Abstract
Background
Ubiquitin-specific proteases (UBPs) are a large family of deubiquitinating enzymes (DUBs). They are widespread in plants and are critical for plant growth, development, and response to external stresses. However, there are few studies on the functional characteristics of the UBP gene family in the important staple crop, maize (Zea mays L.).
Results
In this study, we performed a bioinformatic analysis of the entire maize genome and identified 45 UBP genes. Phylogenetic analysis indicated that 45 ZmUBP genes can be divided into 15 subfamilies. Analysis of evolutionary patterns and divergence levels indicated that ZmUBP genes were present before the isolation of dicotyledons, were highly conserved and subjected to purifying selection during evolution. Most ZmUBP genes exhibited different expression levels in different tissues and developmental stages. Based on transcriptome data and promoter element analysis, we selected eight ZmUBP genes whose promoters contained a large number of plant hormones and stress response elements and were up-regulated under different abiotic stresses for RT-qPCR analysis, results showed that these genes responded to abiotic stresses and phytohormones to varying degrees, indicating that they play important roles in plant growth and stress response.
Conclusions
In this study, the structure, location and evolutionary relationship of maize UBP gene family members were analyzed for the first time, and the ZmUBP genes that may be involved in stress response and plant growth were identified by combining promoter element analysis, transcriptome data and RT-qPCR analysis. This study informs research on the involvement of maize deubiquitination in stress response.
Similar content being viewed by others
Background
Ubiquitin-specific proteases (UBPs), the largest subfamily of plant deubiquitinating enzymes (DUBs), are involved in diverse physiological processes such as plant growth and development [1,2,3,4,5], as well as stress response [6,7,8]. According to previous studies, eukaryotic DUBs can be classified into five subfamilies and two types based on their catalytic domains: cysteine proteases and metalloproteinases [9, 10], with UBPs falling into the cysteine protease category [11]. UBP contains the Ub carboxy-terminal hydrolase (UCH) domain, which is mainly composed of two conserved motifs, cysteine cysteine (Cys) and histidine histidine (His) boxes, and plays an important role in the deubiquitination of plants [12]. Notably, the number of UBP genes varies significantly across species. For instance, the Arabidopsis (Arabidopsis thaliana) genome encodes 27 UBP genes [13], 25 genes in rice (Oryza sativa) [14], 48 genes in moso bamboo (Phyllostachys edulis) [15], and 97 in wheat (Triticum aestivum) [16]. The functions of Arabidopsis UBP genes have been extensively studied. Previous research has shown that AtUBP3 and AtUBP4 play crucial roles in male gametophyte development [1]. AtUBP14 plays a role in the early embryonic development of plants [17]. AtUBP26, on the other hand, is involved in the ubiquitination modification process of histones and is essential for seed development [2, 18]. Furthermore, a recent study has revealed that UBP15 plays a significant role in regulating seed development in both Arabidopsis and rice. UBP15 modulates organ development and seed size in an opposing manner to the ubiquitin receptor DA1, and it positively regulates seed size by promoting cell proliferation in the maternal bead tissues [19]. OsUBP15 directly interacts with OsDA1 to positively regulate the length and width of rice seeds [20]. Loss of AtUBP1 and AtUBP2 function results in hypersensitivity of plants to the amino acid analogue canavaline (CAN) and severe dwarfing, short root development, and yellowing of leaves [13]. Overexpression of UBP12/UBP13 can increase the NAC domain transcription factor ORE1 level and positively regulate leaf senescence induced by nitrogen deficiency [4]. In addition, the deubiquitination enzymes UBP12 and UBP13 regulate the growth process of plants under nitrogen deficiency and positively regulate the recovery process after carbon starvation by regulating the stability of Arabidopsis BES1. In addition, UBP12 and UBP13 directly interact with RGF1 receptors to counteract RGF1-induced ubiquitination and promote root meristem development [21]. UBP12 and UBP13 also play an important role in the regulation of plant flowering time and the biological clock of plants [5]. UBPs have been shown to play an important role in stress responses, such as the plant immune response, drought response mediated by the ABA signaling pathway, salt response and other biological processes [6, 8, 22,23,24].
Maize is one of the world’s leading crops and is of considerable value to feed, food, pharmaceutical, and other industries [25, 26]. However, the UBP gene family in maize has not been extensively studied. To date, only three ZmUBP genes (ZmUBP15, ZmUBP16 and ZmUBP19) have been characterized in maize [27]. ZmUBP15, ZmUBP16 and ZmUBP19 are the three homologous genes of Arabidopsis UBP16 in maize, which play similar functions to Arabidopsis UBP16 in response to salt stress. ZmUBP15, ZmUBP16 and ZmUBP19 expression levels are reduced under salt stress and partially rescue the salt-sensitive phenotype of Arabidopsis ubp16-1 mutants, significantly enhancing the tolerance of ubp16-1 mutants to salt stress [27]. In order to investigate the functions played by members of the maize UBP gene family in plant growth and development and stress response, we identified 45 ZmUBP genes in maize genome-wide, and their conserved motifs, gene structures, chromosome distributions, and expression patterns were analyzed. To understand their evolutionary relationship with other plants, a phylogenetic tree was constructed. Furthermore, the expression profiles of the ZmUBP genes under abiotic stresses and hormone conditions were assessed by using RT-qPCR. The findings of our study will help to understand the roles of ZmUBP genes in the stress response and to further identify the functions of this essential gene family in maize.
Methods
Identification of UBP genes in maize
The hidden Markov model (HMM) profile of the UCH domain (PF00443) obtained from the Pfam database (http://pfam.xfam.org/) was used to blast the maize protein sequence file using the local HMMER 3.0 program [28]. The E-value was limited to less than 1 × 10−18. All the identified ZmUBP candidates were verified using the Pfam database (http://pfam.xfam.org/). Proteins that did not have the UCH protein domain with highly conserved Cys residues (Cys-box) as well as His and Asp/Asn residues (His-box) were excluded. We then turned to the NCBI CD search (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) to further annotate these genes. With the essential information on hand, we bioinformatically analyzed the ZmUBP genes using ExPASy (http://www.expasy.ch/tools/pi_tool.html). This analysis allowed us to determine the molecular weight (MW) and isoelectric point (pI) of the ZmUBP proteins. To gain a deeper understanding of their structural features, we predicted transmembrane structural domains using TMHMM (http://www.cbs.dtu.dk/services/TMHMM). Additionally, we utilized Plant-Ploc (http://www.csbio.sjtu.edu.cn/) to predict the hydrophobicity of the ZmUBP proteins.
Analysis of ZmUBP gene structure and protein structure
To analyze the structure of ZmUBP genes, we used TBtools [29, 30] to compare CDS of the ZmUBP gene family with genomic DNA, mapped exon-intron structure, and predicted protein conserved motifs using MEME (https://meme-suite.org/). These motifs were mapped with TBtools [29, 30]. Finally, we combined the Gene Structure, protein conserved motifs, and protein domains based on the Gene Structure View (Advanced) function of Tbtools [29, 30].
Sequence alignment and phylogenetic analysis
In this study, we retrieved 27 Arabidopsis UBP protein sequences from TAIR (https://www.arabidopsis.org/), 25 rice UBP protein sequences from Phytozome (https://phytozome.jgi.doe.gov/pz/portal.html), and 97 wheat UBP protein sequences from a previous study [16]. To investigate the evolutionary relationships among these proteins, we constructed a phylogenetic tree using the neighbor-joining method in MEGA 6.0 and visualized by iTOL (https://itol.embl.de/). To ensure the statistical reliability of our findings, we performed bootstrap testing with 1000 replicates.
Analysis of cis-acting elements of ZmUBP gene promoters
To gain insights into the cis-elements in the promoter region of ZmUBP genes, we extracted the 2000 bp sequence upstream of these genes. We utilized the PlantCARE (http://www.dna.affrc.go.jp/PLACE/) to predict putative cis-regulatory elements in the promoter sequences. Notably, we identified functionally identical cis-acting elements that were uniquely named. To visualize the results of our analysis, we employed the Simple BioSequence Viewer function of TBtools [29, 30].
Chromosome position, collinearity analysis and calculation of Ka/Ks ratios
We downloaded maize gff3 files from Ensembl Plants (https://plants.ensembl.org/) to analyze the annotation of ZmUBP genes on the chromosome. Using the maize genome annotation information, we obtained the relative distance and location of the ZmUBP genes on the chromosome. To visualize this data, we utilized the Gene Locatin Visualize feature of TBtools [29, 30].
The Dual Systeny Plot program in TBtools [29, 30] was utilized to assess the homology of UBP genes among maize and other species, including Arabidopsis, sorghum, soybean, Kinnow Mandarin (Citrus reticulata Blanco), cotton, medicago (Medicago sativa), rice, and wheat. To further analyze the collinearity and Ka/Ks ratio, we employed the One Step MCScanX and simple Ka/Ks calculator (NJ) of TBtools [29, 30], respectively. The results were visualized using the Microlife Letter (http://www.bioinformatics.com.cn/), providing a comprehensive understanding of the homology relationships among these species.
KEGG and GO enrichment analysis
KEGG and GO enrichment analysis were performed using the microbiotics website (http://www.bioinformatics.com.cn/) to investigate the signalling pathways, biological processes, cellular components and molecular functions involved in the 45 ZmUBP genes.
Temporal and spatial expression profiles of ZmUBP genes
We downloaded transcriptome data for different organs and stress responses of maize from the EMBL-EBI database (https://www.ebi.ac.uk/). To integrate the transcriptome data of the same type, we utilized EXCEL. The results were visualized and presented as heatmaps using the Microbiology Letter website (http://www.bioinformatics.com.cn/).
Plant material, growth conditions, and stress treatments
Hybrid Zhengdan 958 was provided by Grain Crops Research Institute, Henan Academy of Agricultural Sciences (Validation No.: Guoshiyu 20000009, Date of Validation: 2000, Selection and Breeding Unit: Grain Crops Research Institute, Henan Academy of Agricultural Sciences, Selected Breeder: Chunxin Du, Variety Source: Zheng 58/Chang 7 − 2). To investigate the response of maize to various stress conditions, uniform-sized maize seeds were selected and sterilized with 75% ethanol for 5 min. The seeds were then rinsed five times with sterile water and placed in an incubator containing double layers of filter paper. The incubator was set to vernalize the seeds under a 12-hour light (25 °C) and 12-hour dark (20 °C) cycle. After 1 day of germination, well-established and uniform seedlings were selected and transferred into pots containing a mixture of vermiculite and soil (3:1 ratio). After 10 days of growth, the roots of the maize seedlings were watered with 100 mL of a 12% PEG 2000 solution to simulate drought stress. The seedlings were then exposed to low temperature (4 °C) and high temperature (42 °C) stress conditions. Additionally, hormone treatments were applied by watering the roots of the seedlings with different types of plant hormone solutions. All experiments were sampled on the whole plant at 0, 6, and 12 h, with at least three replications for each set of experiments.
Reverse-transcription quantitative polymerase chain reaction
In order to study the expression characteristics of ZmUBP gene under different treatments, total RNA was extracted from the leaves of maize seedlings of control and treatment groups using OminiPlant RNA Kit from Beijing Kangwen Biotechnology Co. Subsequently, cDNA libraries were constructed using the Easy Script One-Step gDNA Removal and cDNA Synthesis Supermix from TransGen Biotech. Primers for the ZmUBP genes were designed using Primer 5.0 software. The cDNA from maize under different treatments was diluted 10-fold and used as a template, with the housekeeping gene actin serving as an internal reference. RT-qPCR was then performed to verify the gene expression characteristics under different treatments. The reaction system consisted of 1 µL cDNA, 1 µL Primer-FW, 1 µL Primer-RV, 10 µL 2× brilliant SYBR RT-qPCR master mix, and 7 µL ddH2O. The relative expression of the selected genes was calculated using the 2-ΔΔCT method. RT-qPCR analysis was performed with three biological replicates.
Statistical analysis
GraphPad Prism 9 was used to draw column and line charts, and SPSS 17.0 was used to analyze the significant differences (among the averages, there was no significant difference when there was a same marked letter, and there was significant difference when there were different marked letters, and the screening condition was P < 0.05). Pictures are mainly processed by Photoshop image processing software.
Results
Identification and characterization of the UBP genes in maize
To identify members of the UBP gene family in maize, we first searched relevant databases using the 27 Arabidopsis UBP protein sequences as queries, and this analysis identified 48 putative ZmUBP genes. The Pfam database (http://pfam.xfam.org) and NCBI CD Search (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi) were used to confirm the existence of the conserved domain UCH in these UBP proteins. After removing the unqualified sequences, a total of 45 ZmUBP genes were finally identified from the maize genome and named ZmUBP1 to ZmUBP45 according to their chromosomal locations (Table 1). All 45 ZmUBPs contained the UCH domain. Detailed information of ZmUBP genes were listed in Table 1, including gene length, physical and chemical parameters, and subcellular location of all ZmUBP genes. The length of ZmUBP genes conding sequence ranged from 1393 to 4462 bp, and the protein length of ZmUBPs was between 368-1284 amino acids. The molecular weight (MW) ranged from 41.81 to 187.82 kDa with the predicted isoelectric point (pI) ranging from 4.72 to 9.35. The transmembrane (TM) domains prediction of ZmUBP protein showed that only ZmUBP26 and ZmUBP22 contained one transmembrane domains. Subcellular localization and protein stability prediction showed that all 45 ZmUBP proteins were located in the nucleus and were unstable proteins. The hydrophobicity of 45 ZmUBP proteins was predicted to be less than 0, all ZmUBP proteins were hydrophilic proteins (Table 1).
To better elucidate the association between gene function and evolution, we explored the structural organization and conserved motifs of ZmUBP genes (Fig. 1). The largest number of exons was 30, and they were detected in ZmUBP25 from G5 subfamily, while the smallest number of exons was 2 detected in ZmUBP12 from G1 subfamily. The exon number in other ZmUBP genes was between 3-26. The ZmUBP genes from the same subfamily shared comparable gene structure, thereby suggesting functional conservation among the maize UBP gene family. Additionally, the exon variation in ZmUBP genes may indicate the functional diversity of the maize UBP gene family. The MEME program was used to predict the composition of the ZmUBP protein motifs. A total of twenty conserved motifs were detected (Fig. 1). With few exceptions, the motifs in most ZmUBP proteins are arranged in the same order as motif 1, motif 2, motif 3, motif 15, motif 2, motif 17, motif 5, and motif 13 (Fig. 1).
Gene and protein structures of 45 ZmUBPs. Phylogenetic relationships (a), protein motifs (b), protein structural domains (c) and gene structures (d) of 45 ZmUBPs. G1-G15 represent 45 ZmUBPs distributed in 15 different subfamilies, and the horizontal coordinates represent the length of genes/amino acid sequences
Phylogenetic analysis of the UBP genes in maize
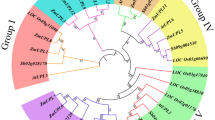
To analyse the phylogenetic organisation of the UBP family, we performed phylogenetic analysis using the protein sequences of 45 maize UBPs, 48 rice UBPs, 27 Arabidopsis UBPs, and 97 wheat UBPs, and generated a phylogenetic tree based on the Neighbour Joining (NJ) method (Fig. 2). Based on their phylogenetic relationships, we divided these UBPs into 15 groups, namely G1 to G15 (Fig. 2). G5 consisted of the most members, namely 14 ZmUBP proteins, followed by G7 including seven ZmUBP proteins. G4, G6, G9, G10, and G15 possessed the fewest members, including only one ZmUBP protein. Similarly, the UBPs of rice, wheat, and Arabidopsis were distributed among 15 groups, indicating that the functions of UBP family members were preserved during species evolution.
Phylogenetic relationships of UBPs from maize, Arabidopsis, wheat and rice. The phylogenetic tree was generated using a neighborhood join method with 1000 duplicates. Group 1-Group 15 on the left indicate that the UBP family members of maize, as well as the UBP family members of rice, wheat and Arabidopsis, are distributed in 15 different subfamilies. The branches of each subfamily are represented by a specific color. The branches of different members in the same subfamily have the same color
Chromosomal location and evolution of ZmUBP genes
To examine the chromosomal distribution of ZmUBP genes, the chromosomal location of each ZmUBP gene was determined. The results showed that the 45 ZmUBP genes were mapped to 10 chromosomes and they were unevenly distributed within each chromosome (Fig. 3). We identified 8 ZmUBP genes on chromosome 2. On chromosomes 1, 3, and 4, 6 ZmUBP genes were identified. On chromosomes 7, 5, 6, 8, 9 and 10, 5, 4, 3, 3, 2, and 2 ZmUBP genes were identified. These results indicate that there is genetic variation in the evolution of maize. Similarly, UBPs were unevenly distributed on different chromosomes in wheat [16], rice [14] and moso bamboo [15]. Obviously, the distribution of UBP genes on different chromosomes of different species is different.
Duplication is the major impetus underlying gene expansion during evolution. Five types of gene duplication may occur in evolution, including singleton, dispersed, tandem, proximal, and segmental duplication [31]. Based on the chromosomal distribution analysis, we conducted gene duplication analysis to reveal the expansion process of the maize UBP genes (Fig. 4). The duplicate gene pairs were identified by comparing the coding sequences of 45 ZmUBP genes, and a total of 10 segmental duplication events (ZmUBP5/ZmUBP16, ZmUBP6/ZmUBP37, ZmUBP9/ZmUBP29, ZmUBP12/ZmUBP21, ZmUBP20/ZmUBP41, ZmUBP24/ZmUBP30, ZmUBP26/ZmUBP28, ZmUBP29/ZmUBP45, ZmUBP33/ZmUBP40, and ZmUBP13/ZmUBP38) were found in the maize genome (Fig. 4). These results indicate that segmental duplication plays an important role in UBP gene expansion in the maize genome. These results are consistent with the reported UBP members of monocotyledons such as wheat [16] and bamboo [15]. To investigate the evolutionary patterns among ZmUBP genes, we calculated the rate of synonymous substitutions between duplicate gene pairs. Our findings indicate that ZmUBP genes were generally under purifying selection, as evidenced by the Ka/Ks ratios of all duplicate gene pairs being less than 1 (Table 2).
Duplication of 45 ZmUBP genes during evolution. Zm1-Zm10 in the innermost circle represents 10 chromosomes of maize, 0-150 represents the distance of genes on chromosomes, the yellow column represents the density of gene distribution on maize chromosomes, and the outermost circle is the distribution of 45 ZmUBP genes on chromosomes. The gray and red lines represent the gene duplication generated during the evolution of all genes in the maize genome and 45 ZmUBP genes, respectively
To elucidate the evolution of the UBP gene families in maize, comparative syntenic maps were constructed with six dicotyledons, Arabidopsis, sorghum (Sorghum bicolor), soybean (Glycine max), Kinnow Mandarin, cotton (Gossypium spp.) and medicago, and two monocotyledons, rice and wheat (Fig. 5). The results of the comparison show that 20, 23, and 25 ZmUBP genes were covalently related to the UBP genes in sorghum, rice and wheat, respectively. This was followed by Kinnow mandarin, soybean, Arabidopsis and cotton, with three ZmUBP genes covalently related to their UBP genes, and only 2 ZmUBP genes were covalently related to the UBP genes of medicago. The ZmUBP genes had the largest number of covariate gene pairs with monocotyledons and much more than dicotyledons. These findings suggest that the UBP gene predates the evolution of dicotyledons and that genetic variation occurred during the transition from dicotyledons to monocotyledons.
Collinearity analysis of UBP genes among maize, Arabidopsis, sorghum, soybean, Kinnow Mandarin, cotton, medicago, rice and wheat. The gray lines indicate the gene duplication that occurred during evolution between all genes in the maize genome and all genes in the genomes of eight different species. The red lines indicate gene duplication between the maize UBP genes and the UBP genes of eight different species that arose during evolution. The green columns represent the chromosomes of different species
To investigate the evolutionary patterns among UBP genes, we calculated the rate of synonymous substitutions between the UBP family in monocotyledon rice/wheat and maize (Fig. 6). The results showed that the Ka/Ks values of 27 homologous genes pairs between the maize UBP family and rice UBP family, as well as 56 homologous genes pairs between the wheat UBP family and maize UBP family were all less than 1 (Fig. 6), indicating that UBP genes were generally under purifying selection during the evolution of monocotyledons. This is consistent with the close evolutionary relationship between monocotyledons, suggesting that the UBP gene may play an important role in the evolutionary process of species.
Ka/Ks ratios of homologous gene pairs in maize and rice (left) and wheat (right). The horizontal coordinates indicate the comparison of UBP genes between different species, and the vertical coordinates indicate Ka/Ks ratio ranging from 0-0.8. Red boxes and dots indicate homologous gene pairs between maize and rice, and blue boxes and triangles indicate homologous gene pairs between maize and wheat
Analysis of ZmUBP genes related to stress response and plant hormone response
KEGG enrichment analysis showed that ZmUBP was mainly involved in ubiquitin-mediated protein hydrolysis and secondary metabolite synthesis processes (Fig. 7a). GO enrichment results showed that ZmUBP was mainly involved in biological processes such as protein deubiquitination, regulation of protein stability, autophagosome organization, jasmonic acid signaling pathway and immune response (Fig. 7b). This suggests that ZmUBP is involved in plant growth and stress response by regulating protein deubiquitination.
KEGG (a) and GO (b) enrichment analysis of the ZmUBP genes. The horizontal coordinates in a indicate the ratio of the number of genes in the pathway to the number of ZmUBP family members, and the vertical coordinates indicate the function of the ZmUBP family members. The colour of the circle indicates the significance of the proportion of genes in the pathway to the total genes (Q-value < 0.05 is significant). The size of the black circle indicates the number of ZmUBP genes in the pathway. The horizontal coordinates in ss b indicate the ratio of the number of genes in the pathway to the number of members of the ZmUBP family,and the vertical coordinates indicate the functions of ZmUBP family members, including cellular components, molecular functions and biological processes. The color of the circle indicates the significance of the proportion of genes in the pathway to the total genes (P < 0.05 is considered significant). After taking -log10(Pvalue), the higher the value is, the higher the significance. The size of the black circle indicates the number of ZmUBP genes in the pathway
The upstream regions of genes contain binding sites for promoters and transcription factors that regulate gene expression [32]. Therefore, the analysis of cis-acting elements contributes to the understanding of gene function and regulatory networks. To identify potential cy-acting elements in the promoter of ZmUBP genes, 2000 bp sequences upstream from the translation initiation codon of ZmUBP genes were retrieved and analyzed in the PlantCARE database. The promoters of the ZmUBP genes contain many elements or sites that respond to environmental factors such as light, temperature, and humidity. The presence of defense and stress response elements suggests that ZmUBPs may play an important role in the plant response to drought and low-temperature stress (Fig. 8). In addition, hormone response elements were found in the ZmUBP gene promoter, including abscisic acid (ABA), salicylic acid (SA), gibberellin (GA), auxin (IAA), and methyl jasmonate (MeJA) response elements. These results suggest that ZmUBP genes may be involved in the plant response to stress by regulating hormone signaling.
Cis-acting element analysis of 45 ZmUBP gene promoters. The 45 ZmUBP genes were classified according to phylogenetic relationships, and light, drought, gibberellin, jasmonic acid, abscisic acid, salicylic acid, auxin, stress, low-temperature, and seed growth response elements are indicated by 10 different colored columns. The horizontal coordinates represent gene length
Expression patterns of ZmUBP genes in different tissues
The expression profiles of 45 genes in different tissues were divided into four groups (Fig. 9a). The first category includes ZmUBP42. Except for the low expression in mature pollen, the expression level of ZmUBP42 in other maize parts was the highest among all ZmUBPs. This expression is similar to AtUBP23 (AT5G57990), which is in the same subfamily as ZmUBP42 in the evolutionary relationship (Fig. 2). AtUBP23 is highly expressed in all tissues of Arabidopsis [33]. The second category includes ZmUBP39, ZmUBP15, ZmUBP3, ZmUBP2, ZmUBP35, ZmUBP25, ZmUBP14, ZmUBP1 and ZmUBP20. Although the expression level of these genes is low in some plant tissues, the overall expression level is high (Fig. 9a). In particular, ZmUBP39, whose high-level expression during embryonic development is similar to AtUBP14 (AT3G20630), which is in the same subfamily as ZmUBP42 in the evolutionary relationship, further validates the conjecture in Fig. 2 that ZmUBP39 may play an important role in early embryonic development. In addition, the expression level of ZmUBP1 in reproductive organs was significantly higher than that in other parts, which was similar to AtUBP3 (AT4G39910) and AtUBP4 (AT2G22310) in Arabidopsis, which were in the same subfamily as ZmUBP1 in the evolutionary relationship. This is the same as the speculation in Fig. 2 that ZmUBP1 plays an important role in plant sexual reproduction. The third category includes ZmUBP30, ZmUBP37, ZmUBP7, ZmUBP4, ZmUBP21, ZmUBP43 and ZmUBP8. These genes have low expression levels in all tissues of plants and may not be involved in plant growth and development (Fig. 9a). The expression levels of other genes in different plant tissues are quite different, ZmUBP41 and ZmUBP6 were highly expressed in mature pollen, ZmUBP19 and ZmUBP26 were highly expressed near the primordium, indicating that they may be involved in plant sexual reproduction. In addition, the expression levels of ZmUBP genes in maize roots and embryos were generally higher than those in other plant parts, indicating that ZmUBP-mediated deubiquitination may be crucial for nutrient uptake and sexual reproduction.
The temporal and spatial expression characteristics of ZmUBP genes. a Expression levels of 45 ZmUBP genes in 17 different tissues of maize. The horizontal coordinates represent different periods and different parts of the maize fetch, and the vertical coordinates indicate the clustering analysis of all ZmUBP genes according to the expression levels at different periods. The red and blue columns indicate the values after z-score normalization of the expression of ZmUBP genes, with red being expression above the mean and considered high expression, while blue being expression below the mean and considered low expression. Groups represent different parts of the plant, including root, stem, leaf, flower and embryo. b 8 ZmUBP genes that may play a role in plant growth and stress responses were selected based on the results of transcriptome analysis. Different colored columns represent different ZmUBP genes, and the horizontal coordinates represent different plant tissues, in which R stands for roots, S for stems, and L for leaves. The vertical coordinates represent the relative expression levels of the 8 ZmUBP genes in different plant tissues (using the expression in roots as control). Significant differences are indicated by the letters a, b, and c labeled at the top of the columns
Up to now, the only study on maize UBP genes is the identification of three Arabidopsis UBP gene homologs in maize, namely, ZmUBP15, ZmUBP16, and ZmUBP19, which only analyzed their expression patterns in roots, leaves, spikes, and seeds, as well as in the presence of metal, salt, and osmotic stresses [25], but there is no in-depth study of the roles that all members of the maize UBP family play in plant growth and in response to high-temperature, low-temperature, and drought stresses. Therefore, we selected eight ZmUBP genes with high expression levels under abiotic stresses using RT-qPCR to investigate their roles in plant growth and abiotic stress response. The results showed that except for ZmUBP17, which was highly expressed in stems and leaves, the other 7 ZmUBP genes were highly expressed in roots (Fig. 9b), and the expression levels in roots were significantly higher than those in stems and leaves (except for ZmUBP14). This is consistent with the transcriptome data in Fig. 9a (the expression levels of ZmUBP genes was significantly higher in roots than in stems and leaves), suggesting that ZmUBP genes were mainly involved in plant root growth.
Expression pattern of ZmUBP genes under abiotic stresses and phytohormone treatment
The expression levels of all ZmUBP genes were analyzed under abiotic stress (high-temperature, drought and low-temperature) based on transcriptomic data (Fig. 10a). The expression levels of all ZmUBP genes except ZmUBP35 and ZmUBP17 decreased significantly when maize taproots were subjected to drought stress (Fig. 10a), which is consistent with the results of promoter element analysis (Fig. 8), that is, most ZmUBP gene promoters contain drought response and ABA response elements, indicating that ZmUBP gene family members may be negative regulators of drought stress. The expression of ZmUBP37, ZmUBP42, ZmUBP7, ZmUBP29, ZmUBP31 and ZmUBP30 increased significantly after high-temperature stress was applied to different genotypes of maize (Fig. 10a). The expression of other ZmUBP genes was significantly downregulated. The expression multiples of ZmUBP37 was most upregulated, which indicated that ZmUBP37 was important for maize to cope with high-temperature stress. ZmUBP16, ZmUBP6, ZmUBP33, ZmUBP11 and ZmUBP17 were significantly induced in different varieties of maize after low-temperature stress. However, ZmUBP24 was only upregulated in the resistant genotype, so ZmUBP24 could be used as an important target for low-temperature resistance breeding of maize. In addition, the promoters of ZmUBP22, ZmUBP1, ZmUBP7, ZmUBP10, ZmUBP34, ZmUBP27, ZmUBP32, ZmUBP35 and ZmUBP15 also contained low-temperature response elements, but their expression levels were significantly reduced under low-temperature stress, indicating that these genes may be negative regulators of low-temperature stress. In addition, no ZmUBP gene was resistant to all three abiotic stresses.
Expression analysis of ZmUBP genes in different tissues by RT-qPCR. a Expression levels of 45 ZmUBP genes under drought, low-temperature and high-temperature treatments. The horizontal coordinates represent the different treatments and treatment times of maize materials, and the vertical coordinates indicate the clustering analysis of all ZmUBP genes according to the expression levels of different treatments. The red and blue columns indicate the multiples of upregulated or downregulated expression of the treatment group compared with the untreated control group (red is upregulated expression, blue is downregulated expression). Group is different treatment methods of maize materials. b-g 8 ZmUBP genes that may play a role in plant growth and stress responses were selected based on the results of transcriptome analysis. Different colored columns represent different ZmUBP genes, and the horizontal coordinates represent abiotic stresses and phytohormones treatments for 0, 6 and 12 hours. The vertical coordinates represent the relative expression levels of the 8 ZmUBP genes under abiotic stresses and phytohormones treatments (using the expression under 0 hour as control). Significant differences are indicated by the letters a, b, and c labeled at the top of the columns
RT-qPCR results showed that ZmUBP14 was significantly induced after low-temperature, high-temperature and drought stresses, and the expression level showed a trend of decreasing and then increasing after IAA and SA treatments (Fig. 10b-d, f and g), suggesting that ZmUBP14 may respond to abiotic stresses by cross-linking multiple phytohormone signaling pathways. In addition, the expression level of ZmUBP35 was significantly increased after high-temperature, low-temperature and ABA treatments (Fig. 10b, c and e), suggesting that ZmUBP35 may be an ABA-mediated temperature regulator. ZmUBP37 was significantly induced by all three phytohormone treatments but did not respond to abiotic stresses (Fig. 10b-g), suggesting that ZmUBP37 may regulate plant growth and development through a cross-linked phytohormone signaling pathway. Interestingly, ZmUBP17, which was highly expressed in roots, stems and leaves, did not respond to to either abiotic stress or phytohormones (Figs. 9 and 10), suggesting that ZmUBP17 may affect plant growth and development through the regulation of other phytohormones. It is well known that ABA is a key phytohormone that regulates drought tolerance in plants, but no ZmUBP genes were found in this study that existed in response to both ABA and drought, suggesting that ZmUBP genes may regulate drought tolerance in plants through other pathways.
Discussion
Identification of UBP gene family members in maize
The eukaryote-specific UBP family is one of the largest DUB families identified to date and plays an important role in plant growth and development [34]. The plant UBP family has different types and amounts of UBP genes. There are 27 UBP members in Arabidopsis [13]. There are 48 UBP members in moso bamboo [15], and they are divided into two major groups (G13-G15, G1). There are 97 UBP members in wheat distributed in 15 subfamilies [16], and 48 UBP members in rice are also distributed in 15 subfamilies [14]. However, the UBP gene family members have not been identified in the maize genome. In this study, 45 putative UBP genes were identified in maize using genome-wide analysis, and were unevenly distributed on 10 chromosomes. According to phylogenetic analysis, the presumed ZmUBP gene family members were divided into 15 subfamilies as in wheat and rice, indicating that no UBP members were deleted during maize evolution. In addition, different members of the same subfamily are generally considered to have similar functions. AtUBP3 (AT4G39910) and AtUBP4 (AT2G22310) in Arabidopsis regulate male gamete development and affect plant sexual reproduction [1]. Therefore, ZmUBP29, ZmUBP1, ZmUBP9 and ZmUBP45, which are in group 2, may also have the same function as AtUBP3 and AtUBP4. Similarly, ZmUBP39 in group 6 may be able to regulate plant early embryonic development like AtUBP14 (AT3G20630). AtUBP12 (AT5G06600) and AtUBP13 (AT3G11910) have been shown to enhance plant drought tolerance through activation of the ABA signalling pathway and to play a role in plant histone debuquitination and organ development [35]. Therefore, ZmUBP15, ZmUBP44, ZmUBP25, ZmUBP35, ZmUBP14, ZmUBP19, ZmUBP32, ZmUBP41, ZmUBP18, ZmUBP17, ZmUBP5, ZmUBP4, ZmUBP23 and ZmUBP8 were also speculated to enhance plant drought resistance and to play a role in plant histone debuquitination and organ development because they also belong to the group 5. A recent study showing that The UBP5 histone H2A deubiquitinase counteracts PRCs-mediated repression to regulate Arabidopsis development informs the study of the histone deubiquitination function of the homologue ZmUBP5 in maize [36]. In addition, OsUBP2 (Os09g0505100) in rice and TaUBP1A.1 in wheat were proved to be resistant to leaf blight and Chinese wheat mosaic virus (CWMV), and both belonged to group 1 [6, 16], indicating that ZmUBP12 and ZmUBP21 belonging to group 1 subfamily may be resistant to biotic stress. In addition, UBP genes of wheat and rice were evolutionarily closer to maize UBP genes than Arabidopsis UBP genes (Fig. 2), as were interspecies covariance analyses (Fig. 5), confirming previously reported relationships between dicotyledons and monocotyledons during evolution [37]. These results indicated that ZmUBP genes existed before the isolation of monocotyledon plants and were highly conserved during plant evolution.
Duplication events of ZmUBP genes during evolution
Gene duplication helps organisms adapt to environmental changes during development and growth and is essential for gene evolution and amplification [38, 39]. Among them, the tandem duplication of genes in the process of genomic DNA duplication and recombination is the key driver of gene family amplification [40]. In the genomes of Arabidopsis and rice, 15–20% of genes consist of tandem repeats of gene clusters thought to be critical for evolution, plant disease resistance, and abiotic stress responses [41]. Two tandem repeat clusters were found in the TaUBP gene family of wheat. The two tandem repeat genes were located on chromosomes 1D and 7D, accounting for only 3.7% of the 54 TaUBP collinear gene pairs, suggesting that tandem repetition may not be the main amplification method during the evolution of the UBP gene family.In the present study, chromosomal localization and gene structure revealed that gene duplication events occurred during genome expansion and evolution in maize. Forty-five members of the ZmUBP gene family produced 10 pairs of segmental duplicated genes without tandem duplication, with the same results as in wheat. It is further suggested that segmental duplication may be the primary method of gene amplification during the evolution of the UBP gene family, rather than tandem duplication. In addition, the Ka/Ks ratio can be used to determine whether selective pressure acts on protein-coding genes. In this study, the Ka/Ks ratio was significantly less than 1 in the intraspecific and interspecific repeat gene pairs of all maize UBP genes, indicating that strong purifying selection plays an important role in the constraint of UBP gene function, which is consistent with the conservation of the UBP gene in the evolutionary process.
ZmUBP genes play an important role in plant growth and stress response
Transcriptional regulation of genes may be influenced by cis-elements in promoter regions that control responses to different stimuli. To investigate the biological function of ZmUBPs, we predicted cis-acting elements in the ZmUBP gene promoter. The results showed that the type of cis-acting element was different for each ZmUBP gene. Therefore, ZmUBPs may be involved in various specific regulatory mechanisms related to the stress response. Cis-acting regulatory elements largely determine tissue-specific gene expression patterns. In this study, we explored the expression profile of ZmUBPs in different tissues (root, stem, leaf, flower and embryo). The expression pattern of ZmUBPs showed that the expression levels of ZmUBPs were different in different plant tissues. The expression levels of ZmUBPs were the highest in maize roots and embryos and the lowest in leaves. This indicates that ZmUBP-mediated deubiquitination may be essential for the development of maize roots and embryos. In addition, some TaUBPs showed tissue-specific expression in maize, such as ZmUBP42, which was not expressed in mature pollen and was expressed at very high levels in all other sites, while ZmUBP1 was only expressed at high levels in the root cortex and was expressed at low levels in other sites. These results indicate that ZmUBPs play different roles in plant growth and development. Cis-acting regulatory elements also largely determine the expression patterns of stress response genes. We found drought, low-temperature, GA, MeJA, ABA, and SA response elements in the promoter region of the ZmUBP genes, suggesting that ZmUBPs play an important role in the hormone-mediated stress response. This was confirmed by transcriptomic results. Except for ZmUBP35 and ZmUBP17, the expression levels of all ZmUBP genes were significantly downregulated after drought treatment, indicating that ZmUBPs responded negatively to drought stress. The expression of ZmUBPs was different under low and high-temperature stress but showed a downward trend, indicating that ZmUBPs negatively regulated the plant stress response to abiotic stress.
Up to now, few studies have been conducted on the function of maize UBP genes. Kong et al. identified three Arabidopsis UBP gene homologs in maize and analyzed their expression patterns in roots, leaves, spikes, and seeds, as well as under metal, salt, and osmotic stresses [25]. However, the functions of ZmUBP genes in phytohormone, high-temperature, low-temperature, and drought responses have not been thoroughly investigated. In present study, we presented a comprehensive investigation of ZmUBP genes expression profiles in different plant tissues and abiotic stresses based on RT-qPCR and transcriptome data. Among them, ZmUBP17 showed high expression levels in roots, stems and leaves, implying that it plays an important role in the growth and development of different parts of plants. Except for ZmUBP17, all other ZmUBP genes were expressed at high levels in roots and lower levels in stems and leaves. The process of deubiquitination has been shown to regulate nutrient uptake and transport in plant roots to maintain optimal root function, so we hypothesised that these seven ZmUBP genes also have the function of regulating nutrient uptake and transport in plant roots [42, 43]. In addition, we found that ZmUBP14 was significantly induced by high-temperature, low-temperature, and drought treatments, and there was a positive response to IAA and SA, suggesting that ZmUBP14 may regulate plant tolerance to abiotic stresses through an ABA-independent pathway. Interestingly, ZmUBP17 showed a negative response to both abiotic stresess and phytohormones despite its high expression level in roots, stems and leaves, suggesting that ZmUBP14 may regulate plant growth and development through other phytohormones besides ABA, IAA, and SA. ABA is a key phytohormone for regulating drought tolerance in plants, and ABA accumulation in the tissues can significantly enhance plant drought tolerance. However, this study did not find any ZmUBP genes that were responsive to both ABA and drought stress, suggesting that ZmUBP genes may regulate plant drought tolerance through an ABA-independent pathway. In addition, ZmUBP37 was responsive to all three phytohormones, ABA, IAA, and SA, and was highly expressed in roots, suggesting that ZmUBP37 may regulate the growth and development of plant roots by cross-linking multiple phytohormone signaling pathways. Combining the transcriptome and RT-qPCR results, we hypothesized that ZmUBP14 may be a core member of the ZmUBP gene family that regulates plant growth and development and stress response.
Conclusion
As the largest subfamily of plant DUBs, UBP is essential for deubiquitination and plays an important role in plant development and stress response. Here, we have conducted a genome-wide analysis of the UBP gene family in maize for the first time. A total of 45 ZmUBP genes were comprehensively identified from the maize genome, and all the maize UBP genes were randomly distributed on the 10 chromosomes of maize and produced 10 duplicate gene pairs in the evolutionary process. Phylogenetic analysis revealed that these maize UBP genes were divided into 15 subfamilies. The protein motifs and gene structures of the ZmUBPs were highly conserved in each group, reflecting their functional conservation. Collinearity analysis showed that a high proportion of the ZmUBP genes might be derived from tandem duplications with purifying selection, providing insights into possible functional divergence among members of the ZmUBP gene family. Furthermore, a large number of stress and hormone response elements are raised on the ZmUBP promoters. ZmUBP14 was highly expressed in roots, stems, and leaves, and there was a positive response to drought, low-temperature, high-temperature, IAA, and SA, which has certain guiding significance for studying the mechanism of ZmUBP genes in response to abiotic stresses and hormones during development mediated by deubiquitination.
Availability of data and materials
Hybrid Zhengdan 958 provided by Grain Crops Research Institute, Henan Academy of Agricultural Sciences (Validation No.: Guoshiyu 20000009, Date of Validation: 2000, Selection and Breeding Unit: Grain Crops Research Institute, Henan Academy of Agricultural Sciences, Selected Breeder: Chunxin Du, Variety Source: Zheng 58/Chang 7-2). The genomic data analyzed in this study are available in the NCBI repository (the accession number of the maize genomic data is CABHLF000000000, the accession number of rice genomic data is JACJVL000000000, the accession number of sorghum genomic data is ABXC00000000, the accession number of soybean genomic data is ACUP00000000, the accession number of Kinnow Mandarin genomic data is NIHA00000000, the accession number of cotton genomic data is VKGJ00000000, the accession number of medicago genomic data is PSQE00000000, the accession number of wheat genomic data is NMPL00000000, The accession number of Arabidopsis genomic data is JAEFBJ000000000). All transcriptome data used in this study are available in the EMBL-EBI database (https://www.ebi.ac.uk/). Website links for all data used in this study are as follows.
Wibsite title | URL |
|---|---|
Ensembl plants | |
Pfam | Pfam: Home page (xfam.org) |
NCBI CD search | NCBI Conserved Domain Search (nih.gov) |
ExPASy | SIB Swiss Institute of Bioinformatics | Expasy |
TMHMM | TMHMM 2.0 - DTU Health Tech - Bioinformatic Services |
Plant-Ploc | Plant-PLoc server (sjtu.edu.cn) |
SWISS MODEL | SWISS-MODEL (expasy.org) |
MEME online tool | MEME - MEME Suite (meme-suite.org) |
TAIR | TAIR - Browse - Gene Families (arabidopsis.org) |
Phytozome | Phytozome (doe.gov) |
iTOL | iTOL: Interactive Tree Of Life (embl.de) |
PlantCARE | https://bioinformatics.psb.ugent.be/webtools/plantcare/html/ |
Microlife Letter | bioinformatics.com.cn |
Tbtools | Releases CJ-Chen/TBtools (github.com) |
STRING | STRING: functional protein association networks (string-db.org) |
EMBL-EBI |
Abbreviations
- maize:
-
Zea mays L.
- UBPs:
-
Ubiquitin-specific proteases
- DUBs:
-
Deubiquitinating enzymes
- ABA:
-
Abscisic acid
- UCH:
-
Ub carboxy-terminal hydrolase
- Cys:
-
Cysteine cysteine
- His:
-
Histidine histidine
- Arabidopsis :
-
Arabidopsis thaliana
- rice:
-
Oryza sativa
- Moso Bamboo:
-
Phyllostachys edulis
- wheat:
-
Triticum aestivum
- CAN:
-
Canavaline
- CWMV:
-
Chinese wheat mosaic virus
- sorghum:
-
Sorghum bicolor
- soybean:
-
Glycine max
- Kinnow Mandarin:
-
Citrus reticulata Blanco
- cotton:
-
Gossypium spp.
- Medicago:
-
Medicago sativa
- Ka:
-
Nonsynonymous substitution rates
- Ks:
-
Synonymous substitution rates
- SA:
-
Salicylic acid
- GA:
-
Gibberellin
- IAA:
-
Auxin
- MeJA:
-
Methyl jasmonate
References
Doelling JH, Phillips AR, Soyler-Ogretim G, Wise J, Chandler J, Callis J, Otegui MS, Vierstra RD. The ubiquitin-specific protease subfamily UBP3/UBP4 is essential for pollen development and transmission in arabidopsis. Plant Physiol. 2007;145(3):801–13.
Luo M, Luo MZ, Buzas D, Finnegan J, Helliwell C, Dennis ES, Peacock WJ, Chaudhury A. Ubiquitin-specific protease 26 is required for seed development and the repression of PHERES1 in arabidopsis. Genetics. 2008;180(1):229–36.
Li WF, Perry PJ, Prafulla NN, Schmidt W. Ubiquitin-specific protease 14 (UBP14) is involved in Root responses to phosphate Deficiency in Arabidopsis. Mol Plant. 2010;3(1):212–23.
Park SH, Jeong JS, Seo JS, Park BS, Chua NH. Arabidopsis ubiquitin-specific proteases UBP12 and UBP13 shape ORE1 levels during leaf senescence induced by nitrogen deficiency. New Phytol. 2019;223(3):1447–60.
Cui X, Lu FL, Li Y, Xue YM, Kang YY, Zhang SB, Qiu Q, Cui XK, Zheng SZ, Liu B, et al. Ubiquitin-specific proteases UBP12 and UBP13 act in circadian clock and photoperiodic flowering regulation in Arabidopsis. Plant Physiol. 2013;162(2):897–906.
Jiang RR, Zhou SC, Da XW, Chen T, Xu JM, Yan P, Mo XR. Ubiquitin-specific protease 2 (OsUBP2) negatively regulates cell death and disease resistance in rice. Plants-Basel. 2022;11(19):2568.
Lim CW, Baek W, Lim J, Hong E, Lee SC. Pepper ubiquitin-specific protease, CaUBP12, positively modulates dehydration resistance by enhancing CaSnRK2.6 stability. Plant J. 2021;107(4):1148–65.
Zhou HP, Zhao JF, Yang YQ, Chen CX, Liu YF, Jin XH, Chen LM, Li XY, Deng XW, Schumaker KS, et al. Ubiquitin-specific protease16 modulates salt tolerance in Arabidopsis by regulating Na+/H+ antiport activity and serine hydroxymethyltransferase Stability. Plant Cell. 2012;24(12):5106–22.
Komander D, Clague MJ, Urbe S. Breaking the chains: structure and function of the deubiquitinases. Nat Rev Mol Cell Biol. 2009;10(8):550–63.
Reyes-Turcu FE, Ventii KH, Wilkinson KD. Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu Rev Biochem. 2009;78:363–97.
Nijman SMB, Luna-Vargas MPA, Velds A, Brummelkamp TR, Dirac AMG, Sixma TK, Bernards R. A genomic and functional inventory of deubiquitinating enzymes. Cell. 2005;123(5):773–86.
Ingvardsen C, Veierskov B. Ubiquitin- and proteasome-dependent proteolysis in plants. Physiol Plant. 2001;112(4):451–9.
Yan N, Doelling JH, Falbel TG, Durski AM, Vierstra RD. The ubiquitin-specific protease family from arabidopsis. AtUBP1 and 2 are required for the resistance to the amino acid analog canavanine. Plant Physiol. 2000;124(4):1828–43.
Wang DH, Song W, Wei SW, Zheng YF, Chen ZS, Han JD, Zhang HT, Luo JC, Qin YM, Xu ZH, et al. Characterization of the ubiquitin C-Terminal hydrolase and ubiquitin-specific protease families in Rice (Oryza sativa). Front Plant Sci. 2018;9:413136.
Wu RH, Shi YR, Zhang Q, Zheng WQ, Chen SL, Du L, Lu CF. Genome-wide identification and characterization of the UBP Gene Family in Moso Bamboo (Phyllostachys edulis). Int J Mol Sci. 2019;20(17):4309.
Xu MZ, Jin P, Liu TT, Gao SQ, Zhang TY, Zhang F, Han XL, He L, Chen JP, Yang J. Genome-wide identification and characterization of UBP gene family in wheat (Triticum aestivum L). PeerJ. 2021;9:e11594.
Doelling JH, Yan N, Kurepa J, Walker J, Vierstra RD. The ubiquitin-specific protease UBP14 is essential for early embryo development in Arabidopsis thaliana. Plant J. 2001;27(5):393–405.
Sridhar VV, Kapoor A, Zhang KL, Zhu JJ, Zhou T, Hasegawa PM, Bressan RA, Zhu JK. Control of DNA methylation and heterochromatic silencing by histone H2B deubiquitination. Nature. 2007;447(7145):735-U718.
Du L, Li N, Chen LL, Xu YX, Li Y, Zhang YY, Li CY, Li YH. The ubiquitin receptor DA1 regulates seed and organ size by modulating the Stability of the ubiquitin-specific protease UBP15/SOD2 in Arabidopsis. Plant Cell. 2014;26(2):665–77.
Shi CL, Ren YL, Liu LL, Wang F, Zhang H, Tian P, Pan T, Wang YF, Jing RN, Liu TZ, et al. Ubiquitin specific protease 15 has an important role in regulating grain width and size in Rice. Plant Physiol. 2019;180(1):381–91.
An ZC, Liu YL, Ou Y, Li J, Zhang BW, Sun DY, Sun Y, Tang WQ. Regulation of the stability of RGF1 receptor by the ubiquitin-specific proteases UBP12/UBP13 is critical for root meristem maintenance. PNAS. 2018;115(5):1123–8.
Zhou Y, Park SH, Chua NH. UBP12/UBP13-mediated deubiquitination of salicylic acid receptor NPR3 suppresses plant immunity. Mol Plant. 2023;16(1):232–44.
Liu GC, Liang JX, Lou LJ, Tian MM, Zhang XY, Liu LJ, Zhao QZ, Xia R, Wu YR, Xie Q, et al. The deubiquitinases UBP12 and UBP13 integrate with the E3 ubiquitin ligase XBAT35.2 to modulate VPS23A stability in ABA signaling. Sci Adv. 2022;8(14):eabl5765.
Zhao JF, Zhou HP, Zhang M, Gao YA, Li L, Gao Y, Li M, Yang YH, Guo Y, Li XY. Ubiquitin-specific protease 24 negatively regulates abscisic acid signalling in Arabidopsis thaliana. Plant Cell Environ. 2016;39(2):427–40.
Islam F, Imran A, Afzaal M, Saeed F, Asghar A, Shahid S, Shams A, Zahra SM, Biswas S, Aslam MA. Nutritional, functional, and ethno-medical properties of sweet corn cob: a concurrent review. Int J Food Sci Technol. 2023;58(5):2181–8.
Trinidad-Calderon PA, Acosta-Cruz E, Rivero-Masante MN, Diaz-Gomez JL, Garcia-Lara S, Lopez-Castillo LM. Maize bioactive peptides: from structure to human health. J Cereal Sci. 2021;100:103232.
Kong JJ, Jin J, Dong Q, Qiu JL, Lii YY, Yang YH, Shi YT, Si WN, Gu LJ, Yang FY, et al. Maize factors ZmUBP15, ZmUBP16 and ZmUBP19 play important roles for heck plants to tolerance the cadmium stress and salt stress. Plant Sci. 2019;280:77–89.
Finn RD, Clements J, Arndt W, Miller BL, Wheeler TJ, Schreiber F, Bateman A, Eddy SR. HMMER web server: 2015 update. Nucleic Acids Res. 2015;43(W1):W30-38.
Chen CJ, Wu Y, Li JW, Wang X, Zeng ZH, Xu J, Liu YL, Feng JT, Chen H, He YH, et al. TBtools-II: a one for all, all for onebioinformatics platform for biological big-data mining. Mol Plant. 2023;16(11):1733–42.
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R. TBtools: an integrative toolkit developed for interactive analyses of big Biological Data. Mol Plant. 2020;13(8):1194–202.
Zheng L, Wu HH, Qanmber G, Ali F, Wang LL, Liu Z, et al. Genome-wide study of the GATL gene family in Gossypium hirsutum L. reveals that GHGATL genes act on pectin synthesis to regulate plant growth and fiber elongation. Genes. 2020;11(1):64.
Liu WS, Stewart CN. Plant synthetic promoters and transcription factors. Curr Opin Biotechnol. 2016;37:36–44.
Liu YF, Wang F, Zhang HY, He H, Ma LG, Deng XW. Functional characterization of Arabidopsis ubiquitin-specific protease gene family reveals specific role and redundancy of individual members in development. Plant J. 2008;55(5):844–56.
March E, Farrona S. Plant deubiquitinases and their role in the control of Gene expression through modification of histones. Front Plant Sci. 2018;8:2274.
Ewan R, Pangestuti R, Thornber S, Craig A, Carr C, O’Donnell L, Zhang C, Sadanandom A. Deubiquitinating enzymes AtUBP12 and AtUBP13 and their tobacco homologue NtUBP12 are negative regulators of plant immunity. New Phytol. 2011;191(1):92–106.
Godwin J, Govindasamy M, Nedounsejian K, March E, Halton R, Bourbousse C, Wolff L, Fort A, Krzyszton M, Corrales JL, et al. The UBP5 histone H2A deubiquitinase counteracts PRCs-mediated repression to regulate Arabidopsis development. Nat Commun. 2024;15(1):667.
Mei C, Liu YW, Dong X, Song QN, Wang HJ, Shi HW, Feng RY. Genome-wide identification and characterization of the potato IQD family during development and stress. Front Genet. 2021;12:693936.
Bowers JE, Chapman BA, Rong JK, Paterson AH. Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature. 2003;422(6930):433–8.
Gu ZL, Steinmetz LM, Gu X, Scharfe C, Davis RW, Li WH. Role of duplicate genes in genetic robustness against null mutations. Nature. 2003;421(6918):63–6.
Moore RC, Purugganan MD. The evolutionary dynamics of plant duplicate genes. Curr Opin Plant Biol. 2005;8(2):122–8.
Flagel LE, Wendel JF. Gene duplication and evolutionary novelty in plants. New Phytol. 2009;183(3):557–64.
Kohli A, Narciso JO, Miro B, Raorane M. Root proteases: reinforced links between nitrogen uptake and mobilization and drought tolerance. Physiol Plant. 2012;145(1):165–79.
Zhou HP, Zhao JF, Cai JQ, Patil SB. Ubiquitin-specific proteases function in plant development and stress responses. Plant Mol Biol. 2017;94(6):565–76.
Acknowledgements
We thank the editors and reviewers for their careful reading and valuable comments. We apologize to researchers whose studies are not cited due to space limitations.
Funding
This work was supported by the Program for Changjiang Scholars and Innovative Research Team in University (grant number: No. IRT17R99) and by Heilongjiang Province Government Postdoctoral Science Foundation (grant number: LBH-Q18008).
Author information
Authors and Affiliations
Contributions
W.C.F. and Y.Y.B. designed the experiments, analyzed the data and wrote the manuscript. W.C.F. and D.L.F. performed the experiments and bioinformatics analysis. S.K.L. supervised the study and critically reviewed the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Fu, W., Fan, D., Liu, S. et al. Genome-wide identification and expression analysis of Ubiquitin-specific protease gene family in maize (Zea mays L.). BMC Plant Biol 24, 404 (2024). https://doi.org/10.1186/s12870-024-04953-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-024-04953-5