Abstract
Cell proliferation is a fundamental process underlying embryogenesis, homeostasis, wound healing, and cancer. The process involves multiple events during each cell cycle, such as cell growth, contractile ring formation, and division to daughter cells, which affect the surrounding cell population geometrically and mechanically. However, existing methods do not comprehensively describe the dynamics of multicellular structures involving cell proliferation at a subcellular resolution. In this study, we present a novel model for proliferative multicellular dynamics at the subcellular level by building upon the nonconservative fluid membrane (NCF) model that we developed in earlier research. The NCF model utilizes a dynamically-rearranging closed triangular mesh to depict the shape of each cell, enabling us to analyze cell dynamics over extended periods beyond each cell cycle, during which cell surface components undergo dynamic turnover. The proposed model represents the process of cell proliferation by incorporating cell volume growth and contractile ring formation through an energy function and topologically dividing each cell at the cleavage furrow formed by the ring. Numerical simulations demonstrated that the model recapitulated the process of cell proliferation at subcellular resolution, including cell volume growth, cleavage furrow formation, and division to daughter cells. Further analyses suggested that the orientation of actomyosin stress in the contractile ring plays a crucial role in the cleavage furrow formation, i.e., circumferential orientation can form a cleavage furrow but isotropic orientation cannot. Furthermore, the model replicated tissue-scale multicellular dynamics, where the successive proliferation of adhesive cells led to the formation of a cell sheet and stratification on the substrate. Overall, the proposed model provides a basis for analyzing proliferative multicellular dynamics at subcellular resolution.
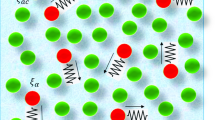
Graphical abstract











Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are partially available from the corresponding author on reasonable request.
References
B. O’Shaughnessy, S. Thiyagarajan, Biophys. Rev. 10, 1667 (2018)
J.A. Knoblich, Nat. Rev. Mol. Cell. Biol. 11, 849 (2010)
D. Ghose, T. Elston, D. Lew, Annu. Rev. Biophys. 51, 431 (2022)
Z. Tang, Y. Hu, Z. Wang, K. Jiang, C. Zhan, W.F. Marshall, N. Tang, Dev. Cell. 44, 313 (2018)
T. Kondo, S. Hayashi, Nature 494, 125 (2013)
B.I. Shraiman, Proc. Natl. Acad. Sci. 102, 3318 (2005)
R. Kikuchi, J. Chem. Phys. 24, 861 (1956)
H. Honda, Y. Ogita, S. Higuchi, K. Kani, J. Morphol. 174, 25 (1982)
T. Nagai, S. Ohta, K. Kawasaki, T. Okuzono, Ph. Trans.: Multinat. J. 28, 177 (1990)
H. Honda, J. Theor. Biol. 72, 523 (1978)
H. Honda, H. Yamanaka, M. Dan-Sohkawa, J. Theor. Biol. 106, 423 (1984)
M. Bock, A.K. Tyagi, J.-U. Kreft, W. Alt, Bull. Math. Biol. 72, 1696 (2010)
J.A. Glazier, F. Graner, Phys. Rev. E 47, 2128 (1993)
T. Hirashima, E.G. Rens, R.M.H. Merks, Dev. Growth Differ. 59, 329 (2017)
F. Graner, J.A. Glazier, Phys. Rev. Lett. 69, 2013 (1992)
A. Moure, H. Gomez, Arch. Comput. Methods Eng. 28, 311 (2021)
M. Nonomura, PLoS ONE 7, 33501 (2012)
M. Akiyama, M. Nonomura, A. Tero, R. Kobayashi, Phys. Biol. 16, 016005 (2018)
B. Lim, P.A.J. Bascom, R.S.C. Cobbold, Biorheology 34, 423 (1997)
K. Tsubota, S. Wada, T. Yamaguchi, Comput. Methods Progr. Biomed. 83, 139 (2006)
J. Mauer, S. Mendez, L. Lanotte, F. Nicoud, M. Abkarian, G. Gompper, D.A. Fedosov, Phys. Rev. Lett. 121, 118103 (2018)
S. Okuda, Y. Inoue, M. Eiraku, Y. Sasai, T. Adachi, Biomech. Model. Mechanobiol. 12, 987 (2013)
P. van Liedekerke, J. Neitsch, T. Johann, E. Warmt, I. Gonzàlez-Valverde, S. Hoehme, S. Grosser, J. Kaes, D. Drasdo, Biomech. Model Mechanobiol. 19, 189 (2020)
A. Torres-Sánchez, M. Kerr Winter, and G. Salbreux, PLoS Comput. Biol. 18, e1010762 (2022).
H. Turlier, B. Audoly, J. Prost, J.-F. Joanny, Biophys. J. 106, 114 (2014)
H.B. da Rocha, J. Bleyer, H. Turlier, J. Mech. Phys. Solids 164, 104876 (2022)
A. Mietke, F. Jülicher, I.F. Sbalzarini, Proc. Natl. Acad. Sci. 116, 29 (2019)
A. Torres-Sánchez, D. Millán, M. Arroyo, J. Fluid. Mech. 872, 218 (2019)
S. Okuda, K. Sato, T. Hiraiwa, Eur. Phys. J. E 45, 69 (2022)
S. Okuda, T. Hiraiwa, Phys. Rev. E 107, 034406 (2023)
M. Belkin, J. Sun, and Y. Wang, in Proceedings of the Twenty-Fourth Annual Symposium on Computational Geometry (2008), pp. 278–287.
S. Okuda, Y. Inoue, M. Eiraku, T. Adachi, Y. Sasai, Biomech. Model Mechanobiol. 14, 413 (2015)
B. Majhy, P. Priyadarshini, A.K. Sen, RSC Adv. 11, 15467 (2021)
G. Salbreux, G. Charras, E. Paluch, Trends Cell. Biol. 22, 536 (2012)
E. Fischer-Friedrich, A.A. Hyman, F. Jülicher, D.J. Müller, J. Helenius, Sci. Rep. 4, 1 (2014)
B. Smeets, M. Cuvelier, J. Pešek, H. Ramon, Biophys. J. 116, 930 (2019)
M. Fritzsche, A. Lewalle, T. Duke, K. Kruse, G. Charras, Mol. Biol. Cell. 24, 757 (2013)
S. Mukhina, Y. Wang, M. Murata-Hori, Dev. Cell. 13, 554 (2007)
A.-C. Reymann, F. Staniscia, A. Erzberger, G. Salbreux, S.W. Grill, Elife 5, e17807 (2016)
Acknowledgements
We would like to thank Hervé Turlier of the Collège de France for his insightful discussions and suggestions. This work was supported by the Japan Science and Technology Agency (JST), CREST Grant No. JPMJCR1921; the Japan Agency for Medical Research and Development (AMED), Grant No. 21bm0704065h0002; the Japan Society for the Promotion of Science (JSPS), KAKENHI Grants No. 21H01209, 21KK0134, 22K18749, and 22H05170; the World Premier International Research Center Initiative, Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan (to SO); and the seed grant of Mechanobiology Institute (to TH).
Author information
Authors and Affiliations
Contributions
SO conceived the project, developed the software, and performed the analyses; TH provided ideas; SO wrote the first draft of the manuscript, and SO and TH reviewed and edited. All authors contributed to the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
There are no conflicts to declare.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary file 1 Video 1. Process of cell growth, contractile ring formation, and division to daughter cells The amplitude of contractile stress is colored with a slope from green to red. The snapshots of this process are shown in Fig. 5.
Supplementary file 2 Video 2. Effects of isotropic stress of contractile ring on cleavage furrow formation The amplitude of contractile stress is colored with a slope from green to red. The snapshots of this process are shown in Fig. 7.
Supplementary file 3 Video 3. Process of tissue growth by successive cell proliferation on substrate Individual cells are shown in different colors. The snapshots of this process are shown in Fig. 9a.
Supplementary file 4 Video 4. Top-view of tissue growth process by successive cell proliferations on substrate. The amplitude of contractile stress is colored with a slope from green to red. The snapshots of this process are shown in Fig. 9b.
Supplementary file 5 Video 5. Proliferation process of a single cell embedded in growing tissue. The amplitude of contractile stress is colored with a slope from green to red. The snapshots of this process are shown in Fig. 9d.
Appendix 1. Energy function for inducing anisotropic stress on triangular surface
Appendix 1. Energy function for inducing anisotropic stress on triangular surface
Anisotropic stress exerted on an arbitrary triangle in the \({{xyz}}\)-orthogonal coordinates is considered. The triangle is composed of vertices 0, 1, and 2 (Fig. 11). Normal vector of the triangle is represented by \({\varvec{n}}\) in the \({{xyz}}\)-orthogonal coordinates. In addition, the \(x^{\prime}y^{\prime}\) orthogonal coordinates can be locally defined in the plane of the triangle, whose basis vectors are represented by \({\varvec{e}}_{{x^{\prime}}}\) and \({\varvec{e}}_{{y^{\prime}}}\) (\(= {\varvec{n}} \times {\varvec{e}}_{{x^{\prime}}}\)). Using \({\varvec{e}}_{{x^{\prime}}}\) and \({\varvec{e}}_{{y^{\prime}}}\), the area of the triangle, represented by \(A\), is given by
where \({\varvec{d}}_{\alpha \beta }\) is the distance vector from the \(\alpha\)- to \(\beta\)- vertices.
To introduce the anisotropic stress on the membrane, we redefine the area of the triangle as a function dependent on direction. This is represented by \(A_{{x^{\prime}}}\) and \(A_{{y^{\prime}}}\), where \({\varvec{\alpha}}\) is a directional vector. Both \(A_{{x^{\prime}}}\) and \(A_{{y^{\prime}}}\) have the same value as \(A\). The partial derivatives of \(A_{{x^{\prime}}}\) and \(A_{{y^{\prime}}}\) with respect to the displacement of a vertex, denoted by \(dA_{{x^{\prime}}}\) and \(dA_{{y^{\prime}}}\), respectively, correspond to the changes in area when the displacement has only \(x^{\prime}\) and \(y^{\prime}\) components. By regarding the components of vertex locations along \({\varvec{e}}_{{y^{\prime}}}\) as constants in Eq. (A1), Eq. (A1) is rewritten as follows
where \(c_{{y^{\prime}1}}\) and \( c_{{y^{\prime}2}}\) are regarded as constants. Similarly, by regarding the components of vertex locations along \({\varvec{e}}_{{x^{\prime}}}\) as constants in (A2), Eq. (A1) is given by
where \(c_{{x^{\prime}1}}\) and \( c_{{x^{\prime}2}}\) are regarded as constants.
Using Eqs. (A2) and (A3), the energy of anisotropic stress in the triangle surface, represented by \(u\), can be given by
where the constants \(\sigma_{{x^{\prime}}}\) and \(\sigma_{{y^{\prime}}}\) are the exerted stress along \({\varvec{e}}_{{x^{\prime}}}\) and \({\varvec{e}}_{{y^{\prime}}}\), respectively. When the exerted tension is isotropic (\(\sigma_{{x^{\prime}}} = \sigma_{{y^{\prime}}} = \sigma /2\)), the energy function becomes \(u = \sigma A\).
By calculating the partial derivatives of Eq. (A4), the forces acting on vertices 0, 1, and 2, represented by \({\varvec{f}}_{0}\), \({\varvec{f}}_{1}\), and \({\varvec{f}}_{2}\), respectively, were given by
respectively. The sum of each of the forces and moments in Eq. (A5) becomes zero. According to the above formulation, the area of the jth triangle in the ith cell is rewritten as a direction-dependent function, where the components of vertex position vectors in the direction normal to \({\varvec{\alpha}}\) is regarded as constants, \(A_{j\left( i \right)}^{\parallel } \left( {\varvec{\alpha}} \right)\).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Okuda, S., Hiraiwa, T. Modelling contractile ring formation and division to daughter cells for simulating proliferative multicellular dynamics. Eur. Phys. J. E 46, 56 (2023). https://doi.org/10.1140/epje/s10189-023-00315-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1140/epje/s10189-023-00315-5