Abstract
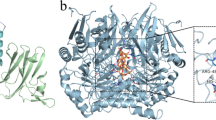
α-Amylases (1,4-α-D-glucanohydrolases) are widely used in starch liquefaction, but their acid stability needs to be continuously explored to reduce the costs of raw material and operation. In this study, to better meet the industrial requirements, the acid stability of Bacillus licheniformis α-amylase (BLA) was further improved by directed evolution using error prone polymerase chain reaction (PCR). The mutant BLA (G81R) was selected with the improved acid stability based on a high-throughput activity assay. After incubating at pH 4.5 for 40 min, G81R still retained 10% of its initial activity, but the wild-type (WT) was already inactive. The kcat/Km value of G81R at pH 4.5 was 1.4-fold higher than that of WT. Combined with the three-dimensional structural modeling analysis, the improved stability of G81R under low pH condition might be due to the interactions of electrostatic, hydrophilicity, and helix propensity. Therefore, these findings would be beneficial for developing BLA with properties suitable for applications in industrial starch processing.






Similar content being viewed by others
Abbreviations
- BAA:
-
Bacillus amyloliquefaciens α-amylase
- BLA:
-
Bacillus licheniformis α-amylase
- CAZy:
-
Carbohydrate-Active enZymes α-amylase
- GSA:
-
Geobacillus stearothermophilus α-amylase
- GTA:
-
Geobacillus thermoleovorans α-amylase
- IPTG:
-
isopropyl β-D-1-thiogalactopyranoside
- LB:
-
Luria-Bertani
- PCR:
-
polymerase chain reaction
- PFA:
-
Pyrococcus furiosus α-amylase
- SDS-PAGE:
-
sodiumdodecyl sulfate-polyacrylamide gel electrophoresis α-amylase
- THA:
-
Thermococcus hydrothermalis α-amylase
- WT:
-
wild-type
References
Alikhajeh J, Khajeh K, Ranjbar B, Naderimanesh H, Lin YH, Liu E, Guan HH, Hsieh YC, Chuankhayan P, Huang YC, Jeyaraman J, Liu MY, Chen CJ (2010) Structure of Bacillus amyloliquefaciens alpha-amylase at high resolution: implications for thermal stability. Acta Crystallogr Sect F 66:121–129. https://doi.org/10.1107/S1744309109051938
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22:195–201. https://doi.org/10.1093/bioinformatics/bti770
Asoodeh A, Emtenani S, Emtenani S, Jalal R, Housaindokht MR (2014) Molecular cloning and biochemical characterization of a thermoacidophilic, organic-solvent tolerant α-amylase from a Bacillus strain in Escherichia coli. J Mol Catal B Enzym 99:114–120. https://doi.org/10.1016/j.molcatb.2013.10.025
Bessler C, Schmitt J, Maurer KH, Schmid R (2003) Directed evolution of a bacterial alpha-amylase: toward enhanced pH-performance and higher specific activity. Protein Sci 12:2141–2149. https://doi.org/10.1110/ps.0384403
Božić N, Lončar N, Slavić MS, Vujčić Z (2017) Raw starch degrading α-amylases: an unsolved riddle. Amylase 1:12–25. https://doi.org/10.1515/amylase-2017-0002
Buchholz K, Kasche V, Bornscheuer UT (2012) Biocatalysts and enzyme technology, 2nd edn. Wiley-Blackwell, Hoboken
Declerck N, Machius M, Chambert R, Wiegand G, Huber R, Gaillardin C (1997) Hyperthermostable mutants of Bacillus licheniformis alpha-amylase: thermodynamic studies and structural interpretation. Protein Eng 10:541–549. https://doi.org/10.1093/protein/10.5.541
Dey TB, Kumar A, Banerjee R, Chandna P, Kuhad RC (2016) Improvement of microbial α-amylase stability: strategic approaches. Process Biochem 51:1380–1390. https://doi.org/10.1016/j.procbio.2016.06.021
Delano, WL (2002) PyMOL: An Open-Source Molecular Graphics tool. https://www.ccp4.ac.uk/newsletters/newsletter40/11_pymol.pdf
Fuwa H (1954) A new method f or microdetermination of amylase activity by the use of amylose as the substrate. J Biochem 41:583–603. https://doi.org/10.1093/oxfordjournals.jbchem.a126476
Gai Y, Chen J, Zhang S, Zhu B, Zhang D (2018) Property improvement of α-amylase from Bacillus stearothermophilus by deletion of amino acid residues arginine 179 and glycine 180. Food Technol Biotechnol 56:58–64. https://doi.org/10.17113/ftb.56.01.18.5448
Hmidet N, Bayoudh A, Berrin JG, Kanoun S, Juge N, Nasri M (2008) Purification and biochemical characterization of a novel α-amylase from Bacillus licheniformis NH1: cloning, nucleotide sequence and expression of amyN gene in Escherichia coli. Process Biochem 43:499–510. https://doi.org/10.1016/j.procbio.2008.01.017
Hwang KY, Song HK, Chang C, Lee J, Lee SY, Kim KK, Choe S, Sweet RM, Suh SW (1997) Crystal structure of thermostable alpha-amylase from Bacillus licheniformis refined at 1.7 Å resolution. Mol Cell 7:251–258. https://doi.org/10.1016/S0921-0423(96)80367-4
Janeček Š, Svensson B, MacGregor EA (2014) α-Amylase: an enzyme specificity found in various families of glycoside hydrolases. Cell Mol Life Sci 71:1149–1170. https://doi.org/10.1007/s00018-013-1388-z
Jiang T, Cai MH, Huang MM, He H, Lu J, Zhou XS, Zhang YX (2015) Characterization of a thermostable raw-starch hydrolyzing α-amylase from deep-sea thermophile Geobacillus sp. Protein Expr Purif 114:15–22. https://doi.org/10.1016/j.pep.2015.06.002
Jones A, Lamsa M, Frandsen TP, Spendler T, Harris P, Sloma A, Xu F, Nielsen JB, Cherry JR (2008) Directed evolution of a maltogenic α-amylase from Bacillus sp. TS-25. J Biotechnol 134:325–333. https://doi.org/10.1016/j.jbiotec.2008.01.016
Jørgensen S, Vorgias CE, Antranikian G (1997) Cloning, sequencing, characterization, and expression of an extracellular a-amylase from the Hyperthermophilic archaeon Pyrococcus furiosus in Escherichia coli and Bacillus subtilis. J Biol Chem 272:16335–16342. https://doi.org/10.1074/jbc.272.26.16335
Joyet P, Declerck N, Gaillardin C (1992) Hyperthermostable variants of a highly thermostable alpha-amylase. Bio/technology 10:1579–1583. https://doi.org/10.1038/nbt1292-1579
Karaki N, Aljawish A, Humeau C, Muniglia L, Jasniewski J (2016) Enzymatic modification of polysaccharides: mechanisms, properties, and potential applications: a review. Enzym Microb Technol 90:1–18. https://doi.org/10.1016/j.enzmictec.2016.04.004
Koropatkin NM, Cameron EA, Martens EC (2012) How glycan metabolism shapes the human gut microbiota. Nat Rev Microbiol 10:183–193. https://doi.org/10.1038/nrmicro2746
Kumar A, Singh S (2013) Directed evolution: tailoring biocatalysts for industrial applications. Crit Rev Biotechnol 33:365–378. https://doi.org/10.3109/07388551.2012.716810
Kuriki T, Imanaka T (1999) The concept of the α-amylase family: structural similarity and common catalytic mechanism. J Biosci Bioeng 87:557–565. https://doi.org/10.1016/S1389-1723(99)80114-5
Lee S, Mouri Y, Minoda M, Oneda H, Inouye K (2006) Comparison of the wild-type α-amylase and its variant enzymes in Bacillus amyloliquefaciens in activity and thermal stability, and insights into engineering the thermal stability of bacillus α-amylase. J Biochem 139:1007–1015. https://doi.org/10.1093/jb/mvj107
Lévêque E, Haye B, Belarbi A (2000) Cloning and expression of an K-amylase encoding gene from the hyperthermophilic archaebacterium Thermococcus hydrothermalis and biochemical characterisation of the recombinant enzyme. FEMS Microbiol 186:67–71. https://doi.org/10.1016/S0378-1097(00)00117-8
Liu YH, Lu FP, Li Y, Wang JL, Gao C (2008a) Acid stabilization of Bacillus licheniformis alpha-amylase through introduction of mutations. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-008-1580-5
Liu YH, Lu FP, Li Y, Yin XB, Wang Y, Gao C (2008b) Characterisation of mutagenised acid-resistant alpha-amylase expressed in Bacillus subtilis WB600. Appl Microbiol Biotechnol 78:85–94. https://doi.org/10.1007/s00253-007-1287-z
Liu YH, Hu B, Xu YJ, Bo JX, Fan S, Wang JL, Lu FP (2012) Improvement of the acid stability of Bacillus licheniformis alpha amylase by error-prone PCR. J Appl Microbiol 113:541–549. https://doi.org/10.1111/j.1365-2672.2012.05359.x
Liu YH, Fan S, Liu XG, Zhang ZM, Wang JL, Wang ZX, Lu FP (2014) A highly active alpha amylase from Bacillus licheniformis: directed evolution, enzyme characterization and structural analysis. J Microbiol Biotechnol 24:898–904. https://doi.org/10.4014/jmb.1402.02004
Liu YH, Huang L, Jia LB, Gui S, Fu Y, Zheng D, Guo W, Lu FP (2017) Improvement of the acid stability of Bacillus licheniformis alpha amylase by site-directed mutagenesis. Process Biochem 58:174–180. https://doi.org/10.1016/j.procbio.2017.04.040
Lombard V, Golaconda RH, Drula E, Coutinho PM, Henrissat B (2013) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:D490–D495. https://doi.org/10.1093/nar/gkt1178
Macgregor EA, Winnipeg M (1993) Relationships between structure and activity in the α-amylase family of starch-metabolising enzymes. Starch-Starke 45:232–237. https://doi.org/10.1002/star.19930450705
McCarter JD, Withers SG (1996) Unequivocal identification of Asp-214 as the catalytic nucleophile of Saccharomyces cerevisiae α-glucosidase using 5-fluoro glycosyl fluorides. J Biol Chem 271:6889–6894. https://doi.org/10.1074/jbc.271.12.6889
Mehta D, Satyanarayana T (2013) Biochemical and molecular characterization of recombinant acidic and thermostable raw-starch hydrolysing α-amylase from an extreme thermophile Geobacillus thermoleovorans. J Mol Catal B Enzym 85-86:229–238. https://doi.org/10.1016/j.molcatb.2012.08.017
Nielsen JE, Beier L, Otzen D, Borchert TV, Frantzen HB, Andersen KV, Svendsen A (1999) Electrostatics in the active site of an alpha-amylase. Eur J Biochem 264:816–824. https://doi.org/10.1046/j.1432-1327.1999.00664.x
Nielsen JE, Borchert TV, Vriend G (2001) The determinants of α-amylase pH–activity profiles. Protein Eng 14:505–512. https://doi.org/10.1093/protein/14.7.505
Park JT, Song HN, Jung TY, Lee MH, Park SG, Woo EJ, Park KH (2013) A novel domain arrangement in a monomeric cyclodextrin-hydrolyzing enzyme from the hyperthermophile Pyrococcus furiosus. Biochim Biophys Acta 1834:380–386. https://doi.org/10.1016/j.bbapap.2012.08.001
Priyadharshini R, Manoharan S, Hemalatha D, Gunasekaran P (2010) Repeated random mutagenesis of α-Amylase from Bacillus licheniformis for improved pH performance. J Microbiol Biotechnol 20:1696–1701. https://doi.org/10.4014/jmb.1008.08013
Pujadas G, Palau J (2001) Evolution of α-amylases: architectural features and key residues in the stabilization of the (β/α)8 scaffold. Mol Biol Evol 18:38–54. https://doi.org/10.1093/oxfordjournals.molbev.a003718
Sharma A, Satyanarayana T (2013) Microbial acid-stable α-amylases: characteristics, genetic engineering and applications. Process Biochem 48:201–211. https://doi.org/10.1016/j.procbio.2012.12.018
Sindhu R, Binod P, Madhavan A, Beevi US, Mathew AK, Abraham A, Pandey A, Kumar V (2017) Molecular improvements in microbial α-amylases for enhanced stability and catalytic efficiency. Bioresour Technol 245:1740–1748. https://doi.org/10.1016/j.biortech.2017.04.098
Uitdehaag JCM, Mosi R, Kalk KH, van der Veen BA, Dijkhuizen L, Withers SG, Dijkstra BW (1999) X-ray structures along the reaction pathway of cyclodextrin glycosyltransferase elucidate catalysis in the α-amylase family. Nat Struct Biol 6:432–436. https://doi.org/10.1038/8235
Vamadevan V, Bertoft E (2015) Structure-function relationships of starch components. Starch-Starke 67:55–68. https://doi.org/10.1002/star.201400188
Wind RD, Uitdehaag JCM, Buitelaar RM, Dijkstra BW, Dijkhuizen L (1998) Engineering of cyclodextrin product specificity and pH optima of the thermostable cyclodextrin glycosyltransferase from Thermoanaerobacterium thermosulfurigenes EM1. J Biol Chem 273:5771–5779. https://doi.org/10.1074/jbc.273.10.5771
Zhang QG, Han Y, Xiao HZ (2017) Microbial α-amylase: a biomolecular overview. Process Biochem 53:88–101. https://doi.org/10.1016/j.procbio.2016.11.012
Acknowledgments
This work was supported by the National Key R&D Program of China (2017YFD0201405-04); the China Postdoctoral Science Foundation (2018 M641660); the Tianjin Natural Science Fund of China (17JCYBJC23700); and the Public Service Platform Program of Tianjin Engineering Research Center of Microbial Metabolism and Fermentation Process Control (17PTGCCX00190).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Huang, L., Shan, M., Ma, J. et al. Directed evolution of α-amylase from Bacillus licheniformis to enhance its acid-stable performance. Biologia 74, 1363–1372 (2019). https://doi.org/10.2478/s11756-019-00262-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-019-00262-7