Abstract
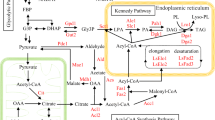
Lipomyces starkeyi is an oil-producing yeast that can produce triacylglycerol (TAG) from glycerol as a carbon source. The TAG was mainly produced after nitrogen depletion alongside reduced cell proliferation. To obtain clues for enhancing the TAG production, cell metabolism during the TAG-producing phase was characterized by metabolomics with 13C labeling. The turnover analysis showed that the time constants of intermediates from glycerol to pyruvate (Pyr) were large, whereas those of tricarboxylic acid (TCA) cycle intermediates were much smaller than that of Pyr. Surprisingly, the time constants of intermediates in gluconeogenesis and the pentose phosphate (PP) pathway were large, suggesting that a large amount of the uptaken glycerol was metabolized via the PP pathway. To synthesize fatty acids that make up TAG from acetyl-CoA (AcCoA), 14 molecules of nicotinamide adenine dinucleotide phosphate (NADPH) per C16 fatty acid molecule are required. Because the oxidative PP pathway generates NADPH, this pathway would contribute to supply NADPH for fatty acid synthesis. To confirm that the oxidative PP pathway can supply the NADPH required for TAG production, flux analysis was conducted based on the measured specific rates and mass balances. Flux analysis revealed that the NADPH necessary for TAG production was supplied by metabolizing 48.2% of the uptaken glycerol through gluconeogenesis and the PP pathway. This result was consistent with the result of the 13C-labeling experiment. Furthermore, comparison of the actual flux distribution with the ideal flux distribution for TAG production suggested that it is necessary to flow more dihydroxyacetonephosphate (DHAP) through gluconeogenesis to improve TAG yield.






Similar content being viewed by others
References
Amaretti A, Raimondia S, Leonardia A, Rossia M (2012) Candida Freyschussii: an oleaginous yeast producing lipids from glycerol. Chem Eng Trans 27:139–144. https://doi.org/10.3303/CET1227024
An PNT, Yamaguchi M, Fukusaki E (2017) Metabolic profiling of Drosophila melanogaster metamorphosis: a new insight into the central metabolic pathways. Metabolomics 13:29. https://doi.org/10.1007/s11306-017-1167-1
Barnes LD, Kuehn GD, Atkinson DE (1971) Yeast diphosphopyridine nucleotide specific isocitrate dehydrogenase. Purification and some properties. Biochemistry 10:3939–3944. https://doi.org/10.1021/bi00797a022
Canelas AB, Ras C, ten Pierick A, van Dam JC, Heijnen JJ, van Gulik WM (2008) Leakage-free rapid quenching technique for yeast metabolomics. Metabolomics 4:226–239. https://doi.org/10.1007/s11306-008-0116-4
Edwards JS, Ibarra RU, Palsson BO (2001) In silico predictions of Escherichia coli metabolic capabilities are consistent with experimental data. Nat Biotechnol 19:125–130. https://doi.org/10.1038/84379
Evans CT, Ratledge C (1985) The role of NAD+: isocitrate dehydrogenase in lipid accumulation by the oleaginous yeast Rhodosporidium toruloides CBS 14. Can J Microbiol 31:845–850. https://doi.org/10.1139/m85-157
Feist AM, Palsson BO (2010) The biomass objective function. Curr Opin Microbiol 13:344–349. https://doi.org/10.1016/j.mib.2010.03.003
Goncalves EC, Wilkie AC, Kirst M, Rathinasabapathi B (2016) Metabolic regulation of triacylglycerol accumulation in the green algae: identification of potential targets for engineering to improve oil yield. Plant Biotechnol J 14:1649–1660. https://doi.org/10.1111/pbi.12523
Habe H, Shimada Y, Yakushi T, Hattori H, Ano Y, Fukuoka T, Kitamoto D, Itagaki M, Watanabe K, Yanagishita H, Matsushita K, Sakaki K (2009) Microbial production of glyceric acid, an organic acid that can be mass produced from glycerol. Appl Environ Microbiol 75:7760–7766. https://doi.org/10.1128/AEM.01535-09
Habe H, Sato S, Fukuoka T, Kitamoto D, Yakushi T, Matsushita K, Sasaki K (2011) Membrane-bound alcohol dehydrogenase is essential for glyceric acid production in Acetobacter tropicalis. J Oleo Sci 60:489–494. https://doi.org/10.5650/jos.60.489
Hasunuma T, Harada K, Miyazawa S, Kondo A, Fukusaki E, Miyake C (2010) Metabolic turnover analysis by a combination of in vivo 13C-labelling from 13CO2 and metabolic profiling with CE-MS/MS reveals rate-limiting steps of the C3 photosynthetic pathway in Nicotiana tabacum leaves. J Exp Bot 61:1041–1051. https://doi.org/10.1093/jxb/erp374
Hirayama A, Kami K, Sugimoto M, Sugawara M, Toki N, Onozuka H, Kinoshita T, Saito N, Ochiai A, Tomita M, Esumi H, Soga T (2009) Quantitative metabolome profiling of colon and stomach cancer microenvironment by capillary electrophoresis time-of-flight mass spectrometry. Cancer Res 69:4918–4925. https://doi.org/10.1158/0008-5472.CAN-08-4806
Ichihashi K, Yuki D, Kurokawa H, Igarashi A, Yajima T, Fujiwara M, Maeno K, Sekiguchi S, Iwata M, Nishino H (2011) Dynamic analysis of phorbol esters in the manufacturing process of fatty acid methyl esters from Jatropha curcas seed oil. J Am Oil Chem Soc 88:851–861. https://doi.org/10.1007/s11746-010-1741-4
Ito T, Tanaka M, Shinkawa H, Nakada T, Ano Y, Kurano N, Soga T, Tomita M (2013) Metabolic and morphological changes of an oil accumulating trebouxiophycean alga in nitrogen-deficient conditions. Metabolomics 9:S178–S187. https://doi.org/10.1007/s11306-012-0463-z
Joyce AR, Palsson BO (2007) Toward whole cell modeling and simulation: comprehensive functional genomics through the constraint-based approach. Prog Drug Res 64:265–309. https://doi.org/10.1007/978-3-7643-7567-6_11
Korenaga H, Naganuma T, Uzuka Y, Tanaka K (1977) Determination of simple defined media for normal growth and lipid production of the yeast Lipomyces starkeyi. Nippon Nōgeikagaku Kaishi 51:449–455. https://doi.org/10.1271/nogeikagaku1924.51.7_449
Kurokawa H, Fukano Y, Ohki T, Naganuma T (2017) Conversion of glycerol to triacylglycerol by Lipomyces yeast. Oleoscience 17:127–133
Kurtzman CP, Boekhout JW (2011) The yeast, a taxonomic study, Fifth Edition. Elsevier
Lin J, Shen H, Tan H, Zhao X, Wu S, Hu C, Zhao ZK (2011) Lipid production by Lipomyces starkeyi cells in glucose solution without auxiliary nutrients. J Biotechnol 152:184–188. https://doi.org/10.1016/j.jbiotec.2011.02.010
Liu L, Zong M, Hu Y, Li N, Lou W, Wu H (2017) Efficient microbial oil production on crude glycerol by Lipomyces starkeyi AS 2.1560 and its kinetics. Process Biochem 58:230–238. https://doi.org/10.1016/j.procbio.2017.03.024
Matsumoto M, Nojima D, Tanaka T (2016) Development of outdoor mass cultivation for green oil production using marine microalgae. Bioindustry 33:10
Meesters PAEP, Huijberts GNM, Eggink G (1996) High-cell-density cultivation of the lipid accumulating yeast Cryptococcus curvatus using glycerol as a carbon source. Appl Microbiol Biotechnol 45:575–579. https://doi.org/10.1007/s002530050731
Nakagawa Y, Shinmia Y, Kosoa S, Tomishige K (2010) Direct hydrogenolysis of glycerol into 1,3-propanediol over rhenium-modified iridium catalyst. J Catal 272:191–194. https://doi.org/10.1016/j.jcat.2010.04.009
OECD/FAO (2011) OECD-FAO Agricultural Outlook 2011–2020, OECD Publishing and FAO. https://doi.org/10.1787/agr_outlook-2011-en. Accessed 19 Jan 2018
Pavel K, Petr S, Ivana B (2005) WO2005/021476. US Patent. https://patentscope.wipo.int/search/en/detail.jsf?docId=WO2005021476&redirectedID=true. Accessed 19 Jan 2018
Ratledge C (2014) The role of malic enzyme as the provider of NADPH in oleaginous microorganisms: a reappraisal and unsolved problems. Biotechnol Lett 36:1557–1568. https://doi.org/10.1007/s10529-014-1532-3
Sauer U, Canonaco F, Heri S, Perrenoud A, Fischer E (2004) The soluble and membrane-bound transhydrogenases UdhA and PntAB have divergent functions in NADPH metabolism of Escherichia coli. J Biol Chem 279:6613–6619. https://doi.org/10.1074/jbc.M311657200
Soga T, Igarashi K, Ito C, Mizobuchi K, Zimmermann HP, Tomita M (2009) Metabolomic profiling of anionic metabolites by capillary electrophoresis mass spectrometry. Anal Chem 81:6165–6174. https://doi.org/10.1021/ac900675k
Spier F, Buffon JG, Burkert CA (2015) Bioconversion of raw glycerol generated from the synthesis of biodiesel by different oleaginous yeasts: lipid content and fatty acid profile of biomass. Indian J Microbiol 55:415–422. https://doi.org/10.1007/s12088-015-0533-9
Sugimoto M, Wong DT, Hirayama A, Soga T, Tomita M (2010) Capillary electrophoresis mass spectrometry-based saliva metabolomics identified oral, breast and pancreatic cancer-specific profiles. Metabolomics 6:78–95. https://doi.org/10.1007/s11306-009-0178-y
Tang W, Zhang S, Wang Q, Tan H, Zhao ZK (2009) The isocitrate dehydrogenase gene of oleaginous yeast Lipomyces starkeyi is linked to lipid accumulation. Can J Microbiol 55:1062–1069. https://doi.org/10.1139/w09-063
Tapia VE, Anschau A, Coradini AL, Franco TT, Deckmann AC (2012) Optimization of lipid production by the oleaginous yeast Lipomyces starkeyi by random mutagenesis coupled to cerulenin screening. AMB Express 2:64–71. https://doi.org/10.1186/2191-0855-2-64
Tchakouteu SS, Kalantzi O, Gardeli C, Koutinas AA, Aggelis G, Papanikolaou S (2015) Lipid production by yeasts growing on biodiesel-derived crude glycerol: strain selection and impact of substrate concentration on the fermentation efficiency. J Appl Microbiol 118:911–927. https://doi.org/10.1111/jam.12736
Theodoulou FL, Holdsworth M, Baker A (2006) Peroxisomal ABC transporters. FEBS Lett 580:1139–1155. https://doi.org/10.1016/j.febslet.2005.12.095
Tsakona S, Kopsahelis N, Chatzifragkou A, Papanikolaou S, Kookos IK, Koutinas AA (2014) Formulation of fermentation media from flour-rich waste streams for microbial lipid production by Lipomyces starkeyi. J Biotechnol 189:36–45. https://doi.org/10.1016/j.jbiotec.2014.08.011
UN (2015) World population prospects: the 2015 revision. United Nations DESA/Population division. https://esa.un.org/unpd/wpp/publications/files/key_findings_wpp_2015.pdf. Accessed 19 Jan 2018
van Winden WA, Wittmann C, Heinzle E, Heijnen JJ (2002) Correcting mass isotopomer distributions for naturally occurring isotopes. Biotechnol Bioeng 80:477–479. https://doi.org/10.1002/bit.10393
Varma A, Palsson BO (1994) Stoichiometric flux balance models quantitatively predict growth and metabolic by-product secretion in wild-type Escherichia coli W3110. Appl Environ Microbiol 60:3724–3731
Wasylenko TM, Ahn WS, Stephanopoulos G (2015) The oxidative pentose phosphate pathway is the primary source of NADPH for lipid overproduction from glucose in Yarrowia lipolytica. Metab Eng 30:27–39. https://doi.org/10.1016/j.ymben.2015.02.007
Xu J, Zhao X, Wang W, Du W, Liu D (2012) Microbial conversion of biodiesel byproduct glycerol to triacylglycerols by oleaginous yeast Rhodosporidium toruloides and the individual effect of some impurities on lipid production. Biochem Eng J 65:30–36. https://doi.org/10.1016/j.bej.2012.04.003
Yamauchi H, Mori H, Kobayashi T, Shimizu S (1983) Mass production of lipids by Lipomyces starkeyi in microcomputer-aided fed-batch culture. J Ferment Technol 61:275–281
Yang F, Hanna MA, Sun R (2012) Value-added uses for crude glycerol–a byproduct of biodiesel production. Biotechnol Biofuels 5:13. https://doi.org/10.1186/1754-6834-5-13
Yang ZK, Niu YF, Ma YH, Xue J, Zhang MH, Yang WD, Liu JS (2013) Molecular and cellular mechanisms of neutral lipid accumulation in diatom following nitrogen deprivation. Biotechnol Biofuels 6:67. https://doi.org/10.1186/1754-6834-6-67
Yang X, Jin G, Gong Z, Shen H, Bai F, Zhao ZK (2014) Recycling biodiesel-derived glycerol by the oleaginous yeast Rhodosporidium toruloides Y4 through the two-stage lipid production process. Biochem Eng J 91:86–91. https://doi.org/10.1016/j.bej.2014.07.015
Zhang H, Zhang L, Chen H, Chen YQ, Ratledge C, Song Y, Chen W (2013) Regulatory properties of malic enzyme in the oleaginous yeast, Yarrowia lipolytica, and its non-involvement in lipid accumulation. Biotechnol Lett 35:2091–2098. https://doi.org/10.1007/s10529-013-1302-7
Zhu Z, Zhang S, Liu H, Shen H, Lin X, Yang F, Zhou YJ, Jin G, Ye M, Zou H, Zhao ZK (2012) A multi-omic map of the lipid-producing yeast Rhodosporidium toruloides. Nat Commun 3:1112. https://doi.org/10.1038/ncomms2112
Acknowledgements
This work was supported by the Japan Science and Technology Agency, Adaptable and Seamless Technology Transfer Program, JST A-STEP through target-driven R&D. We thank Dr. Takafumi Naganuma (Yamanashi University, Kofu, Japan) for providing advice on L. starkeyi culture. We thank Dr. Kenjiro Kami (Human Metabolome Technologies Inc., Tsuruoka, Japan) for providing advice on metabolome data interpretation.
Funding
This study was funded by Adaptable and Seamless Technology Transfer Program through target-driven R&D (A-STEP) from the Japan Science and Technology Agency (JST).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 179 kb)
Rights and permissions
About this article
Cite this article
Maruyama, Y., Toya, Y., Kurokawa, H. et al. Characterization of oil-producing yeast Lipomyces starkeyi on glycerol carbon source based on metabolomics and 13C-labeling. Appl Microbiol Biotechnol 102, 8909–8920 (2018). https://doi.org/10.1007/s00253-018-9261-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9261-5