Abstract
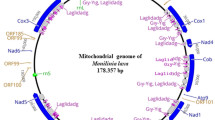
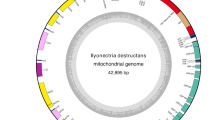
Endophytic fungi (EF) live within plants and have profound impacts on plant communities. They are astonishingly diverse but poorly studied at the genome level. Herein, we assembled the mitochondrial genome (mitogenome) of the EF Pestalotiopsis fici, annotated and compared it with those of other relatives to better understand the evolution of the EF lineage. Except for standard fungal mitochondrial genes, the 69,529-bp circular mitogenome of P. fici harbors 18 introns acquired possibly through lateral transfer from other fungi and nine free-standing open reading frames with some scarcely seen in fungal mitogenomes. BLAST analysis detected no obvious duplication events of large fragments between mitochondrial and nuclear genomes of the fungus. Transcription analyses validated the expression of all mitochondrial genes, while most genes showed higher expression on rice than in two other media. The mitogenome of P. fici is highly syntenic with the Xylariales species Annulohypoxylon stygium and the endophyte Epichloe festucae var. lolii, but lacks synteny with another endophyte Penicillium polonicum. This study reports the first mitogenome of Pestalotiopsis and the third published mitogenome from an EF and provides insights into the evolution of the EF lineage.



Similar content being viewed by others
References
Aguileta G, de Vienne DM, Ross ON, Hood ME, Giraud T, Petit E, Gabaldon T (2014) High variability of mitochondrial gene order among fungi. Genome Biol Evol 6(2):451–465
Bensasson D (2001) Mitochondrial pseudogenes: evolution’s misplaced witnesses. Trends Ecol Evol 16(6):314–321
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27(2):573–580
Brenner S, Johnson M, Bridgham J, Golda G, Lloyd DH, Johnson D, Luo S, McCurdy S, Foy M, Ewan M, Roth R, George D, Eletr S, Albrecht G, Vermaas E, Williams SR, Moon K, Burcham T, Pallas M, DuBridge RB, Kirchner J, Fearon K, Mao J-i, Corcoran K (2000) Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nat Biotech 18(6):630–634
Chen L, Zhang QY, Jia M, Ming QL, Yue W, Rahman K, Qin LP, Han T (2016) Endophytic fungi with antitumor activities: their occurrence and anticancer compounds. Crit Rev Microbiol 42(3):454–473
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42(Database issue):D222–D230
Hazkani-Covo E, Zeller RM, Martin W (2010) Molecular poltergeists: mitochondrial DNA copies (numts) in sequenced nuclear genomes. PLoS Genet 6(2):e1000834
Huang Y, Niu B, Gao Y, Fu L, Li W (2010) CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics 26:680–682
Jaklitsch WM, Gardiennet A, Voglmayr H (2016) Resolution of morphology-based taxonomic delusions: Acrocordiella, Basiseptospora, Blogiascospora, Clypeosphaeria, Hymenopleella, Lepteutypa, Pseudapiospora, Requienella, Seiridium and Strickeria. Persoonia 37:82–105
Jeewon R, Liew ECY, Hyde KD (2002) Phylogenetic relationships of Pestalotiopsis and allied genera inferred from ribosomal DNA sequences and morphological characters. Mol Phylogenet Evol 25(3):378–392
Kang X, Liu C, Liu D, Zeng L, Shi Q, Qian K, Xie B (2016) The complete mitochondrial genome of huperzine A-producing endophytic fungus Penicillium polonicum. Mitochondr DNA Part B 1(1):202–203
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12(4):357–360
Kurtz S, Schleiermacher C (1999) REPuter: fast computation of maximal repeats in complete genomes. Bioinformatics 15(5):426–427
Lanfear R, Calcott B, Ho SY, Guindon S (2012) Partitionfinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29(6):1695–1701
Laslett D, Canbäck B (2004) ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res 32(1):11–16
Laslett D, Canbäck B (2008) ARWEN: a program to detect tRNA genes in metazoan mitochondrial nucleotide sequences. Bioinformatics 24(2):172–175
Liu L (2011) Bioactive metabolites from the plant endophyte Pestalotiopsis fici. Mycology 2(1):37–45
Liu L, Liu S, Jiang L, Chen X, Guo L, Che Y (2008a) Chloropupukeananin, the first chlorinated pupukeanane derivative, and its precursors from Pestalotiopsis fici. Org Lett 10(7):1397–1400
Liu L, Tian RR, Liu SC, Chen XL, Guo LD, Che YS (2008b) Pestaloficiols A-E, bioactive cyclopropane derivatives from the plant endophytic fungus Pestalotiopsis fici. Bioorg Med Chem 16(11):6021–6026
Liu A-R, Chen S-C, Wu S-Y, Xu T, Guo L-D, Jeewon R, Wei J-G (2010) Cultural studies coupled with DNA based sequence analyses and its implication on pigmentation as a phylogenetic marker in Pestalotiopsis taxonomy. Mol Phylogenet Evol 57(2):528–535
Liu L, Bruhn T, Guo LD, Gotz DCG, Brun R, Stich A, Che YS, Bringmann G (2011) Chloropupukeanolides C-E: cytotoxic pupukeanane chlorides with a spiroketal skeleton from Pestalotiopsis fici. Chemi-Eur J 17(9):2604–2613
Liu L, Li Y, Li L, Cao Y, Guo L, Liu G, Che Y (2013) Spiroketals of Pestalotiopsis fici provide evidence for a biosynthetic hypothesis involving diversified Diels-Alder reaction cascades. J Org Chem 78(7):2992–3000
Liu L, Zhao C, Li L, Guo LD, Che YS (2015) Pestalotriols A and B, new spiro[2.5]octane derivatives from the endophytic fungus Pestalotiopsis fici. RSC Adv 5(96):78708–78711
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25(5):955–964
Maharachchikumbura SSN, Guo LD, Chukeatirote E, Bahkali AH, Hyde KD (2011) Pestalotiopsis-morphology, phylogeny, biochemistry and diversity. Fungal Divers 50(1):167–187
Maharachchikumbura SSN, Hyde KD, Groenewald JZ, Xu J, Crous PW (2014) Pestalotiopsis revisited. Stud Mycol 79:121–186
Martins M, Dairou J, Rodrigues-Lima F, Dupret JM, Silar P (2010) Insights into the phylogeny or arylamine N-acetyltransferases in fungi. J Mol Evol 71(2):141–152
Nadimi M, Daubois L, Hijri M (2016) Mitochondrial comparative genomics and phylogenetic signal assessment of mtDNA among arbuscular mycorrhizal fungi. Mol Phylogenet Evol 98:74–83
Rodriguez RJ, White JF Jr, Arnold AE, Redman RS (2009) Fungal endophytes: diversity and functional roles. New Phytol 182(2):314–330
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Hohna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542
Senanayake IC, Maharachchikumbura SSN, Hyde KD, Bhat JD, Jones EBG, McKenzie EHC, Dai DQ, Daranagama DA, Dayarathne MC, Goonasekara ID, Konta S, Li WJ, Shang QJ, Stadler M, Wijayawardene NN, Xiao YP, Norphanphoun C, Li QR, Liu XZ, Bahkali AH, Kang JC, Wang Y, Wen TC, Wendt L, Xu JC, Camporesi E (2015) Towards unraveling relationships in Xylariomycetidae (Sordariomycetes). Fungal Divers 73(1):73–144
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313
Venugopalan A, Srivastava S (2015) Endophytes as in vitro production platforms of high value plant secondary metabolites. Biotechnol Adv 33(6, Part 1):873–887
Wagemaker MJ, Eastwood DC, Welagen J, van der Drift C, Jetten MS, Burton K, Van Griensven LJ, Op den Camp HJ (2007) The role of ornithine aminotransferase in fruiting body formation of the mushroom Agaricus bisporus. Mycol Res 111(Pt 8):909–918
Wang KW, Lei JX, Wei JG, Yao N (2012) Bioactive natural compounds from the plant endophytic fungi Pestalotiopsis spp. Mini-Rev Med Chem 12(13):1382–1393
Wang X, Zhang X, Liu L, Xiang M, Wang W, Sun X, Che Y, Guo L, Liu G, Guo L, Wang C, Yin WB, Stadler M, Zhang X, Liu X (2015) Genomic and transcriptomic analysis of the endophytic fungus Pestalotiopsis fici reveals its lifestyle and high potential for synthesis of natural products. BMC Genomics 16(1):28
Wang B, Zhang ZW, Guo LD, Liu L (2016) New cytotoxic meroterpenoids from the plant endophytic fungus Pestalotiopsis fici. Hel Chim Acta 99(2):151–156
Wu GW, Zhou HC, Zhang P, Wang XN, Li W, Zhang WW, Liu XZ, Liu HW, Keller NP, An ZQ, Yin WB (2016) Polyketide production of pestaloficiols and macrodiolide ficiolides revealed by manipulations of epigenetic regulators in an endophytic fungus. Org Lett 18(8):1832–1835
Zhang YJ, Zhang S, Liu XZ, Wen HA, Wang M (2010) A simple method of genomic DNA extraction suitable for analysis of bulk fungal strains. Lett Appl Microbiol 51(1):114–118
Zhang Y, Zhang S, Zhang G, Liu X, Wang C, Xu J (2015) Comparison of mitochondrial genomes provides insights into intron dynamics and evolution in the caterpillar fungus Cordyceps militaris. Fungal Genet Biol 77:95–107
Zhang YJ, Zhao YX, Zhang S, Chen L, Liu XZ (2017) Reanalysis of the mitochondrial genome of the pneumocandin-producing fungus Glarea lozoyensis. Acta Microbiol Sin 2017 (in press)
Zhu YY, Ai C, Zhang J, Zhang ZW, Zhao CQ (2011) Bioactive secondary metabolites from endophytic fungi in plants. Prog Chem 23(4):704–730
Acknowledgements
This study was funded by the National Science Foundation of China (81102759), the Natural Science Foundation of Shanxi Province (2014021030-2, 201601D011065), the Fund Program for the Scientific Activities of Selected Returned Overseas Professionals in Shanxi Province, and the Special Fund for Large Scientific Instruments and Equipments in Shanxi Province. The authors thank Xiaoqing Yang for her help in drawing the circular mitochondrial map.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Zhang, S., Wang, XN., Zhang, XL. et al. Complete mitochondrial genome of the endophytic fungus Pestalotiopsis fici: features and evolution. Appl Microbiol Biotechnol 101, 1593–1604 (2017). https://doi.org/10.1007/s00253-017-8112-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-017-8112-0