Abstract—
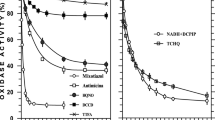
Lacticaseibacillus rhamnosus CM MSU 529 was grown in a batch culture under intensive aeration in the presence of 38 µM hemin and 18 µM vitamin K2 as a source of menaquinone to activate respiratory metabolism. Unsupplemented aerobic culture served as a control. Supplementation of the growth medium with hemin or menaquinone alone had no significant effect on the culture growth. In respiratory conditions (hemin + K2), a biomass production was 2.86 ± 0.05 g dw cells/L for a 24-h culture; the molar growth yield coefficient YP/S after 18 h of cultivation was 25.6 ± 1.5 g dw cells/mol of glucose consumed. Both values were 27% higher compared to those for the aerobic conditions. Spectral analysis revealed the presence of cytochromes b- and d-types in membranes of L. rhamnosus CM MSU 529. The activity of the bacterial electron transport chain was investigated using polarography. Membrane preparations of cells grown aerobically in a hemin-supplemented medium intensively consumed oxygen in the presence of 1 mM NADH. The addition of 0.2 mM menaquinone to the reaction mixture was accompanied by a 4.6-fold increase in NADH oxidation rate. Enzymes presumably involved in NADH oxidation by membranes and identified using MALDI-TOF MS/MS included pyridine nucleotide-disulfide oxidoreductase (Nox-2), NADH dehydrogenase 2 (Ndh-2), and ubiquinol oxidase bd subunit I (CydA). Therefore, in NADH oxidation, 80% of electron transport from NADH to oxygen occurred via Ndh-2, menaquinone, and bd-type quinol oxidase; as little as 20% were transfered via Nox-2. The study reports experimental evidence to support an electron transport chain functioning in L. rhamnosus CM MSU 529 under aerobic cultivation in the presence of hemin and menaquinone. The NADH oxidation rates of membrane preparations of lactic acid bacteria were measured for the first time. Additionally, this is the first time when the property of exogenous menaquinone to transfer electrons from Ndh-2 to bd-type quinol oxidase was demonstrated in vitro in lactic acid bacteria.


Similar content being viewed by others
REFERENCES
Hill, C., Guarner, F., Reid, G., Gibson, G.R., Merenstein, D.J., Pot, B., Morelli, L., Canani, R.B., Flint, H.J., Salminen, S., Calder, P.C., and San-ders, M.E., The international scientific association for probiotics and prebiotics consensus statement on the scope and appropriate use of the term probiotic, Nat. Rev. Gastroenterol. Hepatol., 2014, vol. 11, no. 8, pp. 506–514.
Moslem, P., Hossein, N., Mahdi, R., Seyed, N.H., and Seyed, A.S., Lactobacillus rhamnosus Gorbach-Goldin (GG): A top well-researched probiotic strain, J. Med. Microbiol., 2017, vol. 5, nos. 5–6, pp. 46–59.
Pedersen, M.B., Gaudu, P., Lechardeur, D., Petit, M.A., and Gruss, A., Aerobic respiration metabolism in lactic acid bacteria and uses in biotechnology, Annu. Rev. Food Sci. Technol., 2012, vol. 3, pp. 37–58.
Arioli, S., Zambelli, D., Guglielmetti, S., De Noni, I., Pedersen, M.B., Pedersen, P.D., Dal Bello, F., and Mora, D., Increasing the heme-dependent respiratory efficiency of Lactococcus lactis by inhibition of lactate dehydrogenase, Appl. Environ. Microbiol., 2013, vol. 79, no. 1, pp. 376–380.
Ianniello, R.G., Zotta, T., Matera, A., Genovese, F., Parente, E., and Ricciardi, A., Investigation of factors affecting aerobic and respiratory growth in the oxygen-tolerant strain Lactobacillus casei N87, PLoS One, 2016, vol. 11, no. 11, p. e0164065.
Zotta, T., Parente, E., and Ricciardi, A., Aerobic metabolism in the genus Lactobacillus: Impact on stress response and potential applications in the food industry, J. Appl. Microbiol., 2017, vol. 122, no. 4, pp. 857–869.
Siciliano, R.A. Pannella, G., Lippolis, R., Ricciardi, A., Mazzeo, M.F., and Zotta, T., Impact of aerobic and respirative life-style on Lactobacillus casei N87 proteome, Int. J. Food Microbiol., 2019, vol. 298, pp. 51–62.
Johanson, A., Goel, A., Olsson, L., and Franzén, C.J., Respiratory physiology of Lactococcus lactis in chemostat cultures and its effect on cellular robustness in frozen and freeze-dried starter cultures, Appl. Environ. Microbiol., 2020, vol. 86, no. 6, p. e02785–19.
Klimko, A.I., Cherdyntseva, T.A., Brioukhanov, A.L., and Netrusov, A.I., In vitro evaluation of probiotic potential of selected lactic acid bacteria strains, Probiotics Antimicrob. Proteins, 2020, vol. 12, no. 3, pp. 1139–1148.
Bradford, M.M., A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding, Anal. Biochem., 1976, vol. 72, nos. 1–2, pp. 248–254.
Teteneva, N.A., Mart’yanov, S.V., Esteban-López, M., Kahnt, J., Glatter, T., Netrusov, A.I., Plakunov, V.K., and Sourjik, V., Multiple drug-induced stress responses inhibit formation of Escherichia coli biofilms, Appl. Environ. Microbiol., 2020, vol. 86, no. 21, p. e01113–20.
Wagner, T., Wegner, C.E., Kahnt, J., Ermler, U., and Shima, S., Phylogenetic and structural comparisons of the three types of methyl coenzyme M reductase from Methanococcales and Methanobacteriales, J. Bacteriol., 2017, vol. 199, no. 16, p. e00197–17.
Eng, J.K., McCormack, A.L., and Yates, J.R., An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database, J. Am. Soc. Mass Spectrom., 1994, vol. 5, no. 11, pp. 976–989.
Ianniello, R.G., Zheng, J., Zotta, T., Ricciardi, A., and Gänzle, M.G., Biochemical analysis of respiratory metabolism in the heterofermentative Lactobacillus spicheri and Lactobacillus reuteri, J. Appl. Microbiol., 2015, vol. 119, no. 3, pp. 763–775.
Sakamoto, J., Koga, E., Mizuta, T., Sato, C., Noguchi, S., and Sone, N., Gene structure and quinol oxidase activity of a cytochrome bd-type oxidase from Bacillus stearothermophilus, Biochim. Biophys. Acta, 1999, vol. 1411, no. 1, pp. 147–158.
Averina, O.V., Poluektova, E.U., Marsova, M.V., and Danilenko, V.N., Biomarkers and utility of the antioxidant potential of probiotic lactobacilli and bifidobacteria as representatives of the human gut microbiota, Biomedicines, 2021, vol. 9, no. 10, p. e1340.
Marreiros, B.C., Sena, F.V., Sousa, F.M., Batista, A.P., Manuela, M., and Pereira, M.M., Type II NADH:quinone oxidoreductase family: phylogenetic distribution, structural diversity and evolutionary divergences, Environ. Microbiol., 2016, vol. 18, no. 12, pp. 4697–4709.
Funding
The study was performed as a part of the state assignment on the topic of the Department of Microbiology of Moscow State University “Physiology and Biochemistry of Phototrophic and Chemotrophic Microorganisms,” project no. 121032300094-7.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest. The authors declare that they have no conflicts of interest.
Statement on The Welfare of Animals. The article does not contain any studies involving animals or humans performed by any of the authors.
Additional information
Translated by E. Kuznetsova
About this article
Cite this article
Dinarieva, T.Y., Klimko, A.I., Cherdyntseva, T.A. et al. Vitamin K2 Mediates Electron Transport from NADH Dehydrogenase 2 to bd-type Quinol Oxidase in Lacticaseibacillus rhamnosus CM MSU 529. Moscow Univ. Biol.Sci. Bull. 77, 172–177 (2022). https://doi.org/10.3103/S0096392522030038
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.3103/S0096392522030038