Abstract
Background
Ctenocephalides felis, the cat flea, is the most common ectoparasite of cats and dogs worldwide. As a cause of flea allergy dermatitis and a vector for two genera of zoonotic pathogens (Bartonella and Rickettsia spp.), the effect of the C. felis microbiome on pathogen transmission and vector survival is of substantial medical importance to both human and veterinary medicine. The aim of this study was to assay the pathogenic and commensal eubacterial microbial communities of individual C. felis from multiple geographic locations and analyze these findings by location, qPCR pathogen prevalence, and flea genetic diversity.
Methods
16S Next Generation Sequencing (NGS) was utilized to sequence the microbiome of fleas collected from free-roaming cats, and the cox1 gene was used for flea phylogenetic analysis. NGS data were analyzed for 168 individual fleas from seven locations within the US and UK. Given inconsistency in the genera historically reported to constitute the C. felis microbiome, we utilized the decontam prevalence method followed by literature review to separate contaminants from true microbiome members.
Results
NGS identified a single dominant and cosmopolitan amplicon sequence variant (ASV) from Rickettsia and Wolbachia while identifying one dominant Bartonella clarridgeiae and one dominant Bartonella henselae/Bartonella koehlerae ASV. Multiple less common ASVs from these genera were detected within restricted geographical ranges. Co-detection of two or more genera (Bartonella, Rickettsia, and/or Wolbachia) or multiple ASVs from a single genus in a single flea was common. Achromobacter, Peptoniphilus, and Rhodococcus were identified as additional candidate members of the C. felis microbiome on the basis of decontam analysis and literature review. Ctenocephalides felis phylogenetic diversity as assessed by the cox1 gene fell within currently characterized clades while identifying seven novel haplotypes. NGS sensitivity and specificity for Bartonella and Rickettsia spp. DNA detection were compared to targeted qPCR.
Conclusions
Our findings confirm the widespread coinfection of fleas with multiple bacterial genera and strains, proposing three additional microbiome members. The presence of minor Bartonella, Rickettsia, and Wolbachia ASVs was found to vary by location and flea haplotype. These findings have important implications for flea-borne pathogen transmission and control.
Graphical Abstract

Similar content being viewed by others
Background
Responsible for the transmission of multiple zoonotic pathogens, Ctenocephalides felis, the cat flea, is the most common ectoparasite of cats and dogs worldwide [1]. Despite this standing, few studies have attempted to characterize the microbiome of C. felis. Previous work has identified Bartonella, Rickettsia, and Wolbachia spp. as the dominant genera within the C. felis microbiome, but a more complete understanding of the flea microbiome including important questions related to coinfection of fleas by multiple pathogens and the relationship between microbial diversity and flea phylogeny remain poorly understood [2, 3].
The genera Bartonella and Rickettsia include multiple pathogenic species vectored by C. felis [4,5,6], with a broad range of zoonotic disease presentations that vary in severity [7,8,9,10]. Flea-associated Rickettsia spp. utilize C. felis as a reservoir host via vertical transmission [11, 12], whereas the C. felis-associated Bartonella spp. utilize the cat as a reservoir host, with free-roaming domestic cats (FRDC) serving as a source of maintenance within the environment [13, 14]. Bartonella spp. display geographic patterns consistent with C. felis [15, 16]. The diversity of C. felis-associated Rickettsia spp. has only recently been appreciated; therefore, little is known regarding their geographic distribution [17,18,19,20]. In addition to pathogenic Bartonella and Rickettsia spp., Wolbachia is widely accepted as a C. felis microbial constituent. Wolbachia spp. are closely related to Rickettsia spp. and are estimated to infect 66% of known insect species [21, 22]. Wolbachia spp. can exert a variety of effects on their hosts including manipulating host reproduction and altering pathogen transmission [23,24,25]. Research describing the C. felis-Wolbachia relationship, including the exchange of metabolites, epidemiology, and diversity of Wolbachia infecting C. felis, is in its infancy [26].
Beyond Bartonella, Rickettsia, and Wolbachia, research identifying additional members of the C. felis microbiome is difficult to interpret. As with other low biomass samples, sequencing-based methods for studying the C. felis microbiome suffers from the presence of contaminating DNA contributed from external sources, such as reagents or the laboratory environment [27, 28]. Contamination in low microbial biomass samples obscures community-wide measurements and complicates the identification of true microbiome members [29]. However, the careful use of control samples can help distinguish contaminants from microbes truly present in samples and thereby more accurately identify C. felis microbial genera [27, 28].
The ability of vector microbiome members to exert effects, whether positive or negative, on both the arthropod host and other microbes is well accepted. Because of the potential to influence pathogen transmission, describing these effects is a primary goal of many vector microbiome investigations [30]. The ability of specific members within the C. felis microbiome to alter the growth and reproduction of other microbes remains poorly described. A previous study of Oropyslla spp., a flea genus that primarily infests rodents, indicated a clear negative relationship between Bartonella and Rickettsia spp. [31]. This negative correlation between Bartonella and Rickettsia spp. has not been clearly demonstrated in C. felis, highlighting the need for further characterization of these genera within individual fleas. In ticks, infection with one Rickettsia spp. is known to interfere with transovarial transmission of a second Rickettsia spp. [32]. However, other reports indicate that coinfection with multiple Rickettsia species in ticks and fleas is possible. These coinfections are thought to result in decreased transmissibility, but the impact on transovarial transmission was not reported [33, 34]. Surveying the prevalence, distribution, and variability in Bartonella, Rickettsia, and Wolbachia spp. coinfections in individual fleas is critical to understand their interactions, with important implications for pathogen transmission.
In addition to the relationship between bacterial species carried by vectors, studies examining mosquitos and plant pathogen vectors document the importance of vector genotype on pathogen transmission dynamics [35, 36]. Ctenocephalides felis is a highly diverse species with four bioclimatically limited clusters originating from Africa [37]. Most published studies either have not determined the infection status of fleas surveyed solely for phylogenetic diversity or investigated only a small number of fleas from which both genotype and pathogen carriage was established [37,38,39].
The present study assessed the eubacterial diversity of individual C. felis collected from FRDC in diverse geographic locations via 16S rRNA NGS. The first aim was to use data on known pathogenic and commensal bacteria present in fleas (Bartonella, Rickettsia, and Wolbachia spp.) to remove contaminants and propose additional C. felis microbiome community members. To understand factors influencing pathogen presence in fleas, the second aim was to determine the presence and prevalence of each of these genera, including intra- and inter-genus coinfection, and compare known microbiome members to geographic location and vector phylogenetics. We expected to find location, flea genotype, and coinfection status variations among the three genera known to infect fleas, while identifying previously undescribed C. felis microbial genera. We also expected to find fleas colonized with multiple Bartonella and Wolbachia spp. ASVs, but only single Rickettsia spp. infections. Finally, the third aim was to compare NGS and qPCR for the detection of Bartonella and Rickettsia spp. in fleas, with the hypothesis that NGS would have lower sensitivity than qPCR for both pathogens.
Methods
Study design
This observational study examined fleas collected from FRDCs from six locations in the US and one in the UK: Davis, CA; San Francisco, CA; Sulphur, LA; Gastonia, NC; Harnett, NC; Washington, NC; London, UK. Fleas were collected from FRDCs when examined by veterinarians for spay or neuter through one local Trap-Neuter-Release (TNR) program in each location. All fleas were collected in March through July of 2019. The total numbers of fleas collected and cats sampled from each geographic location are displayed in Table 1.
Data and specimen collection
All cats presenting for TNR were combed for fleas regardless of demographic group or apparent ectoparasite presence. Fleas were then frozen prior to overnight shipment to the Intracellular Pathogens Research Laboratory at North Carolina State University. If more than six fleas were collected from a single cat, a random number generator in R was utilized to select six fleas for inclusion.
Fleas were visually identified to the species level with the assistance of Dr. James Flowers, Clinical Professor of Parasitology at the North Carolina State University College of Veterinary Medicine [40,41,42]. Individual fleas underwent two PBS and two ethanol washes. Washed fleas were then crushed by a magnetic bead beater until samples were fully homogenized. DNA was then extracted with the Qiagen DNeasy Blood and Tissue Kit (Qiagen, Valencia, CA, USA) following the manufacturer’s protocol for tissue extraction. Resulting DNA concentration (ng/μl) and purity (A260/A280) were determined spectrophotometrically (Nanodrop, Thermo Fisher Scientific, Waltham, MA, USA).
Library preparation for 16S rRNA amplicon sequencing
Microbial community 16S rRNA gene amplicon sequencing was performed on DNA samples from individual fleas. DNA was submitted including 10 μl of DNA at 1.5 ng/μl per flea. Library preparation and sequencing were performed in July 2020 by the North Carolina State University Genomic Sciences Laboratory (Raleigh, NC) via the Illumina MiSeq system with primers targeting the 16S V3-V4 region. Sequencing relied on the forward primer 5ʹ-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG-3ʹ and the reverse primer 5ʹ-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC-3ʹ, selected from Klindworth et al. [43]. Ten negative extraction controls were included in sample processing and library preparation. These extraction controls were generated by performing manual DNA extraction of PBS at the same time as flea samples.
Sequence preparation and filtering
Following library preparation, sequence data were prepared and analyzed in R (version 4.0.4) first utilizing DADA2 [44] version 1.20.0 to inspect quality profiles, filter, and trim sequences and then infer amplicon sequence variants (ASVs) and remove chimeras. Taxonomy was assigned using the DADA2 assignTaxonomy() function and the non-redundant Silva taxonomic training database version 138.1 (“silva_nr99_v138.1_train_set.fa”, https://www.arb-silva.de/). Species identity was assigned by the Silva species database version 138.1 (“silva_species_assignment_v138.1.fa.gz”, https://www.arb-silva.de/) based on exact matching.
Filtering was first performed to remove ASVs with fewer than three reads. Decontam [45] was then employed to identify potential members of the microbiome based on the scores of the three genera which are known C. felis microbiome constituents: Bartonella, Rickettsia, and Wolbachia. The decontam prevalence method is built on the premise that contaminant ASVs will be present more often in negative controls than they are in true samples. Decontam generates a score that discriminates between likely contaminants (scores closer to zero) and likely non-contaminants (scores closer to one). All genera with one or more ASV(s) above the score assigned to known microbiome members were considered potential microbiome members and reported. Literature review was utilized to select genera from this list to be considered candidate true microbiome members. To perform the literature review, all potential microbiome genera were searched on PubMed in combination with “microbiome,” “flea,” “tick,” and “mosquito” to identify relevant reports. The identified literature was examined on the location of detection and rigor of filtering to select only the genera which possess considerable evidence to be considered candidate microbiome members. Of those genera selected as candidate microbiome members, ASVs detected in more than three of the extraction control samples were removed.
ASV analysis
Amplicon Sequence Variants (ASVs) are unique sequences generated by NGS after the correction of errors in amplicon sequencing data by the DADA2 method. Each ASV represents a group of organisms that share identical sequences over the amplified region of the 16S rRNA gene. Short-read 16S ASVs, such as those analyzed here, correspond roughly to species-breadth units, but can differentiate sub-species clades within some bacterial species. To compare the phylogenetic relationship between ASVs, the R package ape was utilized to build a neighbor-joining tree to compare the relatedness of ASVs [46, 47]. The Interactive Tree of Life (iTOL) software allowed tree visualization and annotation [48].
Pathogen detection with qPCR
Quantitative polymerase chain reaction (qPCR) was performed for the amplification of Anaplasma, Bartonella, Ehrlichia, Mycoplasma, and Rickettsia spp. DNA. Table 2 lists the targeted genes used to amplify DNA of each pathogen. Each reaction included three controls: molecular-grade water non-template control, negative control from a known negative cat, and positive control plasmid. All positive samples were confirmed by sequencing (GENEWIZ Inc., Raleigh, NC). Sequence cleanup and viewing were performed in SnapGene (Insightful Science, San Diego, CA) and alignment with NCBI BLAST (http://blast.ncbi.nlm.nih.gov/Blast) [49]. Only fleas from which a sequence was acquired were considered positive by qPCR.
Rickettsia spp. detection was improved utilizing species-specific primers for R. felis and Rickettsia asembonensis (Table 2). Each qPCR reaction was performed with a 25 μl reaction volume consisting of 12.5 μl SsoAdvanced Unviersal SYBR Green Supermix 2X (Bio-Rad), 7.1 μl of molecular grade water, 5 μl DNA, and 0.2 μl of each primer. The reaction began with denaturation at 98 ℃ for 3 min, followed by 45 cycles of denaturation at 98 ℃ for 15 s, annealing at 66 ℃ for 15 s, and extension at 72 ℃ for 20 s, followed by a melt curve.
NGS sensitivity and specificity for the detection of Bartonella and Rickettsia spp. DNA were calculated utilizing qPCR as the reference standard.
Flea phylogenetic analysis with cox1 cPCR
Flea phylogenetic group was assigned by sequencing of the cytochrome c oxidase subunit I (cox1) gene amplified by conventional PCR (cPCR) utilizing the Cff-F and Cff-R primers designed by Lawrence et al. [39]. The cPCR reaction was performed with a 25 μl reaction volume consisting of 12.5 μl 2 × MyTaq HS Red Mix (Meridian Bioscience, Cincinnati, OH), 11.1 μl molecular grade water, 1 μl DNA, and 0.2 μl each primer. The reaction began with denaturation at 95 ℃ for 2 min, followed by 40 cycles of denaturation at 95 ℃ for 30 s, annealing at 55℃ for 30 s, and extension at 72 ℃ for 30 s, ending with a final extension step at 72 ℃ for 10 min. Fleas from which a clean sequence was not acquired were retested at 0.01, 0.1, and 4 µl DNA per reaction to achieve successful amplification. DNA extracts from which C. felis DNA could not be amplified via cox1 cPCR were excluded from analysis.
Haplotype networks were constructed based on a randomized minimum spanning tree, both of which were generated using the pegas package [50]. Specific visualizations of these haplotype networks were then developed to compare flea location and the occurrence of individual ASVs.
Analysis of flea phylogenetic sequences required the creation of a neighbor-joining tree. Detected haplotypes were compared to those reported by Lawrence et al. [37]. α-Diversity was analyzed via calculation of net relatedness index (NRI) [51]. This calculation is based on nodal distance with more negative communities being more phylogenetically diverse. The β-diversity measure PhyloSor was calculated to quantify the phylogenetic similarity between communities with more positive values indicating more genetically similar communities [52]. A reference Ctenocephalides canis cox1 sequence (accession number MW136242.1) served as the outgroup for β-diversity calculations.
All analyses were performed in the R statistical computing environment utilizing the packages ape [46], Biostrings [53], dada2 [44], dplyr [54], decontam [45], filesstrings [55], GEOquery [56], ggforce [57], ggplot2 [58], ggsci [59], here [60], janitor [61], PhyloMeasures [62], phyloseq [63], picante [64], magrittr [65], RVAideMemoire [66], and vegan [67].
Results
A total of 182 fleas were collected; 168 of these fleas passed cox1 cPCR control. These fleas were collected from 75 cats with the number of fleas and cats from each geographic location reported in Table 1.
A total of 12,637 ASVs assigned to 457 genera were inferred by DADA2 across the flea and negative control samples; 1569 of these ASVs were removed as they were represented by fewer than three reads across all samples. Peptoniphilus ASVs were found in all 168 fleas, followed by Rhodococcus (167 fleas), Achromobacter (149 fleas), Rickettsia (128 fleas), Wolbachia (111 fleas), and Bartonella (63 fleas). This order was similar to that found when observing the total number of ASVs and proportion of reads for each genus (Fig. 1a). Wolbachia, Rickettsia, and Bartonella, but not Achromobacter, Peptoniphilus, and Rhodococcus, derived a majority of their reads from a single most abundant ASV (Fig. 1a).
a Total number of ASVs (orange) assigned to each of the six genera indicated as most likely to be true microbiome members, as well as the proportion of reads assigned to each genera (black) and the proportion of reads assigned to the most abundant ASV of that genera (gray). b Proportion of fleas from each location with one or more read from the indicated genus. c Proportion of reads from each location assigned to each genus. The total number of fleas from each location was included in parentheses after the location on the x-axis
The decontam prevalence method indicated that the most abundant Wolbachia, Rickettsia, and Bartonella ASVs were assigned a score of 0.97, 0.92, and 0.89, respectively. Therefore, we set a threshold of ≥ 0.89 to consider a genus as a potential microbiome member. Thirteen of 457 (3%) genera exceeded this threshold (Additional file 1: Fig. S1): Achromobacter [68, 69], Anaerococcus [70,71,72,73], Bacteroides [74, 75], Blastomonas [76, 77], Clostridium sensu stricto 5 [78], Lachnoclostridium [79], Methanothermobacter [80], Peptoniphilus [81, 82], Peptostreptococcus [82, 83], and Rhodococcus [84, 85]. A literature search evaluating previous reports of these genera and the rigor of the contaminant filtering in these publications supported the idea that three of these genera (Achromobacter, Peptoniphilus, and Rhodococcus) should be considered candidate C. felis microbiome members.
Anaplasma, Ehrlichia, and hemotropic Mycoplasma DNA was not amplified from any of the fleas by either qPCR or NGS.
When examining the number of fleas with at least one read from Achromobacter, Bartonella, Rickettsia, or Wolbachia, there were geographic location-based differences in prevalence not observed in Peptoniphilus or Rhodococcus (Fig. 1b). There were also differences in the proportion of reads from each location assigned to the identified genera with Washington, NC, and Gastonia, NC, consistently reporting the highest proportion of Bartonella, Rickettsia, or Wolbachia reads (Fig. 1c).
NGS yielded 16 distinct Bartonella spp. ASVs (Fig. 2a). A majority of the Bartonella ASVs detected aligned closely with either B. clarridgeiae (Bc-like ASVs) or B. henselae/B. koehlerae (Bh/Bk-like ASVs) (Fig. 3). A single dominant ASV was found in both groups: ASV9 and ASV55. ASV9 was detected in all locations except for San Francisco, CA, and ASV55 was detected in all locations except for San Francisco, CA, Sulphur, LA, and London, UK. One of 21 fleas from Sulphur, LA, contained ASV4296, a Bh/Bk-like ASV. All minor Bc-like or Bh/Bk-like ASVs were found in a single location or two North Carolina locations. Two outlier Bartonella ASVs (ASV9207 and ASV10434) were detected in a single flea each. Neither of these ASVs aligned with known Bartonella spp.
NGS yielded 42 distinct Rickettsia ASVs. Read abundance was dominated by a single ASV: ASV4 (99.75% R. asembonensis, accession ID JN315973.1) (Fig. 2b). Only one of the 42 ASVs aligned perfectly with a known Rickettsia: ASV38 and ‘Candidatus Rickettsia senegalensis’ (Fig. 4). No ASV aligned perfectly with R. felis. Rickettsia ASVs also displayed geographic location-based differences in detection: ASV4 was detected in all locations, ASV38 and ASV3856 were detected in Washington, NC, and Davis, CA, and all other ASVs were found in a single location or two NC locations (Fig. 4).
NGS yielded 36 distinct Wolbachia spp. ASVs (Fig. 2c). The most abundant Wolbachia ASV (ASV6) aligned perfectly with wCfeT (accession ID NZ_CP051156.1). ASV383, the fourth most abundant Wolbachia ASV, aligned perfectly with wCfeJ (accession ID NZ_CP051157.1) [26]. The full diversity of Wolbachia ASVs compared to wCfeT and wCfeJ is shown in Fig. 5. Wolbachia ASVs displayed geographic location-based differences in detection: ASV6 was detected in all locations, ASV998 was detected in Washington, NC, and Sulphur, LA, and all other ASVs were found in a single location or two NC locations.
Flea phylogenetics
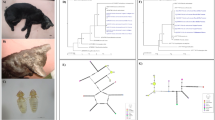
The cox1 sequencing identified 17 flea haplotypes with these haplotypes arising from mutations in 32 base pair locations (Fig. 6b). The phylogenetic tree and haplotype network describing the relationship between these haplotypes are in Fig. 6a and c, respectively. A haplotype network displaying the proportion of fleas from each location is shown in Fig. 6d. The α-diversity metric NRI was used to compare the within-group diversity when fleas were grouped by location. NRI indicated that Washington, NC, was the most phylogenetically diverse location among the seven study sites, while London, UK, was the least phylogenetically diverse. To quantify phylogenetic similarity between local flea communities, the β-diversity measure PhyloSor was calculated [52]. This calculation indicated two location-based groups of phylogenetic similarity: one comprised of Sulphur, LA, and the three North Carolina locations and the other comprised of London, UK, and the two California locations.
a Phylogenetic tree displaying the relationship between identified haplotypes and location at which they were detected. b Specific base pair sites identified as sources of variability between haplotypes. c Haplotype network displaying the relationship between haplotypes and the d location of origin of the fleas from different haplotypes
When comparing these fleas to the haplotypes published by Lawrence et al. [37], our flea haplotypes all fell within previously reported clades: Clade 1 (n = 9), Clade 2 (n = 1), Clade 4 (n = 5), and Clade 6 (n = 2), revealing new haplotypes in Clades 1 (n = 5), 2 (n = 1), 4 (n = 3), and 6 (n = 1) (Additional file 2: Fig. S2).
Haplotype networks comparing specific flea haplotype to Bartonella spp. reported by NGS (dominant ASV) and qPCR sequencing are illustrated in Fig. 7a and b, respectively. Only one minor Bc-like ASV was found in multiple fleas (ASV257). These fleas were from two NC locations and belonged to the same haplotype. A single Bh/Bk-like ASV was detected in multiple fleas: ASV55. ASV55 was detected in only two haplotypes but four locations. All other fleas containing Bh/Bk-like ASVs belonged to other haplotypes, suggesting that B. henselae strains may be associated with specific C. felis haplotype.
Wolbachia ASVs also displayed haplotype-specific occurrence patterns. The dominant Wolbachia ASV (ASV6) was detected in all geographic locations and in fleas from three clades (Fig. 7e). Five minor Wolbachia ASVs were detected in more than one flea. All of these ASVs were detected in only a single flea clade, with a majority (3/5) found in only one flea haplotype despite four of the Wolbachia ASVs occurring in more than one location. ASV998, an ASV detected in Sulphur, LA, and Washington, NC, was detected in two fleas, one from each location. Both of these fleas were assigned to haplotype 3 in clade 6, a haplotype and clade otherwise populated exclusively by Sulphur, LA, fleas. The haplotype network comparing flea haplotype to the dominant Wolbachia ASV is illustrated in Fig. 7e.
Rickettsia strain prevalence by flea genotype was less clear than that of either Bartonella or Wolbachia. A dominant Rickettsia ASV (ASV4), detected in all locations except San Francisco, CA, was present in fleas from all four clades. Six minor Rickettsia ASVs were detected in multiple fleas, all of which were from multiple locations and in fleas from multiple clades. The haplotype network comparing flea haplotype to the dominant Rickettsia ASV and qPCR Rickettsia spp. is illustrated in Fig. 7c and d, respectively.
Coinfection
The majority of fleas in this study were coinfected with more than one of the genera Bartonella, Rickettsia, and Wolbachia (Fig. 8a). The NGS data also provided support for intra-genus coinfection. Among infected fleas, 14% (9/63) contained more than one Bartonella ASV (Fig. 8b), 29% (37/128) had more than one Rickettsia ASV (Fig. 8c), and 23% (26/111) had more than one Wolbachia ASV (Fig. 8d). It is unlikely that intra-genomic variation between different copies of the 16S rRNA gene can explain these results as Rickettsia and Wolbachia only have one gene copy and Bartonella has two copies, relatively few compared to other bacterial genomes.
qPCR comparison with NGS
Combining results from both NGS and qPCR diagnostic methods, Bartonella spp. sequences were found in 47% (79/168) of fleas. A total of 63 fleas were identified as infected with Bartonella spp. by NGS compared to 41 fleas by qPCR. When qPCR is considered the reference standard, NGS had an overall sensitivity of 61% and specificity of 70% for detection of Bartonella spp. in fleas (Table 3). The proportion of qPCR-positive fleas for which NGS also returned a positive detection did not vary by the qPCR species, NGS ASV, or qPCR primer set.
Despite NGS detecting Rickettsia ASV(s) in 76% (128/168) fleas, we were only able to sequence clean Rickettsia spp. amplicons from 20% (34/168) of fleas by qPCR. When qPCR is considered the reference standard, NGS had an overall sensitivity of 94% and specificity of 28% for detection of Rickettsia spp. in fleas (Table 4). The fraction of qPCR-positive fleas for which NGS also returned a positive detection did not vary by the qPCR species, NGS ASV, or qPCR primer set.
Discussion
This study used 16S NGS to analyze the eubacterial microbiome of C. felis fleas removed from free-roaming cats across diverse geographic locations. As expected, we documented widespread infection with known C. felis microbiome members (Bartonella, Rickettsia, and Wolbachia). Comparisons between negative controls and flea samples combined with literature search facilitated the selection of Achromobacter, Peptoniphilus, and Rhodococcus as candidate C. felis microbiome members. Multiple Bartonella, Rickettsia, and/or Wolbachia ASVs were amplified from individual fleas, suggesting that coinfection both between these genera and among species/strains within each of these genera is a common occurrence. Almost all minor ASVs were detected in fleas from only a single location or in multiple locations from the same state suggesting location-based strain variation. Ctenocephalides felis phylogenetic clade and haplotype were associated with geographic location. Combined C. felis phylogenetic and NGS data more strongly supported a relationship between flea phylogeny and infecting Bartonella and Wolbachia ASV than flea phylogeny and Rickettsia ASV.
NGS paired with robust data analysis is a promising approach for identifying true microbiome members in low-biomass or contaminant prone samples. It is critical that stringent controls and comprehensive techniques, such as those used in this study, are utilized to identify vector microbiome members. We did not detect Elizabethkingia or Snodgrassella spp. DNA in this study, as previously reported in a study by Vasconcelos et al. [3]. None of the genera reported by Cohen et al. in Synosternus cleopatrae fleas, other than those noted in our analyses, were deemed microbiome members due to either their absence in flea samples or their presence in negative controls [86]. Our inability to either detect or recognize previously reported genera as C. felis microbiome members may be due to these genera not being present in the fleas in our study, or because those genera are unrecognized contamination.
Our finding that most minor ASVs from the three known C. felis genera (Bartonella, Rickettsia, and Wolbachia) were only detected in a single or geographically close location(s) may imply that diversity within these bacterial genera is associated with geographical location. The association of bacterial diversity with C. felis genotype remains to be elucidated. The full implications of these findings are unknown; however, specific strains of Bartonella spp. are known to exhibit highly varying pathogenicity [87] and display differences in factors relating to the risk and severity of infection. Studies that further characterize diversity within C. felis-associated bacteria are necessary, as the relatively short (402 bp) sequence of the 16S gene amplified by NGS is not designed to resolve strain-based variability in virulence factors or pathogenic potential.
Regarding flea phylogeny, our data reemphasize how limited our knowledge is of C. felis diversity in the US as a sample of 168 fleas identified seven new haplotypes within the previously reported clades [37]. Comparing our flea haplotypes to those reported by Lawrence et al., a majority of the California fleas were assigned to Clade 1, a temperate clade, while Sulphur, LA, fleas were assigned primarily to Clade 6, a tropical II clade and a single flea in Clade 2, a temperate clade. North Carolina fleas were primarily assigned to Clade 4, a tropical I clade, with a limited number of fleas in Clade 1 or Clade 6.
Although not the first study to report both flea genotype and pathogen carriage status [2, 20], this study reports a large sample of C. felis genotype and Bartonella, Rickettsia, and Wolbachia infection status. Our data provide evidence that C. felis genotypic haplotype is most likely to be related to the infecting Bartonella and Wolbachia strain but less likely to be related to Rickettsia strain. The dominant Bh/Bk-like ASV was detected in only two haplotypes despite being present in four locations. Furthermore, no two Bh/Bk-like ASVs infected the same flea haplotype. The single minor Bc-like ASV detected in multiple fleas was detected in multiple geographic locations, but only one flea haplotype. For Wolbachia spp., we determined that the single Washington, NC, flea assigned to clade 6 was infected with Wolbachia ASV998, a clade and ASV only found to be inhabited by or infecting Sulphur, LA, fleas of clade 6. A majority (3/5) of all minor Wolbachia ASVs were detected in only a single clade. These findings indicate that C. felis-associated bacteria may display previously unidentified vector genotype by pathogen genotype relationships or divergent evolution with the fleas they inhabit. Further study is necessary to establish the relationship between geographic location, flea haplotype, microbiome diversity, pathogen prevalence, and strain delineation using markers with higher resolution than the short-read 16S target utilized.
Our analyses also allowed us to compare detection of Bartonella spp. in NGS versus qPCR. Given the lack of a true gold standard, we utilized qPCR, which is not a 100% sensitive Bartonella diagnostic assay [88], a factor that likely contributed to NGS having a specificity of only 68% (89/127). It is likely that the additional 38 fleas positive by NGS were truly infected at a level below qPCR detection. NGS did not detect Bartonella ASVs in 39% (16/41) of the fleas positive by qPCR and sequencing. Our NGS findings indicated that B. clarridgeiae is the most common Bartonella spp. in C. felis, a conclusion that was supported by qPCR findings and previous literature [89, 90]. Comparing the agreement between NGS and qPCR, which was only 68% (114/168), it is likely that the sensitivity of both methods is suboptimal.
The dominant Rickettsia ASV (ASV4) detected in our analysis is most closely related to R. asembonensis, not R. felis, suggesting that C. felis in the US may more frequently harbor Rickettsia spp. other than R. felis. Beyond the dominant ASV, we found a very diverse collection of Rickettsia ASVs that have not been characterized. These findings reinforce the importance of exploring the epidemiology of flea-associated Rickettsia spp. in future surveys [7, 91]. In the absence of a gold standard assay, we again compared detection in qPCR and NGS and found that NGS reported a sensitivity of 94% (32/34) for the detection of Rickettsia spp. Unfortunately, the 28% (38/134) specificity indicates a failure of the NGS or qPCR assay. Due to the large diversity of Rickettsia ASVs detected by NGS, we suspect that the qPCR assays lacked specificity for the larger diversity of Rickettsia present in these fleas. Based on NGS results, Rickettsia co-infection data suggest that contrary to the patterns observed in ticks [32], colonization with one Rickettsia does not inhibit colonization with a second Rickettsia spp. in C. felis.
We detected Wolbachia ASVs in 66% (111/168) of fleas, substantially more than the 21% previously reported by qPCR [22]. The high proportion of fleas with coinfecting Wolbachia ASVs is on par with previous evidence of Wolbachia co-infection in laboratory flea colonies. Dricoll et al. proposed a selection for wCfeT over wCfeJ in nature, which agrees with our finding that ASV6 (100% wCfeT) was the dominant Wolbachia ASV and ASV383 (100% wCfeJ) was observed in only a single flea [26].
An important limitation of the present study was the low biomass of each C. felis sample, which likely allowed contaminant DNA from extraction kits and cross-contamination to dominate data [28]. The impact of this was mitigated by use of decontam [45] and literature search of proposed microbiome members; however, it restricted our ability to perform whole microbial community analysis. Additional limitations included our sampling of C. felis from free-roaming cats, which may not accurately represent the fleas from client-owned cats. Sex was not determined for the sampled C. felis, preventing comparison between male and female bacterial carriage [86]. Rickettsia felis prevalence may be underestimated in this study as the cat is not considered a reservoir host [12]. Instead, current literature implicates dogs, rodents, opossums, or fleas themselves as reservoir hosts for R. felis [92]. Concerning diagnostic techniques, qPCR, while specific as results were confirmed by DNA sequencing, suffers from a lack of sensitivity and therefore likely underestimated pathogen prevalence [88].
Conclusions
The present study reports diverse Bartonella, Rickettsia, and Wolbachia spp. colonizing the C. felis microbiome, which varies by C. felis geographic origin and haplotype. While the read abundance of each of these genera was clearly dominated by a single ASV, multiple other ASVs were also identified, likely representing previously uncharacterized strains or species with unknown pathogenicity. Fleas were found to be phylogenetically diverse with haplotype diversity varying by location of origin and multiple new flea haplotypes detected. This work reinforces the importance of future research investigating the diversity of C. felis and their pathogenic and non-pathogenic microbiome members to guide diagnosis, treatment, and flea-borne pathogen control recommendations.
Availability of data and materials
The datasets supporting the conclusions of this article are available in a Dryad repository (https://doi.org/10.5061/dryad.3ffbg79km).
Abbreviations
- ASV:
-
Amplicon sequence variant
- FRDC:
-
Free-roaming domestic cat
- iTOL:
-
Interactive tree of life
- NGS:
-
Next generation sequencing
- NRI:
-
Net relatedness index
- qPCR:
-
Quantitative polymerase chain reaction
- TNR:
-
Trap-neuter-release
References
Rust MK. The biology and ecology of cat fleas and advancements in their pest management: A review. Insects. 2017;8:51.
Lawrence AL, Hii SF, Chong R, Webb CE, Traub R, Brown G, et al. Evaluation of the bacterial microbiome of two flea species using different DNA-isolation techniques provides insights into flea host ecology. FEMS Microbiol Ecol. 2015;91:11.
Vasconcelos EJR, Billeter SA, Jett LA, Meinersmann RJ, Barr MC, Diniz PPVP, et al. Assessing cat flea microbiomes in northern and southern California by 16S rRNA next-generation sequencing. Vector Borne Zoonotic Dis. 2018;18:491–9.
Chomel BB, Kasten RW, Floyd-Hawkins K, Chi B, Yamamoto K, Roberts-Wilson J, et al. Experimental transmission of Bartonella henselae by the cat flea. J Clin Microbiol. 1996;34:1952–6.
Kordick DL, Hilyard EJ, Hadfield TEDL, Wilson KH, Steigerwalt AG, Brenner DONJ, et al. Bartonella clarridgeiae, a newly recognized zoonotic pathogen causing inoculation papules, fever, and lymphadenopathy (cat scratch disease). J Clin Microbiol. 1997;35:1813–8.
Avidor B, Graidy M, Efrat G, Leibowitz C, Shapira G, Schattner A, et al. Bartonella koehlerae, a new cat-associated agent of culture-negative human endocarditis. J Clin Microbiol. 2004;42:3462–8.
Brown LD, Macaluso KR. Rickettsia felis, an emerging flea-borne rickettsiosis. Curr Trop Med Rep. 2016;3:27–39.
Chomel BB, Kasten RW, Williams C, Wey AC, Henn JB, Maggi RG, et al. Bartonella endocarditis: a pathology shared by animal reservoirs and patients. Ann NY Acad Sci. 2009;1166:120–6.
Breitschwerdt EB. Bartonellosis, one health and all creatures great and small. Vet Dermatol. 2017;28:96–106.
Breitschwerdt E, Sontakke S, Hopkins S. Neurological manifestations of Bartonellosis in immunocompetent patients: a composite of reports from 2005–2012. J Neuroparasitology. 2012;3:15.
Hirunkanokpun S, Thepparit C, Foil LD, Macaluso KR. Horizontal transmission of Rickettsia felis between cat fleas Ctenocephalides felis. Mol Ecol. 2011;20:4577–86.
Wedincamp J, Foil LD. Vertical transmission of Rickettsia felis in the cat flea (Ctenocephalides felis Bouche). J Vector Ecol. 2002;27:96–101.
Manvell C, Ferris K, Maggi R, Breitschwerdt EB, Lashnits E. Prevalence of vector-borne pathogens in reproductive and non-reproductive tissue samples from free-roaming domestic cats in the South Atlantic USA. Pathogens. 2021;10:17.
Koehler JE, Glaser CA, Jordan W. Rochalimaea henselae infection a new zoonosis with the domestic cat as reservoir. J Am Med Assoc. 1994;271:532–5.
Lashnits EW, Dawson DE, Breitschwerdt E, Lanzas C. Ecological and socioeconomic factors associated with Bartonella henselae exposure in dogs tested for vector-borne diseases in North Carolina. Vector Borne Zoonotic Dis. 2019;19:582–95.
Jameson P, Greene C, Regnery R, Dryden M, Marks A, Brown J, et al. Prevalence of Bartonella henselae antibodies in pet cats throughout regions of North America. J Infect Dis. 1995;172:1145–9.
Blanton LS, Quade BR, Bouyer DH. Differentiation of Rickettsia felis and Rickettsia felis-like organisms via restriction fragment length polymorphism analysis. Vector Borne Zoonotic Dis. 2019;19:637–9.
Loyola S, Flores-Mendoza C, Torre A, Kocher C, Melendrez M, Luce-Fedrow A, et al. Rickettsia asembonensis characterization by multilocus sequence typing of complete genes. Peru Emerg Infect Dis. 2014;24:931–3.
Eremeeva ME, Capps D, McBride CL, Williams-Newkirk AJ, Dasch GA, Salzer JS, et al. Detection of Rickettsia asembonensis in fleas (Siphonaptera: Pulicidae, Ceratophyllidae) collected in five counties in Georgia, United States. J Med Entomol. 2020;57:1246–53.
Ferreira FC, Fonseca DM, Hamilton G, Price D. Metagenomic analysis of human - biting cat fleas in urban northeastern United States of America reveals an emerging zoonotic pathogen. Sci Rep. 2020;10(1):1-8. .
Hilgenboecker K, Hammerstein P, Schlattmann P, Telschow A, Werren JH. How many species are infected with Wolbachia?—A statistical analysis of current data. FEMS Microbiol Lett. 2008;281:215–20.
Gorham CH, Fang QQ, Durden LA. Wolbachia endosymbionts in fleas (Siphonaptera). J Parasitol. 2003;89:283–9.
Werren JH, Baldo L, Clark ME. Wolbachia: master manipulators of invertebrate biology. Nat Rev Microbiol. 2008;6:741–51.
Cirimotich CM, Dong Y, Clayton AM, Sandiford SL, Souza-Neto JA, Mulenga M, et al. Natural microbe-mediated refractoriness to Plasmodium infection in Anopheles gambiae. Science. 2011;332:855–8.
O’Neill SL, Ryan PA, Turley AP, Wilson G, Retzki K, Iturbe-Ormaetxe I, et al. Scaled deployment of Wolbachia to protect the community from Aedes transmitted arboviruses. Gates Open Res. 2018;2:36.
Driscoll TP, Verhoeve VI, Brockway C, Shrewsberry DL, Plumer M, Sevdalis SE, et al. Evolution of Wolbachia mutualism and reproductive parasitism: insight from two novel strains that co-infect cat fleas. PeerJ. 2020;8:39.
Salter SJ, Cox MJ, Turek EM, Calus ST, Cookson WO, Moffatt MF, et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 2014;12:12.
Eisenhofer R, Minich JJ, Marotz C, Cooper A, Knight R, Weyrich LS. Contamination in low microbial biomass microbiome studies: issues and recommendations. Trends Microbiol. 2019;27:105–17.
Lejal E, Estrada-peña A, Marsot M, Cosson J, Rué O, Mariadassou M, et al. Taxon appearance from extraction and amplification steps demonstrates the value of multiple controls in tick microbiota analysis. Front Microbiol. 2020;11:10.
Bonnet SI, Binetruy F, Hernández-Jarguín AM, Duron O. The tick microbiome: why non-pathogenic microorganisms matter in tick biology and pathogen transmission. Front Cell Infect Microbiol. 2017;7:14.
Jones RT, Bernhardt SA, Martin AP, Gage KL. Interactions among symbionts of Oropsylla spp. (Siphonoptera: Ceratophyllidae). J Med Entomol. 2012;49:492–6.
Macaluso KR, Sonenshine DE, Ceraul SM, Azad AF. Rickettsial infection in Dermacentor variabilis (Acari: Ixodidae) inhibits transovarial transmission of a second Rickettsia. J Med Entomol. 2002;39:809–13.
Carmichael JR, Fuerst PA. A rickettsial mixed infection in a Dermacentor variabilis tick from Ohio. Ann NY Acad Sci. 2006;1078:334–7.
Noden BH, Radulovic S, Higgins JA, Azad AF. Molecular Identification of Rickettsia typhi and R. felis in co-infected Ctenocephalides felis (Siphonaptera: Pulicidae). J Med Entomol. 1998;35:410–4.
Gloria-Soria A, Armstrong PM, Turner PE, Turner PE. Infection rate of Aedes aegypti mosquitoes with dengue virus depends on the interaction between temperature and mosquito genotype. Proc R Soc. 2017;284:9.
Lambrechts L, Halbert J, Durand P, Gouagna LC, Koella JC. Host genotype by parasite genotype interactions underlying the resistance of anopheline mosquitoes to Plasmodium falciparum. Malar J. 2005;4:8.
Lawrence AL, Webb CE, Clark NJ, Halajian A, Mihalca AD, Miret J, et al. Out-of-Africa, human-mediated dispersal of the common cat flea, Ctenocephalides felis: the hitchhiker’s guide to world domination. Int J Parasitol. 2019;49:321–36.
Chandra S, Forsyth M, Lawrence AL, Emery D, Slapeta J. Cat fleas (Ctenocephalides felis) from cats and dogs in New Zealand : molecular characterization, presence of Rickettsia felis and Bartonella clarridgeiae and comparison with Australia. Vet Parasitol. 2017;234:25–30.
Lawrence AL, Brown GK, Peters B, Spielman DS, Morin-Adeline V, Šlapeta J. High phylogenetic diversity of the cat flea (Ctenocephalides felis) at two mitochondrial DNA markers. Med Vet Entomol. 2014;28:330–6.
National Communicable Disease Center. Pictorial keys: arthropods, reptiles, birds, and mammals of public health significance. US: Public Health Service publication; 1969.
Fox I. Fleas of Eastern United States. Ames: The Iowa State College Press; 1940.
Linardi PM, Santos JLC. Ctenocephalides felis felis vs. Ctenocephalides canis (Siphonaptera:Pulicidae): some issues in correctly identify these species. Rev Bras Parasitol Vet. 2012;21:345–54.
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013;41:11.
Callahan BJ, Mcmurdie PJ, Rosen MJ, Han AW, Johnson AJ, Holmes SP. DADA2: High resolution sample interference from Illumina amplicon data. Nat Methods. 2016;13:581–3.
Davis NM, Proctor DM, Holmes SP, Relman DA, Callahan BJ. Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome. 2018;6:14.
Paradis E, Schliep K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics. 2019;35:526–8.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.
Letunic I, Bork P. Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021;49:293–6.
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, et al. GenBank. Nucleic Acids Res. 2013;41:36–42.
Paradis E. pegas: an R package for population genetics with an integrated-modular approach. Bioinformatics. 2010;26:419–20.
Webb CO. Exploring the phylogenetic structure of ecological communities: an example for rain forest trees. Am Nat. 2000;156:145–55.
Bryant JA, Lamanna C, Lè Ne Morlon H, Kerkhoff AJ, Enquist BJ, Green JL. Microbes on mountainsides: contrasting elevational patterns of bacterial and plant diversity. PNAS. 2008;105:11505–11.
Pagès H, Aboyoun P, Gentleman R, DebRoy S. Biostrings: efficient manipulation of biological strings. 2021.
Wickham H, François R, Henry L, Müller K. dplyr: a grammar of data manipulation. 2021.
Nolan R, Padilla-Parra S. filesstrings: an R package for file and string manipulation. J Open Source Softw. 2017. https://doi.org/10.21105/joss.00260.
Davis S, Meltzer P. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics. 2007;14:1846–7.
Pedersen TL. ggforce: accelerating ‘ggplot2’. 2021.
Wickham H. ggplot2: elegant graphics for data analysis. New York: Springer-Verlag; 2016.
Xiao N. ggsci: scientific journal and sci-fi themed color palettes for ‘ggplot2’. 2018.
Müller K. here: a simpler way to find your files. 2020.
Firke S. janitor: simple tools for examining and cleaning dirty data. 2021.
Tsirogiannis C, Sandel B. PhyloMeasures: fast and exact algorithms for computing phylogenetic biodiversity measures. 2017.
Mcmurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE. 2013;8:e61217.
Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD, et al. Picante: R tools for integrating phylogenies and ecology. Bioinformatics. 2010;26:1463–4.
Bache SM, Wickham H. magrittr: a forward-pipe operator for R. 2020.
Hervé M. RVAideMemoire: testing and plotting procedures for biostatistics. 2021.
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D. et al. Vegan: community ecology package. 2020.
Okla H, Sosnowska M, Jasik KP, Slodki J, Wojtyczka RD. Nonspecific bacterial flora isolated from the body surface and inside Ixodes ricinus ticks. Pol J Microbiol. 2012;61:205–9.
Menetrey Q, Sorlin P, Jumas-Bilak E, Chiron R, Dupont C, Marchandin H. Achromobacter xylosoxidans and stenotrophomonas maltophilia: Emerging pathogens well-armed for life in the cystic fibrosis patients’ lung. Genes. 2021;12:21.
Tall ML, Pham TPT, Bellali S, Ngom II, Delerce J, Lo CI, et al. Anaerococcus marasmi sp. nov., a new bacterium isolated from human gut microbiota. New Microbes New Infect. 2020;35:5.
Veloo ACM, de Vries ED, Jean-Pierre H, van Winkelhoff AJ. Anaerococcus nagyae sp. nov., isolated from human clinical specimens. Anaerobe. 2016;38:111–5.
Muturi EJ, Dunlap C, Smartt CT, Shin D. Resistance to permethrin alters the gut microbiota of Aedes aegypti. Sci Rep Nat Res. 2021;11:8.
Duan DY, Liu GH, Cheng TY. Microbiome analysis of the saliva and midgut from partially or fully engorged female adult Dermacentor silvarum ticks in China. Exp Appl Acarol. 2020;80:543–58.
Keskin A, Bursali A, Snow DE, Dowd SE, Tekin S. Assessment of bacterial diversity in Hyalomma aegyptium, H. marginatum and H. excavatum ticks through tag-encoded pyrosequencing. Exp Appl Acarol. 2017;73:461–75.
Wexler HM. Bacteroides: the good, the bad, and the nitty-gritty. Clin Microbiol Rev. 2007;20:593–621.
Sala-Comorera L, Caudet-Segarra L, Galofré B, Lucena F, Blanch AR, García-Aljaro C. Unravelling the composition of tap and mineral water microbiota: divergences between next-generation sequencing techniques and culture-based methods. Int J Food Microbiol. 2020;334:10.
Cani PD, de Vos WM. Next-generation beneficial microbes: the case of Akkermansia muciniphila. Front Microbiol. 2017;8:8.
Qu M, Liu G, Zhao J, Li H, Liu W, Yan Y, et al. Fate of atrazine and its relationship with environmental factors in distinctly different lake sediments associated with hydrophytes. Environ Pollut. 2020;256:113371.
Li T, Nakatsu G, Chen YX, Yau TO, Chu E, Wong S, et al. A novel faecal Lachnoclostridium marker for the non-invasive diagnosis of colorectal adenoma and cancer. Gut. 2020;69:1248–57.
Wasserfallen A, Nolling J, Pfister P, Reeve J, de Macario EC. Phylogenetic analysis of 18 thermophilic Methanobacterium isolates supports the proposals to create a new genus, Methanothermobacter gen. nov., and to reclassify several isolates in three species, Methanothermobacter thermautotrophicus comb. nov., Methanothermobacter wolfeii comb. nov., and Methanothermobacter marburgensis sp. nov. Int J Syst Evol Microbiol. 2000;50:43–53.
Nakao R, Abe T, Funayama S, Sugimoto C. Horizontally transferred genetic elements in the tsetse fly genome: an alignment-free clustering approach using Batch Learning Self-Organising Map (BLSOM). Biomed Res Int. 2016;2016:8.
Woo PCY, Lau SKP, Teng JLL, Tse H, Yuen KY. Then and now: use of 16S rDNA gene sequencing for bacterial identification and discovery of novel bacteria in clinical microbiology laboratories. Clin Microbiol Infec. 2008;14:908–34.
He XM, Cheng TY. Tick saliva microbiomes isolated from engorged and partially fed adults of Haemaphysalis flava tick females. J Appl Entomol. 2018;142:173–80.
Beard C, Cordon-Rosales C, Durvasula R. Bacterial symbionts of the triatominae and their potential use in control of chagas disease transmission. Annu Rev Entomol. 2002;47:123–41.
Schabereiter-Gurtner C, Lubitz W, Rölleke S. Application of broad-range 16S rRNA PCR amplification and DGGE fingerprinting for detection of tick-infecting bacteria. J Microbiol Methods. 2003;52:251–60.
Cohen C, Toh E, Munro D, Dong Q, Hawlena H. Similarities and seasonal variations in bacterial communities from the blood of rodents and from their flea vectors. ISME J. 2015;9:1662–76.
Breitschwerdt EB, Kordick DL. Bartonella infection in animals: carriership, reservoir potential, pathogenicity, and zoonotic potential for human infection. Clin Microbiol Rev. 2000;13:428–38.
Maggi RG, Richardson T, Breitschwerdt EB, Miller JC. Development and validation of a droplet digital PCR assay for the detection and quantification of Bartonella species within human clinical samples. J Microbiol Methods. 2020;176:106022.
Abdullah S, Helps C, Tasker S, Newbury H, Wall R. Pathogens in fleas collected from cats and dogs: Distribution and prevalence in the UK. Parasit Vectors. 2019;12:10.
Lappin MR, Griffin B, Brunt J, Riley A, Burney D, Hawley J, et al. Prevalence of Bartonella species, haemoplasma species, Ehrlichia species, Anaplasma phagocytophilum, and Neorickettsia risticii DNA in the blood of cats and their fleas in the United States. J Feline Med Surg. 2006;8:85–90.
Richards AL, Jiang J, Omulo S, Dare R, Abdirahman K, Ali A, et al. Human infection with Rickettsia felis. Kenya Emerg Infect Dis. 2010;16:1081–6.
Hii SF, Kopp SR, Abdad MY, Thompson MF, O’Leary CA, Rees RL, et al. Molecular evidence supports the role of dogs as potential reservoirs for Rickettsia felis. Vector Borne Zoonotic Dis. 2011;11:1007–12.
Acknowledgements
We thank the many veterinarians, veterinary technicians, veterinary students, and volunteers who coordinated and carried out trap-neuter-release programs to assist both free-roaming cats and this study, specifically, Paws and Love Inc. (Kelli Ferris), Gastonia County Animal Services (Kristine Blankenship), the San Francisco SPCA (Amie Knieper), Maplewood Animal Hospital (Lori Chang), UC Davis Veterinary Medical Teaching Hospital (Grace Woo), the Orphaned Kitten Project (Karen Vernau), and Beaumont Sainsbury Animal Hospital (Paul Pollard). We would also like to thank Dr. James Flowers for his assistance in flea species identification.
Funding
A portion of this project was completed while CM was supported by the North Carolina State University’s Comparative Medicine Institute Summer Interdisciplinary Research Initiative and the NC State Molecular Biotechnology Training Program of the National Institutes of Health under award number 1T32GM133366. Additionally, portions of this project were also completed while EL’s research was supported by the Comparative Medicine and Translational Research Program of the National Institutes of Health under award number T32OD011130 and by the University of Wisconsin School of Veterinary Medicine Companion Animal Fund. Funding sources had no role in study conceptualization, data collection, data analysis, or manuscript preparation and submission.
Author information
Authors and Affiliations
Contributions
CM, HB, and EL designed the study. CM, RM, EB, WS, and EL acquired study resources and funding. CM, KF, WS, and EL collected samples. CM, HB, BC, and EL conceptualized data analysis. CM and HB performed formal analysis and data visualization. CM and HB performed original manuscript draft preparation. All authors reviewed and edited the final manuscript draft. BC, EB, KF, RM, and EL supervised the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This study was approved by the North Carolina State University IACUC (Protocol #19-003).
Consent for publication
Not applicable.
Competing interests
In conjunction with Sushama Sontakke and North Carolina State University, Edward B. Breitschwerdt, DVM holds US Patent No. 7,115,385: “Media and Methods for Cultivation of Microorganisms,” issued 3 October 2006. He is a co-founder, shareholder, and Chief Scientific Officer for Galaxy Diagnostics, a company that provides advanced diagnostic testing for the detection of Bartonella species infections. Ricardo Maggi is the Chief Technical Officer, share-holder and co-founder of Galaxy Diagnostics. All other authors declare no potential competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1
: Figure S1. Decontam assigned metric score for ASVs from the genera identified to have at least one ASV with a decontam score above the minimum from Bartonella, Rickettsia and Wolbachia (> 0.89). Those identified as candidate microbiome members on the basis of literature search are outlined in green while the known microbiome members are outlined in red.
Additional file 2
: Figure S2. Phylogenetic tree aligning haplotypes detected in the present study to those reported by Lawrence et al.37 Nodes which contained haplotypes reported by this study are highlighted in red and indicated by a capital H.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Manvell, C., Berman, H., Callahan, B. et al. Identification of microbial taxa present in Ctenocephalides felis (cat flea) reveals widespread co-infection and associations with vector phylogeny. Parasites Vectors 15, 398 (2022). https://doi.org/10.1186/s13071-022-05487-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13071-022-05487-1