach that can definitely assess the potential pathogenicity of these changes is still a matter of debate [7, 40,41,42].
Among the four VUS, c.387G>T presented a low allele frequency (allele 1/119896; http://exac.broadinstitute.org/variant/16-68835796-G-T) and the other 3 variants were not described in EXAC. Therefore, the classification of the pathogenicity of these variants might be possible only by studying the other families with the same variants and with family history suggestive of hereditary diffuse gastric cancer syndrome or performing functional tests. All carriers identified in our study were diagnosed with the early onset diffuse gastric adenocarcinoma. However, they did not report family history of stomach and/or breast cancer. Thus, segregation studies were not a viable approach. In addition, these variants have never been previously published or reported in CLINVAR and/or LOVD. Computational algorithms were used to predict their pathogenicity, but the results were discordant among the prediction tools. These findings highlight that in silico predictions should be used with caution, as a complementary tool, and that important clinical decisions regarding the interpretation of variants cannot be made based on the in silico outcomes alone [42, 43]. Functional impact on splicing experiments, which may help in the characterization of newly identified VUS, were performed.
The variant c.387G>T was further tested in mRNA from the patient’s tumor sample, because it is located on the exon/intron boundary (last base exon 3). The splicing prediction tool indicated that the splicing site might be lost and a probable novel splicing site might be located approximately 1460 bases inside the intron (NNsplice: available at https://omictools.com/nnsplice-tool, accessed November 2018). Our results, however, detected only the mRNA transcribed from the normal allele, because even the variant was not present in the amplicon. Based on these results, we still cannot infer the pathogenicity of the missense variant.
Our study was limited to explore the presence of germline variants only in the CDH1 gene and not in other gastric cancer predisposing genes. Although CDH1 is the most relevant gene, explaining about 40% of the cases, other genes may be involved in familial gastric cancer. Recently, new candidates have been identified including CTNNA1, BRCA1, BRCA2, STK11, PRSS1, PALB2, ATM, MSR1, SDHB, RAD51, and MAP3K6 [38, 44,45,46], but the clinical relevance of these findings still requires further validation.
Although the intestinal type of gastric cancer is associated with diet and lifestyle habits, the contribution of known modifiable risk factors to the incidence of diffuse-type gastric cancer is still under investigation. In the report of Continuous Update Project, processed meat intake and alcohol consumption above moderate levels were associated with the increased risk of gastric cancer regardless of histology subtype. In addition, there is limited evidence if the consumption of grilled fish, meat, and fruit affects the risk of developing gastric cancer [47]. In our study, in which the diffuse type was predominant, patients with EOGC reported a significantly higher consumption of red and processed meat, as well as fruits, compared to the eating habits of the Brazilian population. We did not find an association between alcohol intake and gastric cancer; however, alcohol exposure was measured in a very distinctive manner, evaluating any exposure in the last 6 months and not taking into account the amount of daily alcohol consumption (in grams per day) as usually reported [48]. Interestingly, although reports from Europe suggest that consuming little or no fruit increases the risk of gastric cancer [49], our findings showed the opposite effect and, as a consequence, require further investigation. The information about H. pylori infection is missing for the majority of our patients, but it is important to acknowledge that this well-established risk factor may also contribute to diffuse gastric cancer risk. Indeed, gastric cancer risk likely reflects a complex interaction among various diet and lifestyle habits, and H. pylori infection may function as a confounder or potential effect modifier [50].
In conclusion, unequivocal pathogenic germline CDH1 variants did not contribute significantly for EOGC predisposition in our cohort and the assessment of the potential pathogenicity of missense variants still represents a major challenge. In addition, it was observed that the nutrition habits of our patients are inadequate. For neoplasms like gastric cancer, in which the influence of external factors such as diet might increase the risk the disease, this information is relevant and warrants further investigation for the purpose of health promotion in the Brazilian population.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA. 2018;68:394–424.
Park JY, von Karsa L, Herrero R. Prevention strategies for gastric cancer: a global perspective. Clin Endosc. 2014;47:478–89.
INCA. Estimativa 2018: Incidência de Câncer no Brasil. http://www.inca.gov.br/estimativa/2018/: Instituto Nacional do Câncer—Ministério da Saúde; 2018.
Vogelaar IP, van der Post RS, Bisseling TM, van Krieken JH, Ligtenberg MJ, Hoogerbrugge N. Familial gastric cancer: detection of a hereditary cause helps to understand its etiology. Hered Cancer Clin Pract. 2012;10:18.
van Roy F, Berx G. The cell-cell adhesion molecule E-cadherin. Cell Mol Life Sci. 2008;65:3756–88.
Kaurah P, MacMillan A, Boyd N, et al. Founder and recurrent CDH1 mutations in families with hereditary diffuse gastric cancer. JAMA. 2007;297:2360–72.
Suriano G, Yew S, Ferreira P, et al. Characterization of a recurrent germ line mutation of the E-cadherin gene: implications for genetic testing and clinical management. Clin Cancer Res. 2005;11:5401–9.
Moreira-Nunes CA, Barros MB, do Nascimento Borges B, et al. Genetic screening analysis of patients with hereditary diffuse gastric cancer from northern and northeastern Brazil. Hereditary Cancer Clin Pract. 2014;12:18.
El-Husny A, Raiol-Moraes M, Amador M, et al. CDH1 mutations in gastric cancer patients from northern Brazil identified by next-generation sequencing (NGS). Gene Mol Biol. 2016;39:189–98.
Campos ECR, Ribeiro S, Higashi R. Hereditary diffuse gastric cancer: laparoscopic surgical approach associated to rare mutattion of CDH1 gene. Arquivos Brasileiros de Cirurgia Digestiva 2015.
Van Domselaar F, Correa D, Vaccaro C, et al. [Hereditary diffuse gastric cancer (HDGC): presentation of a family with a new mutation of the CDH1 gene]. Acta Gastroenterol Latinoam. 2007;37:158–63.
Corso G, Pedrazzani C, Pinheiro H, et al. E-cadherin genetic screening and clinico-pathologic characteristics of early onset gastric cancer. Eur J Cancer. 2011;47:631–9.
Caldas C, Carneiro F, Lynch HT, et al. Familial gastric cancer: overview and guidelines for management. J Med Genet. 1999;36:873–80.
Brooks-Wilson AR, Kaurah P, Suriano G, et al. Germline E-cadherin mutations in hereditary diffuse gastric cancer: assessment of 42 new families and review of genetic screening criteria. J Med Genet. 2004;41:508–17.
Fitzgerald RC, Hardwick R, Huntsman D, et al. Hereditary diffuse gastric cancer: updated consensus guidelines for clinical management and directions for future research. J Med Genet. 2010;47:436–44.
van der Post RS, Vogelaar IP, Carneiro F, et al. Hereditary diffuse gastric cancer: updated clinical guidelines with an emphasis on germline CDH1 mutation carriers. J Med Genet. 2015;52:361–74.
Landrum MJ, Lee JM, Riley GR, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014;42:5.
Brito L, Yamamoto G, Melo S, et al. Rare variants in the epithelial cadherin gene underlying the genetic etiology of nonsyndromic cleft lip with or without cleft palate. Hum Mutat. 2015;36:1029–33.
Adzhubei IA, Schmidt S, Peshkin L, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
Ng PC, Henikoff S. Predicting deleterious amino acid substitutions. Genome Res. 2001;11:863–74.
Tavtigian SV, Deffenbaugh AM, Yin L. Comprehensive statistical study of 452 BRCA1 missense substitutions with classification of eight recurrent substitutions as neutral. J Med Gene 2006.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 2014.
Choi Y, Sims GE, Murphy S, Miller JR, Chan AP. Predicting the functional effect of amino acid substitutions and indels. PloS One 2012.
Desmet FO, Hamroun D, Lalande M. Human Splicing Finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res 2009.
Reese MG, Eeckman FH, Kulp D. Improved splice site detection in Genie. J Comput Biol. 1997.
Yeo G, Burge CB. Maximum entropy modeling of short sequence motifs with applications to RNA splicing signals. J Computat Biol. 2004.
Brunak S, Engelbrecht J, Knudsen S. Prediction of human mRNA donor and acceptor sites from the DNA sequence. J Mol Biol. 1991.
Richards S, Aziz N, Bale S, Bick D, Das S. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015.
Instituto Brasileiro de Geografia e Estatística (IBGE). Pesquisa Nacional de Saúde 2013. Percepção do estado de saúde, estilo de vida e doenças crônicas. ftp://ftp.ibge.gov.br/PNS/2013/pns2013.pdf2014.
Sistema de Vigilância Alimentar Nutricional. Sisvan Web. http://dabsistemas.saude.gov.br/sistemas/sisvan/relatorios_publicos/relatorios.php.
Instituto Nacional de. Ciência e Tecnologia para Políticas Públicas do Álcool e Outras Drogas. Segundo Levantamento Nacional de Álcool e Drogas (II LENAD). Relatório 2012. 2012. http://inpad.org.br/wp-content/uploads/2014/03/Lenad-II-Relatório.pdf.
Risinger JI, Berchuck A, Kohler MF, Boyd J. Mutations of the E–cadherin gene in human gynecologic cancers. Nat Genet. 1994;7:98–102.
Ascano JJ, Frierson H Jr, Moskaluk CA, et al. Inactivation of the E-cadherin gene in sporadic diffuse-type gastric cancer. Mod Pathol. 2001;14:942–9.
Suriano G, Oliveira C, Ferreira P, et al. Identification of CDH1 germline missense mutations associated with functional inactivation of the E-cadherin protein in young gastric cancer probands. Hum Mol Genet. 2003;12:575–82.
Valente AL, Rummel S, Shriver CD, Ellsworth RE. Sequence-based detection of mutations in cadherin 1 to determine the prevalence of germline mutations in patients with invasive lobular carcinoma of the breast. Hered Cancer Clin Pract. 2014;12:17.
Guilford P, Hopkins J, Harraway J, et al. E-cadherin germline mutations in familial gastric cancer. Nature. 1998;392:402–5.
Guilford PJ, Hopkins JBW, Grady WM. E-cadherin germline mutations define an inherited cancer syndrome dominated by diffuse gastric cancer. Human Mut. 1999.
Hansford S, Kaurah P, Li-Chang H, et al. Hereditary diffuse gastric cancer syndrome: CDH1 mutations and beyond. JAMA Oncol. 2015;1:23–32.
Corso G, Marrelli D, Pascale V, Vindigni C, Roviello F. Frequency of CDH1 germline mutations in gastric carcinoma coming from high- and low-risk areas: metanalysis and systematic review of the literature. BMC Cancer. 2012;12:8.
Suriano G, Oliveira C, Ferreira P, et al. Identification of CDH1 germline missense mutations associated with functional inactivation of the E-cadherin protein in young gastric cancer probands. Human Mol Genet. 2003;12:575–82.
Suriano G, Mulholland D, de Wever O, et al. The intracellular E-cadherin germline mutation V832 M lacks the ability to mediate cell-cell adhesion and to suppress invasion. Oncogene. 2003;22:5716–9.
Melo S, Figueiredo J, Fernandes MS, et al. Predicting the functional impact of CDH1 missense mutations in hereditary diffuse gastric cancer. Int J Mol Sci. 2017;18.
Suriano G, Seixas S, Rocha J, Seruca R. A model to infer the pathogenic significance of CDH1 germline missense variants. J Mol Med (Berl). 2006;84:1023–31.
Sahasrabudhe R, Lott P, Bohorquez M, et al. Germline mutations in PALB2, BRCA1, and RAD51C, which regulate DNA recombination repair, in patients with gastric cancer. Gastroenterology. 2017;152:983–6 e6.
Kupfer SS. Gaining ground in the genetics of gastric cancer. Gastroenterology. 2017;152:926–8.
Gaston D, Hansford S, Oliveira C, et al. Germline mutations in MAP3K6 are associated with familial gastric cancer. PLoS Genet. 2014;10.
World Cancer Research Fund/American Institute for Cancer Research. Food, nutrition, physical activity, and the prevention of cancer: a global perspective. http://wcrf.org/stomach-cancer-2016.2016.
Theodoratou E, Timofeeva M, Li X, Meng X, Ioannidis JPA. Nature, nurture, and cancer risks: genetic and nutritional contributions to cancer. Annu Rev Nutr. 2017;37:293–320.
Fang X, Wei J, He X, et al. Landscape of dietary factors associated with risk of gastric cancer: a systematic review and dose-response meta-analysis of prospective cohort studies. Eur J Cancer. 2015;51:2820–32.
Puculek M, Machlowska J, Wierzbicki R, Baj J, Maciejewski R, Sitarz R. Helicobacter pylori associated factors in the development of gastric cancer with special reference to the early-onset subtype. Oncotarget. 2018;9:31146–62.
Funding
This work was supported by NAP-Biobanco/USP. MCVC is a medical student, recipient of an initiation research PIBIC/CNPq grant. PCAAB received an initiation research grant from Fundação de Amparo à Pesquisa do Estado de Sao Paulo, FAPESP grant 2012/11337-3. RC and MAAKF received a productivity grant from CNPq, agência do Ministério da Ciência, Tecnologia, Inovações e Comunicações (MCTIC). GHB was recipient of a Visiting Professor grant from Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), fundação do Ministério da Educação (MEC). SM received a pos doc grant from CAPES. DMS received a Ph.D. grant from CAPES. LS and GFLP received a Professional Development program grant from Secretaria de Saúde do Estado de São Paulo.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
All procedures were in accordance with the ethical standards of the Institutional and National Ethics Committee and with the Helsinki Declaration of 1964 and later versions.
Informed consent
Informed consent to be included in the study, or the equivalent, was obtained from all patients.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Guindalini, R.S.C., Cormedi, M.C.V., Maistro, S. et al. Frequency of CDH1 germline variants and contribution of dietary habits in early age onset gastric cancer patients in Brazil. Gastric Cancer 22, 920–931 (2019). https://doi.org/10.1007/s10120-019-00945-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10120-019-00945-9