Abstract
Background
This study was carried out to determine the prevalence and the genetic background of extended-spectrum β-lactamase-producing Escherichia coli invasive isolates obtained from a tertiary-care hospital in Budapest, Hungary.
Methods
Between October–November 2018, all invasive ESBL-producing E. coli isolates were collected from Central Hospital of Southern Pest. The antimicrobial susceptibility testing was performed according to the EUCAST guidelines. The possible clonal relationships were investigated by core genome (cg)MLST (SeqSphere +) using whole-genome sequencing (WGS) data of isolates obtained from Illumina 251-bp paired-end sequencing. From WGS data acquired antimicrobial resistance genes, virulence genes and replicon types were retrieved using ResFinder3.1, PlasmidFinder2.1, pMLST-2.0, VirulenceFinder2.0 and Virulence Factors Database online tools.
Results
Overall, six E. coli isolates proved to be resistant to third-generation cephalosporins and ESBL-producers in the study period. Full genome sequence analysis showed that five E. coli isolates belonged to the ST131 clone: two to C1-M27 subclade with blaCTX-M-27 and three to C2/H30Rx subclade with blaCTX-M-15. One isolate belonged to ST1193 with blaCTX-M-27. According to cgMLST, all C2/H30Rx isolates formed a cluster (≤ 6 allele differences), while the blaCTX-M-27-producing C1-M27 isolates differed at least 35 alleles from each other. Both C2/H30Rx and C1-M27 ST131 isolates harbored similar antimicrobial resistance gene sets. However, only C2/H30Rx isolates had the qnrB and aac(3)-IIa. The isolates carried similar extraintestinal virulence gene set but differed in some genes encoding siderophores, protectins and toxins. Moreover, only one C2/H30Rx isolate carried salmochelin siderophore system and showed virotype B. All isolates showed resistance against ceftriaxone, cefotaxime, and ciprofloxacin, and the C2/H30Rx isolates were also resistant to gentamicin, tobramycin, and ceftazidime.
Conclusions
Out of six ESBL-producing E. coli, five belonged to the ST131 clone. This study indicates, that the C2/H30Rx and C1-M27 subclades of the ST131 appear to be the dominant clones collected in a Hungarian hospital.
Similar content being viewed by others
Background
Recently, among the most worrying multidrug resistant bacteria the burden of disease caused by third-generation cephalosporin-resistant Escherichia coli increased the most, in terms of the number of infections and the number of deaths in Europe [1]. Its global spread is associated with the sequence type 131 (ST131) high-risk clonal group. Members of ST131 are primarily extraintestinal pathogenic E. coli (ExPEC) harbor various virulence genes that allow them to cause severe extraintestinal infections, including bloodstream infections, urinary tract infections, pneumonia, and neonatal meningitis [2].
E. coli ST131 high-risk clones connected with extended-spectrum β-lactamase (ESBL) production and fluoroquinolone resistance causing multidrug-resistant infections belong to the most prevalent clade C associated with fimH30. Within C/H30, two subclades were emerging in the 2000s: C1/H30R subclade with fluoroquinolone resistance (FQ-R) and C2/H30Rx subclade with blaCTX-M-15, and FQ-R [3, 4]. In the late 2000s, C1-M27 clade with blaCTX-M-27 of C1/H30R became dominant in Asia and showed global distribution like the C2/H30Rx did before [4]. The expansion of the C1 and C2 subclades was driven by the acquisition of CTX-M genes containing IncF plasmids which evolve with the clone. The F2:A1:B- plasmid carrying the blaCTX-M-15 gene is associated with the C2/H30Rx subclade and the blaCTX-M-27 encoding F1:A2:B20 plasmid is associated with the C1-M27 subclade [5, 6]. The other recently emerging high-risk clones among ESBL-producing E. coli are ST10, ST38, ST69, ST155, ST315, ST405, ST410, ST648, and ST1193 [7,8,9,10].
Previous studies have shown that the proportion of third-generation cephalosporin-resistance among invasive E. coli has risen in Hungary from 5.1% in 2006 to 22.6% in 2018 [11, 12]. However, there are few data available on the incidence of third-generation cephalosporin-resistant invasive E. coli from Hungary, and its genomic epidemiology and no prospective study was made before.
This study was designed to determine the incidence of third-generation cephalosporin-resistant invasive E. coli collected from a Hungarian hospital; and to perform genomic typing of these isolates in order to characterize their genetic background.
Methods
Study design
Central Hospital of Southern Pest National Institute of Hematology and Infectious Diseases is a tertiary-care hospital in Budapest, Hungary, with a capacity of approximately 1400 beds. Between October–November 2018, all invasive consecutive non-duplicate clinical isolates of ESBL-producing E. coli were collected in the Central Hospital of Southern Pest. An isolate was considered invasive if it has been isolated from blood or cerebrospinal fluid [1].
Bacterial isolates and antimicrobial susceptibility testing
The culturing from clinical samples, the preliminary antibiotic susceptibility tests and the identification of isolates were performed in the clinical microbiological laboratory of the hospital. All isolates were identified using matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI Biotyper, Bruker, Bremen, Germany).
The ability of ESBL-production were determined by Double Disc Synergy Test (DDST) for all isolates. The DDST confirmation test was performed by the “ESβL Detection Disc Set” (MAST Diagnostica, Reinfeld, Germany) according to the manufacturer’s instruction.
The antibiotic susceptibility test of putative ESBL-producing isolates was performed to ceftriaxone, ceftazidime, cefotaxime, gentamicin, amikacin, tobramycin, fosfomycin, ceftazidime/avibactam, tigecycline, ertapenem by MIC Test Strips (Liofilchem, Roseto degli Abruzzi, Italy); and to imipenem, meropenem, ciprofloxacin, colistin by broth microdilution and interpreted using EUCAST guidelines (EUCAST v_8.0; [13], EUCAST v_11.0 [14]). Changes have been made between the clinical breakpoints of EUCAST v_8.0 (valid in 2018) and the current EUCAST guideline v_11.0. The results were interpreted according to both versions. The clinical breakpoints for interpretation are shown in the Table 1.
The susceptibility tests were performed using Mueller Hinton Agar (Bio-Rad, Hercules, California, USA) and the cultures were incubated at temperatures of 35–37 °C for 16–20 h.
Statistical analysis
The incidence rate or incidence density of invasive infection caused by E. coli were calculated per 1000 hospital admissions or 1000 patient–days, respectively.
Molecular characterization
The DNA extraction from the bacterial cultures was performed with the DNeasy UltraClean Microbial Kit (Qiagen, Hilden, Germany) according to the manufacturer's instruction. Genomic DNA analysis of the isolates was performed by whole-genome sequencing (WGS) using Illumina MiSeq 251-bp paired-end sequencing. Raw data were processed by Ridom SeqSphere + and assembly was performed using Velvet 1.1.04 [15]. The quality of the sequencing of a given sample was accepted if the genome quality indicators of the assembly were: N50 > 100 kb, approximated assembled genome size: 5 ± 0.3 Mb and average depth of sequencing coverage at least 50-fold. The possible clonal relationships were investigated by core genome (cg)MLST (SeqSphere +). Cluster type (CT) for E. coli was defined as isolates attributed to the same clone with ≤ 10 differing alleles (CT threshold) within a group. The ST131 isolates were compared to three international ST131 E. coli genomes (C1-M27: EC81009 [16] and H105 [17], C2/H30Rx: JJ2434 [18]) obtained from GenBank, to reveal the relatedness to the subclades of the ST131 clone. The replicon types were retrieved using PlasmidFinder2.1 and pMLST-2.0 [19] online tools at the Center for Genomic Epidemiology (CGE, http://genomicepidemiology.org/). In order to determine sequence similarity (coverage and identity), the isolates were aligned to the pEC-81009 (125.7 Kb, Group 1 replicon: IncFII_1, IncFIA_2, IncFIB_20), to the uk_P46212 plasmid (143.7 Kb, Group 2 of replicon: IncFII_2, IncFIA_1) and pU1 (F1:A1:B16) reference plasmid sequences [6]. The reads were trimmed (the quality threshold of 30 reads was filtered for a window of 20 bases based on filtering) and mapped to the reference plasmid sequences by BWA (SeqSphere +) and the mapped sequences were analyzed by BLASTn. The pEC-81009 (C1 clades reference plasmid, Group 1 of replicon: IncF[F1:A2:B20]; CP021180), the uk_P46212 plasmid (C2 clades reference plasmid, Group 2 of replicon: IncF[F2:A1:B-]; CP013657), and the pU1 (MK295825) reference plasmid sequences, and their terminologies and definitions were obtained from a meta-analysis of Kondratyeva et al. [6], and the GenBank. The presence of the M27PP1 prophage-like region characteristic was investigated by comparing it with the M27PP1 prophage-like region sequences of KUN5781 [20]. From WGS data the acquired antimicrobial resistance genes were identified using ResFinder3.1 [21] CGE online tool. The online analysis was performed with the default settings of 30% of the identity threshold and 20% of minimum length overlap. The presence of gene was accepted when the identity and coverage were > 90% identified.
Virulence factor genes were retrieved using VirulenceFinder2.0 [22] and Virulence Factors Database (VFDB [23]) online tools. The virotypes A to D of the ST131 isolates were assigned according to the scheme developed according to Blanco et al. [24]: virotype A (afa positive, iroN negative, ibeA negative, sat positive or negative), virotype B (afa negative, iroN positive, ibeA negative, sat positive or negative), virotype C (afa negative, iroN negative, ibeA negative, sat positive), and virotype D (afa negative, iroN positive or negative, ibeA positive, sat positive or negative).
The raw reads are available on the Sequence Read Archive (SRA) database under the BioProject number PRJNA683640.
Results
Clinical epidemiology
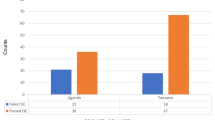
Twenty-five E. coli were isolated from haemoculture in the study period. Six from 25 isolates (24%) proved to be resistant to third-generation cephalosporins and were obtained from patients with bloodstream infection. The incidence density and the incidence rate of third-generation cephalosporin-resistant E. coli were 0.11 per 1,000 patient-days and 0.79 per 1,000 hospital admission, respectively. The phenotypic detection tests showed that all of them were ESBL-producer. Out of the six isolates collected from the Central Hospital of Southern Pest, two isolates originated from the Haematology and Stem Cell Transplantation ward, two from the Infectious Diseases ward, one from the Intensive Care Unit and the last one from the 1st Internal Medicine ward. The median age of the patients was 70 years (range 38–90), and the sex ratio was 1:1.
Antimicrobial susceptibility
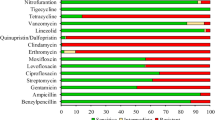
According to the both EUCAST guideline versions the isolates were resistant to ceftriaxone, cefotaxime, and ciprofloxacin but remained susceptible to colistin, fosfomycin, ceftazidime-avibactam, and carbapenems (Table 1). The Ec1, Ec2, and Ec3 showed resistance to ceftazidime, gentamicin, and tobramycin too. Among several revised clinical breakpoints of investigated antibiotics only the change of tigecycline breakpoints has affected the susceptibility interpretations of our isolates.
Molecular characterization
The whole-genome sequencing and genome assembly results met all quality requirements for all isolates. The assembled draft genome size of the isolates was 5 325 328 bp for Ec1, 5 309 727 bp for Ec2, 5 298 859 bp for Ec3, 5 041 503 bp, for Ec4, 5 070 035 bp for Ec5 and 5 115 835 bp for Ec6.
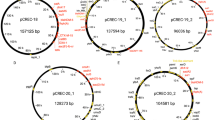
The six ESBL-producing E. coli isolates could be assigned to two different sequence types (ST) by MLST. Of the six isolates, five (Ec1-Ec5) belonged to the globally dominant ST131 clone (O25:H4 serotype), which comprised two distinguished groups of strains with different characteristics. The first group contains two isolates with blaCTX-M-15 and one isolate with blaCTX-M-15 and blaTEM-1B while the two other isolates harbored blaCTX-M-27 ESBL-genes. The remaining isolate belonged to the ST1193 with blaCTX-M-27. Analysis of genetic relatedness between the strains was performed by cgMLST. The five ST131 isolates were compared to one C2/H30Rx (JJ2434) and two C1-M27 (H105, EC81009) international ST131 ESBL-producing E. coli isolates, to determine the subclade-specific relatedness of the isolates within the ST131 (Fig. 1). The JJ2434 isolate and the three CTX-M-15-producing isolates (named the first group below) belonged ST131 C2/H30Rx subclade. The H105 and EC81009 isolates with the two CTX-M-27-producing isolates from this study belonged to the ST131 C1-M27 subclade. The Ec4 and Ec5 harboured exclusively the C1-M27 clade-specific M27PP1 prophage-like genomic island.
The ST131 subclade-specific phylogenetic relationship of the Escherichia coli isolates. The unrooted circular phylogenetic tree of ST131 Escherichia coli isolates based on the Ridom™ SeqSphere + core genome multilocus sequencing typing (cgMLST) including 2517 alleles. The blue colour corresponds to Sequence Type 131 C1-M27 subclade, while the red to Sequence Type 131 C2/H30Rx subclade
According to cgMLST (Fig. 2), all the three Hungarian ST131 E. coli isolates producing blaCTX-M-15 formed a close cluster (≤ 6 allele differences), while the C1-M27 isolates producing blaCTX-M-27 differed at 35 alleles from each other. The ST1193 isolates showed ≥ 2209 allele differences from any others.
Core genome MLST based Minimum Spanning Tree of the ESBL-producing Escherichia coli isolates. Minimum spanning tree of Escherichia coli isolates based on the Ridom™ SeqSphere + core genome multilocus sequencing typing (cgMLST) including 2517 alleles with a cluster threshold ≤ 10. Colors correspond with a different genotype characteristic and sequence type of the group of strains. Numbers of allele differences are indicated between the two nodes
The isolates of each group belonged to ST131 harboured very similar resistance gene sets. The characteristics of resistome and plasmid replicon types of the isolates are shown in Table 2. Although, there are some differences between the two ST131 subclades in terms of resistance genes like the qnrB (FQR) and aac(3)-IIa (aminoglycoside-R) which were present only in C2/H30Rx isolates. The ST1193 isolate harboured L416F mutation in parE, and also had no qnrB and aac(3)-IIa genes. Three plasmid families were detected: IncF, IncQ, and Col-like. The IncF replicons were present in every isolate, but only the C1-M27 ST131 isolates (Ec4, Ec5) carried the Group1 replicons (IncFII_1, IncFIA_2, IncFIB_20). The coverage of the Group1 (F1:A1:B20) reference plasmid by the reads of Ec4 and Ec5 was 96%, and 72%, respectively (Table 3). None of the C2/H30Rx isolates (Ec1-Ec3) co-harboured the two reference replicons of Group2 (IncFII_2, IncFIA_1). The Ec2 did not carry any IncFIA replicon but carried the IncFII_2 replicon. The coverage of the Group2 (F2:A1:B-) reference plasmid by the reads of Ec1, Ec2, and Ec3 was 70%, 47%, and 70%, respectively. The reads of Ec1 and Ec3 covered pU1 plasmid to a higher breadth compared to Group2 reference (≥ 91% vs. 70%). The ST1193 isolates harbored IncFII_4-like, IncF1A_1, IncFIB_10, and the repCol156, and repCol(BS512) replicons. The Col(pHAD28), Col156, and Col8282 replicons were at least present in one of the ST131 isolates.
Table 4 shows the virulence gene sets and corresponding virotype. The virulome of the isolates belonged to ST131 showed high similarities. These isolates belonged to two virotypes: B and C. Four ST131 isolates (Ec1, Ec3-5) showed virotype C and the Ec2 ST131 isolate showed virotype B. The Ec2 carried 63 virulence genes while the other isolates (including the ST1193 isolate) carried 57 ± 1 virulence genes. Difference was found between the iron uptake genes (Ec2 carried exclusively the iroB,iroC, iroD, iroE, iroN genes) and several toxin genes.
Discussion
In our study, the incidence density of invasive infection caused by third-generation cephalosporin-resistant E. coli was 0.11 per 1,000 patient-days and the incidence rate was 0.79 per 1,000 hospital admission. There are no previous data on the incidence of third-generation cephalosporin resistant E. coli invasive infections in Hungary.
De Kraker et al. investigated changes in the trends of blood stream infections for major bacterial pathogens by analysing the European Antimicrobial Resistance Surveillance System database between 2002–2008. In 2008 the average incidence density of invasive infections caused by third-generation cephalosporin resistant E. coli was 0.03/1,000 patient-days in Europe. There was significant difference in incidence density between northern region and southern region of Europe: 0.002/1,000 patient-days and 0.06/1,000 patient-days, respectively. In the eastern region of Europe (including Hungary) the incidence density was similar to the average (0.02/1,000 patient-days) [25]. These findings supported by investigation of Martelius et al. where the incidence density was 0.008/1,000 patient-days in 17 Finnish acute care hospitals [26] and investigation of De Angelis et al. where the incidence density was 0.1/1,000 patient-days in a teaching hospital in Rome, Italy [27] in similar period. The incidence density found in our study has increased significantly compared to incidence density found in eastern region of Europe in 2008 and became similar to that in the study from Rome.
In our study, six from the 25 E. coli isolates (24%) proved to be third-generation cephalosporin-resistant and ESBL-producer. In 2010, Pál T. et al. found 26 ESBL-producers among 117 E. coli isolates (22.2%) from bloodstream infections collected in three university hospitals in Hungary between March and November 2010, where nine of them belonged to ST131 clone carrying exclusively blaCTX-M-15 [14]. The rate of ESBL-producer E. coli was similar in our study, however, the rate of ST131 was higher (83.3 vs. 34.6%). Furthermore, in our study two of the five isolates harbouring blaCTX-M-27 gene belonged to the C1-M27 subclade. The first ESBL-producing invasive E. coli isolates belonged to the C1-M27 subclade were detected in 2015 in Hungary (unpublished data, the National Public Health Center). In 2018, according to an observational study of Jánvári et al., 44.6% (75/168) of invasive ESBL-producing E. coli isolates investigated at the National Public Health Center belonged to the ST131 clone, where the ratio of C2/H30Rx and C1-M27 was 1 to 0.8 [28].
In this study, ST131, including the C1-M27 (2 isolates) and the C2/H30Rx subclades (3 isolates) proved to be the dominant multidrug-resistant E. coli clone in invasive infections. The three ST131 C2/H30Rx isolates formed a relatively close cluster (≤ 6 allele differences), and this could suggest an undetected outbreak affected two hospital wards (Infectious Diseases ward, 1st Internal Medicine ward). However, based on the results of plasmid replicon analysis it was unlikely.
All isolates were resistant to ceftriaxone and cefotaxime. The isolates with CTX-M-15 ESBL were resistant to ceftazidime, while the CTX-M-27 expressing isolates remained susceptible, probably because these enzymes hydrolyze ceftazidime poorly. Moreover, the C2/H30Rx isolates showed resistance to tobramycin and gentamicin probably because they have AAC(3)-IIa aminoglycoside-modifying enzyme [29].
We found one isolate that belonged to the emerging ST1193 global high-risk clone [30]. To the best of our knowledge, this study revealed for the first time the presence of the ST1193 E. coli in Hungary. This clone has been widely spread in Asia [31, 32]. The ST1193 was the second most frequent ST (23.7% of 99 isolates) after ST131 (37.3%) among ciprofloxacin-resistant invasive E. coli isolates in the Tertiary Care University Hospital in Korea [33]. In Europe, it has been described in Germany where the rate of ESBL-producing E. coli ST1193 was 0.6% (3/495) among all E. coli infections [34]. In a South-West England study, 11 of 836 E. coli from urine samples belonged to ST1193 [35].
It has been known that the narrow-host-range IncF type plasmids have been evolved along with fluoroquinolone-resistant ST131 H30 clades. The F2:A1:B- plasmid is associated with the C2/H30Rx subclade, whereas the F1:A2:B20 plasmid is strongly associated with the C1/H30R subclade [5]. The presence of Col-like plasmids has been reported also in ST131, and in ST1193 E. coli clones, however, the exact role of these plasmids is not clear [6]. In our study, each of the isolates had various Inc-type (F, Q) replicons and except Ec5, they had a Col-type (pHAD28, BS512, 156, 8282) replicons as well. The C1-M-27 isolates carried the Group1 plasmid replicons (IncFII_1, IncFIA_2, IncFIB_20) and the reads of Ec4 isolate covered the Group1 reference plasmid to a higher breadth compared to Ec5 (96 vs. 72%). The C2/H30Rx isolates did not co-encode together the standard Group2 plasmid replicons (IncFII_2, IncFIA_1). The reads of Ec1 and Ec3 shared 70% coverage with F2:A1:B- Group2 plasmid and Ec2 shared 47%. Ec1 and Ec3 had identical replicons to the pU1 plasmid (pMLST: F1:A1:B16) [6]. The coverage of pU1 by Ec1 and Ec3 reads was 91 and 92%, respectively. These findings suggest that these two isolates harbored a similar plasmid like the pU1. The ST1193 isolate had different replicons (IncFII_4-like, IncFIB_10) than in Group1 and Group2 plasmids.
It is not completely understood, which key factors determine the successful dissemination of the ST131 C2/H30Rx and C1/H30R (C1-M27) clones worldwide, besides the antibiotic genes, but the virulence armament probably has an important part in it. The ST131 extraintestinal E. coli isolates have a wide range of virulence-coding genes, including adhesins, invasines, iron-uptake factors, protectins, toxins that enable them to bind to the host cells, invade the host tissues and avoid host defense systems [36, 37]. Several studies concerned that there are differences in the content of the ST131 virulence factors, and only a few virulence genes were consistently identified in all strains. The content can vary with their virotypes [24]. We identified 56–63 virulence-associated genes in our ExPEC isolates. Among the virulence set we found, some virulence genes were associated with ST131 isolates, like sat, iha, iucD, ompT, usp [38,39,40]. The ST1193 isolate also carried the sat, iha, vat, iutA, usp which were associated with ST1193 isolate [30, 32, 41]. The isolates carried siderophores which enable them to survive in a low-iron condition: aerobactin (iucABCD), yersiniabactin (irp2), and the Ec2 isolate also carried the salmochelin (iroBCDEN) operon. The Ec2 showed virotype B, while the other isolates of both subclades of ST131 showed virotype C. The global disseminated virotype C has been described as the most prevalent E. coli ST131 virotype [24, 42]. There is only a few data in the literature about significancy of the ST131 isolate with virotype B [24, 43]. Blanco et al. found that virotype B was significantly associated with a lower likelihood of symptomatic infection compared to virotype C [24].
It has been described that ST131 clones can transmit among human and animal hosts [44]. Based on a study from Israel, the CTX-M-27-producing ST131 subclones had a higher transmission rate than CTX-M-15- producing ST131 subclones [45]. At this number of isolates, we cannot estimate if the selection effect of the antibiotic consumption influences the dissemination of the ST131 subclades, especially the third-generation cephalosporins, because it needs further investigation.
Conclusion
This study has several limitations, like small sample size and a two months time limit. However, it gives an insight into the incidence and genomic characteristics of ESBL-producing E. coli isolates in a Hungarian hospital. It highlights the dominance of the ST131 clones and the C1-M27 and C2/H30Rx subclone in invasive infections. Although, further national studies are needed with a larger collection of isolates belonging to the two subclades to determine the possible factors (virulence, antibiotic resistance) that can influence their dissemination.
Availability of data and materials
The data set supporting the results of this article are included within the article.
Abbreviations
- BSI:
-
Bloodstream infection
- cgMLST:
-
Core genome multilocus sequence typing
- CT:
-
Cluster-type
- ESBL:
-
Extended-spectrum β-lactamase
- EUCAST:
-
European Committee on Antimicrobial Susceptibility Testing
- ExPEC:
-
Extraintestinal pathogenic E. coli
- FQ-R:
-
Fluoroquinolone resistance
- MALDI-TOF:
-
Matrix-assisted laser desorption ionization time-of-flight mass spectrometry
- MIC:
-
Minimum inhibitory concentration
- MLST:
-
Multilocus sequence typing
- SRA:
-
Sequence Read Archive
- ST:
-
Sequence typing
- WGS:
-
Whole-genome sequencing
- VFDB:
-
Virulence Factors Database
References
Cassini A, Högberg LD, Plachouras D, Quattrocchi A, Hoxha A, Simonsen GS, et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: a population-level modelling analysis. Lancet Infect Dis. 2019;19:56–66.
Smith JL, Fratamico PM, Gunther NW. Extraintestinal pathogenic Escherichia coli. Foodborne Pathog Dis. 2007;4:134–63.
Pitout JDD, DeVinney R. Escherichia coli ST131: a multidrug-resistant clone primed for global domination. F1000Res. 2017;6:1–7.
Mathers AJ, Peirano G, Pitout JDD. The role of epidemic resistance plasmids and international high- risk clones in the spread of multidrug-resistant Enterobacteriaceae. Clin Microbiol Rev. 2015;28:565–91.
Johnson TJ, Danzeisen JL, Youmans B, Case K, Llop K, Munoz-Aguayo J, et al. Separate F-type plasmids have shaped the evolution of the H 30 subclone of Escherichia coli sequence type 131. MSphere. 2016. https://doi.org/10.1128/mSphere.00121-16.
Kondratyeva K, Salmon-Divon M, Navon-Venezia S. Meta-analysis of pandemic Escherichia coli ST131 plasmidome proves restricted plasmid-clade associations. Sci Rep. 2020;10:1–11.
Peirano G, Van Der Bij AK, Gregson DB, Pitout JDD. Molecular epidemiology over an 11-year period (2000 to 2010) of extended-spectrum β-lactamase-producing Escherichia coli causing bacteremia in a centralized Canadian region. J Clin Microbiol. 2012;50:294–9.
Roer L, Overballe-Petersen S, Hansen F, Schønning K, Wang M, Røder BL, et al. Escherichia coli sequence type 410 is causing new international high-risk clones. MSphere. 2018;3:e00337-e418.
Baquero F, Tedim AP, Coque TM. Antibiotic resistance shaping multi-level population biology of bacteria. Front Microbiol. 2013;4:1–15.
Schaufler K, Semmler T, Wieler LH, Trott DJ, Pitout J, Peirano G, et al. Genomic and Functional Analysis of Emerging Virulent and Multidrug-Resistant Escherichia coli Lineage Sequence Type 648. Antimicrob Agents Chemother. 2019;63:1–12.
Tóth Á, Juhász-Kaszanyitzky É, Mag T, Hajbel-Vékony G, Pászti J, Damjanova I. Characterization of extended-spectrum β -lactamase (ESBL) producing Escherichia coli strains isolated from animal and human clinical samples in Hungary in 2006–2007. Acta Microbiol Immunol Hung. 2013;60:175–85.
European Centre for Disease Prevention and Control. Surveillance of antimicrobial resistance in Europe 2018. Stockholm: ECDC; 2019.
The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 8.0, 2018. http://www.eucast.org [Accessed 1 January 2018]
The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 11.0, 2021. http://www.eucast.org [Accessed 1 January 2021]
Zerbino DR, Birney E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008;18:821–9.
Mutti M, Sonnevend Á, Pál T, Junttila S, Ekker H, Galik B, et al. Complete genome sequence of Escherichia coli 81009, a representative of the sequence type 131 C1–M27 clade with a multidrug-resistant phenotype. Genome Announc. 2018;6:1–2.
Ghosh H, Bunk B, Doijad S, Schmiedel J, Falgenhauer L, Spröer C, et al. Complete genome sequence of blaCTX-M-27-encoding Escherichia coli strain H105 of sequence type 131 lineage C1/H30R. Genome Announc. 2017;5:30–1.
Johnson TJ, Danzeisen JL, Youmans B, Case K, Llop K, Munoz-Aguayo J, et al. Separate F-type plasmids have shaped the evolution of the H30 subclone of Escherichia coli sequence type 131. MSphere. 2016;1:1–15.
Sahare P, Moon A. In silico modelling of β-lactam resistant Enterococcus faecalis PBP4 and its interactions with various phyto-ligands. Int J Pharm Pharm Sci. 2016;8:151–5.
Matsumura Y, Pitout JDD, Gomi R, Matsuda T, Noguchi T, Yamamoto M, et al. Global Escherichia coli sequence type 131 clade with bla CTX-M-27 gene. Emerg Infect Dis. 2016;22:1900–7.
Bortolaia V, Kaas RS, Ruppe E, Roberts MC, Schwarz S, Cattoir V, et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J Antimicrob Chemother. 2020;75:3491–500.
Joensen KG, Scheutz F, Lund O, Hasman H, Kaas RS, Nielsen EM, et al. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J Clin Microbiol. 2014. https://doi.org/10.1128/JCM.03617-13.
Chen L, Zheng D, Liu B, Yang J, Jin Q. VFDB 2016: Hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Res. 2016;44:D694–7.
Blanco J, Mora A, Mamani R, López C, Blanco M, Dahbi G, et al. Four main virotypes among extended-spectrum-β-lactamase-producing isolates of Escherichia coli O25b:H4-B2-ST131: bacterial, epidemiological, and clinical characteristics. J Clin Microbiol. 2013;51:3358–67.
de Kraker MEA, Jarlier V, Monen JCM, Heuer OE, van de Sande N, Grundmann H. The changing epidemiology of bacteraemias in Europe: trends from the European antimicrobial resistance surveillance system. Clin Microbiol Infect. 2013;19:860–8. https://doi.org/10.1111/1469-0691.12028.
Martelius T, Jalava J, Kärki T, Möttönen T, Ollgren J, Lyytikäinen O. Nosocomial bloodstream infections caused by Escherichia coli and Klebsiella pneumoniae resistant to third-generation cephalosporins, Finland, 1999–2013: Trends, patient characteristics and mortality. Infect Dis (Auckl). 2016;48:229–34.
De Angelis G, Fiori B, Menchinelli G, D’Inzeo T, Liotti FM, Morandotti GA, et al. Incidence and antimicrobial resistance trends in bloodstream infections caused by ESKAPE and Escherichia coli at a large teaching hospital in Rome, a 9-year analysis (2007–2015). Eur J Clin Microbiol Infect Dis. 2018;37:1627–36.
Jánvári L, Damjanova I, Topf J, Mag T, Mestyan G, Szolyka G, et al. Countrywide dissemination of CTX-M-27-producing Escherichia coli ST131 subclade C1-M27 in Hungary, 2015-2017. ECCMID 2018 accepted abstract (Poster number P1495)
Shaw KJ, Rather PN, Hare RS, Miller GH. Molecular genetics of aminoglycoside resistance genes and familial relationships of the aminoglycoside-modifying enzymes. Microbiol Rev. 1993;57:138–63.
Johnson TJ, Elnekave E, Miller EA, Munoz-Aguayo J, Flores Figueroa C, Johnston B, et al. Phylogenomic analysis of extraintestinal pathogenic Escherichia coli sequence type 1193, an emerging multidrug-resistant clonal group. Antimicrob Agents Chemother. 2018;63:1–15.
Al-Farsi HM, Camporeale A, Ininbergs K, Al-Azri S, Al-Muharrmi Z, Al-Jardani A, et al. Clinical and molecular characteristics of carbapenem non-susceptible Escherichia coli: a nationwide survey from Oman. PLoS ONE. 2020. https://doi.org/10.1371/journal.pone.0239924.
Wu J, Lan F, Lu Y, He Q, Li B. Molecular characteristics of ST1193 clone among phylogenetic group B2 Non-ST131 fluoroquinolone-resistant Escherichia coli. Front Microbiol. 2017. https://doi.org/10.3389/fmicb.2017.02294.
Kim Y, Oh T, Nam Y, Cho S, Lee H. Prevalence of ST131 and ST1193 among bloodstream isolates of Escherichia coli not susceptible to ciprofloxacin in a Tertiary Care University Hospital in Korea, 2013–2014. Clin Lab. 2017. https://doi.org/10.7754/Clin.Lab.2017.170319.
Valenza G, Werner M, Eisenberger D, Nickel S, Lehner-Reindl V, Höller C, et al. First report of the new emerging global clone ST1193 among clinical isolates of extended-spectrum β-lactamase (ESBL)-producing Escherichia coli from Germany. J Glob Antimicrob Resist. 2019;17:305–8.
Findlay J, Gould VC, North P, Bowker KE, Williams MO, MacGowan AP, et al. Characterization of cefotaxime-resistant urinary Escherichia coli from primary care in South-West England 2017–18. J Antimicrob Chemother. 2020;75:65–71.
Dahbi G, Mora A, Mamani R, López C, Alonso MP, Marzoa J, et al. Molecular epidemiology and virulence of Escherichia coli O16: H5-ST131: comparison with H30 and H30-Rx subclones of O25b: H4-ST131. Int J Med Microbiol. 2014;304:1247–57. https://doi.org/10.1016/j.ijmm.2014.10.002.
Sarowska J, Futoma-Koloch B, Jama-Kmiecik A, Frej-Madrzak M, Ksiazczyk M, Bugla-Ploskonska G, et al. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: Recent reports. Gut Pathog. 2019;11:1–16. https://doi.org/10.1186/s13099-019-0290-0.
Johnson JR, Johnston B, Clabots C, Kuskowski MA, Castanheira M. Escherichia coli sequence type ST131 as the major cause of serious multidrug-resistant E. coli infections in the United States. Clin Infect Dis. 2010;51:286–94.
Mora A, Dahbi G, López C, Mamani R, Marzoa J, Dion S, et al. Virulence patterns in a murine sepsis model of ST131 Escherichia coli clinical isolates belonging to serotypes O25b:H4 and O16:H5 are associated to specific virotypes. PLoS ONE. 2014. https://doi.org/10.1371/journal.pone.0087025.
Coelho A, Mora A, Mamani R, López C, González-López JJ, Larrosa MN, et al. Spread of Escherichia coli O25b:H4-B2-ST131 producing CTX-M-15 and SHV-12 with high virulence gene content in Barcelona (Spain). J Antimicrob Chemother. 2011;66:517–26.
Johnson JR, Johnston BD, Porter SB, Clabots C, Bender TL, Thuras P, et al. Rapid emergence, subsidence, and molecular detection of Escherichia coli sequence type 1193-fimH64, a new disseminated multidrug-resistant commensal and extraintestinal pathogen. J Clin Microbiol. 2019. https://doi.org/10.1128/JCM.01664-18.
Nicolas-Chanoine MH, Bertrand X, Madec JY. Escherichia coli st131, an intriguing clonal group. Clin Microbiol Rev. 2014;27:543–74.
Lanza VF, de Toro M, Garcillán-Barcia MP, Mora A, Blanco J, Coque TM, et al. Plasmid flux in Escherichia coli ST131 sublineages, analyzed by plasmid constellation network (PLACNET), a new method for plasmid reconstruction from whole genome sequences. PLoS Genet. 2014. https://doi.org/10.1371/journal.pgen.1004766.
Kidsley AK, White RT, Beatson SA, Saputra S, Schembri MA, Gordon D, et al. Companion animals are spillover hosts of the multidrug-resistant human extraintestinal escherichia coli pandemic clones ST131 and ST1193. Front Microbiol. 2020;11:1–10.
Adler A, Gniadkowski M, Baraniak A, Izdebski R, Fiett J, Hryniewicz W, et al. Transmission dynamics of ESBL-producing Escherichia coli clones in rehabilitation wards at a tertiary care centre. Clin Microbiol Infect. 2012. https://doi.org/10.1111/j.1469-0691.2012.03999.x.
Acknowledgements
Dr. Ivelina Damjanova provided helpful suggestions and assisted by critically reading the manuscript.
Funding
Open access funding provided by Semmelweis University. This study was financially supported by OTKA Hungarian Scientific Fund, Grant Number: 108481.
Author information
Authors and Affiliations
Contributions
KT and ÁT contributed to the study design. KT performed the experiments and wrote the manuscript. KK and VN collected the samples. TK, ÁT, and DSz drafted and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This is a retrospective study, not directly associated with patients and it was consistent with the principles of the Helsinki Declaration.
Consent for publication
All authors approve the publication of this work.
Competing interests
All authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Tóth, K., Tóth, Á., Kamotsay, K. et al. Population snapshot of the extended-spectrum β-lactamase-producing Escherichia coli invasive strains isolated from a Hungarian hospital. Ann Clin Microbiol Antimicrob 21, 3 (2022). https://doi.org/10.1186/s12941-022-00493-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12941-022-00493-8