Abstract
Recent developments in water resource monitoring have increased the demand for the reliable identification of faecal pollution sources, also defined as microbial (faecal) source tracking (MST). Standardized faecal indicator bacteria (SFIB) enumeration does not directly support MST, as SFIB occur in animal and human sources. The aim of this study was to rigorously evaluate the applicability of host-associated faecal genetic MST markers detected by quantitative PCR (qPCR) at representative Austrian water resources (ground-, surface-, raw and treated wastewater, n = 196 samples) with high importance for the water management sector. Groundwater covered a gradient of non- (i.e., deep wells) to surface influenced resources (i.e., karst and shallow wells). In addition, single faecal excreta from humans as well as representative livestock and wildlife species were collected to evaluate the faecal source-specificity and -sensitivity of the MST assays. Genetic MST marker resistance against UV irradiation was evaluated in on-site ground and wastewater treatment installations. Bacteroides-based human- (HF183II, BacHum), ruminant- (BacR), and pig-associated (Pig2Bac) MST marker qPCR quantification was performed in concert with cultivation of E. coli, intestinal enterococci, and Clostridium perfringens (SFIB diagnostics). The selected MST makers revealed high faecal source identification capacity for the Austrian water compartments and quantitatively reflected the selected faecal pollution gradient. The study also demonstrated that SFIB data can efficiently be combined with MST data to solve previously unanswered questions in water safety monitoring and management (e.g., support pollution source-targeted catchment protection, hazard assessment, and health risk management). Further research and development needs are discussed to exploit the full power of MST technology. In conclusion, this study illustrates the capacity of molecular faecal pollution diagnostics to revolutionize water quality testing in the decades to come.
Zusammenfassung
Steigende Ansprüche im Management der mikrobiologisch-hygienischen Wasserqualität verlangen zunehmend nach Methoden, die die Bestimmung des Ausmaßes fäkaler Einträge sowie der korrespondierenden Verschmutzungsquellen, auch als mikrobielles Source-Tracking (MST) bezeichnet, ermöglichen. Seit mehr als 130 Jahren kommt dem Nachweis fäkaler Verschmutzungen durch Fäkalindikatorbakterien mit standardisierten Methoden (SFIB) eine fundamentale Rolle zu. Die Diagnostik mittels SFIB ermöglicht jedoch keine unmittelbaren MST Anwendungen, da sie sowohl in tierischen als auch menschlichen Quellen vorkommen. Ziel dieser Studie war es, die Anwendbarkeit von Verursacher-assoziierten genetischen Fäkalmarkern (MST-Marker) an repräsentativen österreichischen Wasserressourcen zu evaluieren. Hierfür wurden 196 Grund‑, Oberflächen‑, Roh- und gereinigte Abwasserproben mittels qPCR, basierend auf dem molekularbiologischen Nachweis von hoch abundanten Darmbakterien (Bacteroidetes) mit den MST Markern HF183II und BacHum (Mensch-assoziiert), BacR (Wiederkäuer-assoziiert) und Pig2Bac (Schwein-assoziiert), sowie mittels den kultivierungsbasierten Standardmethoden auf SFIB (E. coli, intestinalen Enterokokken, Clostridium perfringens) analysiert. Für die Bewertung spezieller MST-Leistungskriterien der ausgewählten qPCR Methodik (fäkale Spezifität/Sensitivität), wurden zusätzlich fäkale Ausscheidungen von Menschen und repräsentativen Vieh- und Wildtierbeständen in der Region gesammelt und analysiert. Darüber hinaus wurde die Resistenz von MST Markern gegenüber UV-Licht in der Trinkwasserdesinfektion, sowie der Abwasserreinigung erhoben. Die ausgewählten molekularen Nachweissysteme zeigten für die österreichischen Wasserressourcen eine hohe Zuverlässigkeit und Leistungsfähigkeit zur Identifizierung fäkaler Quellen und spiegelten eindrucksvoll die ausgewählten fäkalen Verschmutzungsgradienten wider. Darüber hinaus zeigte die Studie, dass SFIB- sehr effizient mit MST-Daten kombiniert werden können, um bisher unbeantwortbare Fragestellungen zu lösen, wie z. B. die Identifizierung und zielgerichtete Elimination von SFIB-Quellen im Einzugsgebiet oder eine Gefährdungsidentifizierung und Abschätzung fäkaler Verschmutzungsrisiken. Die Studie demonstriert das Potenzial molekularbiologischer Methoden das Fachgebiet zu revolutionieren. Der weitere FE Bedarf wird diskutiert, um die Leistungsfähigkeit von MST-Technologien in den kommenden Jahren voll auszuschöpfen.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
1.1 Faecal pollution of water and its significance for human health
Impairment of health-related microbiological water quality is a critical issue since it can cause severe outbreaks of infection or contribute to the background rate of endemic disease (WHO 2017). Faecal material frequently contains significant numbers of intestinal pathogens and is thus of paramount importance as a contaminating agent of water (WHO 2017). Detection of all relevant and potentially occurring intestinal pathogens is not possible for routine monitoring and routine detection is generally restricted to a few selected pathogens for microbial risk assessment (WHO 2017). The determination of faecal pollution in water thus has a longstanding tradition of different monitoring applications, and a multitude of different guidelines, standards or directives have been developed for different purposes (Tallon et al. 2005; WHO 2017). Practical determination of faecal pollution still relies on the selective growth of standard faecal indicator bacteria (SFIB), including Escherichia coli and intestinal enterococci (WHO 2017). Without a doubt, water quality testing based on this principle has contributed to a fundamental improvement in water safety management during the last century and is currently still a crucial part of management (Tallon et al. 2005). Novel molecular biological diagnostics, developed during the last two decades, promise to add significant new perspectives.
1.2 Cultivation-based SFIB analysis and the need for extended possibilities
SFIB detection indicates the level of general faecal pollution of water, which can originate from human, livestock or wildlife populations (Farnleitner et al. 2010; Frick et al. 2018), as these microbes occur in high numbers in their faecal excreta. Under certain situations, relevant numbers of SFIBs may also originate from non-enteric systems, such as algae (Power et al. 2005) or sediments (Ishii et al. 2007). However, recent developments in water resource monitoring have increasingly demanded the reliable identification and differentiation of the major contributing sources of faecal pollution, also defined as microbial (faecal) source tracking (MST). For example, in the case of faecal pollution of a recreational environment, such as a lake area dedicated to swimming activities, it is crucial to know the pollution source, to specifically address the problem. In general, MST technologies are needed i) to guide efficient and cost-effective management strategies to minimise faecal contamination in watersheds (Santo-Domingo et al. 2005), ii) to evaluate the effectiveness of best management practices, iii) to support hazard assessment and health risk management (Demeter et al. 2021; Schijven et al. 2015), and iv) to enable the comparison of microbiological diagnostics when different pollution sources are to be considered (e.g., for wastewater-based epidemiology). Hence, there is a growing need for MST technologies to complement standardized faecal pollution monitoring to fulfil future requirements of the water management sector.
1.3 Advent of “intelligent” molecular faecal pollution diagnostics
In recent years, culture-independent investigations of intestinal microbiota have shown that only a very small part of the bacterial diversity can be detected or analysed by standard cultivation procedures. In fact, SFIB, such as E. coli, constitute a minuscule subpopulation (0.1%) of the total faecal bacterial community in the intestine, and the whole diversity of faecal bacteria can only be reasonably covered by molecular-biological detection methods, without being dependent on cultivation (Wuertz et al. 2011). Many of the new insights into population structure were provided by cloning and sequencing 16S rRNA genes from DNA from faecal bacteria (Turnbaugh et al. 2007; Wuertz et al. 2011). For example, studies have shown that the human intestinal microbiota is dominated by the anaerobic phyla Firmicutes and Bacteroidetes (Wuertz et al. 2011). Abundant members of intestinal microbial communities—such as Bifidobacteria or Bacteroides—have been proposed as alternative water quality indicators for many years (Evison and James 1975). However, difficulties in anaerobic cultivation techniques and low environmental persistence limit their use (Mara and Oragui 1983). In the course of these studies, it became also evident that some bacterial populations are associated with certain hosts, such as humans (Resnick and Levin 1981). This insight led to the “first generation” of microbial source tracking (MST) methods based on selective cultivation of source- or host-associated bacterial lineages (Mara and Oragui 1983). The advent of polymerase chain reaction (PCR)-based molecular biological techniques has enabled the investigation and direct environmental detection of abundant intestinal bacterial populations (Kreader 1995). In the field of PCR analysis, this has initiated a surge in the development of “second generation” methods, such as the detection of genetic markers for general and host-associated faecal pollution by quantitative PCR (qPCR) (Reischer et al. 2007, 2006; Wuertz et al. 2011).
1.4 A toolbox for future water safety monitoring
Although there is still the capacity to further improve MST technology, it has already become evident that molecular faecal pollution diagnostics based on genetic MST marker detection is about to revolutionise water quality testing (Boehm et al. 2013; Mayer et al. 2018; USEPA 2019; Zhang et al. 2019). Several case studies and applications have successfully demonstrated the power of qPCR-based genetic faecal MST marker enumeration in recent years (e.g., Ballesté et al. 2020; Nshimyimana et al. 2017; Staley and Edge, 2016). Host-associated MST markers and qPCR detection systems have been developed for important faecal pollution sources, including human and communal faecal pollution, along with livestock and wildlife contamination sources. However, there is still a need for the development of new MST markers with improved diagnostic performance characteristics to widen the identification spectrum of faecal pollution sources (Reischer et al. 2013). To date, most applications rely on bacterial MST; however, other assays, including viral and mitochondrial targets, are increasingly used (Farkas et al. 2020; Malla and Haramoto 2020). To fully exploit the pioneering capacities of molecular diagnostics in the future, the best choice of available and characterised methods must be offered as a “toolbox approach” for the water management sector. The applications of genetic faecal marker diagnostics have focused on lakes and rivers (Ballesté et al. 2020; Frick et al. 2020), recreational waters (Brown et al. 2017; Wyer et al. 2010), shellfish production (Symonds et al. 2017), and on maximum daily load and communal wastewater characterisation (Mayer et al. 2018, 2016). MST is also increasingly used for microbial hazard and risk analysis/modelling to support decision making in water management (Cao et al. 2018; Demeter et al. 2021; Schijven et al. 2015). Except for the characterisation of karst springs (Diston et al. 2018; Reischer et al. 2011), MST has hardly been used for the analysis of groundwater resources.
1.5 Evaluating genetic MST technologies for Austrian water resources
The aim of this study was to evaluate the applicability of host-associated faecal genetic markers by qPCR enumeration for selected water resources, reflecting the diversity and relevance of the Austrian water management sector. The basic water sample set included raw and conventionally treated communal wastewater, various surface water locations, and a broad set of porous and karstic ground water covering a gradient of non- (i.e., deep wells) to surface influenced resources (i.e., karst and shallow wells). In addition, single faecal excreta from humans as well as representative livestock and wildlife species were collected to evaluate genetic MST marker faecal source-specificity and -sensitivity. MST marker resistance against ultraviolet light was pre-evaluated at on-site UV irradiation installations for groundwater (water supply) and treated wastewater (recreational water resource protection). Bacterial human-, ruminant-, and pig-associated faecal genetic marker qPCR quantification was evaluated in concert with E. coli, intestinal enterococci, and Clostridium perfringens cultivation-based SFIB enumeration (see Fig. 1). Based on the evaluated methods, two case studies are presented in addition to demonstrate the complementation of SFIB data. The results are of equal relevance for the disciplines of water supply, irrigation, recreation and wastewater management as well as water pollution control.
Visualization of the two pathways used for the detection of faecal pollution. Left path: Cultivation based standardized methods for enumeration of faecal indicator bacteria (SFIB). Sampling and filtration is followed by incubation on selective media for a defined time to enable colony formation (propagable bacteria). Visual colony counting gives the estimated faecal indicator bacteria concentration in the water sample. These methods do not allow for sample storage, as bacteria concentrations can alter due to proliferation and die-off processes. Right path: DNA-quantification methods for enumeration of genetic faecal markers. Sampling and filtration is followed by a DNA extraction step. Quantitative PCR assays (qPCR) enable the enumeration of host-associated and general genetic MST marker. DNA extracts can be stored for long term at −80 °C for later examination
2 Materials and methods
2.1 Study design and sample set up
A total of 196 water samples from twenty-three different sampling sites in the region of eastern Austria were collected. The basic sample set (n = 181) included groundwater (shallow and deep groundwater), water from karstic springs, surface water from rivers and treated/untreated wastewater from wastewater treatment plants. The two investigated WWTPs (50,000 and 250,000 population equivalents) included mechanical and biological (activated sludge) treatment without further disinfection. Sampling was performed on a monthly basis from June 2018 to March 2019, with single samples per site and date. Three additional samples from two groundwater wells before on-site UV irradiation (Type Wedeco B 300; Visades T2500L-300) were taken. The sample set was further expanded with fifteen samples from a wastewater treatment plant (50,000 population equivalents) equipped with an UV irradiation unit (Type UV-3000 plus, Trojan Technologies, Canada). Samples of the effluent wastewater before and after UV irradiation were collected monthly from May to December 2015. The additional sample set was only used for genetic marker UV irradiation resistance evaluation. Detailed information concerning sampling sites, their grouping, as well as the number of specific samples per site is provided in the additional material, Table 2. Faecal samples were collected from January to April 2018 at different farms, households, and the surrounding area of Krems an der Donau (eastern Austria). The final data set included a total of 75 faecal samples from individual specimens of livestock (n = 51), humans (n = 10), pets (n = 12) and wildlife (n = 2). All human faecal samples were anonymously donated by an equal number of healthy men and women of varying age (18 to 52 years) having mostly a mixed diet. A detailed list of individual specimens of source groups is given in additional material, Table 1.
2.2 Field sampling
Surface water and wastewater samples were collected in sterile 1 l sampling bottles (Nalgene, United Kingdom), and groundwater samples were collected in sterile 4 l sampling bottles (Nalgene, United Kingdom). All samples were stored in the dark in cooling boxes at 4 °C during transport and were processed within 6 h after collection. For the DNA extraction from water samples, suspended biomass of a specified volume of 10 ml for influent wastewater, 50 ml for effluent wastewater, 100–700 ml for surface water, 2–4 l for groundwater and 3–4 l for spring water was filtered through polycarbonate membrane filters (0.2 µm Millipore, Isopore Membrane Filter–GTTP) directly after sampling. For each filtration day a blank value was carried out as a process control. All filters were frozen immediately after filtration and stored at −80 °C until nucleic acid extraction. All faecal samples were collected in sterile 50 ml stool-sampling vials (Greiner, Austria) and transported in the dark on ice to the laboratory. Samples were stored at −20 °C within 6 h after sampling, and DNA extraction was performed within two months after sampling.
2.3 Supporting chemical and physical parameters
Measurement of turbidity (NTU, nephelometric turbidity) was performed using a nephelometer (Turbidimeter, Turb 430 IR, WTW, Germany) following ISO 7027 (measurement angle 90°, wavelength 860 nm) (ISO 2019). Determination of conductivity (µS cm−1) followed ISO 7888 using the Cond 315i/SET-TetraCon 325 device (WTW, Germany) (ISO 1985). The temperature was measured with a Testo 110 device (Testo, Austria) according to DIN 38404‑4 (DIN 1976), pH was measured following ISO 10523 (ISO 2008) and total organic carbon (TOC, mg l−1) following EN 1484 (DIN 2019). Temperature, conductivity and pH were also measured directly in the field during sampling and/or prior to filtration using the Multi3630 IDS SET F device (WTW, Germany).
2.4 Standard faecal indicator bacteria and total cell count
Analysis of standard faecal indicator bacteria (SFIB), including Escherichia coli, intestinal enterococci and presumptive Clostridium perfringens spores, was performed in volumes of 100 ml for surface and wastewater samples and 1000 ml for groundwater samples. Karstic spring water samples were analysed for E. coli using Colilert-18 (IDEXX, Germany) according to the manufacturer’s instructions (http://www.idex.com.water). All other samples were investigated for the presence of E. coli according to ISO 16649‑2 (ISO 2018) on TBX agar. Enterococci were enumerated on Slanetz Bartley agar in accordance with ISO 7899‑2 (ISO 2000). Presumptive Clostridium perfringens spores were investigated following ISO 14189 (ISO 2013) on TSC agar and colony confirmation was performed by an acid phosphatase test. Following the ISO guidelines, concentrations of faecal indicator bacteria were expressed as colony forming units (CFUs) per 100 ml. Total cell counts (TCCs) for groundwater and surface water samples were determined via flow cytometric analysis with an Attune NxT flow cytometer (Thermo Fisher Scientific) after staining with SYBR Green (Schrammel et al. 2018) and by epifluorescence microscopy following a previously described protocol (van Driezum et al. 2018).
2.5 Sample processing and DNA extraction for molecular biological analysis
DNA extraction from water samples was performed using bead-beating and phenol/chloroform as described previously (Griffiths et al. 2000; Mayer et al. 2018; Reischer et al. 2006). In brief, cell lysis was achieved by the addition of CTAB buffer and glass beads in a FastPrep 24 benchtop homogenizer (MP Biomedicals Inc., Irvine, CA) at a speed setting of 6 m s−1 for 30 s. Polycarbonate filters were dissolved completely, and DNA was subsequently purified. Precipitation of the DNA was achieved by the addition of isopropanol, followed by a washing step with ethanol and subsequent elution of the dried DNA in 10 mmol l−1 TRIS buffer (pH 8.0). DNA extraction from faecal samples was performed using the Ultra Clean Soil DNA kit (MolBio Laboratories, Carlsbad, CA) according to the manufacturer’s instructions. Per sample, 0.25 g of faeces (fresh weight) was used. For each extraction batch, an extraction control using only reagents was prepared. All DNA extracts were stored at −80 °C until qPCR analysis.
2.6 Quantitative PCR measurement and quality control
All samples were analysed by microbial source tracking (MST) assays for quantification of human-associated genetic faecal markers: i) HF183II, which is a modified version of a TaqMan HF183 qPCR assay, namely, HF183/BacR287 (Green et al. 2014), and ii) BacHum (Kildare et al. 2007); samples were also screened for quantification of iii) the ruminant-associated genetic faecal marker BacR (Reischer et al. 2006), iv) the pig-associated faecal marker Pig2Bac (Mieszkin et al. 2009) and v) a general Bacteroidetes marker AllBac (Layton et al. 2006). All qPCR reactions were performed in duplicate in a 15 µL volume on a Rotor-Gene Q thermocycler (Qiagen Inc.). The reaction mixture for the AllBac duplex assay with the ntb2 fragment as an internal amplification control (IAC, non-competitive) was composed of 7.5 µL Rotor-Gene Multiplex PCR master mix (Qiagen Inc.), 2.5 µL sample DNA dilution, 600 nM AllBac296f forward primer, 600 nM AllBac412r reverse primer, 25 nM AllBac375Bhqr TaqMan MGB probe, 500 nM ntb2‑f forward primer, 500 nM ntb2‑r reverse primer, 200 nM ntb2‑p probe and 400 ng μL−1 bovine serum albumin. For the HF183II (HF183/BacR287), BacHum, BacR and Pig2Bac assays, the respective reaction mixture was composed of 7.5 µL Rotor-Gene Multiplex PCR mastermix (Qiagen Inc.), 2.5 µL sample DNA dilution and 400 ng μL−1 bovine serum albumin, while the originally published primer and probe concentrations were maintained. The IAC template (plasmid containing the ntb2 gene fragment) was spiked at a concentration of 103 copies per reaction. The cycling parameters were 3 min or 5 min at 95 °C for denaturation and 45 cycles of 30 s or 15 s at 95 °C followed by 45 s or 60 s at 60 °C for the AllBac or all other assays, respectively. Quality assessment of qPCR data was performed as previously described (Mayer et al. 2018; Reischer et al. 2006, 2011). In brief, the AllBac and IAC duplex assay was run first in at least two 4‑fold DNA dilution steps to assess the general ability to amplify extracted DNA and to monitor qPCR amplification inhibition (Anderson et al. 2011). The selection of ideal dilution for further measurements was performed according to a procedure described in (Mayer et al. 2018). All controls, including no-template controls and filtration and DNA extraction blanks, were consistently negative. For each run, qPCR standard dilutions ranging from 101 to 106 targets per reaction were used in a linear regression model for calculation of the qPCR calibration curve with reaction efficiencies ranging from 95–105%. All qPCR data are expressed as log10 (x + 1), where x is the concentration calculated from the standard curve before applying the logarithm to it. Quantitative results of the marker concentrations in the water and faecal samples are given as marker equivalents (ME) 100 ml−1 or marker equivalents (ME) g−1, since extraction efficiency was not explicitly evaluated (Reischer et al. 2006).
2.7 Data analysis
Data analysis was performed using R version 4.0.3 in RStudio Version 1.3 (RCoreTeam 2013) supported by Microsoft Excel. Grouping of groundwater sampling sites of the principal set of water samples was based on similar AllBac concentration values and the well depth. The data set from WWTP 7 (n = 15) was only used for analysis of the influence of UV irradiation on marker concentrations and was not included for other analyses. Correlation and corresponding significance between parameters (zero values excluded) were calculated in R using functions cor, cor.mtest and p.adjust, with the following parameter settings: cor(data, use = “pairwise.complete.obs“, method = “spearman”); cor.mtest(data, conf.level = 0.95, method = “spearman”); p.adjust(pvalue, method = “bonferroni”). All plots were designed using package ggplot2. Descriptive statistics: i) maximum (max), ii) upper quartile (Q75), iii) median, iv) lower quartile (Q25) and v) minimum (min) were also calculated in R using corresponding functions. Formulas for calculating the binary assay source-specificity and -sensitivity are given in Table 1, additional materials.
3 Results
3.1 Good faecal specificity/sensitivity of selected MST markers for local pollution sources
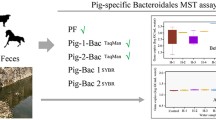
Testing the faecal discrimination capacity of the selected genetic MST marker with individual faecal excreta from target and non-target sources for the selected Austrian locations demonstrated high performance characteristics (Fig. 2). The highest discrimination performance was found for the animal MST markers covering the ruminant (BacR) and pig (Pig2Bac) faecal sources, showing median target concentrations of log10 9.1 ME (marker equivalents + 1) and log10 9.2 ME per g of respective target faecal specimen. Except for outliers, no systematic false positive detections from non-target sources were obvious for the animal markers. Still acceptable but somewhat lower performance characteristics were found for the genetic human faecal MST markers, with median target concentrations of log10 5.6 and log10 7.9 ME per g faeces for HF183II and BacHum, respectively. Interestingly, the human MST marker assays revealed a trade-off between faecal source-specificity and -sensitivity, indicating no systematic false positive detectable target concentrations for the less sensitive HF183II assay, whereas revealing a 75% false positive target concentration of log10 3.2 ME per g faeces for BacHum (Fig. 2). In addition, variability in target concentrations of the human-associated MST markers was extremely high compared to the ruminant and pig markers (Fig. 2). Binary assay source-specificity and -sensitivity achieved were as follows: i) HF183II assay, 78 and 90%, ii) BacHum assay, 60 and 100%, iii) BacR assay, 86 and 100% and iv) Pig2Bac assay, 91 and 100%, respectively (Table 1, additional material).
Faecal source discrimination of the selected human-associated (HF183II, BacHum), the ruminant-associated (BacR) and pig-associated (Pig2Bac) MST marker qPCR assays. Marker concentrations in target and non-target faecal specimen (log10 (ME + 1) per gram faeces) are given. Target faecal specimen were those of either humans, ruminants or pigs and non-target faecal specimen those of all other individuals. n number of analysed samples, max highest value, Q75 75% quantile, median median, Q25 25% quantile, min lowest value
3.2 Absence of human- and animal-associated genetic MST markers in protected groundwater
The absence of detectable genetic MST markers in environments free from faecal contamination is a very basic requirement for their reliable application as faecal indicators in water quality testing. Neither human- nor animal-associated faecal MST markers could be detected in the various deep and protected groundwater wells (Fig. 3b, n = 51 samples), where bacterial surface influence was assumed to be absent. This shows that the genetic targets HF183II and BacHum do not occur as part of the investigated natural soil- and groundwater microbial community. The assumed absence of bacterial faecal pollution was verified by SFIB analysis based on E. coli, intestinal enterococci and Clostridium perfringens (Fig. 3a, n = 49).
Occurrence of a SFIB (E. coli, intestinal enterococci, Clostridium perfringens) and b human-associated MST markers (HF183II and BacHum) in the investigated water resources of the basic sample set: i) deep well, ii) well/spring, iii) surface influenced well/spring, iv) river, v) effluent wastewater and vi) influent wastewater. n number of analyzed samples, max highest value, Q75 75% quantile, median median, Q25 25% quantile, min lowest value, n. d. not detected
3.3 High faecal indication capacity of human MST markers at selected water resources
Human-associated faecal pollution dominated the pollution patterns at the selected Austrian surface water resources. Median concentrations of human-associated MST markers for summarized river samples were log10 4.3 and log10 4.8 ME per 100 ml water sample for HF183II and BacHum, respectively (Fig. 3b), whereas median animal MST marker concentrations ranged from not detectable (pig faecal pollution) to 2.3 ME per 100 ml (ruminant faecal pollution, Figure S‑1 additional materials). The established data set was thus used to focus on the human-associated faecal pollution markers. When looking at the overall picture, including all selected water resources (n = 181 samples), SFIB analysis demonstrates that the fully achievable faecal pollution gradient was covered by the selected sampling locations. E. coli, Enterococci and Clostridium perfringens concentrations ranged from not detectable in protected groundwater up to 107 colony forming units (CFU) per 100 ml in raw communal wastewater (Fig. 3a). The median reduction for the three SFIB parameters in biological-based wastewater treatment plants (WWTPs) were 2.6 log10, 2.9 log10 and 2.1 log10, respectively. SFIB pollution levels for river water locations ranged between groundwater resources and WWTP effluents (Fig. 3a). The human-associated MST markers reflected the observed faecal pollution gradient in the various habitats, as outlined by the SFIB analysis (Fig. 3b). Impressively, HF183II and BacHum concentration levels ranged from not detectable up to 109 ME per 100 ml of sample volume. The MST marker thus showed more than a 100-fold higher concentration range than SFIB (compare Fig. 3a vs. Fig. 3b).
3.4 Creating the whole picture: combining SFIB and MST marker analysis
Correlation analysis for the whole data set demonstrated high statistical associations amongst the fraction of the SFIB (i.e., E. coli, enterococci, C. perfringens) and between the SFIB and the human genetic MST markers (Fig. 4a). This further supports the fact that selected water resources were mainly impacted by human-associated faecal pollution. In addition, a very strong correlation became apparent between the two genetic human MST marker assays, revealing a correlation coefficient of rho = 0.98. This high association proves the precise method of qPCR determination of human-associated faecal pollution by the two different Bacteroidetes PCR assays (Fig. 4b). Only at very low concentration ranges marker levels did not correspond due to statistical stochasticity effects. In contrast to the tight associations between the indicators of general faecal pollution (SFIB) and the human-associated MST faecal markers, correlation analysis indicated only moderate or no detectable statistical associations between SFIB and animal MST markers (Fig. 4a).
a Correlation analysis of SFIB (E. coli, intestinal enterococci, Clostridium perfringens) and the host-associated genetic MST markers (HF183II, BacHum, BacR, Pig2Bac). Spearman rank correlation table with correlation coefficients (rho) given in black. Crossed out values are non-significant values after Bonferroni correction with a significance level of alpha = 0.05. n giving the number of correlated values. b Scatterplot showing the correlation between the two human-associated MST markers BacHum (y-axis) and HF183II (x-axis) with a correlation coefficient of rho = 0.98 and a significance level of p < 0.001. Different water resources are indicated with different colors
3.5 Assessing genetic MST marker resistance during UV irradiation: the “AllBac Story”
As genetic human or animal MST markers occur at very low concentrations in groundwater, assessment of resistance during on-site UV irradiation is not a suitable option for such compartments. For this purpose, a comparable genetic Bacteroides target occurring at measurable concentrations, also in the absence of faecal pollution, was needed. The genetic AllBac marker, targeting Bacteroidetes populations, which can originate from soil, surfaces and faeces, suggested good potential for this purpose. In-depth evaluation for all the selected Austrian habitats demonstrated a concentration range of almost 11 orders of magnitude (Fig. 5, i.e., 0 to 1011 ME per 100 ml), thus qualifying as a surrogate marker for the quantification of UV resistance. From deep to surface influenced wells, AllBac was observed in the range of 101 to 104 ME per 100 ml (Fig. 5). Concerning concentrations of the genetic AllBac marker before and after UV irradiation of groundwater, a median 0.3 log10 reduction was identified (Fig. 6a). During UV irradiation of a WWTP effluent (mechanical and biological treatment), AllBac demonstrated a median 0.3 log10 reduction whereas, HF183II and BacHum revealed a median reduction of 0.7 log10 and 0.5 log10 in the WWTP (Fig. 6b). In conclusion, genetic qPCR targets covering AllBac and human MST markers demonstrated 1 log10 reduction during on-site UV irradiation, including groundwater (i.e., drinking water supply) and WWTP effluent treatment (i.e., protection of sensitive catchments).
Occurrence of the general Bacteriodetes marker (AllBac), measured by qPCR and total cell counts (TCC) measured by flow cytometry for the different water resources of the basic sample set: i) deep well, ii) well/spring, iii) surface influenced well/spring, iv) river, v) effluent wastewater and vi) influent wastewater. For effluent and influent wastewater TCC was not analysed. NA not available, other abbreviations: see Fig. 3
Comparison of genetic marker concentrations (AllBac, HF183II, BacHum) before and after on-site UV irradiation for a groundwater (pooled samples from two wells) and b effluent wastewater (WWTP 7). n number of analysed samples, max highest value, Q75 75% quantile, median median, Q25 25% quantile, min lowest value
3.6 Illustrating the possibilities of genetic MST to complement SFIB monitoring data
To demonstrate the capacity of genetic MST marker analysis to extend SFIB monitoring, two case studies were compiled from the presented data set. Case study I demonstrates the possibility of specifically tracing the pollution impact of WWTP effluents in larger and smaller river settings (Fig. 7). A WWTP faecal pollution impact at the larger river site was indicated by an up to median log10 1.2 concentration factor increase in SFIB downstream of the effluent. Molecular MST strongly suggested the pollution source to be of human origin, as both human-associated markers showed an identical median increase as SFIB. In contrast, the concentrations of the animal MST markers did not change (Fig. 7a). The SFIB concentration increase downstream of a WWTP effluent at the smaller river site (up to a median log10 1 concentration increase factor) could also be explained by human impact (up to a median log10 0.8 concentration factor increase of the human MST markers). In contrast to ruminant faecal pollution, which remained unchanged downstream of the WWTP, the pig-associated MST marker also increased significantly (up to a median log10 1.8 concentration factor increase, Fig. 7b). Analysing the WWTP influent in detail indeed revealed that part of the faecal pollution was contributed by pig emissions (Pig2Bac median log10 6.9 ME per 100 ml), although the dominant type of pollution was of human origin (BacHum median log10 9.2 ME per 100 ml). While case study I was selected to evaluate the tracing capacity of human MST at two locations with obvious human pollution (except the pig faecal pollution fraction), case study II demonstrates more tricky “real life” pollution problems (Fig. 8). The latter illustrates SFIB pollution at two groundwater resources with relevance to the public water supply. Based on SFIB data alone, it was not clear whether the faecal pollution problems were caused by human (e.g., communal sewage) or animal faecal pollution (e.g., by livestock or wildlife). However, extension of SFIB data with information from animal and human genetic MST markers clearly indicated that the karstic spring was mainly polluted with ruminant faecal pollution, whereas for porous well water, human pollution was the dominant contamination source (municipal sewage) (compare Fig. 8b vs. Fig. 8a).
Case Study I: Tracing of human faecal pollution in river systems, by analysis of water samples upstream and downstream of WWTP effluent points with the combination of SFIB and MST data: a larger river example and b smaller river example with: i) human-associated MST markers HF183II and BacHum; ii) ruminant-associated MST marker BacR and pig-associated MST marker Pig2Bac; and iii) SFIB: E. coli, intestinal enterococci and Clostridium perfringens. n number of analysed samples, max highest value, Q75 75% quantile, median median, Q25 25% quantile, min lowest value
Case study II: Demonstrating molecular MST to elucidate the source of faecal pollution in groundwater resources: a karstic spring (pooled samples) and b a surface-influenced well with i) human-associated MST markers HF183II and BacHum; ii) ruminant-associated MST marker BacR and pig-associated MST marker Pig2Bac; iii) SFIB: E. coli, intestinal enterococci and Clostridium perfringens. n number of analysed samples, max highest value, Q75 75% quantile, median median, Q25 25% quantile, min lowest value
4 Discussion
4.1 Good applicability of genetic MST markers for Austrian water resources
A comprehensive field-evaluation study on human and animal genetic microbial source tracking (MST) markers by quantitative PCR (qPCR) detection for faecal pollution source identification at representative Austrian water resources was successfully realised. To the best of our knowledge, this is the most comprehensive field evaluation study available to date, considering a variety of water compartments and treatments with importance for water management (i.e., wastewater, surface water, porous and karstic groundwater, waste- and groundwater UV irradiation). Genetic MST markers targeting host-associated intestinal Bacteroides populations were selected to identify human and relevant animal faecal pollution sources (ruminant and pig faecal pollution). Both selected animal genetic MST markers cover potentially important livestock (e.g., cattle, sheep, feeding and rearing pigs) and wildlife (e.g., roe deer, chamois, stag, wild boar) faecal pollution sources. Selected genetic MST markers proved good faecal source identification capacity for the selected Austrian water compartments. Genetic MST markers also impressively reflected the faecal pollution levels at the investigated water and wastewater locations. For example, MST markers showed concentrations of 109 genetic targets per 100 ml in raw wastewater, exceeding the standardized faecal indicator bacteria (SFIB) of E. coli and intestinal enterococci concentrations by 100-fold. In the absence of faecal pollution (protected ground water), genetic MST markers were not detectable. Importantly, the combination of MST markers with data from SFIB allowed the characterisation of the extent and origin of SFIB pollution for the investigated lotic aquatic systems. SFIB standard monitoring applications, such as for drinking, recreation, bathing or irrigation water quality testing, can be therefore extended with genetic MST diagnostics when needed. Since processed filters and extracted nucleic acids (DNA or RNA) can be stored (Fig. 1), MST diagnostics can still be realized for long periods after sampling. Building up sample banks for post hoc molecular diagnostics proved very useful for several MST and genetic investigations (Mayer et al. 2018; Savio et al. 2015). It must be highlighted that such an analysis strategy differs fundamentally from procedures applied for cultivation-based microbiological diagnostics, where analysis must be performed as soon as possible after sampling activities.
4.2 Opening the “black box” of faecal pollution to support water quality management
Enumeration of E. coli and intestinal enterococci in water quality testing provides information on the extent of general faecal pollution. Questions concerning the responsible contamination sources most often remain unsolved. As demonstrated, applying molecular MST can open the “black box” of faecal pollution to support source-targeted water quality management. The selected case studies illustrate how genetic MST markers can contribute source-specific information to conventional water quality monitoring by SFIB. Case study I evaluated the ability to trace faecal pollution specifically from communal WWTPs in river environments. Case study II demonstrated the ability to elucidate the relevant sources of faecal pollution in karstic and porous groundwater in the case of contamination problems.
Austrian research together with a leading national water supplier played a pioneering international role in the development of genetic MST markers and their application strategy (Savio et al. 2018). One of the first MST marker quantifications by qPCR that became globally available was designed for alpine Austrian karst spring habitats to differentiate human from ruminant faecal pollution fifteen years ago (Reischer et al. 2007, 2006). By applying these markers in combination with hydrologically guided sampling (“nested sampling”), it became possible to trace back the origin of SFIB pollution at alpine karst springs to the responsible contamination sources (Reischer et al. 2008b, 2011). Using developed MST diagnostics, allowed verifying that human wastewater disposal measures were efficiently working at the investigated very large and hardly accessible alpine karstic catchments and human wastewater was not leading to SFIB contamination of the water resources (Reischer et al. 2011). Based on the developed methods, an integrated strategy to guide health-related microbial quality management for karstic drinking water resources was established (Farnleitner et al. 2018; Savio et al. 2018). The suggested framework can be applied to any headwater catchment and is not limited to karstic and fracture water resources (Farnleitner et al. 2018; Savio et al. 2018). The ruminant MST marker assay (“BacR”), initially developed for the Austrian spring environment, is still one of the best performing and frequently used qPCR assays to enumerate ruminant faecal pollution around the world (Linke et al. 2020; Raith et al. 2013; Reischer et al. 2013).
Genetic MST markers were also applied to determine the origin of faecal pollution along the whole stretch of the navigable Danube River during the Joint Danube Surveys initiated by the ICPDR (Kirschner et al. 2008, 2017, 2020; Reischer et al. 2008a, 2015). Integrating SFIB data with information from molecular MST diagnostics allowed to demonstrate that human faecal pollution (communal wastewater) was the main driver of pollution during the time of the investigations (Kirschner et al. 2008, 2015, 2017). By combining the concept of hydrological connectivity in concert with genetic MST markers, it could also be demonstrated that external (i.e., communal wastewater effluents) and internal (i.e., animal faecal excreta) faecal pollution can play a role in water resource contamination in backwater areas (Frick et al. 2020). Molecular MST applications are not restricted to lotic environments, they have also been successfully applied at stagnant water resources. For example, in concert with other microbiological and chemical parameters, it became possible to evaluate the potential influence of human wastewater on Lake Neusiedl, which is one of the most important recreational water resources in Austria. Important insights could be gathered by MST to guide problem-targeted water quality management (Blaschke et al. 2018; Hatvani et al. 2018; Herzig et al. 2019).
Genetic MST markers have also been successfully used in hydrological models to simulate microbial faecal pollution levels and to assess microbial infection risks for different pathogen contamination scenarios. The simulation tool QMRAcatch was initially programmed for an Austrian section of the Danube River, including its associated backwater areas (Schijven et al. 2015). QMRAcatch can specifically be calibrated and verified by genetic MST markers (Derx et al. 2016). For example, QMRAcatch could be specifically calibrated by human genetic MST markers for the situation of communal wastewater pollution in the Danube River, as observed upstream of Vienna. The calibrated model could then be used to simulate different future scenarios until 2050, including climate change, population growth and wastewater infrastructure improvements, to assess the required management measures to sustainably produce safe drinking water (Demeter et al. 2021).
Information from genetic MST markers may also be combined with efforts on the assessment and monitoring of the ecological quality and status of groundwater. For example, the D‑A‑C index approach, as recently tested for the River Mur Valley (Retter et al. 2021), awaits future evaluation on the ability to integrate MST information such as human versus life stock faecal pollution pressure.
4.3 Current status and future perspectives in MST technology
Without doubt, the application of genetic MST markers for molecular faecal pollution diagnostics and source tracking offers very valuable and fascinating new possibilities for the water management sector. However, this field is still in its exponential development stage, and further major innovations are to be expected in the future (Farnleitner and Reischer 2013; Farnleitner and Sommer 2015; Holcomb and Stewart 2020). Current genetic MST marker detection systems are not yet standardized and show varying performance characteristics, including both analytical (i.e., related to technical detection) and diagnostic (i.e., related to faecal indication interpretation) criteria. For example, as demonstrated in this study, the faecal sensitivity and specificity differed significantly among the evaluated genetic MST markers (cf. Fig. 2, Table 1 additional material). Furthermore, standard qPCR detection technology is not designed to discriminate between viable, dormant and dead cell populations. The PCR targets are small DNA segments in the genome of the selected microbes, which is the reason to designate them as genetic MST markers. The characteristics of the targets are in line with the observed low reduction in genetic MST markers during UV irradiation (1 log10, Fig. 6). In contrast, the reduction in genetic MST markers during mechanical and biological wastewater treatment was much more pronounced (HF183II, 3.1 log10; BacHum, 3.2 log10, (Mayer et al. 2016)). In addition to the mentioned differences in resistance of the genetic MST markers (i.e., fate and reduction during technical treatment), the understanding of the persistence of the genetic MST markers (i.e., fate and reduction in the natural aquatic system) is a fundamental parameter in MST application. This knowledge becomes especially important when hydrological retention times in aquatic systems are expected to be very long.
The realization of MST applications requires comprehensive background knowledge for choosing the optimal study design and to enable the correct interpretation of the results. This study focuses on human, ruminant and pig faecal pollution targets. Nonetheless, other potentially relevant faecal sources also exist for practical water management, such as birds and poultry. However, bird and poultry markers have not yet reached the same level of performance characteristics as achieved for human, ruminant and pig sources and need further technical assay improvements. Additionally, other faecal pollution sources are of increasing concern. For example, animals that live abundantly in urban areas (e.g., rats, cats, dogs) or poikilotrophs (e.g., fish) and invertebrates (e.g., snails) occurring in natural systems (Frick et al. 2018; Wuertz et al. 2011). It is thus very likely that further genetic MST targets and enumeration assays will become available soon for extended MST applications.
In addition to bacterial MST targets (this evaluation study), alternative biological targets are increasingly being applied (Farkas et al. 2020; Malla and Haramoto 2020). Viral MST targets will be applied in concert with bacterial MST targets to reflect the full diversity of microbial faecal pollution characteristics in water resources in future applications (i.e., persistence, resistance, mobility, (Farkas et al. 2020; Kitajima et al. 2018; Mayer et al. 2016)).
Finally, the current “gold” standard of genetic MST marker detection is qPCR enumeration (Harwood et al. 2018; Layton et al. 2013; Raith et al. 2013; Wuertz et al. 2011). Additionally, digital droplet PCR is increasingly applied (Nshimyimana et al. 2019). Both molecular diagnostic quantification technologies require advanced equipment and experienced personnel. In addition, sample processing and DNA/RNA extraction can be very laborious. Future technologies thus have to provide more straightforward solutions to be applicable not only at the university or specialist level but also for conventional water quality laboratories (Farnleitner and Sommer 2015; Martzy et al. 2019). Technologies covering straightforward isothermal amplification procedures, such as LAMP (loop-mediated isothermal amplification) or HAD (helicase-dependent amplification) detection and easy extraction and sample processing systems, are thus the focus of current development (Kolm et al. 2017; Martzy et al. 2019). Nonetheless, expert systems in molecular MST will also be further developed. For example, high-throughput sequencing coupled with appropriate bioinformatic analysis pipelines may also add information as a complementary tool to MST (Savio et al. 2015; Tan et al. 2015; Vierheilig et al. 2015).
References
Anderson, A., Pietsch, K., Zucker, R., Mayr, A., Müller-Hohe, E., Messelhäusser, U., Sing, A., Busch, U. and Huber, I. (2011): Validation of a Duplex Real-Time PCR for the Detection of Salmonella spp. in Different Food Products. Food Analytical Methods 4(3), 259–267.
Ballesté, E., Demeter, K., Masterson, B., Timoneda, N., Sala-Comorera, L. and Meijer, W.G. (2020): Implementation and integration of microbial source tracking in a river watershed monitoring plan. Science of The Total Environment 736, 139573.
Blaschke, A.P., Farnleitner, A.H., Kirschner, A.K.T., Sommer, R. (2018): Nachhaltiges Gesamtkonzept zum Management des mikrobiologisch-hygienischen Zustandes des Neusiedler Sees unter Berücksichtigung der Siedlungswasser-Wirtschaft der Umlandgemeinden, Phase 1: „Pilotstudie“. Bundesministerium für Nachhaltigkeit und Tourismus, Wien.
Boehm, A.B., Van De Werfhorst, L.C., Griffith, J.F., Holden, P.A., Jay, J.A., Shanks, O.C., Wang, D. and Weisberg, S.B. (2013): Performance of forty-one microbial source tracking methods: a twenty-seven lab evaluation study. Water Res 47(18), 6812–6828.
Brown, K.I., Graham, K.E. and Boehm, A.B. (2017): Risk-Based Threshold of Gull-Associated Fecal Marker Concentrations for Recreational Water. Environmental Science Technology Letters 4(2), 44–48.
Cao, Y., Sivaganesan, M., Kelty, C.A., Wang, D., Boehm, A.B., Griffith, J.F., Weisberg, S.B. and Shanks, O.C. (2018): A human fecal contamination score for ranking recreational sites using the HF183/BacR287 quantitative real-time PCR method. Water Research 128, 148–156.
Demeter, K., Derx, J., Komma, J., Parajka, J., Schijven, J., Sommer, R., Cervero-Aragó, S., Lindner, G., Zoufal-Hruza, C.M., Linke, R., Savio, D., Ixenmaier, S.K., Kirschner, A.K.T., Kromp, H., Blaschke, A.P. and Farnleitner, A.H. (2021): Modelling the interplay of future changes and wastewater management measures on the microbiological river water quality considering safe drinking water production. Science of The Total Environment 768, 144278.
Derx, J., Schijven, J., Sommer, R., Zoufal-Hruza, C.M., van Driezum, I.H., Reischer, G., Ixenmaier, S., Kirschner, A., Frick, C., de Roda Husman, A.M., Farnleitner, A.H. and Blaschke, A.P. (2016): QMRAcatch: Human-Associated Fecal Pollution and Infection Risk Modeling for a River/Floodplain Environment. Journal of environmental quality 45(4), 1205–1214.
DIN 1976 DIN 38404-4:1976-12 German Standard Methods for Analysing of Water, Waste Water and Sludge; Physical and Physical-chemical Parameters (Group C); Determination of Temperature (C4) DIN.
DIN 2019 DIN EN 1484:2019-04 Water analysis—Guidelines for the determination of total organic carbon (TOC) and dissolved organic carbon (DOC); German version EN 1484:1997 DIN.
Diston, D., Robbi, R., Baumgartner, A. and Felleisen, R. (2018): Microbial source tracking in highly vulnerable karst drinking water resources. Journal of Water and Health 16(1), 138–149.
van Driezum, I.H., Chik, A.H.S., Jakwerth, S., Lindner, G., Farnleitner, A.H., Sommer, R., Blaschke, A.P. and Kirschner, A.K.T. (2018): Spatiotemporal analysis of bacterial biomass and activity to understand surface and groundwater interactions in a highly dynamic riverbank filtration system. Science of the Total Environment 627, 450–461.
Evison, L. and James, A. (1975): Bifidobacterium as an indicator of faecal pollution in water. Progress in Water Technology 7, 57–66.
Farkas, K., Walker, D.I., Adriaenssens, E.M., McDonald, J.E., Hillary, L.S., Malham, S.K. and Jones, D.L. (2020): Viral indicators for tracking domestic wastewater contamination in the aquatic environment. Water Research 181, 115926.
Farnleitner, A.H. and Reischer, G. (2013): New perspectives in water quality testing. International Innovation. Disseminating Science, Research and Technology. Health Partnership, 18–20.
Farnleitner, A.H. and Sommer, R. (2015): Wasserqualität und Gesundheit: zukünftige Herausforderungen? Schriftenreihe des Österreichischen Wasser- und Abfallwirtschaftsverbandes (ÖWAV) Heft 170 „Zukunft Denken“ – Wasserwirtschaft 2035, 35–45.
Farnleitner, A.H., Ryzinska-Paier, G., Reischer, G.H., Burtscher, M.M., Knetsch, S., Kirschner, A.K.T., Dirnbock, T., Kuschnig, G., Mach, R.L. and Sommer, R. (2010): Escherichia coli and enterococci are sensitive and reliable indicators for human, livestock and wildlife faecal pollution in alpine mountainous water resources. Journal of Applied Microbiology 109(5), 1599–1608.
Farnleitner, A.H., Savio, D., Sommer, R., Reischer, G.H., Linke, R., Kirschner, A., Zerobin, W. and Stadler, H. (2018): Karst Groundwater Contamination and Public Health. W. B. White, J.S.H., E. K. Herman, and M. Rutigliano (ed), Springer.
Frick, C., Vierheilig, J., Linke, R., Savio, D., Zornig, H., Antensteiner, R., Baumgartner, C., Bucher, C., Blaschke, A.P., Derx, J., Kirschner, A.K.T., Ryzinska-Paier, G., Mayer, R., Seidl, D., Nadiotis-Tsaka, T., Sommer, R. and Farnleitner, A.H. (2018): Poikilothermic Animals as a Previously Unrecognized Source of Fecal Indicator Bacteria in a Backwater Ecosystem of a Large River. Applied and environmental microbiology 84(16), 15.
Frick, C., Vierheilig, J., Nadiotis-Tsaka, T., Ixenmaier, S., Linke, R., Reischer, G.H., Komma, J., Kirschner, A.K.T., Mach, R.L., Savio, D., Seidl, D., Blaschke, A.P., Sommer, R., Derx, J. and Farnleitner, A.H. (2020): Elucidating fecal pollution patterns in alluvial water resources by linking standard fecal indicator bacteria to river connectivity and genetic microbial source tracking. Water Research 184, 12.
Green, H.C., Haugland, R.A., Varma, M., Millen, H.T., Borchardt, M. A., Field, K.G., Walters, W.A., Knight, R., Sivaganesan, M., Kelty, C.A. and Shanks, O.C. (2014): Improved HF183 Quantitative Real-Time PCR Assay for Characterization of Human Fecal Pollution in Ambient Surface Water Samples. Applied and environmental microbiology 80(10), 3086–3094.
Griffiths, R.I., Whiteley, A.S., O’Donnell, A.G. and Bailey, M.J. (2000): Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA- and rRNA-based microbial community composition. Appl Environ Microbiol 66(12), 5488–5491.
Harwood, V.J., Shanks, O., Koraijkic, A., Verbyla, M. E., Ahmed, W. and Iriarte, M. (2018): Global Water Pathogen Project. Rose, J.B. and Jimenez Cisneros, B.E. (eds), Michigan State University, Lansing, MI; UNESCO.
Hatvani, I.G., Kirschner, A.K.T., Farnleitner, A.H., Tanos, P. and Herzig, A. (2018): Hotspots and main drivers of fecal pollution in Neusiedler See, a large shallow lake in Central Europe. Environ. Sci. Pollut. Res. 25(29), 28884–28898.
Herzig, A., Hatvani, I.G., Tanos, P., Blaschke, A.P., Sommer, R., Farnleitner, A.H. and Kirschner, A.K.T. (2019): Mikrobiologisch-hygienische Untersuchungen am Neusiedler See – von der Einzeluntersuchung zum Gesamtkonzept. Österreichische Wasser- und Abfallwirtschaft 71(11), 537–555.
Holcomb, D.A. and Stewart, J.R. (2020): Microbial Indicators of Fecal Pollution: Recent Progress and Challenges in Assessing Water Quality. Curr. Environ. Health Rep., 7, 311–324. https://doi.org/10.1007/s40572-020-00278-1
Ishii, S., Hansen, D.L., Hicks, R.E. and Sadowsky, M.J. (2007): Beach Sand and Sediments are Temporal Sinks and Sources of Escherichia coli in Lake Superior. Environmental Science Technology 41(7), 2203–2209.
ISO 1985 ISO 7888:1985 Water quality—Determination of electrical conductivity, ISO.
ISO 2000 ISO 7899-2:2000 Water quality—Detection and enumeration of intestinal enterococci—Part 2: Membrane filtration method.
ISO 2008 ISO 10523:2008 Water quality—Determination of pH, ISO.
ISO 2013 ISO 14189:2013 Water quality—Enumeration of Clostridium perfringens—Method using membrane filtration, p. 12.
ISO 2018 ISO 16649-1:2018: Microbiology of food and animal feeding stuffs—Horizontal method for the enumeration of beta-glucuronidase-positive Escherichia coli—Part 1: Colony-count technique at 44 degrees C using membranes and 5‑bromo-4-chloro-3-indolyl beta-D-glucuronide, p. 9.
ISO 2019 ISO 7027-2:2019 Water quality—Determination of turbidity—Part 2: Semi-quantitative methods for the assessment of transparency of waters, ISO.
Kildare, B.J., Leutenegger, C.M., McSwain, B.S., Bambic, D.G., Rajal, V.B., Wuertz, S. (2007): 16S rRNA-based assays for quantitative detection of universal human- cow- and dog-specific fecal Bacteroidales: A Bayesian approach. Water Research 41(16), 3701–3715. https://doi.org/10.1016/j.watres.2007.06.037
Kirschner, A., Kavka, G., Velimirov, B., Reischer, G., Mach, R. and Farnleitner, A. (2008): Joint Danube Survey 2. Final Scientific Report. Igor Liška, F.W., Jaroslav Slobodník (ed), ICPDR—International Commission for the Protection of the Danube River, Vienna, Austria.
Kirschner, A.K.T., Schachner, I., Jakwerth, S., Savio, D., Kolarevic, S., Kracun-Kolarevic, M., G., Z., Kittinger, C., Koller, M., Linke, R. and Farnleitner, A.H. (2020): Extent and origin of microbial fecal pollution. Joint Danube Survey 4—A comprehensive Analysis of Danube Water Quality, Final scientific report.
Kirschner, A.K.T., Kavka, G., Reischer, G.H., Sommer, R., Blaschke, A.P., Stevenson, M., Vierheilig, J., Mach, R.L. and Farnleitner, A.H. (2015): The Danube River Basin. I., L. (ed), Springer-Verlag Berlin Heidelberg.
Kirschner, A.K.T., Reischer, G.H., Jakwerth, S., Savio, D., Ixenmaier, S., Toth, E., Sommer, R., Mach, R.L., Linke, R., Eiler, A., Kolarevic, S. and Farnleitner, A.H. (2017): Multiparametric monitoring of microbial faecal pollution reveals the dominance of human contamination along the whole Danube River. Water Research 124, 543–555.
Kitajima, M., Sassi, H.P. and Torrey, J.R. (2018): Pepper mild mottle virus as a water quality indicator. NPJ Clean Water 1, 9.
Kolm, C., Martzy, R., Brunner, K., Mach, R.L., Krska, R., Heinze, G., Sommer, R., Reischer, G.H. and Farnleitner, A.H. (2017): A Complementary Isothermal Amplification Method to the U.S. EPA Quantitative Polymerase Chain Reaction Approach for the Detection of Enterococci in Environmental Waters. Environ. Sci. Technol. 51(12), 7028–7035.
Kreader, C.A. (1995): Design and evaluation of Bacteroides DNA probes for the specific detection of human fecal pollution. Appl Environ Microbiol 61(4), 1171–1179.
Layton, A., McKay, L., Williams, D., Garrett, V., Gentry, R., Sayler, G. (2006): Applied and Environmental Microbiology 72(6), 4214–4224. https://doi.org/10.1128/AEM.01036-05
Layton, B.A., Cao, Y., Ebentier, D.L., Hanley, K., Balleste, E., Brandao, J., Byappanahalli, M., Converse, R., Farnleitner, A.H., Gentry-Shields, J., Gidley, M.L., Gourmelon, M., Lee, C.S., Lee, J., Lozach, S., Madi, T., Meijer, W.G., Noble, R., Peed, L., Reischer, G.H., Rodrigues, R., Rose, J.B., Schriewer, A., Sinigalliano, C., Srinivasan, S., Stewart, J., Van De Werfhorst, L.C., Wang, D., Whitman, R., Wuertz, S., Jay, J., Holden, P.A., Boehm, A.B., Shanks, O. and Griffith, J.F. (2013): Performance of human fecal anaerobe-associated PCR-based assays in a multi-laboratory method evaluation study. Water Res 47(18), 6897–6908.
Linke, R.B., Kebede, G., Mushi, D., Lakew, A., Hayes, D.S., Graf, W. and Farnleitner, A.H. (2020): Assessing the faecal source sensitivity and specificity of ruminant and human genetic microbial source tracking markers in the central Ethiopian highlands. Letters in Applied Microbiology 72, 458–466. https://doi.org/10.1111/lam.13436
Malla, B. and Haramoto, E. (2020): Host-specific mitochondrial DNA markers for tracking the sources of fecal pollution. Curr. Opin. Environ. Sci. Health 16, 34–46.
Mara, D. and Oragui, J. (1983): Sorbitol-fermenting bifidobacteria as specific indicators of human faecal pollution. Journal of Applied Bacteriology 55, 349–357.
Martzy, R., Kolm, C., Krska, R., Mach, R.L., Farnleitner, A.H. and Reischer, G.H. (2019): Challenges and perspectives in the application of isothermal DNA amplification methods for food and water analysis. Anal. Bioanal. Chem. 411(9), 1695–1702.
Mayer, R.E., Reischer, G.H., Ixenmaier, S.K., Derx, J., Blaschke, A.P., Ebdon, J.E., Linke, R., Egle, L., Ahmed, W., Blanch, A.R., Byamukama, D., Savill, M., Mushi, D., Cristobal, H.A., Edge, T.A., Schade, M. A., Aslan, A., Brooks, Y.M., Sommer, R., Masago, Y., Sato, M.I., Taylor, H.D., Rose, J.B., Wuertz, S., Shanks, O.C., Piringer, H., Mach, R.L., Savio, D., Zessner, M. and Farnleitner, A.H. (2018): Global Distribution of Human-Associated Fecal Genetic Markers in Reference Samples from Six Continents. Environ Sci Technol 52(9), 5076–5084.
Mayer, R.E., Sofill-Mas, S., Egle, L., Reischer, G.H., Schade, M., Fernandez-Cassi, X., Fuchs, W., Mach, R.L., Lindner, G., Kirschner, A., Gaisbauer, M., Piringer, H., Blaschke, A.P., Girones, R., Zessnet, M., Sommer, R. and Farnleitner, A.H. (2016): Occurrence of human-associated Bacteroidetes genetic source tracking markers in raw and treated wastewater of municipal and domestic origin and comparison to standard and alternative indicators of faecal pollution. Water research 90, 265–276.
Mieszkin, S., Furet, J.P., Corthier, G. and Gourmelon, M. (2009): Estimation of pig fecal contamination in a river catchment by real-time PCR using two pig-specific Bacteroidales 16S rRNA genetic markers. Appl Environ Microbiol 75(10), 3045–3054.
Nshimyimana, J.P., Cruz, M.C., Thompson, R.J. and Wuertz, S. (2017): Bacteroidales markers for microbial source tracking in Southeast Asia. Water Research 118, 239–248.
Nshimyimana, J.P., Cruz, M.C., Wuertz, S. and Thompson, J.R. (2019): Variably improved microbial source tracking with digital droplet PCR. Water Research 159, 192–202.
Power, M.L., Littlefield-Wyer, J., Gordon, D.M., Veal, D.A. and Slade, M.B. (2005): Phenotypic and genotypic characterization of encapsulated Escherichia coli isolated from blooms in two Australian lakes. Environmental microbiology 7(5), 631–640.
Raith, M.R., Kelty, C.A., Griffith, J.F., Schriewer, A., Wuertz, S., Mieszkin, S., Gourmelon, M., Reischer, G.H., Farnleitner, A.H., Ervin, J.S., Holden, P.A., Ebentier, D.L., Jay, J.A., Wang, D., Boehm, A.B., Aw, T.G., Rose, J.B., Balleste, E., Meijer, W.G., Sivaganesan, M. and Shanks, O.C. (2013): Comparison of PCR and quantitative real-time PCR methods for the characterization of ruminant and cattle fecal pollution sources. Water Res 47(18), 6921–6928.
RCoreTeam (2013): R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/.
Reischer, G., Kavka, G., Kasper, D., Winter, C., Mach, R. and Farnleitner, A. (2008a): Applicability of DNA based quantitative microbial source tracking (QMST) evaluated on a large scale in the Danube River and its important tributaries. Fundamental and Applied Limnolog, Suppl. Large Rivers 162, 117–125.
Reischer, G., Kirschner, A.K.T., Schnitzer, G., Savio, D., Mach, R.L., Bahlmann, A., Schulze, T., Brack, W. and Farnleitner, A.H. (2015): Joint Danube Survey 3. A comprehensive analysis of Danube water quality. Igor Liška, F.W., Manfred Sengl, Karin Deutsch, Jaroslav Slobodník (ed), ICPDR—International Commission for the Protection of the Danube River, Vienna, Austria.
Reischer, G.H., Haider, J.M., Sommer, R., Stadler, H., Keiblinger, K.M., Hornek, R., Zerobin, W., Mach, R.L. and Farnleitner, A.H. (2008b): Quantitative microbial faecal source tracking with sampling guided by hydrological catchment dynamics. Environmental microbiology 10(10), 2598–2608.
Reischer, G.H., Kasper, D.C., Steinborn, R., Mach, R.L. and Farnleitner, A.H. (2006): Quantitative PCR method for sensitive detection of ruminant fecal pollution in freshwater and evaluation of this method in alpine karstic regions. Applied and environmental microbiology 72(8), 5610–5614.
Reischer, G.H., Kasper, D.C., Steinborn, R., Farnleitner, A.H. and Mach, R.L. (2007): A quantitative real-time PCR assay for the highly sensitive and specific detection of human faecal influence in spring water from a large alpine catchment area. Letters in Applied Microbiology 44(4), 351–356.
Reischer, G.H., Kollanur, D., Vierheilig, J., Wehrspaun, C., Mach, R.L., Sommer, R., Stadler, H. and Farnleitner, A.H. (2011): Hypothesis-Driven Approach for the Identification of Fecal Pollution Sources in Water Resources. Environmental Science Technology 45(9), 4038–4045.
Reischer, G.H., Ebdon, J.E., Bauer, J.M., Schuster, N., Ahmed, W., Astrom, J., Blanch, A.R., Bloeschl, G., Byamukama, D., Coakley, T., Ferguson, C., Goshu, G., Ko, G., Husman, A.M.d.R., Mushi, D., Poma, R., Pradhan, B., Rajal, V., Schade, M. A., Sommer, R., Taylor, H., Toth, E.M., Vrajmasu, V., Wuertz, S., Mach, R.L. and Farnleitner, A.H. (2013): Performance Characteristics of qPCR Assays Targeting Human- and Ruminant-Associated Bacteroidetes for Microbial Source Tracking across Sixteen Countries on Six Continents. Environmental Science Technology 47(15), 8548–8556.
Resnick, I.G. and Levin, M. A. (1981): Assessment of bifidobacteria as indicators of human fecal pollution. Appl Environ Microbiol 42(3), 433–438.
Retter, A., C. Griebler, Haas, J., S. Birk, C. Stump, H. Brielmann and L Fillinger (2021): Application of the D‑A-(C) index as a simple tool for microbial-ecological characterization and assessment of groundwater ecosystems—a case study of the Mur River Valley, Austria. Österreichische Wasser und Abfallwirtschaft, Heft 11–12/2021: „Mikrobiologie und Wasser“
Santo-Domingo, J.S., Edge, T., Griffith, J., Hansel, J., Harwood, V.J., Jenkins, M., Layton, A., Marirosa, M., Nakatsu, C., Oshiro, R., Sadowsky, M., Shanks, O., Stelma, G., Stewart, J., Stoeckel, D., Wiggins, B. and Wilbur, J. (2005): Microbial source tracking guide document. Agency, U.E.P. (ed), p. 135, US Environmental Protection Agency.
Savio, D., Sinclair, L., Ijaz, U.Z., Parajka, J., Reischer, G.H., Stadler, P., Blaschke, A.P., Bloschl, G., Mach, R.L., Kirschner, A.K.T., Farnleitner, A.H. and Eiler, A. (2015): Bacterial diversity along a 2600km river continuum. Environmental microbiology 17(12), 4994–5007.
Savio, D., Stadler, P., Reischer, G.H., Kirschner, A.K.T., Demeter, K., Linke, R., Blaschke, A.P., Sommer, R., Szewzyk, U., Wilhartitz, I.C., Mach, R.L., Stadler, H. and Farnleitner, A.H. (2018): Opening the black box of spring water microbiology from alpine karst aquifers to support proactive drinking water resource management. Wiley Interdisciplinary Reviews-Water. https://doi.org/10.1002/wat2.1282
Schijven, J., Derx, J., Husman, A.M.d.R., Blaschke, A.P. and Farnleitner, A.H. (2015): QMRAcatch: Microbial Quality Simulation of Water Resources including Infection Risk Assessment. Journal of environmental quality 44(5), 1491–1502.
Schrammel, B., Cervero-Aragó, S., Dietersdorfer, E., Walochnik, J., Lück, C., Sommer, R. and Kirschner, A. (2018): Differential development of Legionella sub-populations during short- and long-term starvation. Water Research 141, 417–427.
Staley, Z.R. and Edge, T.A. (2016): Comparative microbial source tracking methods for identification of fecal contamination sources at Sunnyside Beach in the Toronto region area of concern. Journal of Water and Health 14(5), 839–850.
Symonds, E.M., Young, S., Verbyla, M. E., McQuaig-Ulrich, S.M., Ross, E., Jimenez, J.A., Harwood, V.J. and Breitbart, M. (2017): Microbial source tracking in shellfish harvesting waters in the Gulf of Nicoya, Costa Rica. Water Research 111, 177–184.
Tallon, P., Magajna, B., Lofranco, C. and Leung, K.T. (2005): Microbial Indicators of Faecal Contamination in Water: A Current Perspective. Water, Air, Soil Pollution 166(1), 139–166.
Tan, B., Ng, C., Nshimyimana, J.P., Loh, L.L., Gin, K.Y.H. and Thompson, J.R. (2015): Next-generation sequencing (NGS) for assessment of microbial water quality: current progress, challenges, and future opportunities. Front. Microbiol. 6, 20.
Turnbaugh, P.J., Ley, R.E., Hamady, M., Fraser-Liggett, C.M., Knight, R. and Gordon, J.I. (2007): The Human Microbiome Project. Nature 449(7164), 804–810.
USEPA (2019): Method 1696: Characterization of Human Fecal Pollution in Water by HF183/BacR287 TaqMan® Quantitative Polymerase Chain Reaction (qPCR) Assay, USEPA.
Vierheilig, J., Savio, D., Ley, R.E., Mach, R.L., Farnleitner, A.H. and Reischer, G.H. (2015): Potential applications of next generation DNA sequencing of 16S rRNA gene amplicons in microbial water quality monitoring. Water Science and Technology 72(11), 1962–1972.
WHO (ed) (2017): Guidelines for drinking-water quality: Fourth edition incorporating the first addendum, World Health Organisation.
Wuertz, S., Wang, D., Reischer, G.H. and Farnleitner, A.H. (2011): Microbial Source Tracking: Methods, Applications, and Case Studies, pp. 61–112.
Wyer, M.D., Kay, D., Watkins, J., Davies, C., Kay, C., Thomas, R., Porter, J., Stapleton, C.M. and Moore, H. (2010): Evaluating short-term changes in recreational water quality during a hydrograph event using a combination of microbial tracers, environmental microbiology, microbial source tracking and hydrological techniques: a case study in Southwest Wales, UK. Water Res 44(16), 4783–4795.
Zhang, Q., Gallard, J., Wu, B., Harwood, V.J., Sadowsky, M.J., Hamilton, K.A. and Ahmed, W. (2019): Synergy between quantitative microbial source tracking (qMST) and quantitative microbial risk assessment (QMRA): A review and prospectus. Environment International 130, 104703. https://doi.org/10.1016/j.envint.2019.03.051
Acknowledgements
This research was funded by the Niederösterreichische Forschungs- und Bildungsgesellschaft (NFB) Science Call 2015 “Ressource und Lebensgrundlage Wasser” (project SC15-016 Aquasafe). Further support came also from the “Aquascreen” project (WST3-F-5031298/001-2017) funded by the FTI-program of the province of Lower Austria by providing DNA extracts from distinct ground water wells. We thank EVN Wasser GmbH for their great support during the sampling campaign and for the provision of data on the investigated water resources. Besides, we also want to thank the operators of the wastewater treatment plants participating in this study for their great support and the provision of samples from their institutions. We also acknowledge the project ‘Vienna Water Resources Systems 2020+’, a research cooperation between Wiener Wasser and the ICC Water and Health, for providing samples from alpine karst springs. Additional support came from the ‘Bundesministerium für Land- und Forstwirtschaft, Umwelt und Wasserwirtschaft’ represented by the Kommunalkredit Public Consulting GmbH in the frame of the project GZ B401184 to realize the investigation of UV-resistence of genetic MST markers in waste water. This work represents a joint effort of the Interuniversity Cooperation Center for Water Health (www.waterandhealth.at).
Funding
Open access funding provided by TU Wien (TUW).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The authors S.D. Steinbacher and D. Savio contributed equally to the manuscript.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Steinbacher, S.D., Savio, D., Demeter, K. et al. Genetic microbial faecal source tracking: rising technology to support future water quality testing and safety management. Österr Wasser- und Abfallw 73, 468–481 (2021). https://doi.org/10.1007/s00506-021-00811-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00506-021-00811-y
Keywords
- Faecal pollution
- Microbial source tracking
- Faecal indicator bacteria
- Genetic faecal marker
- Bacterial viral intestinal pathogens
- Water quality
- Austria