Abstract
Background
Antimicrobial resistance affects the treatment of several bacterial infections, including pneumonia. This subsequently increased the morbidity and mortality rates of patients with bacterial pneumonia, especially in resource-limited settings. In this study, we aimed to determine the patterns of multidrug-resistant (MDR) bacteria isolated from the sputum samples of patients with bacterial pneumonia attending a tertiary hospital in Tanzania.
Methodology
A retrospective cross-sectional study was conducted. It involved reviewing the laboratory sputum data in the laboratory information system at Muhimbili National Hospital in Tanzania. The sputum samples were previously processed using standard methods (culture, Gram staining, and biochemical tests) to isolate and identify the bacteria. At the same time, antibiogram profiles were determined using antimicrobial susceptibility tests. Bacterial isolates that expressed MDR patterns were identified. Demographic information was collected from patients' medical records. We used the chi-square test to determine factors associated with MDR. A p-value < 0.05 was considered significant.
Results
We retrieved and analysed 169 laboratory records of patients with a provisional clinical diagnosis of bacterial pneumonia confirmed in the microbiology laboratory. Nearly 98% of the records were from adult patients. The patients’ mean age was 48.3 years and 17.3 standard deviations. About 84% of the isolated bacteria were Gram-negative; the most predominant was Klebsiella pneumoniae (59/142; 41.5%). The predominant Gram-positive bacteria was Staphylococcus aureus (25/27; 92.6%). Furthermore, 80 out of 169 (47.3%) bacteria were MDR; Klebsiella pneumoniae (32.5%) was predominant. In addition, 50% of Staphylococcus aureus was methicillin resistance. MDR bacterial pneumonia was highly observed in patients admitted to the Intensive Care Unit (p < 0.05).
Conclusions
Although our study was limited by variations in the number of bacterial isolates subjected to the same antibiotic drugs and a lack of information on risk factors such as occupation, smoking history, and marital status, we observed that a high proportion of bacterial pneumonia is caused by MDR Gram-negative bacteria in our local setting. These results inform the need to improve infection prevention control measures in hospitals to reduce the burden of MDR bacteria in our settings and other similar resource-limited settings.
Similar content being viewed by others
Background
Pneumonia is one of the diseases associated with high morbidity and mortality globally. It may result in severe complications such as sepsis, coagulopathy, respiratory failure, lung fibrosis, and abscess (Mani 2018). Bacterial pneumonia, especially caused by multidrug-resistant (MDR) bacteria, is usually associated with severe complications (Ferreira-Coimbra et al. 2020). Generally, infections due to MDR bacteria are an increasing threat to public health (Bassetti and Righi 2013; Catalano et al. 2022; Tanwar et al. 2014). This effect is observed in several bacterial infections, such as bacterial pneumonia, whereby the intensifying rates of antimicrobial resistance (AMR) have increased morbidity and mortality (Salam et al. 2023). MDR-associated infections are usually hard to treat and sometimes are associated with poor clinical outcomes and prolonged hospitalisation (Parmanik et al. 2022). It is estimated that globally, 4·95 million deaths occurred due to bacterial AMR in 2019. The highest death rate at 27.3 deaths per 100,000 (20.9–35.3) was reported in the western region of sub-Saharan Africa (Antimicrobial Resistance Collaborators 2022).
Thus, MDR bacteria that cause pneumonia are also difficult to treat and often lead to prolonged hospitalisation and treatment, which may sometimes require the prolonged use of multiple broad-spectrum antibiotics. Prolonged hospital stays and treatment of MDR-associated pneumonia subsequently lead to an increase in healthcare costs (Hanberger et al. 2019; Rodriguez and Surani 2017).
In Tanzania, bacterial pneumonia has been investigated and reported (Kishimbo et al. 2020; Moremi et al. 2016; Wilfred et al. 2021). However, there is limited information on the burden of MDR bacteria in the context of bacterial pneumonia, specifically the specific prevalence and types of MDR bacteria that cause pneumonia. Therefore, the present study aimed to determine the trends of MDR bacteria isolated from the sputum samples of patients presenting with pneumonia attending a tertiary hospital in Tanzania. Evaluation of the contribution of MDR bacteria causing pneumonia in our local settings is important to guide suitable public health interventions to curb AMR and appropriate treatment options in resource-limited settings where empirical treatment is often practised.
Methods
Study design
A retrospective cross-sectional study was conducted. It involved reviewing the medical records and laboratory results of patients whose sputum samples were processed at the Central Pathology Laboratory (CPL) at Muhimbili National Hospital (MNH).
Study setting
CPL is accredited according to ISO 15189 by the Southern African Development Community Accreditation Service. At CPL, sputum samples from patients suspected of bacterial pneumonia are collected using a clean container and transported for processing within two (2) hours of collection. The samples are usually processed at the bacteriology unit for culture and antimicrobial susceptibility test (AST). The unit receives an average of 60 sputum samples monthly. The samples are processed effectively to detect and identify potential pathogens and the antimicrobial susceptibility patterns of the isolated bacteria. The sputum samples are screened for quality using Bartlett’s grading system. Sputum samples with more than 25 polymorphonuclear leukocytes and less than ten squamous epithelial cells are considered active infections. These samples are usually inoculated on MacConkey agar and sheep blood agar, then incubated at standard conditions (37 °C for 24 h) and in 5% carbon dioxide on Chocolate agar. Colony morphology, Gram staining, and conventional biochemical tests are applied to identify the isolated bacteria. Biochemical tests for Gram-positive bacteria, catalase and coagulase tests, and Gram-negative bacteria, glucose/sucrose fermentation, hydrogen sulphide production, indole production, urease production, citrate utilisation, and motility are performed for bacteria identification. An analytical profile index test (API20E) is usually used to identify Gram-negative bacteria that had not been identified using conventional techniques.
Routinely, AST is performed using the Kirby–Bauer disc diffusion technique on Mueller–Hinton agar as per the Clinical and Laboratory Standards Institute (CLSI) (CLSI 2022). Briefly, all antibiotic categories are included as per CLSI, including amoxicillin/clavulanic acid (20/10 μg), ciprofloxacin (5 μg), trimethoprim/sulfamethoxazole (1.25/23.75 μg), gentamycin (10 μg), erythromycin (15 μg), clindamycin (2 μg), amikacin (30 μg), meropenem (10 μg), ceftriaxone (30 μg), and ceftazidime (30 μg). The cefoxitin disc diffusion test detects methicillin-resistant S. aureus (MRSA) as per CLSI 2022 M100 32nd edition (CLSI 2022). ESBL production was screened using ceftazidime (30 μg) or cefotaxime (30 μg) antibiotic discs, and phenotypic confirmation was done using a double-disc synergy test (Dhara et al. 2012). The culture and susceptibility results are reported through the laboratory information system (LIS).
Study population
The study involved patients who attended MNH between 4th April and 30th June 2022 and met the inclusion criteria of having a provisional diagnosis of bacterial pneumonia. Their sputum samples were collected for culture and antimicrobial susceptibility testing at CPL.
Sample size estimation
The sample size for our retrospective cross-sectional study was calculated based on an expected proportion of MDR bacteria, estimated at 46.3% (Kalita et al. 2021), with a margin of error set at 8%. Using the Kish and Leslie formula provided a minimum sample size of 149. However, during the data collection from April 4th to June 30th, 2022, 169 eligible sputum records were identified that met the inclusion criteria. Given that these records were all suitable for inclusion and provided a larger dataset for robust analysis, we decided to include all 169 records. This decision was driven by the ethical use of data, which promotes using all available data to contribute to scientific knowledge without additional cost or participant burden.
Data collection and procedures
Records of 169 patients with provisional pneumonia diagnoses whose sputum samples were processed for culture and AST to identify the pathogenic bacteria during the study period were analysed. Data collection involved reviewing the patient’s medical records and laboratory results. First, the eligible laboratory sputum data from 4th April to 30th June 2022 were retrieved from the LIS. Then, the patient’s medical records were reviewed for data completeness: whether all the study variables were complete. We also checked whether the participants were provisionally diagnosed clinically with bacterial pneumonia (having fever, tachypnoea, tachycardia, and productive/non-productive cough). Thereafter, the individually selected data was extracted from the LIS Jeeva software. The socio-demographic and clinical data (age, gender, ward admitted, signs and symptoms of pneumonia) of these participants whose laboratory records were selected were retrieved from participants’ medical records.
Data processing and analysis
The participants’ laboratory, socio-demographic, and clinical data were entered and merged into Microsoft Excel software version 2019. Thereafter, the data were cleaned, and data quality control was conducted. Next, the data were exported into SPSS software version 26 for statistical analysis, whereby independent and dependent variables were tabulated. Data such as socio-demographic characteristics and distribution of bacterial isolates from sputum samples were presented in proportions, charts and tables. We used the chi-square test to determine the association between participants’ factors and MDR. The antimicrobial resistance rate was calculated by dividing the number of specific bacterial isolates that resisted a specific drug over the sum of isolates of the same species. Bacterial isolates with intermediate AST results were considered resistant. MDR was determined when the bacteria isolate was resistant to one or more antibiotic agents in three or more antibiotic classes.
Results
Socio-demographic characteristics
The study analysed the records of 169 patients who met the selection criteria by having a provisional diagnosis of bacterial pneumonia microbiologically confirmed at the laboratory at MNH. The majority, 97.6% (165/169), were adults aged 18 years old and above; the mean age of adults was 48.3 years and standard deviation (SD) of 17.3, while the mean age of children was 10.3 years, SD ± 8.4. Most participants, 54.4% (92/169), were female. The majority, 74%, were inpatients; > 30% of the inpatients were admitted to the general medical ward. The inpatient participants had an average of 5 days of hospital stay. We observed that 5.3% (9/169) were in the intensive care unit (ICU), only 2 out 9 participants were on mechanical ventilators, and their duration of hospital stay ranged between two weeks and three months. In this study, the patients who attended the outpatient department (OPD) contributed to 20.7% (35/169) of the study participants (Table 1).
Bacteria isolated from sputum samples
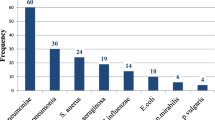
In this study, among the 169 laboratory records that were analysed, we observed that most of the isolated bacteria, 142/169 (84%), were Gram-negative. Among the Gram-negative bacteria, the most predominant was Klebsiella pneumoniae (59/142; 41.5%), followed by Pseudomonas aeruginosa (27/142; 19%) and Klebsiella oxytoca (13/142; 9.2%). In contrast, among the Gram-positive bacteria, the predominant was Staphylococcus aureus (25/27; 92.6%) and Streptococcus pneumonia attributed to (2/27; 7.4%). The spectrum of bacterial species isolated from sputum samples among the patients with bacterial pneumonia who attended MNH is summarised in Fig. 1.
Antimicrobial resistance patterns
Among the Gram-negative bacteria, resistance to gentamicin was observed in 22.9% (8/35) of Klebsiella spp and 16.7% (3/18) of P. aeruginosa isolates, while 14.3% (6/42) and 14.3% (2/14) of Klebsiella spp and P. aeruginosa isolated respectively, were amikacin resistant. Generally, the Gram-negative bacteria showed high resistance of at least 40.0% and above for co-trimoxazole, augmentin and meropenem. Klebsiella spp and P. aeruginosa displayed minimal resistance to ciprofloxacin, 11.5% (12/62) and 12.5% (3/24), respectively. Other Gram-negative bacteria analysed have been summarised in Table 2.
S. aureus was dominant among the Gram-positive bacteria, at 92.6% (25/27). S. aureus also displayed high resistance to penicillin 82.4% (14/17), erythromycin 68.4% (13/19) and clindamycin 66. 7% (10/15) (Table 2). In this study, only 22 out of 25 (88%) S. aureus isolates were screened for MRSA using cefoxitin disc diffusion test. Out of 22 S. aureus isolates, 50% (11/22) were MRSA. Among the MRSA isolates, 8 out of 11 were tested for susceptibility to vancomycin, and we observed that 75.0% (6/8) were resistant to vancomycin.
Proportion of MDR bacteria
The proportion of MDR bacteria accounted for 47.3% (80/169) of the total bacteria isolated (Fig. 2a). Among MDR bacteria, the predominant bacteria were Klebsiella pneumoniae (32.5%) and Staphylococcus aureus (27.5%) (Fig. 2b). Further stratification of MDR proportions according to patients’ characteristics revealed that patients admitted to the ICU had more MDR bacteria (88.9%) than other wards 42.9–45.6%) (p < 0.05), and we observed that all were Gram-negative bacteria with P. aerugonisa being the predominant bacteria (3/8). We did not observe statistically significant differences in the proportion of MDR with the age and sex of participants (Table 1).
Discussion
Understanding the aetiology of bacterial pneumonia and the bacteria resistance patterns is crucial in planning for public interventions and managing bacterial pneumonia during this era, where AMR is rising globally. The present study aimed to determine the trends of MDR bacteria isolated from the sputum samples and their antimicrobial susceptibility patterns among patients with bacterial pneumonia at a tertiary hospital in Tanzania. Our study has revealed that the aetiology of bacterial pneumonia is mainly (84.0%) attributed to Gram-negative bacteria in our setting. Our study has shown that K. pneumoniae accounted for the most isolated bacteria from the sputum samples, followed by P. aeruginosa among Gram-negative bacteria. This trend is similar to findings obtained from other studies in Tanzania and elsewhere that have reported Gram-negative bacteria, specifically Klebsiella spp, which are frequently associated with bacterial pneumonia in both community and hospital-acquired pneumonia (Kishimbo et al. 2020; Yang et al. 2023). In the present study, we have observed that S. aureus was predominant among Gram-positive bacteria; this trend concurs with the findings reported in a study that assessed the aetiology of community-acquired pneumonia in the adult population in a similar setting in Mwanza, Tanzania (Aliberti et al. 2013; Self et al. 2016). Our study's predominant Gram-negative bacteria aetiology is expected as more than half of the patients were inpatients. Previous studies have linked this with the shift of pathogenic bacteria trends and selective pressure of antibiotics due to increased antibiotic use (Arancibia et al. 2002; Kishimbo et al. 2020; Minch and Regasa 2014; Mussema et al. 2023). Furthermore, the majority of participants in our study were adults; hence, the aetiology of pneumonia observed in our study may also be influenced by the organisms colonising the oropharynx and gastrointestinal tract because aspiration from the upper respiratory tract to the lungs is the most common transport route (Siegel and Weiser 2015). In addition, an exogenous source of bacteria such as MDR microbial biota contracted during hospitalisation contributes significantly to the aetiology of bacterial pneumonia in adult patients in our settings (Assefa 2022; Tozzo et al. 2022).
The present study has also revealed that 47.3% of bacteria causing bacterial pneumonia are MDR. This finding concurs with other studies that have reported a high proportion of MDR bacteria causing community and hospital-acquired pneumonia in similar settings (Falcone et al. 2015, 2021; Reynolds et al. 2022; Temesgen et al. 2019). These findings indicate that AMR continues to rise, hence becoming more of a public health concern that subsequently impacts morbidity and mortality and increases treatment costs. This calls for urgent action, including implementing antimicrobial stewardship programmes to counteract AMR in our settings (Guclu et al. 2021; World Health Organization (WHO) 2021). Our study has shown that many participants admitted to the ICU had MDR bacteria. This finding correlates with another study done in Mwanza, Tanzania, and other settings elsewhere, where MDR bacteria isolated from critical care units were shown to be highly prevalent (Ferreira et al. 2019; Han et al. 2022; Silago et al. 2020; Wu et al. 2023). The overall prevalence of MDR and specifically high MDR prevalence in ICU could be because more than 90% of the participants were adults and inpatients, and adults normally have increased exposure to antibiotics. Also, the hospital environment increases the risk of MDR bacteria exposure (Assefa 2022; Bassetti and Righi 2013). In addition, in the ICU, frequent use of antibiotics and surface contaminations could have contributed to MDR, as was reported in a study that assessed the predominance of MDR Gram-negative bacteria on contaminated hospital surfaces in Tanzania and elsewhere (Joachim et al. 2023; Waele et al. 2020).
Our study has shown that most K. pneumoniae isolated are MDR bacteria. This trend is also similar to what has been reported previously in Tanzania and other parts of the world, where studies have shown that most K. pneumoniae isolated have shown to be MDR(Awoke et al. 2021; Hao et al. 2022; Kalita et al. 2021). These findings further support other studies showing evidence of an emergence and global expansion of hypervirulent and MDR clones of K. pneumoniae, which have been implicated in both community and acquired nosocomial infections (Arcari and Carattoli 2023; Raro et al. 2023; Shankar et al. 2020). Furthermore, our study has shown that 50.0% of S. aureus isolated from sputum samples of patients with bacterial pneumonia are MRSA. This finding concurs with other similar studies that have reported MRSA is among the frequent causes of pneumonia (Francis et al. 2005; Moran et al. 2012; Rubinstein et al. 2008). A high proportion of 77.7% of MRSA was also reported in developed countries such as the USA among inpatients with pneumonia (Pharmd et al. 2016). The present study suggests the need for routine surveillance of bacterial infection and resistance to antibiotics. Larger sample size studies are warranted to inform clinicians about the local antibiogram and sensitise them to adhere to an evidence-based approach in treating patients based on local data. This will aid in avoiding unnecessary antibiotic use and optimising the duration of therapy, which will decrease morbidity and mortality caused by MDR bacteria.
Our study has some limitations. The bacterial isolates were not all tested to the same panel of antibiotic agents; therefore, we could not collectively report the antimicrobial susceptibility data of all the isolated bacteria. We were also not able to capture some important information such as occupation, smoking history and even marital status from both the LIS and participants’ medical records. Further studies are warranted to investigate these factors' contribution to bacterial pneumonia and determine clone linage of the MDR Klebsiella pneumoniae and MRSA.
Conclusions
Our study found a significant prevalence of MDR bacteria in patients with typical bacterial pneumonia, primarily caused by K. pneumoniae and S. aureus. We observed that MDR bacterial pneumonia was common among patients admitted to the ICU. These results inform the need to improve infection prevention and control measures to curb the burden of MDR.
Availability of data and materials
Data generated in this study are available from the corresponding author upon a reasonable request.
Abbreviations
- AMR:
-
Antimicrobial resistance
- AST:
-
Antimicrobial susceptibility test
- CPL:
-
Central Pathology Laboratory
- ICU:
-
Intensive Care Unit
- LIS:
-
Laboratory information system
- MDR:
-
Multidrug resistant
- MNH:
-
Muhimbili National Hospital
- MRSA:
-
Methicillin resistance Staphylococcus aureus
References
Aliberti S, Cilloniz C, Chalmers JD, Zanaboni AM, Cosentini R, Tarsia P, Pesci A, Blasi F, Torres A (2013) Multidrug-resistant pathogens in hospitalised patients coming from the community with pneumonia: a European perspective. Respir Infect 68:997–999. https://doi.org/10.1136/thoraxjnl-2013-203384
Antimicrobial Resistance Collaborators (2022) Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis, vol 399, pp 629–55
Arancibia F, Bauer TT, Ewig S, Mensa J, Gonzalez J, Niederman MS, Torres A (2002) Community-acquired pneumonia due to gram-negative bacteria and Pseudomonas aeruginosa: incidence, risk, and prognosis. Arch Intern Med 162(16):1849–1858. https://doi.org/10.1001/archinte.162.16.1849
Arcari G, Carattoli A (2023) Global spread and evolutionary convergence of multidrug-resistant and hypervirulent Klebsiella pneumoniae high-risk clones. Pathog Glob Health 117(4):328–341. https://doi.org/10.1080/20477724.2022.2121362
Assefa M (2022) Multi-drug resistant gram-negative bacterial pneumonia: etiology, risk factors, and drug resistance patterns. Pneumonia 1(14):4. https://doi.org/10.1186/s41479-022-00096-z
Awoke T, Teka B, Seman A, Sebre S, Yeshitela B (2021) High prevalence of multidrug-resistant klebsiella pneumoniae in a tertiary care hospital in Ethiopia. Antibiotics 10(1007):1–9. https://doi.org/10.3390/antibiotics10081007
Bassetti M, Righi E (2013) Multidrug-resistant bacteria: What is the threat? Am Soc Hematol 1(2013):428–432. https://doi.org/10.1182/asheducation-2013.1.428
Catalano A, Iacopetta D, Ceramella J, Scumaci D, Giuzio F, Saturnino C, Aquaro S, Rosano C, Sinicropi MS (2022) Multidrug resistance (MDR): a widespread phenomenon in pharmacological therapies. Molecules 3(27):1–18. https://doi.org/10.3390/molecules27030616
CLSI (2022) Performance standards for antimicrobial susceptibility testing, 32nd edn
De Waele JJ, Boelens J, Leroux-roels I (2020) Multidrug-resistant bacteria in ICU: fact or myth. Curr Opin Anesthesiol 33(2):156–161. https://doi.org/10.1097/ACO.0000000000000830
Dhara M, Disha P, Sachin P, Manisha J, Seema B, Vegad MM (2012) Comparison of various methods for the detection of extended spectrum beta lactamase in Klebsiella pneumoniae isolated from neonatal Intensive Care Unit. Ahmedabad Natl J Med Res 2(3):348–353
Falcone M, Russo A, Giannella M, Cangemi R, Scarpellini G, Bertazzoni G, Alarcón JM, Taliani G (2015) Individualizing risk of multidrug-resistant pathogens in community-onset pneumonia. PLoS ONE 10(4):1–16. https://doi.org/10.1371/journal.pone.0119528
Falcone M, Tiseo G, Menichetti F (2021) Community-acquired pneumonia owing to multidrug-resistant pathogens: a step toward an early identification. Ann Am Thorac Soc 2(18):211–213. https://doi.org/10.1513/AnnalsATS.202009-1207ED
Ferreira RL, da Silva BC, Rezende GS, Nakamura-Silva R, Pitondo-Silva A, Campanini EB, Brito MC (2019) High prevalence of multidrug-resistant klebsiella pneumoniae harboring several virulence and β-lactamase encoding genes in a brazilian intensive care unit. Front Microbiol. https://doi.org/10.3389/fmicb.2018.03198
Ferreira-Coimbra J, Sarda C, Rello J (2020) Burden of community-acquired pneumonia and unmet clinical needs. Adv Ther 37(4):1302–1318
Francis JS, Doherty MC, Lopatin U, Johnston CP, Sinha G, Ross T, Cai M, Hansel NN, Perl T, Ticehurst JR, Carroll K, Thomas DL, Nuermberger E, Bartlett JG (2005) Severe community-onset pneumonia in healthy adults caused by methicillin-resistant Staphylococcus aureus carrying the panton-valentine leukocidin genes. Clin Infect Dis 40(1):100–107. https://doi.org/10.1086/427148
Guclu AU, Kocak AA, Ok MA, Tutluoglu B, Basustaoglu A (2021) Antibacterial resistance in lower respiratory tract bacterial pathogens: a multicenter analysis from Turkey. J Infect Dev Ctries 15(2):254–262. https://doi.org/10.3855/JIDC.12599
Han Y, Zhang J, Zhang H, Zhang X, Wang Y (2022) Multidrug-resistant organisms in intensive care units and logistic analysis of risk factors. World J Clin Cases 10(6):1795–1806. https://doi.org/10.12998/wjcc.v10.i6.1795
Hanberger H, Hoang NTB, Tran DM, Larsson M (2019) Multiple antibiotic resistance as a risk factor for mortality and prolonged hospital stay: a cohort study among neonatal intensive care patients with hospital-acquired infections caused by gram-negative bacteria in Vietnam. PLoS ONE 5(14):1–18. https://doi.org/10.1371/journal.pone.0215666
Hao Y, Jiang Y, Ishaq HM, Liu W, Zhao H, Wang M, Yang F (2022) Molecular characterization of klebsiella pneumoniae isolated from sputum in a tertiary hospital in Xinxiang, China. Infect Drug Resist 15:3829–3839
Joachim A, Manyahi J, Issa H, Lwoga J, Msafiri F, Majigo M (2023) Predominance of multidrug-resistant gram-negative bacteria on contaminated surfaces at a tertiary hospital in Tanzania: a call to strengthening environmental infection prevention and control measures statistical packages for social sciences. Curr Microbiol 80(5):148. https://doi.org/10.1007/s00284-023-03254-8
Kalita D, Sarma R, Sharma K, Deka S (2021) High proportion of drug-resistant isolates in adult community-acquired pneumonia from Northeast India: a hospital-based study. Lung India 38(5):460. https://doi.org/10.4103/LUNGINDIA.LUNGINDIA_978_20
Kishimbo P, Sogone NM, Kalokola F, Mshana SE (2020) Prevalence of gram negative bacteria causing community acquired pneumonia among adults in Mwanza City, Tanzania. Pneumonia 12:1–9. https://doi.org/10.1186/s41479-020-00069-0
Mani CS (2018) Acute pneumonia and its complications. Princ Pract Pediatr Infect Dis 2018(4):238–249. https://doi.org/10.1016/B978-0-323-40181-4.00034-7
Minch A, Regasa B (2014) Medical microbiology & diagnosis drug resistance patterns of bacterial pathogens from adult patients with. J Med Microbiol Diagn 3(4):151. https://doi.org/10.4172/2161-0703.1000151
Moran GJ, Krishnadasan A, Gorwitz RJ, Fosheim GE, Albrecht V, Limbago B (2012) Prevalence of methicillin-resistant Staphylococcus aureus as an etiology of community-acquired pneumonia. Clin Infect Dis 54(8):1126–1133. https://doi.org/10.1093/cid/cis022
Moremi N, Claus H, Mshana SE (2016) Antimicrobial resistance pattern: a report of microbiological cultures at a tertiary hospital in Tanzania. BMC Infect Dis 16:1–7. https://doi.org/10.1186/s12879-016-2082-11
Mussema A, Beyene G, Gudina EK, Alelign D (2023) Bacterial etiology, antimicrobial resistance and factors associated with community acquired pneumonia among adult hospitalized patients in Southwest Ethiopia. Iran J Microbiol 15(4):492–502. 10.18502%2Fijm.v15i4.13503
Parmanik A, Das S, Kar B, Bose A, Dwivedi GR, Pandey MM (2022) Current treatment strategies against multidrug-resistant bacteria: a review. Curr Microbiol 79(12):1–15. https://doi.org/10.1007/s00284-022-03061-7
Pharmd DMJ, Ba AS, Words K (2016) Prevalence of and outcomes from Staphylococcus aureus pneumonia among hospitalized patients in the United States, 2009–2012. Am J Infect Control 45(4):404–409. https://doi.org/10.1016/j.ajic.2016.11.014
Raro HF, Otávio A, Nordmann P, Pino MD, Findlay J, Poirel L (2023) Emergence of carbapenemase-producing hypervirulent klebsiella pneumoniae in Switzerland. Antimicrob Agents Chemother 67(3):1–7. https://doi.org/10.1128/aac.01424-22
Reynolds D, Burnham JP, Guillamet CV, McCabe M, Yuenger V, Betthauser K, Micek ST, Kollef MH (2022) The threat of multidrug-resistant/extensively drug-resistant Gram-negative respiratory infections: another pandemic. Eur Respir Rev. https://doi.org/10.1183/16000617.0068-2022
Rodriguez M, Surani SR (2017) Multidrug-resistant gram-negative pneumonia and infection in Intensive Care Unit. Contemp Top Pneumonia. https://doi.org/10.5772/intechopen.69377
Rubinstein E, Kollef MH, Nathwani D (2008) Pneumonia caused by methicillin-resistant Staphylococcus aureus. Clin Infect Dis 46(Suppl 5):S378–S385. https://doi.org/10.1086/533594
Salam A, Al-amin Y, Salam MT, Pawar JS (2023) Antimicrobial resistance: a growing serious threat for global. Threat Glob Public Health 11(13):1–20. https://doi.org/10.3390/healthcare11131946
Self WH, Wunderink RG, Williams DJ, Zhu Y, Anderson EJ, Balk RA, Fakhran SS, Chappell JD, Casimir G, Courtney DM, Trabue C, Waterer GW, Bramley A, Magill S, Jain S, Edwards KM, Grijalva CG (2016) Staphylococcus aureus community-acquired pneumonia : prevalence, clinical characteristics, and outcomes. Clin Infect Dis 63:300–309. https://doi.org/10.1093/cid/ciw300
Shankar C, Jacob JJ, Vasudevan K, Biswas R (2020) Emergence of multidrug resistant hypervirulent ST23 Klebsiella pneumoniae: multidrug resistant plasmid acquisition drives evolution. Front Cell Infect Microbiol 10:1–13. https://doi.org/10.3389/fcimb.2020.575289
Siegel SJ, Weiser JN (2015) Mechanisms of bacterial colonization of the respiratory tract. Annu Rev Microbiol 16:425–444. https://doi.org/10.1146/annurev-micro-091014-104209.Mechanisms
Silago V, Kovacs D, Msanga DR, Seni J, Matthews L, Oravcová K, Zadoks RN, Lupindu AM, Hoza AS, Mshana SE (2020) Bacteremia in critical care units at Bugando Medical Centre, Mwanza, Tanzania: the role of colonization and contaminated cots and mothers’ hands in cross-transmission of multidrug resistant Gram-negative bacteria. Antimicrob Resist Infect Control 9(58):1–14. https://doi.org/10.1186/s13756-020-00721-w
Tanwar J, Das S, Fatima Z, Hameed S (2014) Multidrug resistance: an emerging crisis. Interdiscip Perspect Infect Dis 2014:1–7. https://doi.org/10.1155/2014/541340
Temesgen D, Bereded F, Derbie A (2019) Bacteriology of community acquired pneumonia in adult patients at Felege Hiwot Referral Hospital, Northwest Ethiopia: a cross-sectional study. Antimicrob Resist Infect Control 8:1–8. https://doi.org/10.1186/s13756-019-0560-0
Tozzo P, Delicati A, Caenazzo L (2022) Human microbiome and microbiota identification for preventing and controlling infections: a systematic review. Front Public Health. https://doi.org/10.3389/fpubh.2022.989496
Wilfred A, Sandi E, Lubinza C, Kahwa A, Petrucka P, Mfinanga S (2021) Antibiotic susceptibility patterns of bacterial isolates from routine clinical specimens from referral hospitals in Tanzania: a prospective hospital-based observational study. Infect Drug Resist 14:869–878
World Health Organization (WHO) (2021) Antimicrobial stewardship interventions: a practical guide
Wu C, Lu J, Ruan L, Yao J (2023) Tracking epidemiological characteristics and risk factors of multi-drug resistant bacteria in intensive care units. Infect Drug Resist 16:1499–1509
Yang W, Yao H, Xi C, Ye X, Chen Q (2023) Prevalence and clinical characteristics of bacterial pneumonia in neurosurgical emergency center patients: a retrospective study spanning 13 years at a tertiary center. Microorganisms. https://doi.org/10.3390/microorganisms11081992
Acknowledgements
The authors acknowledge the Central Pathology Laboratory and MNH staff for their technical support during this study.
Funding
This study did not receive any funding.
Author information
Authors and Affiliations
Contributions
DK and EE developed the study concept and design. DK, DR, AMM, EE and VS performed data entry cleaning and analysis. DK, DR, AMM, EE, VS, and SSM prepared the manuscript. UOK, JM, AJ and MM critically have reviewed the manuscript. All the authors read and approved the final manuscript for submission.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The Senate Research and Publications Committee of the Muhimbili University of Health and Allied Sciences awarded the ethical approval to conduct the study (DA.282/298/01L/Reg. No. 2019-04-13280). We requested permission to access patients’ information from the Executive Director of Muhimbili National Hospital. Our study involved review and analysis of secondary data and did not require direct interaction with participants, hence we did not request the participants’ consent. We ensured the participants’ information was handled with confidentiality and privacy, by using special identification codes instead of the patients’ names.
Consent for publication
Is not applicable.
Competing interests
All the authors declare that they have no commercial or other associations that may pose a conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kamori, D., Renatus, D., Mwandigha, A.M. et al. Prevalence and patterns of multidrug-resistant bacteria isolated from sputum samples of patients with bacterial pneumonia at a tertiary hospital in Tanzania. Bull Natl Res Cent 48, 73 (2024). https://doi.org/10.1186/s42269-024-01228-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s42269-024-01228-9