Abstract
Sister chromatid cohesion is mediated by cohesin. At the onset of mitosis, most cohesin dissociates from chromatin with the exception of a small population, present along chromosome arms and enriched at centromeres. A protein known as shugoshin (Sgo) is essential to maintain arm and centromeric cohesion until the onset of anaphase in transformed human cells, but not in other organisms like Drosophila or mouse. We have used Xenopus egg extracts to further explore this issue. Chromosomes assembled in extracts depleted of Sgo have little or no cohesin at centromeres and display centromeric cohesion defects. Unlike transformed human cells, however, arm cohesion is maintained in the absence of Sgo. Furthermore, Sgo depletion impairs the prophase dissociation of cohesin. This phenotype can be rescued by inhibition of PP2A. The protein phosphatase interacts with Sgo and accumulates at centromeres in mitosis in a Sgo-dependent manner. We propose that Sgo drives relocalization of PP2A from arms to centromeres and, in this way, coordinates release of arm cohesin with protection of centromeric cohesin in mitosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sister chromatid cohesion is essential for proper chromosome segregation. It entails the creation and maintenance of a physical linkage between the two copies of a chromosome that arise from the duplication process until their separation at the metaphase to anaphase transition. Cohesin, a ring-shaped multiprotein complex conserved from yeast to human, is a major mediator of this linkage (Losada and Hirano 2005; Nasmyth and Haering 2005). It consists of a heterodimer of SMC1 and SMC3 and two regulatory subunits, Scc1/Rad21 and Scc3/SA. In vertebrate cells, cohesin is loaded on chromatin in early G1 phase and establishes cohesion during S phase. Most cohesin is released from chromatin during prophase, and only a small population remains between the sister chromatids enriched in the pericentric region (Losada et al. 1998; Waizenegger et al. 2000). This population is essential in preventing precocious separation of the sister chromatids and allowing proper chromosome alignment (Vass et al. 2003; Vagnarelli et al. 2004; Losada et al. 2005). The dissociation of cohesin in prophase requires Polo and Aurora B kinases as well as a number of additional factors that include Wapl (Losada et al. 2002; Sumara et al. 2002; Hauf et al. 2005; Gandhi et al. 2006; Kueng et al. 2006). At the onset of anaphase, cleavage of Scc1/Rad21 by separase drives dissociation of the remaining cohesin and sister chromatid separation ensues (Hauf et al. 2001).
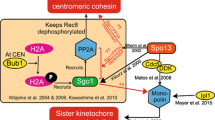
Sequential loss of cohesion is essential also in meiosis (Marston and Amon 2004). Arm cohesion is released in anaphase I to allow resolution of chiasmata and segregation of homologous chromosomes. Centromeric cohesion persists until anaphase II, when it is dissolved to allow segregation of the sister chromatids. In both anaphase I and anaphase II, loss of cohesion involves separase-mediated cleavage of cohesin (Terret et al. 2003; Brar et al. 2006; Kudo et al. 2006). A family of proteins known as shugoshins (Sgo), orthologues of Drosophila melanogaster MEI-S332, protect centromeric cohesin from separase cleavage in meiosis I (Kerrebrock et al. 1995; Kitajima et al. 2004; Rabitsch et al. 2004). One or two orthologues of this protein can be found in most eukaryotes, and they function both in meiosis and mitosis. It has been proposed that one specific variant of the protein phosphatase PP2A associates with Sgo and protects centromeric cohesin in meiosis I by counteracting Rec8 phosphorylation by Polo (Riedel et al. 2006). A second report showed that PP2A associates with Sgo proteins also in HeLa cells and most likely counteracts SA2 phosphorylation by Polo (Kitajima et al. 2006). In Saccharomyces cerevisiae, an organism in which there is no prophase release of cohesin, Sgo1 is not required for cohesion but for sensing tension between sister chromatids (Indjeian et al. 2005). Human Sgo2 also acts as a sensor of tension (Huang et al. 2007; Lee et al. 2008), whereas a number of studies in HeLa cells show that knock down of Sgo1 by small interfering RNA (siRNA) leads to premature separation of the sister chromatids and prometaphase arrest (Salic et al. 2004; McGuinness et al. 2005). Therefore, in these transformed human cells, Sgo1 appears to protect a small population of cohesin both at arms and at centromeres from the prophase pathway (Nakajima et al. 2007). In contrast, the pioneer studies of D. melanogaster mei-S332 mutants indicated the absence of cohesion defects in mitotic cells (Kerrebrock et al. 1995). After removing arm cohesion by treatment with colchicine, sister chromatid separation was observed in 18% of the mutant cells compared with 9% of the wild-type cells, suggesting that shugoshin MEI-S332 was specifically involved in centromeric cohesion (LeBlanc et al. 1999). A similar conclusion can be drawn from the study of Bub1-null mouse embryonic fibroblasts (MEFs; Perera et al. 2007). Bub1 is a component of the spindle assembly checkpoint, and it is also required for proper localization of Sgo at centromeres (Tang et al. 2004; Kitajima et al. 2005; Vaur et al. 2005; Boyarchuk et al. 2007). In Bub1-null MEFs treated with the APC/C inhibitor MG132 to prevent anaphase entry, there is no precocious separation of sister chromatids even though there is no Sgo1 at centromeres. Thus, while the role of Sgo in the protection of centromeric cohesion in meiosis is conserved in most species, its role in regulation of mitotic cohesion is less clear.
We have used Xenopus laevis egg cell-free extracts to further understand how Sgo regulates the behavior of cohesin in mitosis. To date, a single Sgo protein has been identified in X. laevis (Salic et al. 2004). We report that chromosomes assembled in extracts without Sgo show increased distance between sister centromeres, consistent with a centromeric cohesion defect caused by the lack of cohesin enrichment at this region. However, these chromosomes have paired chromosome arms. In fact, cohesin release from chromosome arms is partially defective in extracts without Sgo, most likely due to mislocalization of PP2A.
Materials and methods
Antibodies
Two fragments of Xenopus Bub1 (p1, aminoacids 1–331, and p2, aminoacids 568–828, a gift from J. Maller, University of Colorado) were expressed and purified from bacteria as 6× His-tagged fusion proteins and used to produce rabbit polyclonal and mouse monoclonal antibodies, respectively. Mouse monoclonal and rabbit polyclonal antibodies were also raised against a maltose binding protein fusion of full-length Xenopus Sgo protein (MBP-XSgo, a gift from A. Salic). Rabbit polyclonal sera against Xenopus CENP-A were obtained using a synthetic peptide as immunogen (MRPGSTPPSRRKSRPPRRVC). Other antibodies used in this study were as follows: anti-SMC1, anti-SMC3, and anti-Scc1 (Losada et al. 1998); anti-SA1 (Losada et al. 2000); anti-XCAP-G (Hirano et al. 1997); anti-XINCENP and anti-XAurora B (MacCallum et al. 2002); and anti-Plx1 (Losada et al. 2002). A mouse monoclonal antibody against the catalytic subunit of PP2A (clone 1D6) was purchased from Upstate.
Preparation and depletion of Xenopus egg extracts
Cytostatic factor (CSF)-arrested, low-speed supernatants (LSS) of Xenopus eggs were prepared in XBE2 buffer [10 mM K-Hepes (pH 7.7), 0.1 M KCl, 2 mM MgCl2, 0.1 mM CaCl2, 5 mM EGTA, and 50 mM sucrose] as described (Losada et al. 1998). Interphase LSS were generated by addition of 100 μg/ml cycloheximide and 0.4 mM CaCl2 to CSF-arrested LSS followed by incubation at 22°C for 30 min. To deplete 50 μl of extract, the following antibodies were bound to 25 μl of Dynabeads protein A (Dynal): for Bub1 depletion, 8 μg of rabbit polyclonal anti-Xenopus Bub1 (40 min); for cohesin depletion, 4 μg of anti-SMC3 plus 2.5 μg of anti-SMC1 (two rounds of 40 min); for Aurora B depletion, 3.5 μg of anti-XINCENP plus 3.5 μg of anti-XAurora B (two rounds of 40 min); and for Polo depletion, 8 μg of anti-Polo (two rounds of 40 min). For the depletion of Sgo, 50 μl of rabbit polyclonal serum was bound to 25 μl of Affi-prep protein A Support (BIO-RAD). In all cases, mock depletions were performed using beads coated with pre-immune rabbit IgG (Sigma).
Chromosome assembly and analysis
Sperm nuclei (800–1,000 nuclei per microliter) were incubated at 22°C for 2 h with freshly depleted interphase LSS supplemented with energy mix (1 mM MgATP, 10 mM creatine phosphate, and 50 μg/ml creatine kinase). The extracts were driven into mitosis by addition of an equal volume of CSF-arrested extract and incubated at 22°C for another 2 h. For biochemical analysis of chromatin-bound proteins, samples were diluted tenfold with XBE2 buffer containing 0.25% Triton X-100. The chromatin fractions were isolated by centrifugation through a sucrose cushion (30% sucrose in XBE2) in an Eppendorf 5417R centrifuge at 10,000×g for 15 min. For immunofluorescence, the assembly mixtures were fixed with ten volumes of 2% paraformaldehyde in XBE2 containing 0.5% Triton X-100 for 10 min and centrifuged onto coverslips. Primary antibodies were used at 1–2 μg/ml, whereas Cy3 or fluorescein isothiocyanate-conjugated donkey anti-rabbit or anti-mouse secondary antibodies (Jackson ImmunoResearch) were used at 1:200. DNA was counter-stained with 1 μg/ml 4′,6-diamidino-2-phenylindole (DAPI). In some cases, double immunostaining with two rabbit polyclonal antibodies was performed with one of the antibodies directly labeled with a fluorophore, as described previously (Ono et al. 2004). A Leica DM6000 microscope was used to obtain grayscale images, which were later pseudo-colored and merged using Adobe Photoshop.
Dephosphorylation assay
Soluble cohesin was purified from HeLa cell nuclear extracts with anti-SMC3 as described (Losada et al. 2000) and was first incubated with a Polo kinase fraction (or an Aurora B kinase fraction as control) immunoprecipitated from a CSF extract on Dynabeads protein A. After 20 min, the Polo-incubated cohesin was incubated with a Sgo fraction immunoprecipitated from a CSF extract in the presence or absence of 1 μM okadaic acid. To detect phosphorylation/dephosphorylation of cohesin, the mobility of the SA1 and SA2 subunits were analyzed by immunoblotting.
Results
Xenopus Sgo binds throughout chromatin in interphase but accumulates at centromeres in mitosis
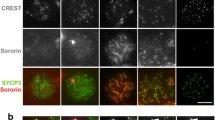
We developed a rabbit polyclonal and a mouse monoclonal antibody against Xenopus Sgo. Both recognize a single band of approximately 100 kDa by Western blot analysis of interphase egg extracts (Fig. 1a). We used these antibodies to study Sgo localization by immunofluorescence in nuclei and chromosomes assembled in vitro in the egg extracts from sperm DNA. Time course analyses show that Sgo distributes all over chromatin in interphase, whereas it quickly accumulates at the centromeric regions upon entry in mitosis (Fig. 1b). Binding to interphase chomatin is not necessary for subsequent centromeric localization in mitosis, since Sgo is found at the centromeres of unreplicated chromosomes assembled directly in CSF extracts (data not shown). Consistently, we also found that neither DNA replication nor the presence of cohesin is required for proper recruitment of Sgo to centromeres in mitosis (Electronic supplementary material Fig. 1).
Sgo localization throughout the cell cycle. a Immunoblot analysis of an egg extract with a mouse monoclonal (Ms) and a rabbit polyclonal (Rb) affinity-purified antibody against Xenopus Sgo. b Sperm chromatin was incubated with an interphase extract to assemble nuclei, and after 120 min, CSF addition triggered entry in mitosis. Samples taken at the times indicated were analyzed by immunofluorescence with antibodies against Sgo (red) and CENP-A (green). DNA was counterstained with DAPI (blue). The insets on the right show the progressive resolution of a single pair of sister centromeres. Bars, 1 μm (insets) and 10 μm
Centromeric cohesion is loosened in chromosomes assembled in the absence of Sgo
To test the effect of removing Sgo on sister chromatid cohesion, replicated chromosomes were assembled in mock-depleted extracts and extracts immunodepleted of more than 95% of Sgo or its targeting kinase, Bub1 (Fig. 2a). First, we checked that both depletions resulted in the absence of Sgo from centromeres (Fig. 2b). To evaluate centromeric cohesion, we measured the distance between sister centromeres labeled with anti-CENP-A. To be sure that we were looking at a centromere pair within a chromosome and not at two centromeres from different chromosomes, these were also labeled with anti-condensin I to reveal the axes of the sister chromatids (Fig. 2c). Our analysis shows that chromosomes with increased distances between sister centromeres are more often found in Sgo-depleted extracts or Bub1-depleted extracts than in control extracts. Thus, we conclude that Sgo function is important for proper centromeric cohesion in chromosomes assembled in Xenopus extracts.
Defects in centromeric cohesion upon Sgo depletion. a Extracts were immunodepleted with control IgG (lanes 1–5, mock-dep), anti-Bub1 (lane 6, Bub1-dep), or anti-Sgo (lane 7, Sgo-dep). To estimate the efficiency of depletion, 1-μl aliquots of the depleted extracts were analyzed by immunoblotting alongside different amounts of the mock-depleted extract (expressed as percentage of 1 μl). b Replicated chromosomes assembled in the extracts were fixed and stained with antibodies against Sgo (red) and Bub1 (green) and counterstained with DAPI (blue). c Replicated chromosomes assembled in the same extracts were stained with antibodies against condensin I subunit XCAP-G (red) and CENP-A (yellow). The distance between sister centromeres was measured for more than 200 chromosomes for each condition (in arbitrary units) and plotted. Bars, 1 μm (insets) and 10 μm
Arm cohesion is not affected by depletion of Sgo
We next analyzed the morphology of individual chromosomes assembled in extracts immunodepleted of Sgo. For comparison, we included in our analyses chromosomes isolated from extracts depleted of cohesin (Fig. 3a). In all cases, condensin I staining was used to highlight the axis of each sister chromatid. Close observation of these chromosomes showed no indication of arm cohesion defects in Sgo-depleted extracts. In contrast, the chromosomes assembled in the absence of cohesin in which two sister chromatids could be recognized showed increased distances between sisters (Fig. 3b). Quantitative analysis of the distance between sister chromatids in individual chromosomes assembled in each condition confirmed that Sgo is not required for arm cohesion in Xenopus extracts, similar to what has been described in Drosophila and mouse and opposite to the results in HeLa cells (Fig. 3c). BLAST searches of the yet incompletely sequenced genomes of Xenopus laevis and Xenopus tropicalis do not show evidence for additional Sgo proteins with redundant functions in cohesion that could explain the lack of arm cohesion defects.
Arm cohesion is maintained in the absence of Sgo. a Extracts were immunodepleted with control IgG (lanes 1–4, mock-dep) or a mixture of anti-SMC1 and anti-SMC3 (lane 5) and the extent of depletion analyzed as in Fig. 2 with an antibody against cohesin subunit Scc1 and with anti-Aurora B (as loading control). b Chromosomes from the depleted extracts were stained with anti-XCAP-G. For the sake of the presentation, pictures of individual chromosomes found in different fields of the same coverslip have been assembled together. Bar, 10 μm. c The distance between the axes of the sister chromatids was measured along the length of several chromosomes (n ≥ 5 for each condition) and plotted
Cohesin distribution is affected by Sgo depletion
To investigate whether the effects that we see on centromeric cohesion in the absence of Sgo or Bub1 are directly related to cohesin, chromosomes assembled under these conditions were stained with a cohesin antibody. A small amount of cohesin is distributed over the arms of the mitotic chromosomes isolated from mock-depleted extracts, and some accumulation of cohesin can be detected at centromeric regions co-stained with CENP-A (Fig. 4a). This accumulation is never seen in the chromosomes from Sgo- or Bub1-depleted extracts. Thus, the defects in centromeric cohesion observed in these chromosomes are most likely the result of the lack of enrichment of cohesin at centromeres in the absence of Sgo. On the other hand, Kitajima et al. (2005) have reported that depletion of Bub1 by siRNA in HeLa cells results in centromeric cohesion defects while arm cohesion is maintained. These authors suggested that the presence of increased amounts of Sgo along chromosome arms provoked ectopic protection of cohesin. In Xenopus extracts, however, there is little Sgo on mitotic chromatin after Bub1 depletion, either at arms or at centromeres (Fig. 2b; Boyarchuk et al. 2007), and we did not detect more cohesin on chromosome arms (Fig. 4a).
Sgo depletion prevents centromeric accumulation of cohesin and the CPC. a Replicated chromosomes were stained with anti-SA1 cohesin subunit (green), anti-CENP-A (red), and DAPI (blue). Cohesin accumulates at centromeres only in the control chromosomes (arrowheads). b Chromosomes assembled in the presence or absence of Sgo were stained with antibodies against INCENP (green), Sgo (red), and DAPI (blue). Bars, 10 μm
Unexpectedly, we observed a modest increase in the staining of cohesin in Sgo-depleted chromosomes. This increase is not due to abnormal loading of cohesin in interphase (compare the amount of cohesin Scc1 in lanes 3 and 6 of Fig. 6e). Instead, it must be the consequence of impaired cohesin dynamics in mitosis. Polo and Aurora B contribute independently to the prophase release of cohesin. We therefore asked whether the effects seen on this pathway upon depletion of Sgo could be specifically linked to any of these two kinases. Aurora B is present in Xenopus extracts in the so-called chromosomal passenger complex (CPC), together with INCENP, survivin, and Dasra/Borealin (Sampath et al. 2004). The CPC is required for proper localization of Sgo and Bub1 in early mitosis (Vigneron et al. 2004; Resnick et al. 2006; Boyarchuk et al. 2007). Importantly, we found that depletion of Sgo also alters the localization of CPC, but not the localization of Polo (Fig. 4b and data not shown). Next, chromosomes were assembled in extracts depleted of Sgo, Aurora B, Polo, or the corresponding pairwise combinations (Fig. 5c). Quantitation of the immunofluorescent signals observed in the chromosomes assembled in each condition shows that co-depletion of Sgo and Aurora B or Polo increases the fraction of cohesin left on mitotic chromosomes with respect to either single depletion (Fig. 5a,b). Thus, although Aurora B and Sgo require each other for proper localization, this interdependence alone does not explain the role of either one in cohesin dissociation.
Cohesin dissociation in mitosis is affected by Sgo depletion. a Chromosomes assembled in the indicated extracts were stained with anti-Sgo (red), anti-SA1 cohesin subunit (green), and DAPI (blue). b Fluorescence intensity of SA1 staining (in arbitrary units) was measured for ten nuclei (masses of chromosomes) in each condition using ImageJ software (http://rsbweb.nih.gov/ij/index.html). c One-μl aliquots of each depleted extract were analyzed by immunoblotting and compared with a mock-depleted extract to estimate the extent of depletion. Bars, 10 μm
Anomalous PP2A distribution in the absence of Sgo impairs cohesin release
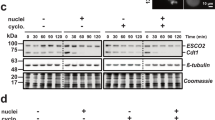
Sgo associates with PP2A in yeast and human cells, and this association is important for cohesin protection (Kitajima et al. 2006; Riedel et al. 2006; Tang et al. 2006). In HeLa cells depleted of Sgo1 by siRNA, cohesion is completely lost even though the centromeric localization of PP2A is preserved, a result that led Kitajima et al. (2006) to suggest that Sgo could have an additional role and provide protection by itself, maybe through physical association with cohesin. We have found no evidence for a direct physical interaction of cohesin and Sgo in Xenopus extracts. We do observe co-immunoprecipitation of the catalytic subunit of PP2A (PP2Ac) and Sgo (Fig. 6a). Furthermore, this Sgo-immunoprecipitated fraction can reverse the phosphorylation by Polo of the SA1 and SA2 subunits of intact cohesin complexes, a reaction that is inhibited by okadaic acid (Fig. 6b). Depletion of 95% of Sgo does not remove an appreciable fraction of PP2Ac from the extract (Fig. 6c). However, the phosphatase accumulates at centromeres in chromosomes assembled in mock-depleted extracts, and unlike HeLa cells, this accumulation is prevented in the absence of Sgo (Fig. 6d). The fact that the same amount of PP2Ac is found in the chromatin fraction of mitotic chromosomes assembled in mock-depleted and Sgo-depleted extracts argues for the presence of increased amounts of PP2Ac along chromosome arms in the absence of Sgo (Fig. 6e). We reasoned that under this condition, PP2A could hinder cohesin release. Consistent with this hypothesis, treatment of the chromosomes assembled in Sgo-depleted extracts with okadaic acid results in total dissociation of cohesin (Fig. 6f). Thus, we envision that PP2A associates with Sgo on chromatin and together relocate from chromosome arms to centromeres at the onset of mitosis (Electronic supplementary material Fig. 2). In the absence of Sgo, PP2A is left along chromosome arms and impairs the dissociation of cohesin. On the other hand, the fact that inhibition of PP2A by okadaic acid treatment results in complete removal of cohesin independently of Sgo supports the critical role of the phosphatase in cohesin protection. It is most likely that PP2A counteracts the phosphorylation of cohesin SA subunits in mitosis in vivo, as suggested by our in vitro results.
PP2A mediates the role of Sgo in cohesin protection and release. a Immunoprecipitation reactions from Xenopus extract with control IgG (lane 2) or anti-Sgo (lane 3) were analyzed by immunoblotting with anti-SMC3 and anti-PP2Ac. An aliquot of the extract was also loaded (lane 1). b A cohesin fraction purified from human cell extracts was incubated first with Aurora B (lane 1) or Polo (lanes 2–4) isolated on beads from mitotic extracts and then with Sgo immunoprecipitated also from mitotic extracts in the absence (lane 3) or presence (lane 4) of okadaic acid. The soluble fraction containing cohesin was then analyzed by immunoblotting with anti-SA1 and anti-SA2. The mobility shift evidences phosphorylation. c Aliquots of a mock-depleted (lane 1) and a Sgo-depleted extract (lane 2) were analyzed by immunoblotting with Sgo and PP2A antibodies. d Chromosomes assembled in mock-depleted and Sgo-depleted extracts were stained with anti-PP2Ac (red) and anti-CENP-A (green) and DAPI (blue). e Samples of assembly mixtures prepared in mock and Sgo-depleted extracts were taken after 2 h in interphase (lanes 3 and 5). Mitosis was induced by addition of CSF extract and samples were taken after 2 h (lanes 4 and 6). Chromatin fractions were analyzed by immunoblotting with anti-Scc1 cohesin subunit, anti-PP2Ac, and anti-phospho H3. Topoisomerase II (topo II) was used as loading control. An aliquot of extract (lane 1) and a mock chromatin fraction without sperm (lane 2) were also analyzed. f Chromosomes assembled in mock and Sgo-depleted extracts were treated with or without okadaic acid for 1 h before being isolated and analyzed by immunofluorescence with anti-SA1 cohesin subunit (green) and anti-Sgo (red). Bars in d and f, 10 μm
Discussion
The role of Sgo in the regulation of mitotic cohesion has been addressed in a number of recent studies, mainly in yeast and human cells (Salic et al. 2004; McGuinness et al. 2005; Vaur et al. 2005; Kitajima et al. 2006; Tang et al. 2006; Kawashima et al. 2007; Vanoosthuyse et al. 2007; Lee et al. 2008). However, this is the first time that the Xenopus egg cell free system, in which vertebrate cohesin was initially characterized, is used. Our results show that Sgo is a protector of cohesin around centromeres in chromosomes assembled in these extracts, as we see a perfect correlation between increased distance between sister centromeres and decreased cohesin staining in the absence of Sgo or its targeting kinase Bub1. This is important because it corroborates the previously proposed role of Sgo as a protector of centromeric cohesin from the prophase dissociation pathway in a “clean” system (Xenopus extracts) without interference of the spindle assembly checkpoint in which Sgo also participates (Indjeian et al. 2005; Huang et al. 2007). In contrast, the role of Sgo as a protector of cohesin along chromosome arms, proposed for human cells, seems not to be relevant in this system (Nakajima et al. 2007). It is possible that mechanistic differences in the regulation of cohesion exist between species or even between primary and transformed cells. These differences could affect cohesin per se or some other aspect of cohesion, e.g., the contribution of catenation. However, since arm cohesion is defective in the absence of cohesin in Xenopus egg extracts (Losada et al. 1998), it is clear that catenation alone cannot account for arm cohesion in the absence of Sgo, and cohesin must play a major role in keeping the sister chromatids together along the entire length of the chromosome. Regarding regulation of cohesin, the prophase dissociation pathway could be less efficient in Xenopus extracts compared with transformed human cells, perhaps due to increased amount and/or activity of the phosphatase PP2A. It is even conceivable that a fraction of PP2A is present along chromosome arms independently of Sgo and contributes to maintain arm cohesion in the chromosomes assembled in these extracts.
It has been previously shown that Aurora B is required for relocalization of Sgo from arms to centromeres (Resnick et al. 2006; Boyarchuk et al. 2007). Our results suggest that conversely, Sgo also contributes to the proper accumulation of the CPC at the inner centromere region. Both in S. cerevisiae and Schizosaccharomyces pombe, Sgo proteins have been reported to regulate at least some CPC components (Kawashima et al. 2007; Vanoosthuyse et al. 2007; Yu and Koshland 2007). The interdependence of Aurora B and Sgo localization at centromeres remains to be investigated in detail, but it is likely to underlie the role of both proteins as sensors of tension across the sister kinetochores (Goulding and Earnshaw 2005). This task appears to be more specific of Sgo2 in human and fission yeast (Kawashima et al. 2007). We suspect that a single Sgo protein in Xenopus may carry out the tasks assumed by Sgo1 and Sgo2 in other organisms.
On the other hand, we unexpectedly found increased amounts of cohesin on chromosome arms after Sgo depletion. This could be due to a problem in cohesin dissociation or could result from redistribution of cohesin from centromeres to arms in the absence of centromeric Sgo. We favor the first possibility. No report to date has shown that cohesin is enriched at centromeres prior to mitosis in metazoa, although this is the case in yeast (e.g., Lengronne et al. 2004). Thus, centromeric cohesin enrichment may be the result of preferential dissociation of cohesin from chromosome arms and protection at centromeres. In this scenario, redistribution is unlikely. In addition, depletion of Bub1 does not result in a similar phenotype of increased arm cohesin, even though Sgo is absent from centromeres also in this case. A fraction of PP2A left along the arms of chromosomes assembled in the absence of Sgo most likely counteracts cohesin phosphorylation by Polo and thereby prevents its release. Consistent with this interpretation, co-depletion of both Polo and Sgo increases only modestly the amount of cohesin left on chromosomes with respect to the single depletion of the kinase. In the case of Aurora B, Sgo appears not to be the only relevant target of the kinase in this prophase pathway. Other possible targets could be condensin, heterochromatin protein 1 (HP1), or some other protein yet to be identified (Fischle et al. 2005; Hirota et al. 2005; Lipp et al. 2007).
References
Boyarchuk Y, Salic A, Dasso M, Arnaoutov A (2007) Bub1 is essential for assembly of the functional inner centromere. J Cell Biol 176:919–928
Brar GA, Kiburz BM, Zhang Y, Kim JE, White F, Amon A (2006) Rec8 phosphorylation and recombination promote the step-wise loss of cohesins in meiosis. Nature 441:532–536
Fischle W, Tseng BS, Dormann HL, Ueberheide BM, Garcia BA, Shabanowitz J, Hunt DF, Funabiki H, Allis CD (2005) Regulation of HP1-chromatin binding by histone H3 methylation and phosphorylation. Nature 438:1116–1122
Gandhi R, Gillespie PJ, Hirano T (2006) Human Wapl is a cohesin-binding protein that promotes sister-chromatid resolution in mitotic prophase. Curr Biol 16:2406–2417
Goulding SE, Earnshaw WC (2005) Shugoshin: a centromeric guardian senses tension. Bioessays 27:588–591
Hauf S, Waizenegger IC, Peters JM (2001) Cohesin cleavage by separase required for anaphase and cytokinesis in human cells. Science 293:1320–1323
Hauf S, Roitinger E, Koch B, Dittrich CM, Mechtler K, Peters JM (2005) Dissociation of cohesin from chromosome arms and loss of arm cohesion during early mitosis depends on phosphorylation of SA2. PLoS Biol 3:e69
Hirano T, Kobayashi R, Hirano M (1997) Condensins, chromosome condensation protein complexes containing XCAP-C, XCAP-E and a Xenopus homolog of the Drosophila barren protein. Cell 89:511–521
Hirota T, Lipp JJ, Toh BH, Peters JM (2005) Histone H3 serine 10 phosphorylation by Aurora B causes HP1 dissociation from heterochromatin. Nature 438:1176–1180
Huang H, Feng J, Famulski J, Rattner JB, Liu ST, Kao GD, Muschel R, Chan GK, Yen TJ (2007) Tripin/hSgo2 recruits MCAK to the inner centromere to correct defective kinetochore attachments. J Cell Biol 177:413–424
Indjeian VB, Stern BM, Murray AW (2005) The centromeric protein Sgo1 is required to sense lack of tension on mitotic chromosomes. Science 307:130–133
Kawashima SA, Tsukahara T, Langegger M, Hauf S, Kitajima TS, Watanabe Y (2007) Shugoshin enables tension-generating attachment of kinetochores by loading Aurora to centromeres. Genes Dev 21:420–435
Kerrebrock AW, Moore DP, Wu JS, Orr-Weaver TL (1995) Mei-S332, a Drosophila protein required for sister-chromatid cohesion, can localize to meiotic centromere regions. Cell 83:247–256
Kitajima TS, Kawashima SA, Watanabe Y (2004) The conserved kinetochore protein shugoshin protects centromeric cohesion during meiosis. Nature 427:510–517
Kitajima TS, Hauf S, Ohsugi M, Yamamoto T, Watanabe Y (2005) Human Bub1 defines the persistent cohesion site along the mitotic chromosome by affecting shugoshin localization. Curr Biol 15:353–359
Kitajima TS, Sakuno T, Ishiguro K, Iemura S, Natsume T, Kawashima SA, Watanabe Y (2006) Shugoshin collaborates with protein phosphatase 2A to protect cohesin. Nature 441:46–52
Kudo NR, Wassmann K, Anger M, Schuh M, Wirth KG, Xu H, Helmhart W, Kudo H, McKay M, Maro B, Ellenberg J, de Boer P, Nasmyth K (2006) Resolution of chiasmata in oocytes requires separase-mediated proteolysis. Cell 126:135–146
Kueng S, Hegemann B, Peters BH, Lipp JS, Schleiffer A, Mechtler K, Peters JM (2006) Wapl controls the dynamic association of cohesin with chromatin. Cell 127:955–967
LeBlanc HN, Tang TT, Wu JS, Orr-Weaver TL (1999) The mitotic centromeric protein MEI-S332 and its role in sister-chromatid cohesion. Chromosoma 108:401–411
Lee J, Kitajima TS, Tanno Y, Yoshida K, Morita T, Miyano T, Miyake M, Watanabe Y (2008) Unified mode of centromeric protection by shugoshin in mammalian oocytes and somatic cells. Nat Cell Biol 10:42–52
Lengronne A, Katou Y, Mori S, Yokobayashi S, Kelly GP, Itoh T, Watanabe Y, Shirahige K, Uhlmann F (2004) Cohesin relocation from sites of chromosomal loading to places of convergent transcription. Nature 430:573–578
Lipp JJ, Hirota T, Poser I, Peters JM (2007) Aurora B controls the association of condensin I but not condensin II with mitotic chromosomes. J Cell Sci 120:1245–1255
Losada A, Hirano T (2005) Dynamic molecular linkers of the genome: the first decade of SMC proteins. Genes Dev 19:1269–1287
Losada A, Hirano M, Hirano T (1998) Identification of Xenopus SMC protein complexes required for sister chromatid cohesion. Genes Dev 12:1986–1997
Losada A, Yokochi T, Kobayashi R, Hirano T (2000) Identification and characterization of SA/Scc3p subunits in the Xenopus and human cohesin complexes. J Cell Biol 150:405–416
Losada A, Hirano M, Hirano T (2002) Cohesin release is required for sister chromatid resolution, but not for condensin-mediated compaction, at the onset of mitosis. Genes Dev 16:3004–3016
Losada A, Yokochi T, Hirano T (2005) Functional contribution of Pds5 to cohesin-mediated cohesion in human cells and Xenopus egg extracts. J Cell Sci 118:2133–2141
MacCallum DE, Losada A, Kobayashi R, Hirano T (2002) ISWI remodeling complexes in Xenopus egg extracts: identification as major chromosomal components that are regulated by INCENP-aurora B. Mol Biol Cell 13:25–39
Marston AL, Amon A (2004) Meiosis: cell-cycle controls shuffle and deal. Nat Rev Mol Cell Biol 5:983–997
McGuinness BE, Hirota T, Kudo NR, Peters JM, Nasmyth K (2005) Shugoshin prevents dissociation of cohesin from centromeres during mitosis in vertebrate cells. PLoS Biol 3:e86
Nakajima M, Kumada K, Hatakeyama K, Noda T, Peters JM, Hirota T (2007) The complete removal of cohesin from chromosome arms depends on separase. J Cell Sci 120:4188–4196
Nasmyth K, Haering CH (2005) The structure and function of SMC and kleisin complexes. Annu Rev Biochem 74:595–648
Ono T, Fang Y, Spector DL, Hirano T (2004) Spatial and temporal regulation of condensins I and II in mitotic chromosome assembly in human cells. Mol Biol Cell 15:3296–3308
Perera D, Tilston V, Hopwood JA, Barchi M, Boot-Handford RP, Taylor SS (2007) Bub1 maintains centromeric cohesion by activation of the spindle checkpoint. Dev Cell 13:566–579
Rabitsch KP, Gregan J, Schleiffer A, Javerzat JP, Eisenhaber F, Nasmyth K (2004) Two fission yeast homologs of Drosophila Mei-S332 are required for chromosome segregation during meiosis I and II. Curr Biol 14:287–301
Resnick TD, Satinover DL, MacIsaac F, Stukenberg PT, Earnshaw WC, Orr-Weaver TL, Carmena M (2006) INCENP and Aurora B promote meiotic sister chromatid cohesion through localization of the shugoshin MEI-S332 in Drosophila. Dev Cell 11:57–68
Riedel CG, Katis VL, Katou Y, Mori S, Itoh T, Helmhart W, Galova M, Petronczki M, Gregan J, Cetin B, Mudrak I, Ogris E, Mechtler K, Pelletier L, Buchholz F, Shirahige K, Nasmyth K (2006) Protein phosphatase 2A protects centromeric sister chromatid cohesion during meiosis I. Nature 441:53–61
Salic A, Waters JC, Mitchison TJ (2004) Vertebrate shugoshin links sister centromere cohesion and kinetochore microtubule stability in mitosis. Cell 118:567–578
Sampath SC, Ohi R, Leismann O, Salic A, Pozniakovski A, Funabiki H (2004) The chromosomal passenger complex is required for chromatin-induced microtubule stabilization and spindle assembly. Cell 118:187–202
Sumara I, Vorlaufer E, Stukenberg PT, Kelm O, Redemann N, Nigg EA, Peters JM (2002) The dissociation of cohesin from chromosomes in prophase is regulated by Polo-like kinase. Mol Cell 9:515–525
Tang Z, Sun Y, Harley SE, Zou H, Yu H (2004) Human Bub1 protects centromeric sister-chromatid cohesion through Shugoshin during mitosis. Proc Natl Acad Sci U S A 101:18012–18017
Tang Z, Shu H, Qi W, Mahmood NA, Mumby MC, Yu H (2006) PP2A is required for centromeric localization of Sgo1 and proper chromosome segregation. Dev Cell 10:575–585
Terret ME, Wassmann K, Waizenegger I, Maro B, Peters JM, Verlhac MH (2003) The meiosis I-to-meiosis II transition in mouse oocytes requires separase activity. Curr Biol 13:1797–1802
Vagnarelli P, Morrison C, Dodson H, Sonoda E, Takeda S, Earnshaw WC (2004) Analysis of Scc1-deficient cells defines a key metaphase role of vertebrate cohesin in linking sister kinetochores. EMBO Rep 5:167–171
Vanoosthuyse V, Prykhozhij S, Hardwick KG (2007) Shugoshin 2 regulates localization of the chromosomal passenger proteins in fission yeast mitosis. Mol Biol Cell 18:1657–1669
Vass S, Cotterill S, Valdeolmillos AM, Barbero JL, Lin E, Warren WD, Heck MM (2003) Depletion of Drad21/Scc1 in Drosophila cells leads to instability of the cohesin complex and disruption of mitotic progression. Curr Biol 13:208–218
Vaur S, Cubizolles F, Plane G, Genier S, Rabitsch PK, Gregan J, Nasmyth K, Vanoosthuyse V, Hardwick KG, Javerzat JP (2005) Control of Shugoshin function during fission-yeast meiosis. Curr Biol 15:2263–2270
Vigneron S, Prieto S, Bernis C, Labbe JC, Castro A, Lorca T (2004) Kinetochore localization of spindle checkpoint proteins: who controls whom? Mol Biol Cell 15:4584–4596
Waizenegger IC, Hauf S, Meinke A, Peters JM (2000) Two distinct pathways remove mammalian cohesin from chromosome arms in prophase and from centromeres in anaphase. Cell 103:399–410
Yu HG, Koshland D (2007) The Aurora kinase Ipl1 maintains the centromeric localization of PP2A to protect cohesin during meiosis. J Cell Biol 176:911–918
Acknowledgments
We thank A. Salic (Harvard Med. School) for Xenopus Sgo cDNA and the antibody used in our initial studies, J. Maller (University of Colorado Med. School) for Bub1 cDNAs, M. Rodríguez for excellent technical help, and J. Méndez (CNIO) for critically reading the manuscript. We are funded by the Spanish Ministry of Science and Education (grants BFU2007-66627 and CSD-Inesgen and a Ramón y Cajal Contract for A.L.), the Fundación Caja Madrid and the EU (Epigenome NoE and MIRG-31126).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by F. Uhlmann
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Sgo targeting does not require replication and does not depend on cohesin. a Sperm chromatin was incubated in an interphase egg extract in the presence of biotin-16-dUTP and allowed to replicate. An equal volume of CSF was added to trigger entry into mitosis and chromosomes were isolated and stained with anti-Sgo (green) and avidin-Cy3 (red). DNA was counterstained with DAPI (blue). Although most nuclei were replicated, as judged by intense Cy3 staining, some nuclei within the sample did not, allowing us to compare the targeting of Sgo in the presence/absence of replication. No differences were found. b Chromosomes were assembled as described in a in a mock-depleted extract and extract depleted of cohesin and stained with anti-Sgo (green), anti-Bub1 (red), and DAPI (blue). Sgo localized at the centromeres in both cases. The lack of cohesion between sister centromeres in the absence of cohesin resulted in a clear splitting and separation of the Sgo and Bub1 signals corresponding to sister centromeres. Bars, 10 μm. (GIF 430 KB)
Supplementary Fig. 2
Partial co-localization of Sgo and PP2Ac in nuclei and chromosomes assembled in egg extracts. Sperm chromatin was incubated with an interphase extract to assemble nuclei, and after 120 min, CSF addition triggered entry in mitosis. Samples taken at the times indicated were analyzed by immunofluorescence with antibodies against PP2Ac (red) and Sgo (green). DNA was counterstained with DAPI (blue). Both proteins relocalize to the centromeres in mitosis. Bar, 10 μm. (GIF 573 KB)
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Rivera, T., Losada, A. Shugoshin regulates cohesion by driving relocalization of PP2A in Xenopus extracts. Chromosoma 118, 223–233 (2009). https://doi.org/10.1007/s00412-008-0190-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-008-0190-4