Abstract
The recently revised Geastrum minimum species complex in sect. Geastrum subsect. Quadrifida revealed that the name G. minimum is a nomen ambiguum and dubium and was collectively used for at least two European species (viz. G. granulosum and G. marginatum). During the morphological revision of the Hungarian materials labelled as G. minimum, different crystal structures were found on the endoperidial body of some specimens than those of characteristic for G. granulosum and G. marginatum. These collections were exclusively found on open rocky grassy vegetation on dolomite bedrock in Hungary. Multigene phylogenetic analyses involving nrITS, nrLSU, rpb1, atp6 and tef1-α sequences of the collections with unique crystal morphology and ecology revealed that these form a distinct clade in close relationship with G. granulosum s.l. (i.e. specimens from Europe and North America). Based on molecular evidence, macro- and micromorphology as well as X-ray Powder Diffraction (XRD) characterisation of the mesoperidial crystals, here we propose the new species Geastrum dolomiticum.
Similar content being viewed by others
Introduction
The worldwide distributed genus Geastrum Pers. is one of the largest genera of gasteroid fungi, encompassing ca. 130 species (He et al. 2019). The taxonomy of Geastrum was intensively studied in the recent years and several new species were described from many parts of the world (e.g. Hemmes and Desjardin 2011; Zamora et al. 2015; Accioly et al. 2019; Crous et al. 2019), although most of the novel species have been found in South America, viz. Brazil’s semi-arid region or in the Amazonas region (Silva et al. 2013, Cabral et al. 2014a, b, 2017, Sousa et al. 2015, 2019, Crous et al. 2016, 2017, 2018a, b, Assis et al. 2019). In contrast to the tropical regions, the genus Geastrum is considered as well-studied in Europe (e.g. Sunhede 1989; Calonge and Zamora 2003; Zamora and Calonge 2007). Taxonomy and systematics of European earthstars (Geastrum and Myriostoma Desv.) have been reviewed by Jeppson et al. (2013), who accepted 30 Geastrum s. str. species for the old continent based on morphological observations and multigene analyses. Zamora et al. (2014) revised the section Schmidelia, and proposed Geastrum senoretiae J.C. Zamora as a new species from Spain. In a later study on phylogeny and classification of Geastrum sect. Geastrum, Zamora et al. (2015) described three additional European species, two from Spain (G. benitoi J.C. Zamora and G. meridionale J.C. Zamora) and one from the United Kingdom (G. britannicum J.C. Zamora). Therefore, Geastrum s. str. currently encompasses 34 species in Europe.
Among the European earthstars, 25 Geastrum species were known from Hungary (Central Europe), which have indicated an exceptional species richness of the genus (e.g. Jeppson 2013; Finy and Jeppson 2021). Taxonomic studies of Geastrum have a long tradition in Hungary. At the beginning of the twentieth century in his monographic book, Hollós (1903) already reported 21 Geastrum species from Hungary. Besides the valuable chorological data, Hollós (1901) also described new earthstar species from Hungary, such as G. hungaricum Hollós, which grows in steppe and dry grassland habitats, and produces an extremely small, hygrometric basidiome. The species has gained legal protection in Hungary since 2006 (Siller et al. 2006). Lately, intensive fieldwork has been taken place devoted to Hungarian earthstars from the sandy forest steppe region of the Carpathian Basin (Rimóczi et al. 2011). In 2015, a peculiar Geastrum species was found in Hungary growing exclusively on dolomite bedrock. Macroscopically it resembled to G. granulosum Fuckel, but based on microscopic and molecular phylogenetic data it differs from all known European species.
In this study we aimed to clarify the taxonomy of those Hungarian collections found on dolomite, based on macro- and micromorphological features of the basidiome, X-ray Powder Diffraction (XRD) characterisation of the mesoperidial crystals and multigene phylogenetic analyses.
Materials and methods
Morphological study
In this study, altogether nine specimens collected from autumn to spring (Table 1) were examined. Type specimens including holotype and paratypes were deposited in the herbarium of the Hungarian Natural History Museum, Budapest (BP) under the accession numbers BP111140–BP111144. All other examined specimens were deposited in the Department of Plant Anatomy, Eötvös Loránd University (abbreviated further as ELTE). Dried mature fruiting bodies were used for macro- and microscopic examination. For light microscopy, samples were mounted in water or in Lactophenol-cotton blue and heated to boiling temperature. The samples were examined with Reichert Polyvar and Olympus BH-2 microscopes. Spore dimensions were inclusive of spore wall ornamentation. Terminology mostly followed Sunhede (1989) and Zamora et al. (2015). Small pieces of peridium and gleba from dried basidiomes were prepared, fixed to stubs, coated with gold and examined under a Hitachi S2460N (Hitachi Ltd., Tokyo, Japan) scanning electron microscope at 22 kV accelerating voltage.
Molecular phylogenetic study
For molecular identification, ITS (internal transcribed spacer) region of the nrDNA, the universal fungal barcode region (Schoch et al. 2012) was amplified using the Phire® Plant Direct PCR Kit (Thermo Scientific, USA) and the primer pairs ITS1F/ITS4 (White et al. 1990; Gardes and Bruns 1993) as described in Papp and Dima (2018). For amplifying further four loci a prior total DNA extraction with the E.Z.N.A. SP Fungal DNA Mini Kit was applied. The primers LR0R (Rehner and Samuels 1994) and LR5 (Vilgalys and Hester 1990) were used to amplify the partial 28S nrRNA gene (LSU) of the nrDNA operon region. The partial RNA polymerase II largest subunit (rpb1) was amplified with RPB1-Af and RPB1-Cr (Matheny et al. 2002) and part of the mitochondrial ATPase subunit 6 (atp6) using the primers atp6-2 and atp6-3 (Kretzer and Bruns 1999). The primers EF1-983F and EF1-2218R (Rehner and Buckley 2005) were used to amplify part of the translation elongation factor 1α (tef1-α). Sequencing of the amplicons was carried out with the primers used for amplification by LGC Genomics (Berlin, Germany). The sequences were compiled from electropherograms using the Staden software package (Staden et al. 2000). Sequences of each locus were aligned separately with sequences of respective species from GenBank (Table 1) using E-INS-i method of the online MAFFT version 7 (Katoh and Standley 2013). The alignments were checked and edited in MEGA7 (Kumar et al. 2016).
Two datasets were used in the phylogenetic analyses. The ‘subsection-level’ dataset was used to gain information about the phylogenetic position of our sequences among those of Geastrum sequences representing subsect. Quadrifida and subsect. Hungarica sensu Zamora et al. (2015). The second dataset represented sequences of only G. granulosum related specimens. For the two datasets, multi-locus Bayesian analysis (BI) were performed with MrBayes 3.1.2 (Ronquist and Huelsenbeck 2003) using the GTR + G nucleotide substitution model. Four Markov chains were run for 10,000,000 generations sampled every 1,000 generations with a burn-in value set at 4,000 sampled trees. Maximum Likelihood (ML) phylogenetic analysis was carried out with the raxmlGUI v. 1.3 (Silvestro and Michalak 2012) implementation of RAxML (Stamatakis 2014). The GTR + G nucleotide substitution model was used with ML estimation of base frequencies and a ML bootstrap analysis with 1,000 replicates was used to test the support of the branches. The phylogenetic trees were visualized and edited using MEGA7 (Kumar et al. 2016) and deposited in TreeBASE (www.treebase.org) as submission 27948.
Following Zamora et al. (2015), the ITS, LSU, rpb1 and atp6 sequences were used for ‘subsection-level’ phylogenetic analyses and considered as separate partitions. These four regions of a selected subset of species were supplemented with additional two partitions (tef1-α, and indels from ITS region) to determine the phylogenetic relationships of G. granulosum related species and to improve phylogenetic resolution. The indels in the ITS region were coded (Nagy et al. 2012) using the simple indel coding algorithm (Simmons et al. 2001; Young and Healy 2003) with the program FastGap (Borchsenius 2009). In the BI analysis, the two-parameter Markov (Mk2 Lewis) model was used for the indel partition of the dataset and the GTR + G model for the nucleotide partitions. In the ML analyses, in addition to the nucleotide partitions (GTR + G), the indel data were treated as binary data (BIN).
X-Ray powder diffraction (XRD) measurement of calcium-oxalate
The measurements of the calcium-oxalate samples were carried out with Bruker D8 Advance instrument. The following parameters were configured: (i) Sample holder: Si low background sample holder (PMMA), (ii) Rotation: 30/min, (iii) Range: 5–80° (two theta), (iv) Mode: continuous scan, (v) Detector type: LYNXEYE XE (energy dispersive), (vi) X-Ray source: Cu-anode (Kα: 1.54184 Å), (vii) X-Ray optics: Bragg–Brentano, (viii) Generator power: 1600 W (40 mA, 40 kV). Samples were prepared using 1–5 mg sample that was gently homogenized in an achate mortar with a pestle to make fine powder. The grinded powder was mounted in a round sample holder (Si low background PMMA) and smooth surface was prepared by pressing it with a glass plate.
Results
Multigene phylogenetic analyses were carried out using two datasets comprising 47 strains and 3513 characters, and 18 strains and 4557 characters including gaps. According to the results, the studied specimens from Hungary belong to Geastrum sect. Geastrum subsect. Quadrifida representing different lineages (Fig. 1). The nine specimens from Hungary comprised in this study (marked with an asterisk in the Taxonomy part) were grouped together with G. granulosum specimens collected in Europe and in the USA. Four of the Hungarian specimens (FP01, FP03, FP11 and FP37) belong to the clade consist of various G. granulosum collection from Europe (Fig. 1). Five of our specimens (FP05, FP07, FP09, FP17 and FP38) represent a well-supported, relatively heterogeneous clade beside the G. granulosum lineages (Fig. 2). Within the novel lineage represented by the five samples, FP07 is grouped with FP17, and FP09 is with FP38, which two form together a sister clade with FP05 (Fig. 2). The difference in the sequences of the novel clade compared with the two groups comprising G. granulosum specimens from Europe, and G. granulosum specimens from the USA were 1.9 and 1.0% for ITS, 0.3% and 0.5% for rpb1, and 1.0% and 0.9% for atp6. Although, tef1-α sequences of G. granulosum specimens from the USA were not available, this locus showed remarkable distance between the European G. granulosum clade and the novel clade (3.5%) with relatively low intragroup heterogeneity (0.1% and 0.6%, respectively).
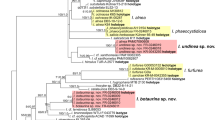
Maximum Likelihood (RAxML) tree of ITS, LSU, rpb1 and atp6 sequences of Geastrum dolomiticum and other Geastrum specimens representing subsect. Quadrifida and subsect. Hungarica sensu Zamora et al. (2015). RAxML bootstrap support values (≥ 70) are shown above branches and before slashes, Bayesian posterior probabilities (≥ 0.90) are shown below branches and after slashes. Materials collected by P. Finy (FP) in Hungary are bold. Holotypes are marked with T. Specimens from the subsect. Hungarica were served as multiple outgroup. The scale bar indicates 0.01 expected changes per site per branch
Maximum Likelihood (RAxML) tree of ITS, LSU, rpb1, atp6, tef1-α sequences and binary data from indel coding of ITS of Geastrum dolomiticum and G. granulosum related specimens in subsect. Quadrifida. RAxML bootstrap support values (≥ 70) are shown above branches and before slashes, Bayesian posterior probabilities (≥ 0.90) are shown below branches and after slashes. The studied Hungarian materials are bold. The holotype of G. dolomiticum is marked with T. Geastrum hungaricum was served as outgroup. The scale bar indicates 0.01 expected changes per site per branch
The XRD analysis of the samples prepared from the studied specimens (viz. G. granulosum and other Hungarian specimens growing on dolomite) showed characteristic peaks that can verify the presence of calcium-oxalate crystals. However, on G. granulosum we observed calcium oxalate dihydrate (COD) crystals, but on the specimens found on dolomite, calcium oxalate monohydrate (COM) crystals were found (Fig. 3).
Results of the molecular phylogenetic analyses and the calcium-oxalate investigation reinforce our hypothesis that the lineage comprising five of the specimens with unique morphological characteristics (for comparison with G. granulosum, G. marginatum and G. austrominimum, see Table 2) collected in Hungary, represent a novel species within Geastrum subsection Quadrifida.
Discussion
The new species, Geastrum dolomiticum, is mainly characterized by the small fruiting body, the lack of big COD crystals, the crested spore ornamentation and the habitat. Geastrum granulosum may also occur in calcareous rocky grasslands, but it has a wider ecological range, most common in open sandy steppe areas.
The recent integrative taxonomic study including morphological, molecular, ecological, and chorological data by Zamora et al. (2015) proposed that the collectively used Geastrum minimum Schwein. name is better to treat as nomen ambiguum and dubium since it includes at least four cryptic species from which two of them occur in Europe (i.e. G. granulosum Fuckel and G. marginatum Vittad.), furthermore the protologue of G. minimum is not enough detailed to know which species was described by Schweinitz (1822). During the taxonomic revision of all material labelled in the Hungarian National History Museum (BP) as well as in private herbaria under the name Geastrum minimum, we found that the collections deposited in BP represent G. granulosum (= G. queletii Hazsl., see Zamora et al. 2015) due to the presence of large and regular COD crystals on the basidiomes. Among our private collections we discovered specimens with different, irregular or when regular than smaller, predominantly COM crystals. These samples originated from calcareous open rocky grasslands on dolomite bedrock. Phylogenetic analyses of the nrITS, nrLSU, rpb1, atp6, and tef1-α sequences revealed that these specimens with the unique crystal morphology and habitat belong to a separate lineage in Geastrum sect. Geastrum subsect. Quadrifida which we suggest as novel species and described here as G. dolomiticum. On the other hand, based on solely the crystal morphology, there is another species in Europe (G. marginatum) having small crystals on the endoperidial body (< 70(–95) µm), however, G. dolomiticum when having regular crystals they are even smaller (< 50 µm) than those of G. marginatum. Furthermore, the stalk of G. marginatum is dark compared to G. dolomiticum which has white stalk. The former species was only verified by two collections in Hungary and grows on grassy habitats on more or less acidic and only rarely on slightly calcareous bedrocks, while the latter species prefers calcareous dolomite bedrocks.
Geastrum granulosum, the most similar species to G. dolomiticum, based on our analyses, seems to be a widespread species in Hungary occurring in dry steppe-like sandy grasslands as well as rocky habitats in the Hungarian mountain ranges on limestone and dolomite. There are examples that sandy grassland species occurs also on calcareous rocky habitats (e.g. Infundibulicybe glareosa (Röllin & Monthoux) Harmaja, Gastrosporium simplex Mattir.), and Tulostoma calcareum Jeppson, Altés, G. Moreno & E. Larss. In contrast, G. dolomiticum has so far been found exclusively on dry rocky grassland on dolomite.
Based on our phylogenetic analyses (Figs. 1, 2), a closely related North American species seems to belong in the G. granulosum–G. dolomiticum lineage too. This North American clade was collectively treated under G. granulosum in Zamora et al. (2015), according to the available phylogenetic data, it might belong to another species, viz. G. minimum s. str. However, this name is currently not in use as discussed above and in Zamora et al. (2015). The morphological delimitation of the North American lineage of G. granulosum and the clarification of the name G. minimum needs further investigations.
Taxonomic treatment
Geastrum dolomiticum Finy, Dima & V. Papp, sp. nov.—TYPE: Hungary, Fejér County, near Csór, in open grassland on dolomite, among mosses and grasses in Seseli leucospermi-Festucetum pallentis, 8 Sep 2015 P. Finy FP20150908 (holotype: BP 111140, FP05*; isotype: ELTE). [MycoBank # MB 835789]. GenBank ITS (MT569463), LSU (MT569455), rpb1 (MT572903), atp6 (MT572900), tef1-α (MT593358) (Figs. 4, 5).
Morphological characteristics of Geastrum dolomiticum (a–c) and G. granulosum (d–f). a Basidioma of G. dolomiticum (FP20150908, holotype); b basidiospores (FP20150908, holotype—BP111140); c crystalline aggregates of calcium oxalate monohydrate (COM) (FP20151227, paratype—BP111144); d basidioma of G. granulosum (FP20150214); e basidiospores (FP20141213); f crystalline aggregates of calcium oxalate dihydrate (COD) (FP20141213). Scale bars: 5 mm (a, d). Photos: K. Bóka (b, c, e, f), P. Finy and V. Papp (a, d)
Etymology: The epithet refers to the habitat requirement of the species, in open rocky grasslands on dolomite bedrock.
Description: Exoperidium 8–22 mm in diam, arched, splitting to the middle in 6–12 non-hygroscopic rays, sometimes they roll towards the endoperidial body. Fibrous layer thin, papyraceous, whitish coloured when denuded. Pseudoparenchymatous layer pale cream, ochraceous to brownish, 1–2 mm thick, covered by a dense mesoperidial crystalline matter, not persisting in old basidiomes. Mycelial layer persisting, intermixed with debris from the substrate. Endoperidial body 3–9 mm diam, greyish, greyish white, cream, brownish, more colourful on younger specimens, covered by a whitish mesoperidial crystalline matter. Peristome fibrillose, flat to broadly conical, up to 1 mm high, white, mostly well-delimited, with a whitish delimitation line. Lighter than the endoperidial body, except for the older, discoloured basidiomes. Stalk more or less stout, round, sometimes ellipsoid in cross section, 0.2–1.0 mm high, whitish to cream. Apophysis present, 0.5–1 mm high, concolorous with the endoperidial body, darker in the lower part, remains coloured. Columella cylindrical, intruding to the half or more into the gleba. Gleba chocolate brown. Basidiospores globose, (4.34)4.66–5.07(5.54) × (4.10)4.46–4.87(5.29) µm, L = 4.83, W = 4.7, Q = (1.00)1.01–1.08(1.12), Qav = 1.04, n = (100/2), with 0.2–0.5 µm high warts, ornamentation isolated or coalescing crest-like warts. Basidia not observed. Capillitial hyphae max. 3.0–5.0 µm wide, light brown, thick-walled with sparse surface debris, no lumen or very narrow. Endoperidial body composed of thick-walled up to 4 µm wide hyaline hyphae with narrow lumen. Peristomal hyphae up to 4 µm wide, light brown. Mesoperidial crystalline aggregates of COM 5–50 µm in diam, irregular shaped or flaky, sometimes mixed with bipyramidal COD which 10–50 µm in diam, covering the endoperidial surface and the pseudoparenchymatous layer (Figs. 5a, c). Pseudoparenchymatous layer thick-walled (1–2 µm thick) composed of 25–50 × 10–25 µm variously shaped elongated cells. Hyphae of the fibrous layer hyaline, thick-walled up to 5 µm wide, lumen visible. Columella hyphae thick-walled up to 5 µm wide.
Diagnosis: Exoperidium 8–22 mm in diam, endoperidial body 3–9 mm in diam, Mesoperidial crystalline aggregates of COM irregularly shaped or flaky, 5–50 µm in diam, sometimes mixed with bipyramidal COD, 10–50 µm in diam. Basidiospores globose, 4.7–5.1 × 4.5–4.9 µm, with isolated or coalescing crest-like warts. Pseudoparenchymatous layer with thick-walled (1–2 µm thick) cells.
Ecology and distribution: Geastrum dolomiticum grows in small groups in calcareous open rocky grassland on dolomite bedrock. The habitat is characterized by heliophilous vegetation e.g. grasses such as Festuca pallens Host and Stipa eriocaulis Borbás as well as cryptogams like mosses and lichens (Bölöni et al. 2011). On approximately horizontal places among the rocks, the undeveloped dolomite soils are mixed up with rubbles and powdered stones. The strong edaphical stress prevents the vegetation from closure and succession (Kun et al. 2005), the soil surface is covered mainly by mosses and lichens, where mature fruiting bodies occur. Unexpanded fruiting bodies develop deep in the moss layer. Accompanying macrofungi on the sites were Lycoperdon lividum Pers., Tulostoma brumale Pers., T. calcareum, and T. kotlabae Pouzar. These habitats are host of many endemic and relict plant species too. Geastrum dolomiticum is only known from Hungarian transdanubial localities in Bakony Mts and Vértes Mts (NW Hungary).
Additional specimens examined: HUNGARY. Fejér County, near Csákvár, Lóállás-tető, in locis graminosis, 21 Oct 1955, L. Baksay, Szujkóné (BP23235); near Csákvár, in declivo graminoso, 20 Jul 1961, J. Ujhelyi (BP31425); near Csákvár, Haraszt-hegy, in Seseli leucospermi-Festucetum pallentis, 12 Aug 2016, P. Finy FP20160812 near Csákberény, in Seseli leucospermi-Festucetum pallentis, 17 Dec 2016, P. Finy FP20161217, ibid., 18 Nov 2017, P. Finy FP20171118, ibid., 25 Nov 2017, P. Finy FP20171125; Iszkaszentgyörgy, in Seseli leucospermi-Festucetum pallentis, 16 Dec 2017, P. Finy FP20171216; Lovasberény, in calcareous pasture, 23 Feb 2014, P. Finy FP20140223 (BP111141, FP38*, paratype); Veszprém County, near Öskü, in Seseli leucospermi-Festucetum pallentis, 15 Oct 2015, P. Finy FP20151015 (BP111142, FP07*, paratype), ibid., 27 Oct 2015, P. Finy FP20151027 (BP111143, FP17*, paratype), ibid., 19 Sep 2017, P. Finy FP20170919; near Várpalota, Baglyas, in Seseli leucospermi-Festucetum pallentis, 11 Feb 2017, P. Finy FP20170211; near Várpalota, Tési-fennsík, in Seseli leucospermi-Festucetum pallentis, 27 Dec 2015, P. Finy FP20151227 (BP111144, FP09*, paratype).
Availability of data and material
Not Applicable.
Code availability
Not applicable.
References
Accioly T, Sousa JO, Moreau P-A, Lécuru C, Silva BDB, Roy M, Gardes M, Baseia IG, Martín MP (2019) Hidden fungal diversity from Neotropics: Geastrum hirsutum, G. schweinitzii (Basidiomycota, Geastrales) and their allies Hidden fungal diversity. PLoS ONE 14:e0211388. https://doi.org/10.1371/journal.pone.0211388
Assis NM, Freitas-Neto JF, Sousa JO, Barbosa FR, Baseia IG (2019) Geastrum hyalinum (Basidiomycota, Geastraceae), a new species from Brazilian Southern Amazon. Stud Fungi 4:83–89. https://doi.org/10.5943/sif/4/1/11
Bölöni J, Molnár Zs, Kun A (eds) (2011) Magyarország élőhelyei. A hazai vegetációtípusok leírása és határozója. ÁNÉR 2011. MTA ÖBKI, Vácrátót
Borchsenius F (2009) FastGap 1.2. Department of Bio-sciences, Aarhus University, Aarhus. Available at: https://www.aubot.dk/FastGap_home.htm
Cabral TS, Silva BDB, Marinho P, Baseia IG (2014a) Geastrum rusticum (Geastraceae, Basidiomycota), a new earthstar fungus in the Brazilian Atlantic rainforest—a molecular analysis. Nova Hedwigia 98:265–272. https://doi.org/10.1127/0029-5035/2013/0158
Cabral TS, Silva BDB, Ishikawa NK, Alfredo DA, Braga-Neto R, Clement CR, Baseia IG (2014b) A new species and new records of gasteroid fungi (Basidiomycota) from Central Amazonia Brasil. Phytotaxa 183:239–253. https://doi.org/10.11646/phytotaxa.183.4.3
Cabral T, Sousa JO, Silva BDB, Martín MP, Clement CR, Baseia IG (2017) A remarkable new species of Geastrum with an elongated branched stipe. Mycoscience 58:344–350. https://doi.org/10.1016/j.myc.2017.03.004
Calonge FD, Zamora JC (2003) Geastrum arenarium, encontrado en España y nuevo para Europa. Bol Soc Micol Madrid 27:59–61
Crous PW, Wingfield MJ, Burgess TI, Hardy GEStJ, Crane C, et al (2016) Fungal Planet description sheets: 469–557. Persoonia 37:218–403. https://doi.org/10.3767/003158516X694499
Crous PW, Wingfield MJ, Burgess TI, Carnegie AJ, Hardy GEStJ, et al (2017) Fungal Planet description sheets: 625–715. Persoonia 39:270–467. https://doi.org/10.3767/persoonia.2017.39.11
Crous PW, Wingfield MJ, Burgess TI, Hardy GEStJ, Gené J, et al (2018a) Fungal Planet description sheets: 716–784. Persoonia 40:239–392. https://doi.org/10.3767/persoonia.2018.40.10
Crous PW, Luangsa-ard JJ, Wingfield MJ, Carnegie AJ, Hernández-Restrepo M et al (2018b) Fungal Planet description sheets: 785–867. Persoonia 41:238–417. https://doi.org/10.3767/persoonia.2018.41.12
Crous PW, Wingfield MJ, Lombard L, Roets F, Swart WJ et al (2019) Fungal Planet description sheets: 951–1041. Persoonia 43:223–425
Finy P, Jeppson M (2021) Noteworthy European records of the xerothermic earthstar Geastrum xerophilum from the Hungarian natural inland sand dunes. Mikol Közlem, Clusiana (in press)
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes—application to the identification of mycorrhizae and rusts. Molec Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
He MQ, Zhao R-L, Hyde KD, Begerow D, Kemler M et al (2019) Notes, outline and divergence times of Basidiomycota. Fungal Divers 99:105–367
Hemmes DE, Desjardin DE (2011) Earthstars (Geastrum, Myriostoma) of the Hawaiian Islands including two new species, Geastrum litchiforme and Geastrum reticulatum. Pacific Sci 65:477–496
Hollós L (1901) Új Gasteromyceta-Fajok Magyarországból. Math Term Értes 19:504–512
Hollós L (1903) Magyarország Gasteromycetái. Franklin társulat, Budapest
Jeppson M (2013) Jordstjärnor – en bestämningsguide. – Mykologiska Publikationer 6. Sveriges Mykologiska Förening, Göteborg
Jeppson M, Nilsson RH, Larsson E (2013) European earthstars in Geastraceae (Geastrales, Phallomycetidae)—a systematic approach using morphology and molecular sequence data. Syst Biodivers 11:437–465. https://doi.org/10.1080/14772000.2013.857367
Katoh K, Standley DM (2013) MAFFT: multiple sequence alignment software version 7: improvements in performance and usability. Molec Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kretzer A, Bruns TD (1999) Use of atp6 in fungal phylogenetics: an example from the Boletales. Molec Phylogen Evol 13:483–492. https://doi.org/10.1006/mpev.1999.0680
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis Version 7.0 for bigger datasets. Molec Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kun A, Tóth T, Szabó B, Koncz J (2005) A dolomitjelenség: kőzettani, talajtani és növényzeti összefüggések (kőzet-, talaj- és növényelemzések magyarországi mészkő- és dolomit sziklagyepekben). Bot Közlem 92:1–25
Matheny PB, Liu YJJ, Ammirati JF, Hall BD (2002) Using RPB1 sequences to improve phylogenetic inference among mushrooms (Inocybe, Agaricales). Amer J Bot 89:688–698. https://doi.org/10.3732/ajb.89.4.688
Nagy LG, Kocsubé S, Csana Z, Kovács GM, Petkovits T, Vágvölgyi C, Papp T (2012) Remind the gap! Insertion–deletion data reveal neglected phylogenetic potential of the nuclear ribosomal internal transcribed spacer (ITS) of fungi. PLoS ONE 7:e49794. https://doi.org/10.1371/journal.pone.0049794
Rehner SA, Buckley E (2005) A Beauveria phylogeny inferred from nuclear ITS and EF1-α sequences: evidence for cryptic diversification and links to Cordyceps teleomorphs. Mycologia 97:84–98. https://doi.org/10.3852/mycologia.97.1.84
Rehner SA, Samuels GJ (1994) Taxonomy and phylogeny of Gliocladium analysed from nuclear large subunit ribosomal DNA sequences. Mycol Res 98:625–634. https://doi.org/10.1016/s0953-7562(09)80409-7
Rimóczi I, Jeppson M, Benedek L (2011) Characteristic and rare species of Gasteromycetes in Eupannonicum. Fungi non delineati 56–57. Edizioni Candusso, Alessio
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, and Fungal Barcoding Consortium (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA 109:6241–6245. https://doi.org/10.1073/pnas.1117018109
Schweinitz LD (1822) Synopsis fungorum Carolinae superioris secundum observations. Schriften Naturf Ges Leipzig 1:20–130
Silva BDB, Cabral TS, Marinho P, Ishikawa NK, Baseia IG (2013) Two new species of Geastrum (Geastraceae, Basidiomycota) found in Brazil. Nova Hedwigia 96:445–456. https://doi.org/10.1127/0029-5035/2013/0089
Silvestro D, Michalak I (2012) raxmlGUI: a graphical front-end for RAxML. Organisms Diversity Evol 12:335–337. https://doi.org/10.1007/s13127-011-0056-0
Simmons MP, Ochoterena H, Carr TG (2001) Incorporation, relative homoplasy, and effect of gap characters in sequencebased phylogenetic analysis. Syst Biol 50:454–462. https://doi.org/10.1080/10635150120427
Sousa JO, Baracho GS, Baseia IG (2015) Geastrum laevisporum: a new earthstar fungus with uncommon spores. Mycosphere 6:501–507. https://doi.org/10.5943/mycosphere/6/4/12
Sousa JO, Baracho GS, Martín MP, Baseia IG (2019) Contribution to Neotropical data of Geastrum section Corollina (Basidiomycota): Two new earth-stars from Caatinga vegetation, Brazil. Nova Hedwigia 109:161–175. https://doi.org/10.1127/nova_hedwigia/2019/0524
Staden R, Beal KF, Bonfield JK (2000) The Staden package, 1998. Meth Molec Biol 132:115–130. https://doi.org/10.1385/1-59259-192-2:115
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Sunhede S (1989) Geastraceae (Basidiomycotina): Morphology, ecology and systematics with special emphasis on the North European species. Syn Fungorum 1:1–534
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246. https://doi.org/10.1128/jb.172.8.4238-4246.1990
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Young ND, Healy J (2003) GapCoder automates the use of indel characters in phylogenetic analysis. BMC Bioinformatics 4:6. https://doi.org/10.1186/1471-2105-4-6
Zamora JC, Calonge FD (2007) Geastrum parvistriatum, una nueva especie encontrada en España. Bol Soc Micol Madrid 31:139–149
Zamora JC, Calonge FD, Martín MP (2014) Combining morphological and phylogenetic analyses to unravel systematics in Geastrum sect. Schmidelia. Mycologia 106:1199–1211. https://doi.org/10.3852/14-072
Zamora JC, Calonge FD, Martin MP (2015) Integrative taxonomy reveals an unexpected diversity in Geastrum section Geastrum (Geastrales, Basidiomycota). Persoonia 34:130–165. https://doi.org/10.3767/003158515X687443
Acknowledgements
We are grateful to Zsolt Sütő for his help in X-ray Powder Diffraction (XRD) characterisation of the mesoperidial crystals. We thank the anonymous reviewers for their comments on the manuscript. This research was supported by the National Research, Development and Innovation Office, Hungary (NKFIH KH-130401), the ELTE Thematic Excellence Programme 2020 supported by the National Research, Development and Innovation Office (TKP2020-IKA-05) and the János Bolyai Research Scholarship of the Hungarian Academy of Sciences (D.G. Knapp). The support of Bolyai+ New National Excellence Program of the Ministry for Innovation and Technology is highly appreciated (D.G. Knapp). We thank Csaba Locsmándi for arranging the loan of Geastrum specimens deposited in Hungarian National History Museum (BP).
Funding
Open access funding provided by Eötvös Loránd University. National Research, Development and Innovation Office, Hungary (NKFIH KH-130401), ELTE Thematic Excellence Programme 2020 supported by the National Research, Development and Innovation Office (TKP2020-IKA-05), János Bolyai Research Scholarship of the Hungarian Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
There is no conflict of interest.
Ethics approval
Not applicable.
Consent to participate
All authors approved the participation as co-authors.
Consent for publication
Not applicable.
Additional information
Handling editor: Miroslav Kolařík.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Information on Electronic Supplementary Material
Information on Electronic Supplementary Material
Online Resource 1. Specimen data of Geastrum granulosum and G. marginatum examined for comparison with G. dolomiticum.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Finy, P., Papp, V., Knapp, D.G. et al. Geastrum dolomiticum, a new earthstar species from Central Europe. Plant Syst Evol 307, 43 (2021). https://doi.org/10.1007/s00606-021-01766-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00606-021-01766-z