Abstract
Background
In paddy fields, the noxious weed barnyard grass secretes 2,4-dihydroxy-7-methoxy-2H-1,4-benzoxazin-3(4H)-one (DIMBOA) to interfere with rice growth. Rice is unable to synthesize DIMBOA. Rice cultivars with high or low levels of allelopathy may respond differently to DIMBOA.
Results
In this study, we found that low concentrations of DIMBOA (≤ 0.06 mM) promoted seedling growth in allelopathic rice PI312777, while DIMBOA (≤ 0.08 mM) had no significant influence on the nonallelopathic rice Lemont. DIMBOA treatment caused changes in the expression of a large number of glutathione S-transferase (GST) proteins, which resulting in enrichment of the glutathione metabolic pathway. This pathway facilitates plant detoxification of heterologous substances. The basal levels of GST activity in Lemont were significantly higher than those in PI312777, while GST activity in PI312777 was slightly induced by increasing DIMBOA concentrations. Overexpression of GST genes (Os09g0367700 and Os01g0949800) in these two cultivars enhanced rice resistance to DIMBOA.
Conclusions
Taken together, our results indicated that different rice accessions with different levels of allelopathy have variable tolerance to DIMBOA. Lemont had higher GST activity, which helped it tolerate DIMBOA, while PI312777 had lower GST activity that was more inducible. The enhancement of GST expression facilitates rice tolerance to DIMBOA toxins from barnyard grass root exudates.
Similar content being viewed by others
Background
Weeds are a serious biotic stressor that reduces rice yields, and barnyard grass is a noxious weed in rice fields that causes a significant loss of yield. Studies have documented that barnyard grass invasion leads to losses ranging from 1.5 to 55.2% of rice yield annually [1], which has led to an increase in herbicide application, resulting in enhanced herbicide resistance in weeds and potential herbicide residues remaining in the environment [2].
In the natural system of rice and weed, there exist rice germplasm materials that can suppress weed growth through allelopathy, which arises from their high capacity for secondary metabolite synthesis and secretion. These compounds include phenolic acids, diterpenoids, and flavonoids, which are recognized as allelochemicals, and these rice accessions are referred to as allelopathic rice [3]. The presence of weeds activates higher transcriptional levels of phenylalanine ammonia-lyase (OsPAL), cinnamate-4-hydroxylase (OsC4H), cinnamyl-alcohol dehydrogenase (OsCAD), momilactone A synthase (OsMAS), and ent-kaurene synthase-like 4 (OsKSL4), which enhances the biosynthesis of allelochemicals. Sufficient concentrations of these compounds from root exudates are then released into the rhizosphere, resulting in allelopathic inhibition of the surrounding weeds [4,5,6,7,8].
Fang et al. (2015) indicated that phenolic acid allelochemicals promoted the proliferation of Myxococcus sp. in the rhizosphere, and the combination of ferulic acid and Myxococcus xanthus led to strong growth inhibition of barnyard grass, which was due to the increase in apurinic/apyrimidinic (AP) sites and the decrease in IAA contents in barnyard grass roots. The increase in DNA damage and reduction in hormone contents suppressed the growth of barnyard grass [9]. Studies have also focused on the allelopathic potential of diterpenoids (momilactone A and momilactone B) and flavonoids against this weed and their impact on soil microorganisms [5, 7]. These characteristics of allelopathic rice cultivars enable their biointerference with weed growth and are considered to be natural survival strategies of rice. In contrast, the weed (barnyard grass) has also evolved unique characteristics to restrict rice growth. Genome sequencing and assembly along with gene annotation of barnyard grass (Echinochloa crus-galli L.) showed that a gene cluster for DIMBOA synthesis was found in the genome of E. crus-galli, but the gene cluster was absent from the rice genome. The presence of the DIMBOA gene cluster in E. crus-galli enables the biosynthesis of these compounds in barnyard grass, and a bioassay of the effects of DIMBOA on rice showed that 0.08 mM DIMBOA significantly suppresses rice growth [10].
DIMBOA, 2,4-dihydroxy-7-methoxy-2H-1,4-benzoxazin-3(4H)-one, is a compound that widely exists in gramineous crops and is a secondary metabolite with antibacterial activity and pest resistance [11]. In wheat and maize, DIMBOA is a vital allelochemical [12], and the concentrations of DIMBOA in wheat can be induced by co-cultured weeds, i.e., Abutilon theophrasti Medicus, Alopecurus Xaponicas Steud, and Avena fatua L.; thus, DIMBOA is regarded as an important and dominant allelochemical against weeds of wheat [13]. The absence of the DIMBOA synthesis gene cluster in rice has resulted in the loss of DIMBOA synthesis; in contrast, the DIMBOA from coexisting barnyard grass is a particularly toxic compound that suppresses rice growth and acts as a biointerference molecule.
Pathways involved in the detoxification of xenobiotics in plants can be classified into three phases. These phases require the involvement of Cytochrome P450s (CYP450s), Glutathione S-transferases (GSTs), and ATP-binding cassette (ABC) transporters, also known as the multixenobiotic resistance (MXR) system, in phases I, II, and III respectively [14]. CYP450s catalyze a range of oxidation and reduction reactions to degrade toxic xenobiotics, transforming them into more polar and water-soluble compounds [15, 16]. Meanwhile, GST enzymes enhance the conjugation of xenobiotics with glutathione (GSH), facilitating their further detoxification within plant cells. Through this conjugation process, polar groups are added to the xenobiotics, aiding in their detoxification [15]. In addition to these steps, an essential aspect of detoxification is the excretion of xenobiotics and/or their metabolites by the MXR system. This system helps transport these compounds out of the plant, further contributing to the detoxification process [14].
Since different allelopathic rice accessions have different levels of allelopathy against the weed, whether there are different levels of tolerance or resistance to DIMBOA toxicity remains unknown. This study compared the different levels of DIMBOA tolerance between the allelopathic rice accession PI312777 and the nonallelopathic rice accession Lemont. PI312777 was developed in the Philippines from indica parents (T65*2/TN 1), both of which originated from Taiwan, China [17, 18]. This particular variety has shown strong suppression against barnyard grass; Lemont, on the other hand, was developed through a cross made in 1974 between ‘Lebonnet’ and the F1 resulting from crossing CI 9881 with PI 331,581 [19]. The proteomic expression profile of these two rice accessions was elucidated, and the function of key proteins and their main pathway was illuminated.
Results
Effects of DIMBOA on the growth of PI312777 and Lemont
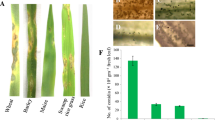
The root and stem lengths of PI312777 and Lemont treated with 0.02 mM, 0.04 mM, 0.06 mM, 0.08 mM and 0.10 mM DIMBOA and those of their control groups without DIMBOA were determined. The results showed that PI312777 and Lemont have different levels of tolerance to DIMBOA toxicity. The growth of roots and stems was promoted in PI312777 under 0.02 mM, 0.04 mM and 0.06 mM DIMBOA treatment and peaked at 0.04 mM DIMBOA treatment with a significant promoting effect, whereas 0.08 mM and 0.10 mM DIMBOA significantly inhibited the root and stem length of PI312777. In contrast, DIMBOA at concentrations up to 0.08 mM did not significantly affect the growth of roots and stems of Lemont, while 0.10 mM DIMBOA significantly inhibited root growth but did not significantly affect stem growth in Lemont (Fig. 1). The different levels of tolerance of the allelopathic rice PI312777 and nonallelopathic rice Lemont to DIMBOA toxicity suggest different strategies of high and low allelopathic potential in response to the biointerference of barnyard grass.
Growth effect of different DIMBOA concentrations on the allelopathic rice accession PI312777 and nonallelopathic rice Lemont. Germinated seeds of PI312777 (A, C) and Lemont (B, D) were soaked in 0.02 mM, 0.04 mM, 0.06 mM, 0.08 mM, and 0.10 mM DIMBOA, while germinated seeds cultured in sterilized ddH2O were used as control groups. After 4 days, rice root and shoot lengths from the control and treatment groups were measured. Error bars indicate standard deviation. Significant differences (p < 0.05) in rice roots and shoots between different treatments are indicated by different lowercase letters according to analysis of variance (ANOVA). Lowercase letters with bold correspond to the root length comparison, and those letters with italics correspond to the shoot length comparison. Bar, 3 cm
Differentially expressed proteins (DEPs) in PI312777 and Lemont
Seedlings of PI312777 and Lemont were treated with 0.04 mM and 0.06 mM DIMBOA, respectively (Fig. S1), and iTRAQ quantitative proteomics was conducted to explore differentially expressed proteins (DEPs) between the DIMBOA-treated rice and the control group without DIMBOA treatment. The results showed that 231 proteins were upregulated and 135 proteins were downregulated in the roots of PI312777 treated with 0.04 mM DIMBOA, while 211 proteins were upregulated and 450 proteins were downregulated in the roots of PI312777 treated with 0.06 mM DIMBOA compared to the control group for PI312777. In Lemont, 0.04 mM DIMBOA treatment resulted in 236 proteins being upregulated and 144 proteins being downregulated in the root, and 0.06 mM DIMBOA treatment resulted in 268 proteins being upregulated and 175 proteins being downregulated in the root when compared with the control group for Lemont (Fig. S2). Prediction of the protein‒protein interactions of the DEPs showed that most of the DEPs from the treatment group and control group closely interacted, especially the DEPs from PI312777 treated with 0.06 mM DIMBOA (Fig. S3). Among these DEPs, the expression of 41 and 40 proteins was significantly upregulated (fold change ≥ 1.5, the same below) in PI312777 under 0.04 mM and 0.06 mM DIMBOA treatment, in comparison with control group respectively, while the expression of 17 and 52 proteins was significantly downregulated. In Lemont, the expression of 38 and 53 proteins was significantly upregulated in the 0.04 mM and 0.06 mM DIMBOA-treated groups, respectively (Fig. 2).
Volcano plot demonstrating the differentially expressed proteins (DEPs) from PI312777 and Lemont under DIMBOA treatment. The differentially expressed proteins between the treatment and control groups from PI312777 and Lemont were selected to draw the volcano map by using fold change and p values. The pairwise comparisons were as follows: 0.04 mM DIMBOA-treated PI312777 compared to its control group without DIMBOA addition (A), 0.06 mM DIMBOA-treated PI312777 compared to its control group (B), 0.04 mM DIMBOA-treated Lemont compared to its control group (C), and 0.06 mM DIMBOA-treated Lemont compared to its control group (D). The red dot represents significantly upregulated proteins, the blue dot represents significantly downregulated proteins (fold change ≥ 1.5), and the grey dot represents the proteins without a significant fold change
Pathway enrichment of DEPs
DEPs from each individual treatment were annotated with relevant pathways to explore KEGG pathway enrichment. The results showed that the DEPs from 0.04 mM DIMBOA-treated PI312777 (PI-TR1) were mainly enriched in glutathione metabolism, plant‒pathogen interaction, sesquiterpenoid and triterpenoid biosynthesis, and photosynthesis (Fig. 3A), while the DEPs from 0.06 mM DIMBOA-treated PI312777 (PI-TR2) were mainly enriched in ribosomes, glutathione metabolism, oxidative phosphorylation, plant‒pathogen interaction, protein export, biological clock, SNARE interactions in vesicular transport and plant circadian rhythm (Fig. 3B). When comparing PI-TR2 and PI-TR1, most DEPs were enriched in glutathione metabolism, oxidative phosphorylation, diterpenoid biosynthesis, SNARE interactions in vesicular transport and nitrogen metabolism. Taken together, glutathione metabolism and the relevant DEPs participating in this pathway were involved in the regulation of PI312777 tolerance to DIMBOA (Fig. 3C).
KEGG enrichment of DEPs in PI312777 and Lemont under DIMBOA treatment. The DEPs between the treatment and control groups for PI312777 and Lemont were annotated with KEGG pathways to determine pathway enrichment. The pairwise comparisons were as follows: 0.04 mM DIMBOA-treated PI312777 compared to its control group without DIMBOA addition (A), 0.06 mM DIMBOA-treated PI312777 compared to its control group (B), 0.06 mM DIMBOA-treated PI312777 compared to 0.04 mM DIMBOA-treated PI312777 (C), 0.04 mM DIMBOA-treated Lemont compared to its control group (D–E), 0.06 mM DIMBOA-treated Lemont compared to its control group, and 0.06 mM DIMBOA-treated Lemont compared to 0.04 mM DIMBOA-treated Lemont (F). The arrows point to the uniformly enriched pathway (glutathione metabolism) among all pairwise comparisons
The DEPs in the Lemont treated with 0.04 mM DIMBOA (LE-TR1) were mainly enriched in glutathione metabolism, oxidative phosphorylation, nitrogen metabolism, monoterpenoid synthesis, carbon fixation in photosynthetic organisms, GPI-anchor biosynthesis, and SNARE interactions in vesicular transport (Fig. 3D). Those in Lemont treated with 0.06 mM DIMBOA (LE-TR2) were significantly enriched in glutathione metabolism, plant hormone signal transduction, anthocyanin synthesis, RNA polymerase, glucosinolate biosynthesis, plant‒pathogen interaction, and carbon fixation in photosynthetic organisms. (Fig. 3E). The DEPs between LE-TR2 and LE-TR1 were also mainly enriched in glutathione metabolism (Fig. 3F). The results of iTRAQ quantitative proteomics for PI312777 and Lemont indicated that glutathione metabolism is indispensable in rice tolerance to DIMBOA.
Changes in glutathione metabolic pathway-related protein expression levels
The expression levels of DEPs involved in glutathione metabolism were analysed. The results showed that the expression levels of several glutathione S-transferases (GSTs) involved in this pathway were changed in PI312777 and Lemont treated with DIMBOA. The expression patterns of PI312777 and Lemont were also different. In PI312777, treatment with 0.04 mM and 0.06 mM DIMBOA resulted in the upregulation of 8 proteins, LOC_Os03g44170.1 (Os03g0643700), LOC_Os03g57200.1 (Os03g0785900), LOC_Os09g20220.1 (Os09g0367700), LOC_Os10g38189.1 (Os10g0525800), LOC_Os01g27210.1 (Os01g0369700), LOC_Os05g03820.1 (Os05g0129000), LOC_Os10g38590.1 (Os05g0129000), and LOC_Os01g72150.1 (Os01g0949900). The relative protein expression increased with increasing DIMBOA concentration (0.04 mM and 0.06 mM) (Fig. 4A). In Lemont, seventeen proteins, LOC_Os10g38340.1 (Os10g0527400), LOC_Os01g72150.1 (Os01g0949900), LOC_Os01g72140.1 (Os01g0949800), LOC_Os09g20220.1 (Os09g0367700), LOC_Os03g57200.1 (Os03g0785900), LOC_Os03g17480.1 (Os03g0283200), LOC_Os01g72170.1 (Os01g0950300), LOC_Os01g49720.1 (Os01g0692100), LOC_Os10g38600.1 (Os10g0529500), LOC_Os10g38590.1 (Os10g0529400), LOC_Os07g07320.1 (Os07g0168300), LOC_Os10g38189.1 (Os10g0525800), LOC_Os01g27210.1 (Os01g0369700), LOC_Os10g38740.1 (Os10g0530900), LOC_Os10g38360.1 (Os10g0527800), LOC_Os10g38160.1, and LOC_Os10g22070.1 (Os10g0365200), were all upregulated in the 0.04 mM DIMBOA and 0.06 mM DIMBOA treatment groups, and the relative protein expression increased with increasing concentration of exogenous DIMBOA (Fig. 4B). These results suggested that glutathione metabolism plays a dominant role in the regulation of rice tolerance.
Expression changes in GSTs participating in glutathione metabolism in the roots of PI312777 (A) and Lemont (B) under DIMBOA treatment. The differentially expressed GSTs in PI312777 and Lemont that participate in glutathione metabolism were selected for a comparison of their fold changes in expression and heatmap analysis. The data in each column represent the fold change in independent proteins based on pairwise comparisons of PI312777 (A) and Lemont (B), including 0.04 mM DIMBOA-treated PI312777 compared to its control group without DIMBOA addition (PI-TR1 vs. PI-CT), 0.06 mM DIMBOA-treated PI312777 compared to its control group (PI-TR2 vs. PI-CT), 0.06 mM DIMBOA-treated PI312777 compared to 0.04 mM DIMBOA-treated PI312777 (PI-TR2 vs. PI-TR1), 0.04 mM DIMBOA-treated Lemont compared to its control group (LE-TR1 vs. LE-CT), 0.06 mM DIMBOA-treated Lemont compared to its control group (LE-TR2 vs. LE-CT), and 0.06 mM DIMBOA-treated Lemont compared to 0.04 mM DIMBOA-treated Lemont (LE-TR2 vs. LE-TR1)
Changes in GST gene expression in PI312777 and lemont roots under DIMBOA treatment
The transcriptional levels of Os01g0949800 and Os09g0367700, two genes encoding glutathione S-transferase (GST), in PI312777 and Lemont roots treated with DIMBOA at different concentrations were detected. The results showed that compared with the control group, the Os01g0949800 gene in PI312777 roots was upregulated 22-fold under 0.02 mM DIMBOA treatment. When the exogenous concentration of DIMBOA was increased to 0.04 mM, 0.06 mM and 0.08 mM, the Os01g0949800 gene was downregulated in the treated group compared with the control group. The expression of the Os09g0367700 gene was upregulated in PI312777 treated with 0.02–0.10 mM DIMBOA, and compared with the control group, this gene was upregulated 544-fold in the 0.02 mM DIMBOA-treated group (Fig. 5A).
In Lemont, the expression of Os01g0949800 was upregulated 1.7-fold under 0.02 mM DIMBOA treatment, whereas it was downregulated when the DIMBOA concentration was increased to 0.04 mM or higher. Os09g0367700 gene expression was upregulated 3.02- and 3.47-fold under 0.02 mM and 0.04 mM DIMBOA treatment, respectively. Gene expression was downregulated in Lemont when the DIMBOA concentration was increased to 0.06 mM or higher (Fig. 5B).
When comparing the expression changes in Os01g0949800 and Os09g0367700 in PI312777 and Lemont under DIMBOA treatment, it was found that GST gene expression in PI312777 was more strongly induced by low concentrations of DIMBOA.
GST activity in the roots of PI312777 and lemont
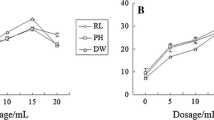
The GST activity in the roots of PI312777 was enhanced with increasing DIMBOA concentrations. The GST activity of PI312777 peaked at 1.88 U/g fresh weight under the 0.08 mM DIMBOA treatment. When the concentration of DIMBOA reached 0.10 mM, the GST activity was 1.02 U/g fresh weight, which was still significantly higher than that of the control group. In comparison with that of PI312777, the GST activity of Lemont rice roots under normal conditions (1.55 U/g fresh weight) was significantly higher than that of PI312777 roots under the same conditions (0.41 U/g fresh weight). Under 0.06 mM DIMBOA treatment, the GST activity of Lemont roots was significantly increased to 2.03 U/g fresh weight, while that under the other treatments showed no significant change compared with the control group (Fig. 6). A comparison of the GST activities in PI312777 and Lemont rice roots showed that the nonallelopathic rice Lemont had higher GST activity, which helped relieve the physiological toxicity of DIMBOA. The GST activity of the allelopathic rice PI312777 was lower than that of Lemont, but the GST activity of PI312777 was induced by DIMBOA treatment.
GST activity in the roots of PI312777 and Lemont under DIMBOA treatment. GST activities in the roots of PI312777 and Lemont treated with 0.02 mM-0.10 mM were determined and compared to the control groups. Error bars indicate standard deviation. Significant differences (p < 0.05) among different treatments are indicated by different lowercase letters according to analysis of variance (ANOVA). Lowercase letters with bold correspond to the PI312777 treatment comparison, and those letters with italics correspond to the Lemont treatment comparison
GST overexpression in transgenic rice affects tolerance to DIMBOA
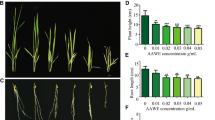
To further validate the function of GST in the regulation of rice tolerance to DIMBOA, we overexpressed Os01g0949800 and Os09g0367700 in PI312777 and Os01g0949800 in Lemont (Fig. S4). Evaluation of the GST-OX rice tolerance to DIMBOA showed that increased Os01g0949800 and Os09g0367700 expression in PI312777 facilitated tolerance, and the root and shoot lengths of Os01g0949800-OX transgenic PI312777 were significantly longer than those of WT PI312777 when these two rice lines were treated with 0.05 mM DIMBOA (Fig. 7A). Overexpression of Os01g0949800 in Lemont also resulted in enhanced rice tolerance to DIMBOA toxicity, and significantly longer root and shoot lengths were observed in Os01g0949800-OX transgenic Lemont than in its WT. In addition, the root length of Os09g0367700-OX transgenic PI312777 was also significantly longer than that of its WT (Fig. 7B).
Growth effect of DIMBOA on the GST-OX and wild type lines. Germinated seeds of PI312777 (A) and Lemont (B) and their GST-OX lines were soaked in 0.05 mM DIMBOA. After 10 days, rice root and shoot lengths from the GST-OX and wild-type groups were measured. Error bars indicate standard deviation. Significant differences between the GST-OX line and its wild type are indicated by double asterisks (p < 0.01) according to Student’s t test
Protein regulatory networks of GST
Proteins that interacted with GST were separated and identified (Fig. 8), and the results showed that the interacting proteins of GST (Os09g0367700) included two glutathione transferases, as well as thioredoxin-dependent peroxiredoxin, glutamate-1-semialdehyde 2,1-aminomutase, quinol cytochrome c reductase, protein-methionine-S-oxide reductase, serine/threonine-protein kinase SNT7, lipoxygenase, (S)-2-hydroxy-acid oxidase, cysteine desulfurase, and chloroplast-associated proteins (Table 1, Dataset S2). These proteins were enriched in metabolic pathways, secondary metabolite synthesis, carbon metabolism, amino acid synthesis, and glutathione metabolism (Fig. S5A).
GST-interacting proteins from GST-OX transgenic rice. Proteins from the leaves of Os01g0949800-OX PI312777 (Lane 1), Os09g0367700-OX PI312777 (Lane 2) and Os01g0949800-OX Lemont (Lane 3) were extracted and incubated with GFP-Trap agarose beads to precipitate the proteins interacting with GST (Os01g0949800 and Os09g0367700). These rice lines were placed into two groups, without (A) and with DIMBOA treatment (B). The bands with a MW of approximately 55 kDa in the Input lanes represent GST protein fused with eYFP. Other bands in other lanes are the proteins interacting with GST
GST (Os01g0949800) also interacted with several glutathione transferases, together with geranylgeranyl reductase, thioredoxin-dependent peroxiredoxin, S-(hydroxymethyl) glutathione dehydrogenase, serine hydroxymethyltransferase, peroxidase, S-adenosylmethionine synthase, peroxisomal membrane protein 11 − 1, sulfate adenylyltransferase, protein disulfide-isomerase, cytochrome b-c1 complex subunit, cytochrome f, and other proteins that interacted with Os09g0367700 (Table 1, Dataset S3). These proteins were enriched in metabolic pathways, secondary metabolite synthesis, carbon metabolism, amino acid synthesis, glutathione metabolism, and oxidative phosphorylation pathways (Fig. S5B).
Transcript level of proteins interacting with GST
The transcript levels of GST-interacting protein-coding genes in PI312777 and Lemont, including L-ascorbate peroxidase 4 (Os08g0549100), phosphoglycerate kinase (Os05g0496200), fructose-6-phosphate-2-kinase (Os05g0164100), GST (Os10g0530500), peroxidase precursor (Os02g0240300), dehydrogenase (Os02g0815500), aminotransferase (Os07g0108300), and glutaredoxin-dependent peroxiredoxin (Peroxiredoxin-2E-2, Os02g0192700) were determined. Among these genes, the expression of Os02g0240300 and Os10g0530500 was upregulated in PI312777 when the rice was treated with 0.02 mM-0.10 mM DIMBOA. The fold changes of these genes were found to be the highest under 0.10 mM DIMBOA treatment, and Os10g0530500 presented a 10.8-fold upregulation in comparison with the control group.
For Lemont, Os10g0530500 and Os02g0815500 were mostly upregulated in these treatment groups, and Os05g0164100 was upregulated in all treatment groups (Fig. 9).
Gene transcript levels of GST-interacting proteins in the roots of PI312777 and Lemont under DIMBOA treatment. The transcript levels of GST (Os01g0949800, Os09g0367700)-interacting proteins in PI312777 and Lemont cultivars treated with concentrations ranging from 0.02 mM to 0.10 mM were determined and compared with the corresponding control groups
Discussion
Weeds are unavoidable limiting factors in crop production because they release inhibitory compounds that can disturb crop growth and act as competitors for resources with crops [10, 20]. Among them, barnyard grass (Echinochloa crus-galli) is one of the most noxious weeds in paddy fields, seriously affecting the yield and quality of rice. Currently, chemical herbicides are mainly used to control Echinochloa crus-galli [21]. Studies have shown that within a certain dose range, allelopathic rice exhibits an enhancement of allelopathic potential when subjected to an increased exogenous addition of barnyard grass extract. However, if the dose of weed extract is too high, it inhibits rice growth [22]. DIMBOA, a root exudate, has been reported as a severely toxic compound that interferes with rice growth. A gene cluster for DIMBOA biosynthesis has been identified in the genome of barnyard grass but is absent in the rice genome [10]. Thus, DIMBOA is specifically released from the roots of barnyard grass to suppress rice growth in paddy fields during rice-barnyard grass competition. Therefore, the tolerance and detoxification capacity of rice towards DIMBOA are vital for the survival of the crop but may vary across different rice accessions.
In this study, the allelopathic rice PI312777 and nonallelopathic rice Lemont showed different levels of tolerance to exogenous DIMBOA. Tolerance in PI312777 was inducible under an appropriate concentration of DIMBOA (≤ 0.06 mM in this study), while Lemont showed more consistent tolerance, with no significant effect observed on Lemont rice growth when the exogenous DIMBOA concentration was lower than 0.08 mM. The results suggest that nonallelopathic rice lacks strong “allelochemicals weapon” to suppress barnyard grass growth [4, 6, 23], nevertheless, this rice variety has evolved to be more tolerant to DIMBOA exudates from barnyard grass. This characteristic benefits the nonallelopathic rice in its survival within the natural rice and weed system.
According to the iTRAQ quantitative proteomics results in rice, GST proteins participating in glutathione metabolism were speculated to be relevant to regulating rice tolerance to DIMBOA. In barnyard grass, genome sequencing and gene annotation revealed that a large number of genes that function in detoxification are distributed in the barnyard grass genome, such as GST and cytochrome P450 monooxygenase (CYP450), and these genes likely play positive roles in the detoxification of rice allelochemicals [24]. GSTs are important defence enzymes that are mainly cytosolic proteins, and these enzymes usually play active roles in the form of dimers to detoxify xenobiotics and reduce oxidative damage and endogenous metabolism, which results in diverse catalytic and noncatalytic activities in metabolism [25]. In the plant response to environmental stress factors, GSTs function in protecting cells from damage by invading organisms, heavy metals, oxidative stress and pathogens [26,27,28]. As a multifunctional hypergene family, there are at least 79 genes encoding GST in rice [29], and plant GST genes are divided into fourteen classes, including tau (U), phi (F), lambda (L), dehydroascorbate reductase (DHAR), theta (T), and zeta (Z), depending on their nucleotide composition [30]. It was documented that phi GSTs in black grass (Alopecurus myosuroides) and annual rye grass (Lolium rigidum) can participate in the detoxification metabolism of herbicides [31].
Two GSTs showed expression changes in both PI312777 and Lemont treated with DIMBOA, and these differentially expressed GSTs resulted in significant enrichment in glutathione metabolism, indicating the tight correlation of this pathway with rice tolerance. Several of these enzymes showed increased fold changes with an increasing DIMBOA dosage, suggesting a positive reaction to the tolerance to DIMBOA. In particular, the transcript levels of Os01g0949800 and Os09g0367700 increased in rice under DIMBOA treatment, especially the expression of Os09g0367700 in DIMBOA-treated PI312777. The enhanced expression of GST leads to increased enzyme activity during detoxification. In PI312777, lower GST activity was found in the roots without DIMBOA treatment; however, GST activity was slightly induced by DIMBOA, resulting in a higher tolerant ability. In contrast to PI312777, the nonallelopathic rice Lemont has a higher GST activity in the roots under natural conditions, and the activity was more stable under low concentrations of DIMBOA, which enabled the rice to tolerate the toxicity of DIMBOA.
A total of 29 DEPs from PI312777 treated with 0.06 mM DIMBOA were closely related to each other, indicating the systemic response of PI312777 to DIMBOA treatment and the cross-talk of the defence network in rice. Here, we overexpressed Os01g0949800 and Os09g0367700 in PI312777 and Os01g0949800 in Lemont. The results showed that increasing the level of GST gene expression in rice enhanced its tolerance to DIMBOA. The root and shoot lengths of Os01g0949800-OX transgenic rice were both significantly longer than those of their corresponding WT when treated with 0.05 mM DIMBOA. The results indicated that the GST-OX transgenic rice lines had a higher tolerance to relieve the toxic effects of DIMBOA, which also validates the role of GST in the transformation or degradation of xenobiotics. Overexpression of the OsGSTL1 gene in rice enhanced tolerance to chlorosulfuron and glyphosate in rice seedlings [32].
During the process of GST activation, Os01g0949800 (GST) interacted with other GSTs, as well as L-ascorbate peroxidase, peroxisomal membrane protein 11 − 1, protein disulfide-isomerase, and many proteins involved in porphyrin and chlorophyll metabolism, oxidative phosphorylation, the pentose phosphate pathway, and the main pathways from plant photosynthesis and respiration. These results suggested that the detoxification process was associated with energy metabolism and substrate metabolite transformation. The increased transcriptional expression of these interacting proteins in PI312777 was consistent with the expression pattern of the target GST, and the results indicated the cooperation of these particular proteins.
In Arabidopsis, O-glucosylation of allelochemical benzoxazolin-2(3 H)-one (BOA) was identified as the predominant detoxification to the chemical [33]. Meanwhile, upregulation of UDP-glucosyltransferase would also facilitate rice detoxification of DIMBOA.
In this study, the different responses of the allelopathic rice PI312777 and nonallelopathic rice Lemont to DIMBOA were mainly dependent on the basal and inducible levels of GST activity. The sensitivity to the induction of GST activity in PI312777 enables the plant to effectively react to xenobiotics, which allows allelopathic rice to lessen the toxic effects on cellular metabolism and to maintain plant growth. However, in the nonallelopathic rice Lemont, a low level of induction but high basal GST activity helped the rice line tolerate the toxicity of DIMBOA, which prevented major reductions in plant growth. The different strategies of allelopathic rice and nonallelopathic rice in the tolerance of the barnyard grass exudate DIMBOA provide additional directions for breeding weed-tolerant rice.
Methods
Plant materials
The allelopathic rice accession PI312777 and nonallelopathic accession Lemont (Oryza sativa japonica) were used as the study materials [34, 35]; PI312777 seeds were obtained from the International Rice Research Institute, The Philippines. Lemont seeds were obtained from Texas A&M University Agricultural Research and Extension Center, Beaumont, Texas.
Evaluation of the DIMBOA tolerance of PI31277 and Lemont
The seeds of PI312777 and Lemont were disinfected with 2.5% sodium hypochlorite (NaClO) solution for 30 min, washed with sterile water to clear the NaClO residue on the seeds, and soaked overnight at 30 °C in a thermostatic incubator to promote germination. DIMBOA (Toronto Research Chemicals Inc., Canada) was dissolved in the methanol to prepare a 100 mM storage solution. This storage solution was then further diluted with sterile water to create different concentrations of a working solution. The treatment groups were prepared with a series of concentration gradients: 0.02 mM, 0.04 mM, 0.06 mM, 0.08 mM, and 0.10 mM DIMBOA. The experiment was conducted in six-well plates, with each well containing 10 ml of solution. In each well, 10 germinated seeds of either PI312777 or Lemont rice were soaked. The corresponding control groups for PI312777 and Lemont consisted of germinated seeds cultured in sterile water with 10 µL of methanol solvent. Both the treatment groups and the control groups were tested in triplicate. The plates were placed in an artificial climate chamber (MGC-300 H, Shanghai Yiheng Scientific Instrument Co., LTD) set at a temperature of 26 °C, humidity of 85%, light intensity of 12,000 lx, and a light/dark cycle of 14 h of light and 10 h of darkness. After 4 days of culture, the root length and plant height of the control group and each treatment group were measured.
Quantitative proteomics analysis of DIMBOA-treated rice seedlings
Seeds of PI312777 and Lemont were disinfected and germinated following the description above. The germinated seeds were sown in a net and floated in a 1 L rectangular container fully filled with a 1/2 concentration of optimized rice hydroponic nutrient solution [36, 37]. The containers were placed in the climate chamber at 28 °C for 14 h/22°C for 10 h for 10 days (when the rice seedlings grew to the 3-leaf stage). Seedlings of PI312777 and Lemont were cultivated in complete hydroponic nutrient solution containing 0.04 mM and 0.06 mM DIMBOA, respectively. Seedlings of both cultivars grown in the nutrient solution without DIMBOA were used as controls. Rice roots from the treatment and control groups were independently sampled after treatment for 7 days and kept in liquid nitrogen.
Rice root protein extraction
One gram of rice roots with 5% PVPP were ground into powder in liquid nitrogen and resuspended in Tris-saturated phenol (Beijing Solarbio Science & Technology Co., Ltd.) at a 5:1 ratio. The mixture was then vortexed at 4 °C for 15 min. After centrifugation at 25,000 g and 4 °C for 20 min, the supernatant was treated by adding 5 volumes of 0.1 M ammonium acetate/methanol with 10 mM DTT to precipitate proteins at -20 °C for 2 h, and then centrifugation at 25,000 g and 4 °C for 15 min. The precipitation step was repeated twice. After that, pre-cooled acetone with 10 mM DTT was added in the precipitation and fully mixed, and kept at at -20 °C for 2 h, the supernatant was discarded after centrifugation at 25,000 g and 4 °C for 20 min, and the precipitation step was repeated once. The proteins were air dried at 4 °C and resuspended in lysis buffer (8 M urea and 40 mM Tris-HCl containing 1 mM PMSF, 2 mM EDTA, 10 mM DTT and 1×Protease Inhibitor Cocktail, pH 8.5) again and ultrasonicated on ice for 5 min (sonication for 2 s/pause for 3 s) to improve protein dissolution. After centrifugation, DTT (10 mM final concentration, FC) was added to the supernatant, which was incubated at 56 °C for 1 h for reduction and then alkylated with 55 mM iodoacetamide (IAM) in the dark at room temperature for 45 min. Five volumes of acetone was used to precipitate proteins at -20 °C overnight. Lysis buffer was used to dissolve the proteins with the help of sonication on ice for 5 min (2 s/3 sec). Proteins were quantitated with a Bradford assay and separated and qualified by SDS‒PAGE for the Isobaric Tags for Relative and Absolute Quantitation (iTRAQ) proteomic analysis. The experiment was conducted by BGI Co., LTD (Shenzhen, China) [38, 39]. The protocol details are available in Supplementary Dataset 1.
Bioinformatics analysis of differentially expressed proteins and their related pathways
The raw MS/MS data were converted into MGF format by the corresponding tool, and the exported MGF files were searched against the database described above via the local Mascot server. In addition, quality control (QC) was performed to determine if a reanalysis step was needed. Automated software named Iquant was applied to the quantification results of the proteins [40]. All the proteins with a false discovery rate (FDR) less than 1% were subjected to downstream analysis, including Gene Ontology (GO), COG/KOG and pathway analyses. Proteins with an expression fold change between the treatment and control of more than 1.5-fold were regarded as differentially expressed proteins (DEPs). Furthermore, analysis of the GO enrichment and KEGG pathway enrichment of DEPs was performed to investigate their relevant pathways.
Determination of glutathione S-transferase (GST) gene expression
Total RNA from the roots of PI312777 and Lemont treated with 0.02 mM, 0.04 mM, 0.06 mM, 0.08 mM and 0.10 mM DIMBOA and their control groups was extracted by using the TRIzol method. TransScript OneStep gDNA Removal and cDNA Synthesis SuperMix (TransGen Biotech Co., Ltd., Beijing, China) were used to reverse transcribe the mRNA into cDNA. The transcript levels of two genes, Os01g0949800 and Os09g0367700, which both encode glutathione S-transferase (GST), were determined in the DIMBOA-treated rice and the control group according to quantitative PCR (qPCR) with gene-specific primers (Os01g0949800, F: 5 ‘-GGGAGTGGCTTACGAGTTCA-3’, R: 5’-TCGGTGGGTAGGATGGGAC-3’; Os09g0367700, F: 5’-CTCGTCATCCTCGAGTACATC-3’, R: 5’- ACAGCTTCTTGTC GACGTAG-3’). The β-actin gene (F: 5 ‘-CTTCATAGGAATGGAAGCTGCGGGTA-3’, 5 ‘-CGACCACCTTGATCTTCATGCTGCTA-3’) was used as an internal reference for qPCR detection. The reaction mix for gene amplification was prepared following the TransStart Green qPCR SuperMix Kit (TransGen Biotech Co., Ltd., Beijing, China). The PCR program was set as follows: initial denaturation at 94 ℃ for 30 s, denaturation at 94 ℃ for 5 s, annealing at 58 ℃ for 15 s, and extension at 72 ℃ for 10 s. The reaction from denaturation to extension was conducted for 42 cycles; thereafter, the temperature rose from 60 ℃ to 95 ℃ at a rate of 0.2 ℃/s. The reaction was conducted in a realplex4 Eppendorf. The threshold value (CT value) was automatically recorded by Eppendorf instrument software, and the relative gene expression level was calculated by the \(2^{-\Delta\Delta{\rm Ct}}\) method [41] and plotted in GraphPad Prism 5.
Determination of GST activity
The roots of P312777 and Lemont treated with 0.02 mM, 0.04 mM, 06 mM, 0.08 mM and 0.10 mM DIMBOA and their control groups were used to extract the crude enzyme. Both the treatment group and the control group were tested in triplicate, and 0.1 g root tissue of each replicate was used for enzyme extraction. The GST activity of PI312777 and Lemont roots treated with different DIMBOA concentrations was determined by using the Glutathione S-transferase (GST) Activity Assay Kit (Beijing Solarbio Science & Technology Co., Ltd.).
Overexpression of GST in rice
The CDS of OsGST (Os01g0949800 and Os09g0367700) was amplified and inserted into a modified pCambia2300 vector (with the 35S promoter and eYFP gene fused to the target gene) to construct the eYFP recombinant vector for rice transformation [14]. Agrobacterium tumefaciens (EHA105)-mediated transformation was performed on calli of PI312777 and Lemont [42, 43]. Resistant calli were screened by 80µg/ml geneticin G418, and putatively transformed tissue was then differentiated and dedifferentiated to generate T0 transgenic rice seedlings. Fragments containing both GST and eYFP were amplified for sequencing (Os01g0949800-Transdetect-F: 5’-AAGGAGACGAAGGAGAACCTGG-3’; Os01g0949800-Transdetect-R: 5’-GTGGCGGATCTTGAAGTTCACC-3’; Os09g0367700-Transdetect-F: 5’-AGAAGCTGTTCGACTGCCAGAC-3’; Os09g0367700-Transdetect-R: 5’-CTCCTTGAAGTCGATGCCCTTC-3’) to identify positive transgenic lines. The positive GST overexpression (GST-OX) T0 transgenic lines were then retained for harvesting the T1 transgenic seeds.
Evaluation of the DIMBOA tolerance of GST-OX lines
The positive GST-OX T1 transgenic PI312777 and Lemont rice seeds were disinfected and germinated following the steps described above. Uniformly germinated transgenic and wild-type (WT) rice seeds were soaked in sterilized water with exogenous concentrations of 0.05 mM DIMBOA as the treatment group. The germinated transgenic and WT seeds cultured in sterile water without DIMBOA were used as the control groups. The experiment was also conducted in six-well plates, and both the treatment group and the control group were tested in triplicate and placed in an artificial climate chamber at 26 °C and 85% humidity. After 7 days of culture, the root length and plant height of the control group and the treated rice were measured.
Identification of the proteins interacting with GST
Natural leaf proteins were extracted from a T1 generation of GST-overexpression transgenic rice by using Pi-IP buffer (50 mM Tris pH 7.5; 150 mM NaCl; 1 mM EDTA; 1% Triton X-100; 1 mM PMSF; 1× Complete cocktail, Roche; 10 µM MG132) and incubated with GFP-Trap agarose (Chromotek) to collect putative GST-interacting proteins. The protocol details are available in Supplementary Dataset 1. These proteins were identified by using a Q-Exactive Plus mass spectrometer (Thermo Fisher Scientific).
Quantitative PCR to determine gene expression levels
The transcript levels of GST-interacting proteins in PI312777 and Lemont under 0.02 mM, 0.04 mM, 0.06 mM, 0.08 mM and 0.10 mM DIMBOA treatment were determined by qPCR. The transcript levels of the promoter binding proteins were also determined to evaluate their regulatory capacity on GST gene expression. The protocol for qPCR determination was described above.
Statistical methods
SPSS 26.0 was utilized for the statistical analysis of the data. The data were presented as mean ± standard error (SE) values obtained from three replicates for each experiment or determination. The variance in the mean values of root length, shoot length, and GST activity among the different concentrations of DIMBOA treatment on rice was analyzed using one-way ANOVA, followed by Tukey’s honestly significant difference (HSD) test at a significance level of p < 0.05.
Data availability
All data generated during this study are included in this published article and its supplementary information files, and the raw data of iTRAQ quantitative proteomics generated in this study have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository [44, 45] with the dataset identifier PXD041754.
Abbreviations
- DEPs:
-
differentially expressed proteins
- DIMBOA:
-
2,4-dihydroxy-7-methoxy-2 H-1,4-benzoxazin-3(4 H)-one
- GST:
-
glutathione S-transferase
- iTRAQ:
-
Isobaric Tags for Relative and Absolute Quantitation
- qPCR:
-
quantitative PCR
References
Zhang Z, Gu T, Zhao B, Yang X, Peng Q, Li Y, Bai L. Effects of common Echinochloa varieties on grain yield and grain quality of rice. Field Crops Res. 2017;203:163–72.
Osterholt MJ, Webster EP, Blouin DC, McKnight BM. Overlay of residual herbicides in rice for improved weed management. Weed Technol. 2019;33:426–30.
Dilday RH, Yan WG, Moldenhauer KAK, Gravois KA. Allelopathic activity in rice for controlling major aquatic weeds. In: Olofsdotter M, editor. Allelopathy in rice. Los Banos: International Rice Research Institute; 1998. pp. 7–26.
Li JY, Zhang Q, Yang XY, Wu HM, Lin RL, He HB. A reappraisal of the content and the differences of phenolic acids between allelopathic and non-allelopathic rice accessions. Allelopathy J. 2017;40:35–46.
Kato-Noguchi H, Peters RJ. The role of momilactones in rice allelopathy. J Chem Ecol. 2013;39:175–85.
He H, Wang H, Fang C, Wu H, Guo X, Liu C, Lin Z, Lin W. Barnyard grass stress up regulates the biosynthesis of phenolic compounds in allelopathic rice. J Plant Physiol. 2012;169:1747–53.
Kong CH, Zhao H, Xu XH, Wang P, Gu Y. Activity and allelopathy of soil of flavone O-glycosides from rice. J Agric Food Chem. 2007;55:6007–12.
Rimando AM, Duke SO. Studies on rice allelochemicals. I. In: Smith CW, Dilday RH, editors. Rice: origin, history, technology and production. New York: Wiley; 2003. pp. 221–44.
Fang CX, Yang LK, Chen WX, Li LL, Zhang PL, Li YZ, He HB, Lin WX. Identification and comparative analysis of microRNAs in barnyardgrass (Echinochloa crus-galli) in response to rice allelopathy. Plant Cell Environ. 2015;38:1368–81.
Guo L, Qiu J, Ye C, Jin G, Mao L, Zhang H, Yang X, Peng Q, Wang Y, Jia L, Lin Z, Li G, Fu F, Liu C, Chen L, Shen E, Wang W, Chu Q, Wu D, Wu S, Xia C, Zhang Y, Zhou X, Wang L, Wu L, Song W, Wang Y, Shu Q, Aoki D, Yumoto E, Yokota T, Miyamoto K, Okada K, Kim D-S, Cai D, Zhang C, Lou Y, Qian Q, Yamaguchi H, Yamane H, Kong C-H, Timko MP, Bai L, Fan L. Echinochloa crus-galli genome analysis provides insight into its adaptation and invasiveness as a weed. Nat Commun. 2017;8:1031.
Cuevas L, Niemeyer HM, Pérez FJ. Reaction of DIMBOA, a resistance factor from cereals, with α-chymotrypsin. Phytochemistry. 1990;29:1429–32.
Wu H, Haig T, Pratley J, Lemerle D, An M. Allelochemicals in wheat (Triticum aestivum L.): production and exudation of 2, 4-dihydroxy-7-methoxy-1, 4-benzoxazin-3-one. J Chem Ecol. 2001;27:1691–700.
Zhang SZ, Li YH, Kong CH, Xu XH. Interference of allelopathic wheat with different weeds. Pest Manag Sci. 2016;72:172–8.
Bonnafé E, Sroda S, Budzinski H, Valière A, Pedelluc J, Marty P, Geret F. Responses of cytochrome P450, GST, and MXR in the mollusk Corbicula fluminea to the exposure to hospital wastewater effluents. Environ Sci Pollut Res. 2015;22:11033–46.
Nelson DR, Koymans L, Kamataki T, Stegeman JJ, Feyereisen R, Waxman DJ, Waterman MR, Gotoh O, Coon MJ, Estabrook RW, Gunsalus IC. Nebert D W. P450 superfamily: update on new sequences, gene mapping, accession numbers and nomenclature. Pharmacogenetics. 1996;6:1–42.
Rewitz KF, Styrishave B, Løbner-Olsen A. Andersen OMarine invertebrate cytochrome P450: emerging insights from vertebrate and insects analogies. Comp Biochem Physiol C Toxicol Pharmacol. 2006;143:363–81.
[GRIN] Germplasm Resources Information Network. Online Database, USDA, ARS, National Genetic Resources Program, National Germplasm Resources Laboratory, Beltsville, MD. www.ars-grin.gov. 2004.
Gealy DR, Estorninos LE Jr., Gbur EE, Chavez RSC. Interference interactions of two rice cultivars and their F3 cross with barnyardgrass (Echinochloa crus-galli) in a replacement series study. Weed Sci. 2005;53:323–30.
Bollich CN, Webb BD, Marchetti MA, Scott JE. Registration of ‘Lemont’ rice. Crop Sci. 1985;25:883–5.
He HB, Wang HB, Fang CX, Lin ZH, Yu ZM, Lin WX. Separation of allelopathy from resource competition using rice/barnyardgrass mixedcultures. PLoS ONE. 2012;7:e37201.
Gaines TA, Busi R, Küpper A. Can new herbicide discovery allow weed management to outpace resistance evolution? Pest Manag Sci. 2021;77:3036–41.
Zhang Q, Li L, Li J, Wang H, Fang C, Yang X, He H. Increasing rice allelopathy by induction of barnyard grass (Echinochloa crus-galli) root exudates. J Plant Growth Regul. 2018;37:745–54.
Cuevas L, Niemeyer HM, Pérez FJ, Aschehoug. Invasive plants versus their new and old neighbors: a mechanism for exotic invasion. Science. 2000;290:521–3.
Guo L, Qiu J, Li L-F, Lu B, Olsen K, Fan L. Genomic clues for crop–weed interactions and evolution. Trends Plant Sci. 2018;23:1102–15.
Frova C. The plant glutathione transferase gene family: genomic structure, functions, expression and evolution. Physiol Plant. 2003;119:469–79.
Banerjee S, Goswami R. GST profile expression study in some selected plants: in silico approach. Mol Cell Biochem. 2019;336:109–26.
Gullner G, Komives T, Király L, Schröder P. Glutathione S-transferase enzymes in plant-pathogen interactions. Front Plant Sci. 2018;9:1836.
Zhang W, Yin K, Li B, Chen L. A glutathione S-transferase from Proteus mirabilis involved in heavy metal resistance and its potential application in removal of Hg2+. J Hazard Mater. 2013;26:646–52.
Lallement P-A, Brouwer B, Keech O, Hecker A, Rouhier N. The still mysterious roles of cysteine-containing glutathione transferases in plants. Front Pharmacol. 2014;5:192.
Jain M, Ghanashyam C, Bhattacharjee A. Comprehensive expression analysis suggests overlapping and specific roles of rice glutathione S-transferase genes during development and stress responses. BMC Genomics. 2010;11:73.
Cummins I, Wortley DJ, Sabbadin F, He Z, Coxona CR, Strakera HE et al. Key role for a glutathione transferase inmultiple-herbicide resistance in grass weeds. P Natl Acad Sci USA. 2013;110:5812–7.
Hu T, Qv X, Xiao G, Huang X. Enhanced tolerance to herbicide of rice plants by over-expression of a glutathione S-transferase. Mol Breed. 2009;24:409–18.
Baerson SR, Sánchez-Moreiras A, Pedrol-Bonjoch N, Schulz M, Kagan IA, Agarwal AK, Reigosa MJ, Duke SO. Detoxification and transcriptome response in Arabidopsis seedlings exposed to the allelochemical benzoxazolin-2(3H)-one. J Biol Chem. 2005;280:21867–81.
Fang CX, Yang LK, Chen WX, Li LL, Zhang PL, Li YZ, He HB, Lin WX. MYB57 transcriptionally regulates MAPK11 to interact with PAL2;3 and modulate rice allelopathy. J Exp Bot. 2020;71:2127–41.
Zhang Q, Zheng XY, Lin SX, Gu CZ, Li L, Li JY, Fang CX, He HB. Transcriptome analysis reveals that barnyard grass exudates increase the allelopathic potential of allelopathic and non-allelopathic rice (Oryza sativa) accessions. Rice. 2019;12:30.
Yoshida S, Forno DA, Cock JH. Laboratory manual for physiological studies of rice, third edition. Manila: The International Rice Research Institute. 1976.
Fang CX, Wang QS, Yu Y, Li QM, Zhang HL, Wu XC, Chen T, Lin WX. Suppression and overexpression of Lsi1 induce differential gene expression in rice under ultraviolet radiation. Plant Growth Regul. 2011;65:1–10.
Ross PL, Huang YN, Marchese JN, Williamson B, Parker K, Hattan S, Khainovski N, Pillai S, Dey S, Daniels S, Purkayastha S, Juhasz P, Martin S, Bartlet-Jones M, He F, Jacobson A, Pappin DJ. Multiplexed protein quantitation in Saccharomyces cerevisiae using aminereactive isobaric tagging reagents. Mol Cell Proteom. 2004;3:1154–69.
Treumann A, Thiede B. Isobaric protein and peptide quantification: perspectives and issues. Expert Rev Proteomic. 2010;7:647–53.
Wen B, Zhou R, Feng Q, Wang Q, Wang J, Liu S. IQuant: an automated pipeline for quantitative proteomics based upon isobaric tags. Proteomics. 2014;14:2280–5.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2 – ∆∆CT method. Methods. 2001;25:402–8.
Chen QH, Chen TY, Lin YJ, Chen H. Agrobacterium-mediated genetic transformation of japonica rice. In Yuan M, Du H, Li XH, editors. Rice Protocol eBook. Beijing: Bio-protocol. 2018. P. 245–254.
Liu Y, Ling F, Lin YJ, Chen H. 2018. Agrobacterium-mediated transformation of indica rice. In Yuan M, Du H, Li XH, editors. Rice Protocol eBook. Beijing: Bio-protocol. 2018. P. 255–264.
Ma J, Chen T, Wu S, Yang C, Bai M, Shu K, Li K, Zhang G, Jin Z, He F, Hermjakob H, Zhu Y. iProX: an integrated proteome resource. Nucleic Acids Res. 2019;47:D1211–7.
Chen T, Ma J, Liu Y, Chen Z, Xiao N, Lu Y, Fu Y, Yang C, Li M, Wu S, Wang X, Li D, He F, Hermjakob H, Zhu Y. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 2022;50:D1522–7.
Acknowledgements
We thank Wuhan Biorun Biosciences Co., Ltd. for help with rice genetic transformation.
Funding
This work was supported by the National Natural Science Foundation of China (Grant No. 31871556, 32001109), the Natural Science Foundation of Fujian Province (Grant No. 2020J01547), the Open Project Program of Guangdong Provincial Key Laboratory of Eco-circular Agriculture, the Foundation of Institute of Modern Seed Industrial Engineering, the Fujian Provincial Undergraduate Innovation Training Program (Grant No. S202210389075), and the Foundation for the Science and Technology Innovation of Fujian Agriculture and Forestry University (Grant No. KFB23088). The funders had no role in study design, data collection and analysis, data interpretation, or in writing of the manuscript.
Author information
Authors and Affiliations
Contributions
CF, HZ and DM designed the experiments. DM, HZ and YL performed most of experiments and analyzed the data. XL, XY, KL, YJ, JL and HL assisted in experiments and discussed the results. CF and WL wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
All methods were performed in accordance with relevant institutional, national, and international guidelines and legislation.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1: Fig. S1.
Phenotype of PI312777 and Lemont seedlings after treatment with DIMBOA. Bar, 5cm
Supplementary Material 2: Fig. S2.
Amounts of differentially expressed proteins from the roots of PI312777 and Lemont after treatment with DIMBOA
Supplementary Material 3: Fig. S3.
Predicted protein‒protein interactions among differentially expressed proteins from DIMBOA-treated PI312777 and the control group. Red nodes represent upregulated proteins; blue nodes represent downregulated proteins
Supplementary Material 4: Fig. S4.
PCR amplification of the DNA fragment of GST-eYFP fused gene from Os09g0367700-OX and Os01g0949800-OX line for positive transgenic lines identification
Supplementary Material 5: Fig. S5.
KEGG enrichment of the proteins interacting with Os09g0367700 and Os01g0949800
Supplementary Material 6: Dataset S1.
Protocol details of iTRAQ proteomics and Co-IP
Supplementary Material 7: Dataset S2.
Mass spectrum identification of proteins interacting with Os09g0367700 in PI312777
Supplementary Material 8: Dataset S3.
Mass spectrum identification of proteins interacting with Os01g0949800 in Lemont
Supplementary Material 9: Electronic Supplementary Material 1.
Full length gel presents PCR amplification of the DNA fragment of GST-eYFP fused gene from Os09g0367700-OX and Os01g0949800-OX transgenic PI312777 lines
Supplementary Material 10: Electronic Supplementary Material 2.
Full length gel presents PCR amplification of the DNA fragment of GST-eYFP fused gene from Os01g0949800-OX transgenic Lemont line
Supplementary Material 11: Electronic Supplementary Material 3.
Full length gel presents GST-interacting proteins from GST-OX transgenic rice without DIMBOA treatment
Supplementary Material 12: Electronic Supplementary Material 4.
Full length gel presents GST-interacting proteins from GST-OX transgenic rice with DIMBOA treatment
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Zhang, H., Mu, D., Li, Y. et al. Glutathione S-transferase activity facilitates rice tolerance to the barnyard grass root exudate DIMBOA. BMC Plant Biol 24, 117 (2024). https://doi.org/10.1186/s12870-024-04802-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-024-04802-5