Abstract
Anaerobic digestion (AD) uses a range of substrates to generate biogas, including energy crops such as globally abundant rice straw (RS). Unfortunately, RS is high in lignocellulosic material and has high to C:N ratios (~80:1), which makes it (alone) a comparatively poor substrate for AD. Co-digestion with dairy manure (DM) has been promoted as a method for balancing C:N ratios to improve RS AD whilst also treating another farm waste and co-producing a potentially useful fertiliser. However, past co-digestion studies have not directly compared RS AD microbial communities with and without DM additions, which has made it hard to assess all impacts of DM addition to RS AD processes. Here, four RS:DM ratios were contrasted in identical semi-continuous-fed AD bioreactors, and 100% RS was found to produce the highest specific methane yields (112 mL CH4/g VS/day; VS, volatile solids), which is over double yields achieved in the reactor with the highest DM content (30:70 RS:DM by mass; 48 mL CH4/g VS/day). To underpin these data, microbial communities were sequenced and characterised across the four reactors. Dominant operational taxonomic units (OTUs) in the 100% RS unit were Bacteroidetes/Firmicutes, whereas the 30:70 RS:DM unit was dominated by Proteobacteria/Spirochaetes, suggesting major microbial community shifts occur with DM additions. However, community richness was lowest with 100% RS (despite higher specific yields), suggesting particular OTUs may be more important to yields than microbial diversity. Further, ambient VFA and VS levels were significantly higher when no DM was added, suggesting DM-amended reactors may cope better with higher organic loading rates (OLR). Results show that RS AD without DM addition is feasible, although co-digestion with DM will probably allow higher OLRs, resulting in great RS throughput in farm AD units.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Anaerobic digestion (AD) uses a range of substrates to generate biogas, including food waste, energy crops, wastewater sludge and animal slurry. Lignocellulosic material, such as rice straw (RS), is not an ideal substrate as it is comparatively recalcitrant to AD and tends to generate lower biogas yields compared with many other crops (Mussoline et al. 2013). However, the demand for biogas in many parts of the world is increasing because the use of traditional fuels has many negative impacts, including increased deforestation (i.e. wood) and greater air pollution as particulate matter. Local biogas production, such as via RS AD, could reduce such pollution, offering a ‘cleaner’ alternative for lighting, heating and/or electricity generation (Krishania et al. 2013).
Various operating options exist for RS AD, but RS often has very high C:N ratios (Mussoline et al. 2013). High C:N ratios can result in incomplete digestion, whereas lower C:N ratios can lead to ammonia inhibition, making the targeted C:N ratio for AD roughly 25–30:1 (Monnet 2003; Ward et al. 2008). In low nitrogen systems, methanogens struggle to produce optimal biogas levels, whereas in lower C:N systems, excess nitrogen causes ammonia accumulation, pH increase, and inhibition (Monnet 2003). RS has a natural C:N ratio of up to 80:1 (Mussoline et al. 2013) and rebalancing C:N ratios to 25:1 can conditionally improves biogas yields (Lei et al. 2010). Co-digestion with manure is seen as a cost-effective method of balancing the C:N ratio in AD whilst using another large waste stream, potentially producing a rich fertiliser (Li et al. 2014a, 2015) and possibly increasing methane yields (Wang et al. 2014; Li et al. 2015). Manure co-digestion can add moisture and seed microbes to an AD system, potentially improving syntrophic potential and microbial growth (El-Mashad and Zhang 2010; Silvestre et al. 2013).
Therefore, co-digesting RS with manure has benefits, but the impact of manure on AD microbial communities related to specific methane yields and also pathogens in AD digestates is not known. Here, we assessed the effects of dairy manure (DM) co-digestion with RS in bench-scale bioreactors fed at different RS:DM ratios. Reactor performance was monitored, and microbial communities were characterised using 16S rDNA amplicon sequencing to assess how eubacteria and archaea composition and diversity vary as a function of DM additions and RS co-digestion conditions.
Materials and methods
Experimental conditions and analyses
Four 2.5 L reactors with control towers were used as the AD units, each with working volumes of 2.0 L. The glass airtight-sealed reactors consisted of a heating jacket set to 37 °C, a biogas sampling bag and a paddle stirrer. Overall, the reactors were operated for 150 days of which the first 75 days were used to acclimate the reactors to different RS:DM mixes. The same reactors had been operated for over 250 days prior to this experiment at different RS organic loading rates (OLRs); therefore, microbial communities were already acclimated to RS before DM amendments were commenced.
During acclimation with DM, the sludge inoculum was operated in draw-fill mode (digester sludge removal prior to feed addition) with a 25-day hydraulic retention time (HRT) and an OLR of 1.0 g VS/L/day (chosen based on previous assays reported in Zealand et al. (2017)) fed once every 7 days. After three HRTs, pH and VFA levels in the reactors had become consistent with time and the formal experiment was commenced (defined as time 0 here). Operationally, a feeding frequency (FF) of one in 7 days and an OLR of 1 g VS/L/day comprised 560 mL of reactor volume being removed per week, with 14 g VS RS (1.0 mm sieve) being added with 560 mL of distilled water. Four RS:DM ratios were assessed here at the same OLR (see Table 1). Reactors were named ‘RS100’, ‘RS90’, ‘RS70’ and ‘RS30’ based on the percentage of RS (by mass) in each reactor feed mixture.
Analysis
Total solids (TS), volatile solids (VS; combustible solids at 550 °C ignition and presumed organic fraction), moisture content (MC), ash content (AC) and total C and N were analysed using American Public Health Association Standard Methods (APHA 1998). VFA analysis was performed three times per week and consisted of filtering each sample through a 0.2-μm PES syringe filter before mixing with 0.1 M octane sulphonic acid (1:1) and sonicating for 40 min. Once sonicated, samples were analysed using ion chromatography Dionex Aquion system equipped with an AS-AP auto sampler with Chameleon 7 Software. RS composition data are provided in Table 2.
Daily biogas volume was determined using a 100-μL gas tight syringe (SGE, Australia). Biogas samples were collected at the same time of day and analysed immediately. To quantify methane content (% CH4), gas samples were directly injected into a Carlo Erba HRGC 5160 GC-FID fitted with a HP-PLOT Q column maintained at 35 °C with hydrogen as the carrier gas and Atlas software. A seven-point calibration was performed before and after each analysis session by injecting neat standards spanning expected methane concentrations (up to 80% CH4, Scientific Technical Gases, UK). All injections were made in triplicate and the standard calibration required a minimum R2 of 0.99. The volume of biogas was collected over time in 1.0 L Supel-Inert Multi-Layer Foil bags (Sigma Aldrich) before daily extraction using a 1 L gas tight syringe (SGE). Biogas and methane analysis was normalised to specific production (mL/g VS/day), corrected for moisture, standard temperature and pressure (STP) and headspace volume, according to the VDI Standard: 4630 (2006).
Sample collection for sequencing and DNA preparation
Samples for DNA extraction (2 mL each) were collected in triplicate from the reactors after 0, 37 and 75 days of operations after acclimation. Raw DM samples also were collected in triplicate to compliment the reactor samples. Samples were always stored at − 20 °C before further processing. For each sample, DNA was extracted following the instructions of the FastDNA SPIN Kit for Soil (MP Biomedicals, Carlsbad, CA, USA). Extracted DNA was assessed for purity and quantified using a Qubit 2.0 Fluorometer (Invitrogen) to ensure adequate quantity and quality prior to sequencing.
Sequencing analysis
Sequencing of extracted DNA (concentration of 1 and 10 ng/μL and a volume of 10–20 μL) was undertaken by LGC Genomics GmbH in Berlin, Germany. Analysis consisted of PCR amplification using universal forward primer (U341F) CCTAYGGGRBGCASCAG and universal reverse primer (U806R) GGACTACNNGGGTATCTAAT, targeting the V3-V4 16S DNA region (Klindworth et al. 2013). The standard protocol was 10 cycles of touchdown PCR (annealing 61–55 °C, decreasing 0.6 °C per cycle), before 26 standard 2-step PCR cycles at 55 °C. Quality control (agarose gel check), library preparation including tagging, equimolar mixing and clean-up were completed. 16S rDNA amplicon sequencing was then performed on Illumina MiSeq V3 (2 × 300 bp).
Bioinformatics analysis consisted of inline barcode demultiplexing, adaptor clipping and amplicon pre-processing using Mothur (Schloss et al. 2009): pair joining, filtering, alignment against Silva 16S, subsampling 5000–25,000 reads per sample, denoising and chimera removal. OTU picking used Mothur with clustering aligned sequences at 97% identity. Specific OTU analysis included assigning taxonomy on the Greengenes database (version 13_8). Predominant OTUs were defined as having ≥ 0.5% abundance in a sample. Phylogenetic analysis of predominant OTUs was performed with the ARB programme (Ludwig et al. 2004), using neighbour-joining and parsimony methods with 1000 bootstrap replication (McDonald et al. 2012; Kuroda et al. 2016). Additional information is provided in Supporting Information (SI).
Data analyses
Analysis of variance (ANOVA) with Tukey comparison was used to compare mean CH4 content and biogas yields with significance defined as 95% confidence (p ≤ 0.05). All statistical analyses and figure plots were conducted or developed using Microsoft Excel and Minitab 17 (Leadtools Technologies Inc., version 17.1.0, 2014). PRIMER 7 (PRIMER-E, Plymouth, UK) was used to perform principal coordinates analysis (PCoA) and distance-based linear modelling (DistLM) of log-transformed, normalised, RS composition data.
Alpha diversity and beta diversity, based on weighted UniFrac distances, were calculated in QIIME 1.9.1. PRIMER 7 (PRIMER-E, Plymouth, UK) was used for principal component analysis (PCA), metric-multidimensional scaling (MDS), heat map development to visualise beta diversity, permutational analysis of variance (PERMANOVA) and for analysis of similarities (ANOSIM). To compare microbial diversity with performance data, RELATE, BEST and DistLM (distance-based linear model) were used for weighted UniFrac distances, employing Bray-Curtis distance (after square root transformation) (Ling et al. 2016; Mei et al. 2016).
Observed OTUs, Chao1, Simpson’s and Shannon’s Indexes were plotted and compared using ANOVA with Tukey comparison in Minitab 17 (Leadtools Technologies Inc., version 17.1.0, 2014). Group significant differences were compared in STAMP v2.1.3 using the t test. These methods were chosen to identify possible correlations/groupings and significant relationships at each point of analysis. The sequence data obtained in this study are deposited at NCBI GenBank with accession no. MG852175 - MG855654.
Results
Effect of dairy manure on reactor performance
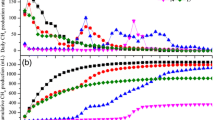
Stable bioreactor operations were confirmed before any samples were collected for microbial community characterisation. Stability was defined as when no statistically significant differences (using ANOVA) in biogas yields were apparent when comparing sequential HRTs of operating time. Biogas production ‘stability’ was achieved after the third HRT, after which specific gas yields, CH4 content, VS, pH and total VFA data were tallied and summarised (see Table 3). Time-course data (after acclimation) for pH, VS, CH4 yields and total VFA are shown in Fig. 1a–d (all time-course data is provided in Figure S1; see SI). Ammonia was always below detection limits throughout the experiment.
Mean specific biogas yields ranged from ~115 to 222 mL/g VS/day across reactors, whereas specific CH4 yields ranged from ~48 to 112 mL CH4/g VS/day. For both parameters, decreasing the RS:DM ratio resulted in significantly lower specific yields (p < 0.001). Further, percent CH4 content within biogas from RS100 and RS90 were ~51 and 48% CH4, respectively, which are significantly higher than in RS30 and RS70 (41 and 43% CH4, respectively).
In contrast to biogas results, RS30 had the highest mean VS% removals (i.e. 38%), significantly higher than the other three reactors, which ranged between 23 to 26% (p = 0.003). RS30 also had the lowest mean VS residuals (4.7 g VS/L), significantly lower than the other three reactors (range from 11 to 13 g VS/L; p < 0.001). Differences in pH among reactors also were significant. RS30 had the highest mean pH (6.3), although all four reactors had pHs > 6.0. Differences in total VFA levels among reactors also were apparent; i.e. RS30 VFA levels were significantly lower than the other three reactors (mean of 97 ppm), which ranged from 401 to 506 ppm (p < 0.003). Although various VFAs were measured throughout the experiment, individual acids were not always present; therefore, only formic, acetic, isobutyric and propionic acid had adequate data to be statistically compared. Of these, propionic acid in RS30 was significantly lower (37 ppm) than the other three acids, which were between 246 and 319 ppm (p < 0.003). Time course data (Fig. 1a–d) show performance parameters were generally consistent during the period when samples were collected for microbial community characterisation.
Beta-diversity and physiochemical parameters
To contrast the richness and evenness of microbial communities between reactors, beta-diversity indices were compared using ‘observed’ and Chao1 OTU numbers, and also Simpson’s and Shannon’s indices (Fig. 2). Based on Bray-Curtis distances, reactor samples clustered according to HRT (Fig. 2a) and RS:DM ratio (Fig. 2b), with RS:DM ratio having the greater effect with an axes variation of > 80%. The relative influence of each operating parameter is reflected by direction and length of the corresponding arrows. The weighted Unifrac distances (Fig. 2c) grouped the samples based on HRT (ellipses) and on RS:DM (shown by coloured lines), which shows RS30 groups separately from the other reactors in terms of reactor performance. For example, lower RS in the feed resulted in lower Total VFA levels, especially acetic, propionic and isobutyric acid (Fig. 2d).
Analyses of beta diversity showing variation of microbial community structure and the influence of physiochemical data. a, b PCO of Bray-Curtis distance, but coloured differently by HRT and RS:DM. c PCO of weighted UniFrac distance. d Boxplot of individual and total VFAs for RS100, RS90, RS70 and RS30 (there was no isovaleric or valeric acid in RS30). Physiochemical data overlaid arrows and dashed elliptical shapes and/or coloured lines indicate sample groupings
To determine whether the groupings and correlations seen in Fig. 2 were statistically significant, RELATE, BEST, DistLM, ANOSIM and PERMANOVA were calculated (see Table 4). ANOSIM and PERMANOVA analysis showed that HRT and RS:DM are significantly different in terms of combined physiochemical data. However, ANOSIM and PERMANOVA showed that while RS:DM was significant in determining differences in microbial communities among reactors (i.e. beta-diversity; 2.2% at p = 0.018), HRT effects are apparent, albeit were not significant (7.3% at p = 0.276). The influence on beta-diversity of physiochemical operating conditions was shown by a significant 0.67 correlation (0.2%; RELATE) with pH (0.725), followed by %CH4, total VFAs, formic and isobutyric acid combined (0.724), providing the BEST correlations. DistLM analysis showed a range of variables were significantly related to beta-diversity, including biogas yield (p = 0.012), %CH4 (p = 0.039) and pH (p = 0.002).
Impact of co-digestion on alpha-diversity
Alpha-diversity indices (Hughes et al. 2001; Lemos et al. 2011) were used to contrast the richness and evenness of microbial communities in the reactors, based on observed and Chao1 OTU numbers (given as Figure S2), and Simpson’s and Shannon’s indices (Fig. 3). Observed OTUs and Chao1 estimations show that raw manure has significantly higher numbers of OTUs compared with any reactor communities (p < 0.001) followed by RS30, RS70 and RS90, although only RS100 had significantly lower OTUs than RS30 (p = 0.041). Chao1 estimations also showed that raw DM had significantly higher OTUs than any reactor (p < 0.001), which means it has a higher number of OTUs only sequenced once. Both these estimates show that as DM addition decreases, reactor community richness decreases (Fig. 3d), although Simpson’s and Shannon’s scores show differences were not always statistically significant (p > 0.05). Shannon’s score considers both richness and evenness, and it was higher for RS30 and DM (5.6 and 5.5). All diversity scores observed here, even in the reactor without DM addition, were quite high (Simpson’s = 0.90–0.95; Shannon’s = 4.8–5.6), especially compared with Zhao et al. (2012).
Predominant OTUs
To assess predominant OTUs, sequencing data were combined for all samples from each reactor because operational data over time did not significantly differ. Predominant OTUs are summarised in Fig. 4 (≥ 0.5% relative abundance, 76 OTUs and ‘others’ that is the cumulative abundance of those > 0.1%), which contrasts differences in major OTUs among reactors. The 76 major OTUs across reactors (not including the cumulative ‘others’) are provided as a phylogenetic tree in the SI (see Figure S3). Further, Fig. 5 shows if any perceived changes among conditions were statistically significant (all RS communities are compared with DM in Figures S4 to S7).
RS100, RS90 and RS70 were generally similar in phyla and dominated by Firmicutes and Bacteroidetes with the remaining 30–40% fraction made up of Proteobacteria, Spirochaetes, Euryarchaeota and Chloroflexi. RS30 differed in that Bacteroidetes was less prevalent and had higher levels of Proteobacteria and Spirochaetes. DM samples were > 65% Firmicutes and had almost no Spirochaetes. The number of predominant OTUs in each reactor was positively related with DM addition, ranging from RS100 and RS90 (32 and 31 OTUs, respectively) through RS70 (35 OTUs) to RS30 (38 OTUs). The DM samples also had a higher number of rare OTUs as their predominant OTU total was the lowest at 26.
Methanogens were present in all reactors, ranging from 4.5 to 6.2% in terms of relative abundance. Highest abundances were seen in RS90 and the lowest in RS100. Methanobacterium and Methanosarcina always were the main methanogen guilds, including OTUs identified as 005 and 044/002, with abundances ranging from 2.2 to 3.5 and 1.1 to 2.4%, respectively. Interestingly, these two Archaea taxa were below detection limits in DM samples. Methanobrevibacter (OTU 052) was dominant in the DM samples (3.4%), but was lower than 0.2% in all the reactors.
Proteiniphilum (OTU 012) was significantly more abundant in RS100 and RS90 (14.4 and 8.9%, respectively) than in RS30 (0.1% at p < 0.001 and 0.008). Anaerolineaceae abundance in RS100 and RS90 was 6.6 and 6.9%, lower in RS70 (4.4%), but significantly lower in RS30 (2.0% at p = 0.031 versus RS100 and 0.019 versus RS90). Christensenellaceae was significantly higher in RS90 (3.4% for OTU 033) than in RS30 (p = 0.007). Ruminococcus (OTU 083) generally had low abundance (~1%), but was significantly higher (p = 0.022) in RS100 (0.9% than RS30 (< 0.1%)). Rhodobacteraceae was highest in RS90 (1.6%) and lowest in RS30 (< 0.1%, p = 0.042) with 0.8 and 1.0% in RS100 and RS70. Christensenellaceae (OTU 023) was 5.0 and 5.9% in RS100 and RS90, but was highest in RS70 (7.8%) and lowest in RS30 (0.1%).
Relative abundance of Spirochaetaceae (OTU 026) was highest in RS30 (11.3%), which was significantly higher than in the other reactors, versus RS100 (> 0.1%), RS90 (1.0%) and RS70 (1.2%) at ≤ 0.006. Clostridium (OTU 027) and Clostridiales (OTU 074) in RS30 (0.7 and 1.2%) were significantly higher than in any other reactor (all p < 0.05), implying that higher DM addition might result in slightly higher putative pathogen levels than at lower DM additions. Peptostreptococcaceae (OTU 015 & 016) were both < 1.0% in RS100, RS90 and RS70, but were significantly higher in RS30 at 2.8 and 2.4% (all p ≤ 0.001).
Discussion
Effect of dairy manure addition on reactor performance
Specific biogas and CH4 yields were significantly greater at higher RS:DM ratios, although actual %CH4 content in the biogas was fairly similar across DM additions. In contrast, increasing DM additions resulted in slightly higher pH, greater VS% removals and lower levels of VS and VFA accumulation. Lower specific CH4 yields were unexpected as co-digestion with manure is touted as a method for enhancing RS AD processes (Marañón et al. 2012).
Based on RS stoichiometry, RS AD appears N-limited and N amendment as DM should improve nutritional balance within RS AD reactors (Wang et al. 2014). However, increasing DM additions significantly reduced specific CH4 yields in our AD reactors. This also was observed by Callaghan et al. (2002) and Dechrugsa et al. (2013) who found increasing levels of manure had a detrimental effect on biogas yields. This contradicts other RS co-digestion studies, such as Estevez et al. (2012), Wang et al. (2014), Sahito and Mahar (2014), Li et al. (2014a), Xavier et al. (2015) and Jiménez et al. (2016), who all found manure additions positively influenced biogas yields. However, none of these positive studies compared specific CH4 yields from RS side-by-side with and without manure addition; i.e. positive conclusions were made without suitable treatment controls. Adding DM to RS AD reactors reduced specific CH4 yields here relative to 100% RS reactors, although our OLRs were comparatively low.
However, DM addition did have positive benefits here that are reflected in non-methane AD performance data, which also was observed by Babaee et al. (2013). Manure addition almost certainly does balance the C:N ratio and provides additional nutrients to improve the AD process (Li et al. 2014a). Further, manure was shown by Li et al. (2015) and Cornell et al. (2012) to allow higher RS AD OLRs (both 6.0 kg VS/m3/day), which are six times higher than those used here. Therefore, at higher OLRs, the benefits of C:N balancing and extra nutrients may overweigh lower degradability of DM used for co-digestion. Therefore, we suspect the relative value of adding DM to RS AD systems depends on many factors, and specific CH4 yields with and without DM may differ dramatically depending on the specific RS and DM. To underpin results herein, 16S rDNA amplicon sequencing was performed to compare raw DM microbial communities with RS AD communities as a function of RS:DM in the reactor feed.
Impact of DM addition on microbial communities in co-digestion systems
As RS content in the reactor feed was decreased from RS100 to RS30, reactor performance moved from higher biogas and VFA production to more stable pH and increased VS% removal in the systems (Fig. 2b). Further, greater accumulation of acetic, formic and propionic acids was evident (Fig. 2d) associated with greater CH4 production, which is consistent with higher RS in an AD unit generating higher gas yields and VFAs whilst lower RS leads to higher pH. In fact, ANOSIM and PERMANOVA show that RS:DM ratio was the dominant factor in determining reactor performance and also beta-diversity of the microbial community.
The addition of manure to RS AD reactors provides additional and varied microbes to the system (El-Mashad and Zhang 2010; Silvestre et al. 2013). We see this here in richness and evenness data on alpha-diversity as indicated by Chao1, OTU numbers and Simpson and Shannon Indices. As DM level in the feed was increased, more diverse communities were apparent (Fig. 3), which is consistent with Mata-Alvarez et al. (2000) who made similar observations on the nature of meta-community.
Relative to specific OTUs, RS100, RS90 and RS70 all were similar in phyla and were dominated by Firmicutes and Bacteroidetes. In contrast, RS30 was dominated by Proteobacteria and Spirochaetes, and the raw DM samples were > 65% Firmicutes with almost no Spirochaetes. The lack of Spirochaetes in the DM is surprising given that Paster and Canale-Parola (1982) found Spirochaetes were important in the rumen, although in their case, they concluded Spirochaetes were less associated with cellulose processing than with the fermentation of plant polymers. Overall, RS AD microbial communities were significantly impacted by RS:DM ratio, especially when DM additions were high, but even with high DM addition, RS:DM were very different than DM itself.
Methanobacterium and Methanosarcina were the dominant methanogens in all our reactors, which also was found by Leite et al. (2015); Ziganshina et al. (2015) and Fontana et al. (2016). This dominance has been explained by the hydrogenotrophic traits of these guilds, allowing them to better cope with increases in hydrogen partial pressure (Goberna et al. 2010; Sun et al. 2015). Further, Methanosarcina dominance in RS100 also may be due to their substrate versatility, acid tolerance and higher specific growth rates (Conklin et al. 2006; Yi et al. 2014; Leite et al. 2015). The fact that Methanosarcina was apparently lower in the other reactors may be due to its limited acetoclastic abilities (FitzGerald et al. 2015); i.e. as acetic acid levels decreased, Methanosarcina also decreased (Fontana et al. 2016). Further, Leite et al. (2015) found that Methanosarcina preferred mono digestion rather than co-digestion environments, which is observed here.
No well-known syntrophic bacteria were observed in the reactors here. High hydrogen production by RS AD, as seen by Kim et al. (2012), potentially reduces the habitable zone for syntrophs that require low hydrogen pressures (Stams and Plugge 2009). A lack of syntrophs was unexpected when compared to other studies on sewage sludge digestion (Mei et al. 2017), although Liu et al. (2017) found that highly active fermenting bacteria could produce inhibiting levels of VFAs and hydrogen. Facultatively syntrophic bacteria, such as Ruminococcus albus, can grow syntrophically with hydrogenotrophic methanogens (Stams and Plugge 2009). However, the increase in acetic and propionic acid with increased RS suggests that syntrophic breakdown of acetate and propionate were overwhelmed (Amani et al. 2011; Banks et al. 2012). Deeper studies are required than those performed here to characterise obligate syntrophic guilds in RS AD, which could potentially further improve the efficiency of RS digestion.
There were a number of significant community changes across the reactors with some bacteria thriving in higher RS conditions. High abundance of Bacteroidetes then Proteiniphilum (OTU 012), which do not use cellulose (Chen and Dong 2005), was unexpected in the high RS reactors. However, these organisms can produce acetic and propionic acid (Krieg et al. 2015), which is consistent with higher VFA levels in RS100 and RS90. Among Chloroflexi, Anaerolineaceae (OTU 019) showed a similar pattern, being more abundant with a higher RS feed. Xia et al. (2016) found that the cellulolytic capacity of Anaerolineaceae was not likely to be its main attribute, although this observation is based on very limited data and its ecological role is uncertain. Spirochaetaceae (OTU 026) was higher in RS90 than RS100 and was significantly higher than in any other condition, although there are few cultivated species of this facultative anaerobe (Paster 2015). Of the four Christensenellaceae that can degrade cellulose (Fontes and Gilbert 2010), Ruminococcus (OTU 083) was in relatively low abundance, but was significantly higher in RS100 than RS30, which may be explained by its association with cellulose hydrolysis and its potential use of cellobiose (Sun et al. 2015).
Microbial communities in RS30 were quite different than the other three reactors. For example, Firmicutes phylum had the greatest number of increases in abundance in RS30 compared with RS100, although Lü et al. (2014) noted 16S rDNA sequence data on the Firmicutes phylum should be made with caution because of interferences and it is extremely complex. Clostridium (OTU 027) and Clostridiales (OTU 074) also were more evident in RS30, which is interesting given they are both higher in raw DM and are prospective pathogens. The presence of Clostridium and Clostridiales in RS30 may simply be legacy of higher DM additions, which may suggest that if DM is too high, strains in DM of potential health concern may prevail in AD digestates. Peptostreptococcaceae (family) (OTU 015 and 016) also were high in RS30 at 2.8 and 2.4%. Li et al. (2014b) found that Peptostreptococcaceae contains a number of genera isolated from manure and Mao et al. (2012) noted that it negatively correlates with VFAs, which may be why it was low in higher RS reactors. Regardless, most of the above genera may also be legacies of higher DM addition.
There were a number of ‘Goldilocks’ bacteria, favouring neither RS100 nor RS30, but something in the middle. Rhodobacteraceae, which is typical of cattle slurry (FitzGerald et al. 2015), was highest in RS90 (1.6%) and Christensenellaceae (OTU 023) was highest in RS70 (7.8%). There is currently only one described species of Christensenellaceae, which was found to favour gut environments (Morotomi et al. 2012; Rosa et al. 2017).
Overall, a greater number of predominant OTUs in reactors with greater DM addition implies that for RS:AD diverse OTUs are not essential for elevated specific CH4 yields. Clearly, greater diversity is related to lower VFA and VS accumulation associated with higher DM additions, although this is not due to greater biomass in higher DM reactors (see Table 3). In fact, increasing levels of DM addition progressively reduced biomass, although diversity increased. Although it is speculation, it is possible that DM additions introduce many diverse species that are not specifically related to methane production from RS, but more related to intermediate fermentation reactions. Therefore, higher DM addition appears to provide greater diversity among species responsible for VFA reduction, but not methanogenesis, which may be more specialised (i.e., Methanobrevibacter dominated DM samples, whereas Methanosarcina and Methanobacterium dominated the RS AD reactors). Overall, this implies co-digestion AD systems with higher DM might cope better at higher OLRs due to greater fermentation versatility, although Zealand et al. (2017) also showed that this capacity is not unlimited because higher OLRs can lead to reactor souring in RS AD units.
In conclusion, the highest specific CH4 yields were observed in the bioreactor without DM addition, whereas lowest yields were observed in the reactor with greatest DM additions. In contrast, as DM additions were increased, both VS and VFA accumulation decreased, and VS% removals increased. Further, increasing DM content in the feed resulted to greater microbial richness compared with reactors with higher levels of RS. Evenness was similar among RS:DM ratios, although the predominant OTUs differed among reactors. Higher RS AD reactors were dominated by Firmicutes and Bacteroidetes, whereas the reactor with the highest DM addition, RS30, was dominated by Proteobacteria and Spirochaetes, and also had detectable Clostridium, which may have implications to the subsequent use digestates as fertilisers.
Methanogen abundances were similar among the reactors, therefore lower abundances of cellulosic hydrolysing bacteria, such as Christensenellaceae and Bacteroidetes, best explain lower specific CH4 production levels when higher DM was in the feed. This hints that carbon short-circuiting may be occurring in the reactor with highest DM additions. However, the overall main benefit of co-digestion with RS and DM appears to be decreased VFA production and higher rates of VS removal, which suggest co-digestion systems can potentially operate at higher OLRs. Conversely, results suggest RS AD without DM also is a viable option, although overall RS throughput in RS only AD systems might be lower due to the need to operate at lower OLRs.
References
Amani T, Nosrati M, Mousavi SM, Kermanshahi RK (2011) Study of syntrophic anaerobic digestion of volatile fatty acids using enriched cultures at mesophilic conditions. Int J Environ Sci Technol 8(1):83–96
APHA (1998) American public health association, 20th edn. American Public Health Assoc, Washington, DC
Babaee A, Shayegan J, Roshani A (2013) Anaerobic slurry co-digestion of poultry manure and straw: effect of organic loading and temperature. J Environ Health Sci Eng 11(1):15
Banks CJ, Zhang Y, Jiang Y, Heaven S (2012) Trace element requirements for stable food waste digestion at elevated ammonia concentrations. Bioresour Technol 104:127–135
Callaghan FJ, Wase DA, Thayanithy K, Forster CF (2002) Continuous co-digestion of cattle slurry with fruit and vegetable wastes and chicken manure. J Biomed Biotechnol 27:71–77
Chen S, Dong X (2005) Proteiniphilum acetatigenes gen. nov., sp. nov., from a UASB reactor treating brewery wastewater. Int J Syst Evol Microbiol 55(Pt 6):2257–2261
Conklin A, Stensel HD, Ferguson J (2006) Growth kinetics and competition between Methanosarcina and Methanosaeta in mesophilic anaerobic digestion. Water Environ Res 78(5):486–496
Cornell M, Banks CJ, Heaven S (2012) Effect of increasing the organic loading rate on the co-digestion and mono-digestion of cattle slurry and maize. Water Sci Technol 66(11):2336–2342
Dechrugsa S, Kantachote D, Chaiprapat S (2013) Effects of inoculum to substrate ratio, substrate mix ratio and inoculum source on batch co-digestion of grass and pig manure. Bioresour Technol 146:101–108
El-Mashad HM, Zhang R (2010) Biogas production from co-digestion of dairy manure and food waste. Bioresour Technol 101(11):4021–4028
Estevez, M.M., Linjordet, R. and Morken, J. (2012) 'Organic loading rate effect on anaerobic digestion: case study on co-digestion of lignocellulosic pre-treated material with cow manure', energy, biomass and biological residues. International conference of agricultural engineering - CIGR-AgEng 2012: agriculture and engineering for a healthier life, Valencia, Spain, 8-12 July 2012, pp. C-0880
FitzGerald JA, Allen E, Wall DM, Jackson SA, Murphy JD, Dobson ADW (2015) Methanosarcina play an important role in anaerobic co-digestion of the seaweed Ulva lactuca: taxonomy and predicted metabolism of functional microbial Communities. PLoS One 10(11):e0142603
Fontana A, Patrone V, Puglisi E, Morelli L, Bassi D, Garuti M, Rossi L, Cappa F (2016) Effects of geographic area, feedstock, temperature, and operating time on microbial communities of six full-scale biogas plants. Bioresour Technol 218:980–990
Fontes CM, Gilbert HJ (2010) Cellulosomes: highly efficient nanomachines designed to deconstruct plant cell wall complex carbohydrates. Annu Rev Biochem 79:655–681
Goberna M, Gadermaier M, García C, Wett B, Insam H (2010) Adaptation of methanogenic communities to the cofermentation of cattle excreta and olive mill wastes at 37°C and 55°C. Appl Environ Microbiol 76(19):6564–6571
Hughes JB, Hellmann JJ, Ricketts TH, Bohannan BJM (2001) Counting the uncountable: statistical approaches to estimating microbial diversity. Appl Environ Microbiol 67(10):4399–4406
Jiménez J, Theuerl S, Bergmann I, Klocke M, Guerra G, Romero-Romero O (2016) Prokaryote community dynamics in anaerobic co-digestion of swine manure, rice straw and industrial clay residuals. Water Sci Technol 74(4):824–835
Kim M, Yang Y, Morikawa-Sakura MS, Wang Q, Lee MV, Lee D-Y, Feng C, Zhou Y, Zhang Z (2012) Hydrogen production by anaerobic co-digestion of rice straw and sewage sludge. Int J Hydrog Energy 37(4):3142–3149
Krieg NR, Ludwig W, Euzéby JP, Whitman WB (2015) Bacteroidetes, in Bergey's manual of systematics of archaea and Bacteria. John Wiley & Sons, Ltd, Hoboken
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glockner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41(1):e1
Krishania M, Kumar V, Vijay VK, Malik A (2013) Analysis of different techniques used for improvement of biomethanation process: a review. Fuel 106:1–9
Kuroda K, Nobu MK, Mei R, Narihiro T, Bocher BTW, Yamaguchi T, Liu W-T (2016) A single-granule-level approach reveals ecological heterogeneity in an Upflow anaerobic sludge blanket Reactor. PLoS One 11(12):e0167788
Lei Z, Chen J, Zhang Z, Sugiura N (2010) Methane production from rice straw with acclimated anaerobic sludge: effect of phosphate supplementation. Bioresour Technol 101(12):4343–4348
Leite AF, Janke L, Lv Z, Harms H, Richnow HH, Nikolausz M (2015) Improved monitoring of semi-continuous anaerobic digestion of sugarcane waste: effects of increasing organic loading rate on methanogenic community dynamics, Int. J Mol Sci 16(10):23210–23226
Lemos LN, Fulthorpe RR, Triplett EW, Roesch LFW (2011) Rethinking microbial diversity analysis in the high throughput sequencing era. J Microbiol Methods 86(1):42–51
Li J, Wei L, Duan Q, Hu G, Zhang G (2014a) Semi-continuous anaerobic co-digestion of dairy manure with three crop residues for biogas production. Bioresour Technol 156:307–313
Li Y-F, Chen P-H, Yu Z (2014b) Spatial and temporal variations of microbial community in a mixed plug-flow loop reactor fed with dairy manure. Microb Biotechnol 7(4):332–346
Li D, Liu S, Mi L, Li Z, Yuan Y, Yan Z, Liu X (2015) Effects of feedstock ratio and organic loading rate on the anaerobic mesophilic co-digestion of rice straw and cow manure. Bioresour Technol 189:319–326
Ling F, Hwang C, LeChevallier MW, Andersen GL, Liu W-T (2016) Core-satellite populations and seasonality of water meter biofilms in a metropolitan drinking water distribution system. ISME J 10(3):582–595
Liu M, Wang S, Nobu MK, Bocher BTW, Kaley SA, Liu W-T (2017) Impacts of biostimulation and bioaugmentation on the performance and microbial ecology in methanogenic reactors treating purified terephthalic acid wastewater, Water Res 122:308-316
Lü F, Bize A, Guillot A, Monnet V, Madigou C, Chapleur O, Mazéas L, He P, Bouchez T (2014) Metaproteomics of cellulose methanisation under thermophilic conditions reveals a surprisingly high proteolytic activity. ISME J 8(1):88–102
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar, Buchner A, Lai T, Steppi S, Jobb G, Förster W, Brettske I, Gerber S, Ginhart AW, Gross O, Grumann S, Hermann S, Jost R, König A, Liss T, Lüßmann R, May M, Nonhoff B, Reichel B, Strehlow R, Stamatakis A, Stuckmann N, Vilbig A, Lenke M, Ludwig T, Bode A, Schleifer K-H (2004) ARB: a software environment for sequence data. Nucleic Acids Res 32(4):1363–1371
Mao S, Zhang R, Wang D, Zhu W (2012) The diversity of the fecal bacterial community and its relationship with the concentration of volatile fatty acids in the feces during subacute rumen acidosis in dairy cows. BMC Vet Res 8(1):237
Marañón E, Castrillón L, Quiroga G, Fernández-Nava Y, Gómez L, García MM (2012) Co-digestion of cattle manure with food waste and sludge to increase biogas production. Waste Manag 32(10):1821–1825
Mata-Alvarez J, Macé S, Llabrés P (2000) Anaerobic digestion of organic solid wastes. An overview of research achievements and perspectives. Bioresour Technol 74(1):3–16
McDonald D, Price MN, Goodrich J, Nawrocki EP, DeSantis TZ, Probst A, Andersen GL, Knight R, Hugenholtz P (2012) An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6(3):610–618
Mei R, Narihiro T, Nobu MK, Liu WT (2016) Effects of heat shocks on microbial community structure and microbial activity of a methanogenic enrichment degrading benzoate. Lett Appl Microbiol 63(5):356–362
Mei R, Nobu MK, Narihiro T, Kuroda K, Muñoz Sierra J, Wu Z, Ye L, Lee PKH, Lee P-H, van Lier JB, McInerney MJ, Kamagata Y, Liu W-T (2017) Operation-driven heterogeneity and overlooked feed-associated populations in global anaerobic digester microbiome. Water Res 124:77–84
Monnet F. (2003) An introduction to anaerobic digestion of organic wastes. Remade Scotland. http://www.biogasmax.co.uk/media/introanaerobicdigestion__073323000_1011_24042007.pdf. Accessed 27 Apr 2018
Morotomi M, Nagai F, Watanabe Y (2012) Description of Christensenella minuta gen. nov., sp. nov., isolated from human faeces, which forms a distinct branch in the order Clostridiales, and proposal of Christensenellaceae fam. nov. Int J Syst Evol Microbiol 62(Pt 1):144–149
Mussoline W, Esposito G, Giordano A, Lens P (2013) The anaerobic digestion of rice straw: a review. Crit Rev Environ Sci Technol 43(9):895–915
Paster BJ (2015) Spirochaetaceae, in Bergey's manual of systematics of archaea and Bacteria. John Wiley & Sons, Ltd, Hoboken
Paster BJ, Canale-Parola E (1982) Physiological diversity of rumen spirochetes. Appl Environ Microbiol 43(3):686–693
Rosa BA, Hallsworth-Pepin K, Martin J, Wollam A, Mitreva M (2017) Genome sequence of Christensenella minuta DSM 22607(T). Genome Ann 5(2):e01451–e01416
Sahito AR, Mahar RB (2014) Enhancing methane production from rice straw co-digested with buffalo dung by optimizing effect of substrate ratio, alkaline doze and particle size. J Anim Plant Sci 24(4):1076–1084
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Silvestre G, Gomez MP, Pascual A, Ruiz B (2013) Anaerobic co-digestion of cattle manure with rice straw: economic & energy feasibility. Water Sci Technol 67(4):745–755
Stams AJM, Plugge CM (2009) Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat Rev Microbiol 7(8):568–577
Sun L, Pope PB, Eijsink VGH, Schnürer A (2015) Characterization of microbial community structure during continuous anaerobic digestion of straw and cow manure. Microb Biotechnol 8(5):815–827
VDI Standard: 4630 (2006) Fermentation of organic materials, characterisation of the substrates, sampling, collection of material data, fermentation test. VDI-Handbuch Energietechnik. http://www.vdi.eu/uploads/tx_vdirili/pdf/2385990.pdf. Accessed 27 Apr 2018
Wang X, Lu X, Li F, Yang G (2014) Effects of temperature and carbon-nitrogen (C/N) ratio on the performance of anaerobic co-digestion of dairy manure, chicken manure and rice straw: focusing on ammonia inhibition. PLoS One 9(5):e97265
Ward AJ, Hobbs PJ, Holliman PJ, Jones DL (2008) Optimisation of the anaerobic digestion of agricultural resources. Bioresour Technol 99(17):7928–7940
Xavier CAN, Moset V, Wahid R, Møller HB (2015) The efficiency of shredded and briquetted wheat straw in anaerobic co-digestion with dairy cattle manure. Biosyst Eng 139:16–24
Xia Y, Wang Y, Wang Y, Chin FYL, Zhang T (2016) Cellular adhesiveness and cellulolytic capacity in Anaerolineae revealed by omics-based genome interpretation. Biotechnol Biofuels 9(1):111
Yi J, Dong B, Jin J, Dai X (2014) Effect of increasing total solids contents on anaerobic digestion of food waste under mesophilic conditions: performance and microbial characteristics Analysis. PLoS One 9(7):e102548
Zealand AM, Roskilly AP, Graham DW (2017) Effect of feeding frequency and organic loading rate on biomethane production in the anaerobic digestion of rice straw, App. En 207:156-165
Zhao G, Ma F, Wei L, Chua H (2012) Using rice straw fermentation liquor to produce bioflocculants during an anaerobic dry fermentation process. Bioresour Technol 113:83–88
Ziganshina EE, Belostotskiy DE, Ilinskaya ON, Boulygina EA, Grigoryeva TV, Ziganshin AM (2015) Effect of the organic loading rate increase and the presence of zeolite on microbial community composition and process stability during anaerobic digestion of chicken wastes. Microb Ecol 70(4):948–960
Funding
This work was partially supported by the Energy from Rice Straw project (EP/L002477/1) funded by EPSRC, DFID and DECC as part of the Energy & International Development: USES programme; the Natural Environment Research Council (NE/L501748/1); the Sir Joseph Swan Centre for Energy Research; and the Xiamen-Newcastle Joint Strategic Partnership Fund. A special acknowledgement goes to Wen-Tso’s group at the University of Illinois at Urbana, USA, without which, this microbial analysis would not have been possible.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 1499 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zealand, A.M., Mei, R., Papachristodoulou, P. et al. Microbial community composition and diversity in rice straw digestion bioreactors with and without dairy manure. Appl Microbiol Biotechnol 102, 8599–8612 (2018). https://doi.org/10.1007/s00253-018-9243-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9243-7