Abstract
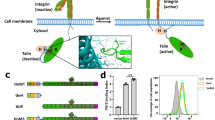
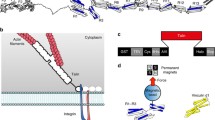
Interaction of the extracellular matrix (ECM) with the cell cytoskeleton is realized via integrin receptors. Adhesion structures formed in the cell after the ECM binding with integrins recruit talin molecules. They are involved both in the regulation of the activity of integrin receptors and binding of these receptors with the actin cytoskeleton. Talin is an adapter protein that contains the head domain, which is an atypical FERM domain, and rod domain, consisting of coiled bundles constructed of four or five α-helices. The pattern of α-helices packing in bundles determines their resistance to tensile forces and their ability to stretch. This review focuses on identifying the relationship between the structural organization of talin and distribution of functions between the head and rod domains. The spatial orientation of subdomains (F0, F1, F2, F3) in the head domain provides accessibility of binding sites in these subdomains for effector molecules and rapid transformation of the domain itself upon talin activation. The linear arrangement of helical bundles (R1–R13) in the rod domain with a predominance of four helical bundles at the N-end of the domain and five helical bundles at its C-end determines: 1) effective interbundle interaction during the formation of an inactive (autoinhibited) form of the talin dimer and 2) the possibility of modifying individual bundle spiralization under the action of physical stimuli. As a result, binding sites with various proteins burried in helical bundles of the talin rod domain are revealed. This means that the N-terminal site of the talin molecule (head domain) transforms some biochemical signals into others. The C-terminal part of the molecule (rod domain) converts physical stimuli into biochemical or physiological signals that regulate the cellular response. The review also considers the interaction of talin with various compounds at the molecular level.

Similar content being viewed by others
REFERENCES
Anthis, N.J., Wegener, K.L., Feng, Y., Kim, C., Goult, B.T., Lowe, E.D., Vakonakis, I., Bate, N., Critchley, D.R., Ginsberg, M.H., and Campbell, I.D., The structure of an integrin/talin complex reveals the basis of inside-out signal transduction, EMBO J., 2009, vol. 28, p. 3623. Atherton, P., Stutchbury, B., Wang, D.Y., Jethwa, D., Tsang, R., Meiler-Rodriguez, E., Wang, P., Bate, N., Zent, R., Barsukov, I.L., Goult, B.T., Critchley, D.R., and Ballestrem, C., Vinculin controls talin engagement with the actomyosin machinery, Nat. Commun., 2015, vol. 6, p. 10038. https://doi.org/10.1038/ncomms10038
Banno, A., Goult, B.T., Lee, H., Bate, N., Critchley, D.R., and Ginsberg, M.H., Subcellular localization of talin is regulated by inter-domain interactions, J. Biol. Chem., 2012, vol. 287, p. 13799.
Barczyk, M., Carracedo, S., and Gullberg, D., Integrins, Cell Tissue Res, 2010, vol. 339, p. 269.
Beaty, B.T., Wang, Y., Bravo-Cordero, J.J., Sharma, V.P., Miskolci, V., Hodgson, L., and Condeelis, J., Talin regulates moesin-NHE-1 recruitment to invadopodia and promotes mammary tumor metastasis, J. Cell Biol., 2014, vol. 205, p. 737.
Berrier, A.L. and Yamada, K.M., Cell-matrix adhesion, J. Cell Physiol., 2007, vol. 213, p. 565.
Bouaouina, M., Lad, Y., and Calderwood, D.A., The N-terminal domains of talin cooperate with the phosphotyrosine binding-like domain to activate β1 and β3 integrins, J. Biol. Chem., 2008, vol. 283, p. 6118.
Bouchet, B.P., Gough, R.E., Ammon, Y.C., van, der, Willige, D., Post, H., Jacquemet, G., Altelaar, A.F.M., Heck, A.J.R., Goult, B.T., and Akhmanova, A., Talin-KANK1 interaction controls the recruitment of cortical microtubule stabilizing complexes to focal adhesions, eLIFE, 2016, vol. 5. e18124. https://doi.org/10.7554/eLife.18124
Bunch, T.A., Integrin AIIbβ3 activation in Chinese hamster ovary cells and platelets increases clustering rather than affinity, J. Biol. Chem., 2010, vol. 285, p. 1841.
Calderwood, D.A., Yan, B., de Pereda, J.M., Garcia-Alvarez, B., Fujioka, Y., Liddington, R.C., and Ginsberg, M.H., The phosphotyrosine binding-like domain of talin activates integrins, J. Biol. Chem., 2002, vol. 277, p. 21749.
Campbell, I.D. and Humphries, M.J., Integrin structure, activation and interactions, Cold Spring Harb. Perspect. -Biol., 2011, vol. 3. a004994. https://doi.org/10.1101/cshperspect.a004994
Chang, Y.C., Zhang, H., Franco-Barraza, J., Brennan, M.L., Patel, T., Cukierman, E., and Wu, J., Structural and mechanistic insights into the recruitment of talin by RIAM in integrin signaling, Structure, 2014, vol. 22, p. 1810.
Changede, R., Xu, X., Margadant, F., and Sheetz, M.P., Nascent integrin adhesions form on all matrix rigidities after integrin activation, Dev. Cell, 2015, vol. 35, p. 614.
Chen, H.C., Appeddu, P.A., Parsons, J.T., Hildebrand, J.D., Schaller, M.D., and Guan, J.L., Interaction of focal adhesion kinase with cytoskeletal protein talin, J. Biol. Chem., 1995, vol. 270, p. 16995.
Chinthalapudi, K., Rangarajan, E.S., and Izard, T., The interaction of talin with the cell membrane is essential for integrin activation and focal adhesion formation, Proc. Natl. Acad. Sci. U. S. A., 2018, vol. 115, p. 10339.
Ciobanasu, C., Wang, H., Henriot, V., Mathieu, C., Fente, A., Csillag, S., Vigouroux, C., Faivre, B., and Clainche, C., Integrin-bound talin head inhibits actin filament barbed-end elongation, J. Biol. Chem., 2018, vol. 293, p. 2586.
Das, M., Ithychanda, S.S., Qin, J., and Plow, E.F., Mechanisms of talin-dependent integrin signaling and crosstalk, Biochim. Biophys. Acta, 2014, vol. 1838, p. 579.
Dedden, D., Schumacher, S., Kelley, C.F., Zacharias, M., Biertümpfel, C., Fässler, R., and Mizuno, N., The architecture of talin1 reveals an autoinhibition mechanism, Cell, 2019, vol. 179, p. 120.
De Pereda, J.M., Wegener, K.L., Santelli, E., Bate, N., Ginsberg, M.H., Critchley, D.R., Campbell, I.D., and Liddington, R.C., Structural basis for phosphatidylinositol phosphate kinase type Iγ binding to talin at focal adhesions, J. Biol. Chem., 2005, vol. 280, p. 8381.
del Rio, A., Perez-Jimenez, R., Liu, R., Roca-Cusachs, P., Fernandez, J.M., and Sheetz, M.P., Stretching single talin rod molecules activates vinculin binding, Science, 2009, vol. 323, p. 638.
Di Paolo, G., Pellegrini, L., Letinie, K., Cestra, G., Zoncu, R., Voronov, S., Chang, S., Guo, J., Wenk, M.R., and De, Camilli, P., Recruitment and regulation of phosphatidylinositol phosphate kinase type Iγ by the FERM domain of talin, Nature, 2002, vol. 420, p. 85.
Elliott, P.R., Goult, B.T., Kopp, P.M., Bate, N., Grossmann, J.G., Roberts, G.C.K., Critchley, D.R., and Barsukov, I.L., The structure of the talin head reveals a novel extended conformation of the FERM domain, Structure, 2010, vol. 18, p. 1289.
Gao, J., Huang, M., Lai, J., Mao, K., Sun, P., Cao, Z., Hu, Y., Zhang, Y., Schulte, M.L., Jin, C., Wang, J., White, G.C., Xu, Z., and Ma, Y.Q., Kindlin supports platelet integrin αIIbβ3 activation by interacting with paxillin, J. Cell Sci., 2017, vol. 130, p. 3764.
Garcia-Alvarez, B., de Pereda, J.M., Calderwood, D.A., Ulmer, T.S., Critchley, D.R., Campbell, I.D., Ginsberg, M.H., and Liddington, R.C., Structural determinants of integrin recognition by talin, Mol. Cell, 2003, vol. 11, p. 49.
Gingras, A.R., Ziegler, W.H., Frank, R., Barsukov, I.L., Roberts, G.C.K., Critchley, D.R., and Emsley, J., Mapping and consensus sequence identification for multiple vinculin binding sites within the talin rod, J. Biol. Chem., 2005, vol. 280, p. 37217.
Gingras, A.R., Bate, N., Goult, B.T., Hazelwood, L., Canestrelli, I., Grossmann, I.G., Liu, H., Putz, N.S.M., Roberts, G.C.K., Volkmann, N., Hanein, D., Barsukov, I.L., and Critchley, D.R., The structure of the C-terminal actin-binding domain of talin, EMBO J., 2008, vol. 27, p. 458.
Gingras, A.R., Bate, N., Goult, B.T., Patel, B., Kopp, P.M., Emsley, J., Barsukov, I.L., Roberts, G.C.K., and Critchley, D.R., Central region of talin has a unique fold that binds vinculin and actin, J. Biol. Chem., 2010, vol. 285, p. 29577.
Goksoy, E., Ma, Y.Q., Wang, X., Kong, X., Perera, D., Plow, E.F., and Qin, J., Structural basis for the autoinhibition of talin in regulating integrin activation, Mol. Cell, 2008, vol. 31, p. 124.
Goult, B.T., Bate, N., Anthis, N.J., Wegener, K.L., Gingras, A.R., Patel, B., Barsukov, I.L., Campbell, I.D., Roberts, G.C.K., and Critchley, D.R., The structure of an interdomain complex that regulates talin activity, J. Biol. Chem, 2009, vol. 284, p. 15097.
Goult, B.T., Bouaouina, M., Elliott, P.R., Bate, N., Patel, B., Gingras, A.R., Grossmann, J.G., Roberts, G.C.K., Calderwood, D.A., Critchley, D.R., and Barsukov, I.L., Structure of a double ubiquitin-like domain in the talin head: a role in integrin activation, EMBO J., 2010, vol. 29, p. 1069.
Goult, B.T., Xu, X.P., Gingras, A.R., Swift, M., Patel, B., Bate, N., Kopp, P.M., Barsukov, I.L., Critchley, D.R., Volkmann, N., and Hanein, D., Structural studies on full-length talin1 reveal a compact auto-inhibited dimer: implications for talin activation, J. Struct. Biol., 2013a, vol. 184, p. 21.
Goult, B.T., Zacharchenko, T., Bate, N., Tsang, R., Hey, F., Gingras, A.R., Elliott, P.R., Roberts, G.C.K., Ballestream, C., Critchley, D.R., and Barsukov, I.L., RIAM and vinculin binding to talin are mutually exclusive and regulate adhesion assembly and turnover, J. Biol. Chem., 2013b, vol. 288, p. 8238.
Goult, B.T., Yan, J., and Schwartz, M.A., Talin as a mechanosensitive signaling hub, J. Cell Biol., 2018, vol. 217, p. 3276.
Han, J., Lim, C.J., Watanabe, N., Soriani, A., Ratnikov, B., Calderwood, D.A., Puzon-McLaughlin, W., Lafuente, E.M., Boussiotis, V.A., Shattil, S.J., and Ginsberg, M.H., Reconstructing and deconstructing agonist-induced activation of integrin αIIbβ3, Curr. Biol., 2006, vol. 16, p. 1796.
Hemmings, L., Rees, D.J.G., Ohanian, V., Bolton, S.J., Gilmore, A.P., Patel, B., Priddle, H., Trevithick, J.E., Hynes, R.O., and Critchley, D.R., Talin contains three actin-binding sites each of which is adjacent to a vinculin-binding sites, J. Cell Sci., 1996, vol. 109, p. 2715.
Hu, X., Jing, C., Xu, X., Nakasawa, N., Cornish, V.W., Margadant, F.M., and Sheetz, M.P., Cooperative vinculin binding to talin mapped by time-resolved super resolution microscopy, Nano Lett., 2016, vol. 16, p. 4062.
Humphries, J.D., Wang, P., Streuli, C., Geiger, B., Humphries, M.J., and Ballestream, C., Vinculin controls focal adhesion formation by direct interactions with talin and actin, J. Cell Biol., 2007, vol. 179, p. 1043.
Hytönen, V.P., and Vogel, V., How force might activate talin’s vinculin binding sites: SMD reveals a structural mechanism, PLoS Comput. Biol., 2008, vol. 4. e24. https://doi.org/10.1371/journal.pcbi.0040024
Kalli, A.C., Campbell, I.D., and Sansom, M.S.P., Conformational changes in talin on binding to anionic phospholipid membranes facilitate signaling by integrin transmembrane helices, PLoS Comput. Biol., 2013, vol. 9. a1003316. https://doi.org/10.1371/journal.pcbi.1003316
Klapholz, B. and Brown, N.H., Talin—the master of integrin adhesions, J. Cell Sci., 2017, vol. 130, p. 2435.
Kopp, P.M., Bate, N., Hansen, T.M., Brindle, N.P.J., Praekelt, U., Debrand, E., Caleman, S., Mazzeo, D., Goult, B.T., Gingras, A.R., Pritchard, C.A., Critchley, D.R., and Monkley, S.J., Studies on the morphology and spreading of human endothelial cells define key inter-and intramolecular interactions for talin1, Eur. J. Cell Biol., 2010, vol. 89, p. 661.
Kumar, A., Ouyang, M, van den Dries, K., McGhee, E.J., Tanaka, K., Anderson, M.D., Groisman, A., Goult, B.T., Anderson, K.I., and Schwartz, M.A., Talin tension sensor reveals novel features of focal adhesion force transmission and mechanosensitivity, J. Cell Biol., 2016, vol. 213, p. 371.
Lau, T.L., Kim, C., Ginsberg, M.H., and Ulmer, T.S., The structure of the integrin αIIbβ3 transmembrane complex explains integrin transmembrane signalling, EMBO J., 2009, vol. 28, p. 1351.
Lawson, C., Lim, S.T., Uryu, S., Chen, X.L., Calderwood, D.A., and Schlaepfer, D.D., FAK promotes recruitment of talin to nascent adhesions to control cell motility, J. Cell Biol., 2012, vol. 196, p. 223.
Lee, H.S., Bellin, R.M., Walker, D.L., Patel, B., Powers, P., Liu, H., Garcia-Alvarez, B., de, Pereda, J.M., Liddington, R.C., Volkmann, N., Hanein, D., Critchley, D.R., and Robson, R.M., Characterization of an actin-binding site within the talin FERM domain, J. Mol. Biol., 2004, vol. 343, p. 771.
Lee, S.E., Kamm, R.D., and Mofrad, M.R., Force-induced activation of talin and its possible role in focal adhesion mechanotransduction, J. Biomech., 2007, vol. 40, p. 2096.
Lee, H.S., Lim, C.J., Puzon-McLaughlin, W., Shattil, S.J., and Ginsberg, M.H., RIAM activates integrins by linking talin to Ras GTPase membrane-targeting sequences, J. B-iol. Chem., 2009, vol. 284, p. 5119.
Legate, K.R. and Fässler, R., Mechanisms that regulate adaptor binding to β-integrin cytoplasmic tails, J. Cell Sci., 2009, vol. 122, p. 187.
Li, G., Du, X., Vass, W.C., Papageorge, A.G., Lowy, D.R., and Qian, X., Full activity of the deleted in liver cancer 1 (DLC1) tumor suppressor depends on an LD-like motif that binds talin and focal adhesion kinase (FAK), Proc. Natl. Acad. Sci. U. S. A., 2011, vol. 108, p. 17129.
Luo, B.H., Springer, T.A., and Takagi, J., A specific interface between integrin transmembrane helices and affinity for ligand, PLoS Biol., 2004, vol. 2. e153. https://doi.org/10.1371/Journal.pbio.0020153
McCann, R.O. and Craig, S.W., The I/LWEQ module: a conserved sequence that signifies F-actin binding in functionally diverse proteins from yeast to mammals, Proc. Natl. Acad. Sci. U. S. A., 1997, vol. 94, p. 5679.
Mehrbod, M., Trisno, S., and Mofrad, M.R.K., On the activation of integrin αIIbβ3: outside-in and Inside-out pathways, Biophys. J., 2013, vol. 105, p. 1304.
Morse, E.M., Brahme, N.N., and Calderwood, D.A., Integrin cytoplasmic tail interactions, Biochemistry, 2014, vol. 53, p. 810.
Pan, L., Zhao, Y., Yuan, Z., and Qin, G., Research advances on structure and biological functions of integrins, SpringerPlus, 2016, vol. 5, 1094. https://doi.org/10.1186/s40064-016-2502-0
Papagrigoriou, E., Gingras, A.R., Barsukov, I.L., Bate, N., Fillingham, I.J., Patel, B., Frank, R., Ziegler, W.H., Roberts, G.C.K., Critchley, D.R., and Emsley, J., Activation of a vinculin-binding site in the talin rod involves rearrangement of a five-helix bundle, EMBO J., 2004, vol. 23, p. 2942.
Rahikainen, R., Öhman, T., Turkki, P., Varjosalo, M., and Hytönen, V.P., Talin-mediated force transmission and talin rod domain unfolding independently regulate adhesion signaling, J. Cell Sci., 2019, vol. 132. jcs226514. https://doi.org/10.1242/jcs.226514
Roberts, G.C.K. and Critchley, D.R., Structural and biophysical properties of the integrin-associated cytoskeletal protein talin, Biophys. Rev., 2009, vol. 1, p. 61.
Rodius, S., Chaloin, O., Moes, M., Schaffner-Reckinger, E., Landriew, I., Lippens, G., Lin, M., Zhang, J., and Kieffer, N., The talin rod IBS2 α-helix interacts with the β3 integrin cytoplasmic tail membrane-proximal helix by establishing charge complementary salt bridges, J. Biol. Chem., 2008, vol. 283, p. 24212.
Saltel, F., Mortier, E., Hytönen, V.P., Jacquier, M.C., Zimmerman, P., Vogel, V., Liu, W., and Wehrle-Heller, B., New PI(4,5)P2- and membrane proximal integrin-binding motifs in the talin head control β3-integrin clustering, J. Cell Biol., 2009, vol. 187, p. 715.
Schiemer, J., Bohm, A., Lin, L., Merrill-Skoloff, G., Flaumenhaft, R., Huang, J.S., Le, Breton, G.C., and Chishti, A.H., Gα13 switch region 2 relieves talin autoinhibition to activate αIIbβ3 integrin, J. Biol. Chem., 2016, vol. 291, p. 26598.
Shattil, S.J., Kim, C., and Ginsberg, M.H., The final steps of integrin activations: the end game, Nat. Rev. Mol. Cell B-iol., 2010, vol. 11, p. 288.
Song, X., Yang, J., Hirbawi, J., Ye, S., Perera, H.D., Goksoy, E., Dwivedi, P., Plow, E.F., Zhang, R., and Qin, J., A novel membrane-dependent on/off switch mechanism talin FERM domain at sites of cell adhesion, Cell Res., 2012, vol. 22, p. 1533.
Sun, N., Critchley, D.R., Paulin, D., Li, Z., and Robson, R.M., Identification of a repeated domain within mammalian α-synemin that interacts directly with talin, Exp. Cell Res., 2008, vol. 314, p. 1839.
Sun, Z., Tseng, H.Y., Tan, S., Senger, F., Kursawa, L., Dedden, D., Mizuno, N., Wasik, A.A., Thery, M., Dunn, A.R., and Fässler, R., Kank2 activates talin, reduces force transduction across integrins and induces central adhesion formation, Nat. Cell Biol., 2016, vol. 18, p. 941.
Tremuth, L., Kreis, S., Melchior, C., Hoebeke, J., Rondé, P., Plancon, S., Takeda, K., and Kieffer, N., A fluorescence cell biology approach to map the second integrin-binding site of talin to a 130-amino acid sequence within the rod domain, J. Biol. Chem., 2004, vol. 279, p. 22258.
Vinogradova, O., Velyvis, A., Velyviene, A., Hu, B., Haas, T.A., Plow, E.F., and Qin, J., A structural mechanism of integrin αIIbβ3 “inside-out” activation as regulated by its cytoplasmic face, Cell, 2002, vol. 110, p. 587.
Wang, S., Watanabe, T., Matsuzawa, K., Katsumi, A., Kakeno, M., Matsui, T., Ye, F., Sato, K., Murase, K., Sugiyama, I., Kimura, K., Mizoguchi, A., Ginsberg, M.H., Collard, J.G., and Kaibuchi, K., Tiam 1 interaction with the PAR complex promotes talin-mediated Rac1 activation during polarized cell migration, J. Cell Biol., 2012, vol. 199, p. 331.
Wegener, K.L., Partridge, A.W., Han, J., Pickford, A.R., Liddington, R.C., Ginsberg, M.H., and Campbell, I.D., Structural basis of integrin activation by talin, Cell, 2007, vol. 128, p. 171.
Wolfenson, H., Lavelin, I., and Geiger, B., Dynamic regulation of the structure and functions of integrin adhesions, Dev. Cell., 2013, vol. 24, p. 447.
Yan, J., Yao, M., Goult, B.T., and Sheetz, M.P., Talin dependent mechanosensitivity of cell focal adhesions, Cell. Mol. Bioeng., 2015, vol. 8, p. 151.
Yang, J., Zhu, L., Zhang, H., Hirbawi, J., Fukuda, K., Dwivedi, P., Liu, J., Byzova, T., Plow, E.F., Wu, J., and Qin, J., Conformational activation of talin by RIAM triggers integrin-mediated cell adhesion, Nat. Commun., 2014, vol. 5, 5880. https://doi.org/10.1038/ncomms6880
Yao, M., Goult, B.T., Chen, H., Cong, P., Sheetz, M.P., and Yan, J., Mechanical activation of vinculin binding to talin locks talin in an unfolded conformation, Sci. Rep., 2014, vol. 4, 4610. https://doi.org/10.1038/srep04610
Yao, M., Goult, B.T., Klapholz, B., Hu, X., Toseland, C.P., Guo, Y., Cong, P., Sheetz, M.P., and Yan, J., The mechanical response of talin, Nat. Commun., 2016, vol. 7, p. 11966. https://doi.org/10.1038/ncomms11966
Ye, X., McLean, M.A., and Sligar, S.G., Conformational equilibrium of talin is regulated by anionic lipids, Biochim. Biophys. Acta, 2016, vol. 1858, p. 1833.
Zacharchenko, T., Qian, X., Goult, B.T., Jethwa, D., Almeida, T.B., Ballestrem, C., Critchley, D.R., Lowy, D.R., and Barsukov, I.L., LD motif recognition by talin: structure of the talin–DLC1 complex, Structure, 2016, vol. 24, p. 1130.
Zaidel-Bar, R., Itzkovitz, S., Ma’ayan, A., Iyengar, R., and Geiger, B., Functional atlas of the integrin adhesome, Nat. Cell Biol., 2007, vol. 9, p. 858.
Zhang, H., Chang, Y.C., Huang, Q., Brennan, M.L., and Wu, J., Structural and functional analysis of a talin triple-domain module suggests an alternative talin autoinhibitory configuration, Structure, 2016, vol. 24, p. 721.
Zhu, L., Yang, J., Bromberger, T., Holly, A., Lu, F., Liu, H., Sun, K., Klapproth, S., Hirbawi, J., Byzova, T.V., Plow, E.F., Moser, M., and Qin, J., Structure of Rap1b bound to talin reveals a pathway for triggering integrin activation, Nat. Commun., 2017, vol. 8, 1744. https://doi.org/10.1038/s41467-017-01822-8
Funding
This work was carried out within the framework of a state order to the Sechenov Institute of Evolutionary Physiology and Biochemistry, Russian Academy of Sciences, state registration no. АААА-А18-118012290371-3.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The author declares that she has no conflict of interest.
The author did not conduct experiments with animals or human beings.
Additional information
Translated by I. Fridlyanskaya
Abbreviations: ECM—extracellular matrix, ABS—actin binding site, DLC1—liver tumor suppressor, FAK—focal adhesion kinase, FERM—family of proteins (protein 4.1, ezrin, radixin, moesin), IBS—integrin binding site, KANK1-2—kidney ankyrin repeat-containing proteins, PIP2—phosphatidylinositol 4,5-bisphosphate, PIPK1γ—phosphatidylinositol 4-phosphate 5-kinase type I, RIAM—Rap1-GTP-interacting adapter molecule, TIAM1—T-cell lymphoma invasion and metastasis inducing factor; VBS,—vinculin binding site.
Rights and permissions
About this article
Cite this article
Ivanova, V.P. Talin: Structural and Functional Relationships. Cell Tiss. Biol. 15, 416–427 (2021). https://doi.org/10.1134/S1990519X21050060
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1990519X21050060