Abstract
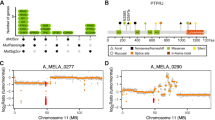
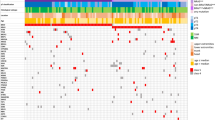
Acral melanoma is one of the most aggressive and fast-growing forms of cutaneous melanoma and is characterized by a predominant location on the palms and feet. Primary tumors, metastases, and normal tissue samples from five acral melanoma patients were examined by massive parallel sequencing, focusing on the coding regions of 4100 genes involved in the origin and progression of hereditary and oncology diseases. Somatic mutations were found in genes related to cell division, proliferation, and apoptosis (BRAF, NRAS, VAV1, GATA1, and GCM2); cell adhesion (CTNND2 and ITGB4); angiogenesis (VEGFA); and the regulation of energy metabolism (BCS1L). Comparisons of target DNA sequences between morphologically normal and primary tumor tissues and between normal and metastatic tissues identified the candidate genes responsible for rapid metastasis in acral melanoma.

Similar content being viewed by others
REFERENCES
Grebennikova O.P., Prilepo V.N. 2009. Melanoma, in Onkologiya dlya praktikuyushchikh vrachei (Oncology for Practicing Physicians), Chistyakov S.S., Ed., Moscow: KMK, pp. 548–563.
Bertolotto C. 2013. Melanoma: From melanocyte to genetic alterations and clinical options. Scientifica (Cairo). 2013, 1–22.
Mazurenko N.N. 2014. Genetic features and markers of skin melanoma. Usp. Mol. Onkol. 2, 26–35.
Feibleman C.E., Stoll H., Maize J.C. 1980. Melanomas of the palm, sole, and nailbed: A clinicopathologic study. Cancer. 46 (11), 2492–2504.
Borisova T.N., Kudryavtseva G.T. 2005. Combined treatment of metastatic acral melanoma. Sib. Oncol. Zh. 7 (14), 39–41.
Curtin J.A., Fridlyand J., Kageshita T., Patel H.N., Busam K.J., Kutzner H., Cho K.H., Aiba S., Bröcker E.B., LeBoit P.E., Pinkel D., Bastian B.C. 2005. Distinct sets of genetic alterations in melanoma. N. Engl. J. Med. 353 (20), 2135–2147.
Bolger A.M., Lohse M., Usadel B. 2014. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics. 30 (15), 2114–2120.
McKenna A., Hanna M., Banks E., Sivachenko A., Cibulskis K., Kernytsky A., Garimella K., Altshuler D., Gabriel S., Daly M., DePristo M.A. 2010. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20 (9), 1297–1303.
Cibulskis K., Lawrence M.S., Carter S.L., Sivachenko A., Jaffe D., Sougnez C., Gabriel S., Meyerson M., Lander E.S., Getz G. 2013. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31 (3), 213–219.
Vaser R., Adusumalli S., Leng S.N., Sikic M., Ng P.C. 2016. SIFT missense predictions for genomes. Nat. Protoc. 11 (1), 1–9.
Adzhubei I.A., Schmidt S., Peshkin L., Ramensky V.E., Gerasimova A., Bork P., Kondrashov A.S., Sunyaev S.R. 2010. A method and server for predicting damaging missense mutations. Nat. Methods. 7 (4), 248–249.
Schwarz J.M., Cooper D.N., Schuelke M., Seelow D. 2014. MutationTaster2: Mutation prediction for the deep-sequencing age. Nat. Methods. 11 (4), 361–362.
Chun S., Fay J.C. 2009. Identification of deleterious mutations within three human genomes. Genome Res. 19 (9), 1553–1561.
Zhu X., Jin H., Xia Z., Wu X., Yang M., Zhang H., Shang X., Cheng R., Zhan Z., Yu Z. 2017. Vav1 expression is increased in esophageal squamous cell carcinoma and indicates poor prognosis. Biochem. Biophys. Res. Commun. 486 (2), 571–576.
Ling T., Crispino J.D., Zingariello M., Martelli F., Migliaccio A.R. 2018. GATA1 insufficiencies in primary myelofibrosis and other hematopoietic disorders: consequences for therapy. Expert Rev. Hematol. 11 (3), 169–184.
Bot S.T., Vermeer S., Buijsman W., Heister A., Voorendt M., Verrips A., Scheffer H., Kremer H.P., Warrenburg B.P., Kamsteeg E.J. 2013. Pure adult-onset spastic paraplegia caused by a novel mutation in the KIAA0196 (SPG8) gene. J. Neurol. 260 (7), 1765–1769.
Bierie B., Pierce S.E., Kroeger C., Stover D.G., Pattabiraman D.R., Thiru P., Liu Donaher J., Reinhardt F., Chaffer C.L., Keckesova Z., Weinberg R.A. 2017. Integrin-β4 identifies cancer stem cell-enriched populations of partially mesenchymal carcinoma cells. Proc. Natl. Acad. Sci. U. S. A. 114 (12), E2337–E2346.
Tahara H., Naito H., Kise K., Wakabayashi T., Kamoi K., Okihara K., Yanagisawa A., Nakai Y., Nonomura N., Morii E., Miki T., Takakura N. 2015. Evaluation of PSF1 as a prognostic biomarker for prostate cancer. Prostate Cancer Prostatic Dis. 18 (1), 56–62.
Zhang J., Wu Q., Wang Z., Zhang Y., Zhang G., Fu J., Liu C. 2015. Knockdown of PSF1 expression inhibits cell proliferation in lung cancer cells in vitro. Tumor Biol. 36 (3), 2163–2168.
Temprano A., Sembongi H., Han G.S., Sebastián D., Capellades J., Moreno C., Guardiola J., Wabitsch M., Richart C., Yanes O., Zorzano A., Carman G.M., Siniossoglou S., Miranda M. 2016. Redundant roles of the phosphatidate phosphatase family in triacylglycerol synthesis in human adipocytes. Diabetologia. 59 (9), 1985–1994.
Beeler N., Riederer B.M., Waeber G., Abderrahmani A. 2009. Role of the JNK-interacting protein 1/islet brain 1 in cell degeneration in Alzheimer disease and diabetes. Brain Res. Bull. 80 (4–5), 274–281.
Sun Y., Wang F., Sun X., Wang X., Zhang L., Li Y. 2017. CX3CR1 regulates osteoarthrosis chondrocyte proliferation and apoptosis via Wnt/β-catenin signaling. Biomed. Pharmacother. 96, 1317–1323.
Wang H., Cai J., Du S., Guo Z., Xin B., Wang J., Wei W., Shen X. 2017. Fractalkine/CX3CR1 induces apoptosis resistance and proliferation through the activation of the AKT/NF-κB cascade in pancreatic cancer cells. Cell Biochem. Funct. 35 (6), 315–326.
Stout M.C., Narayan S., Pillet E.S., Salvino J.M., Campbell P.M. 2018. Inhibition of CX3CR1 reduces cell motility and viability in pancreatic adenocarcinoma epithelial cells. Biochem. Biophys. Res. Commun. 495 (3), 2264–2269.
Everything about cancer: diagnosis, therapy, oncologists’ advice online, OncoForum. http://www.oncoforum.ru/. Accessed November 17, 2018.
Funding
This work was supported by the Russian Science Foundation (project no. 14-35-00107).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests. The authors declare that they have no conflict of interest.
Statement of compliance with standards of research involving humans as subjects. The study was approved by the Ethics Committee at the Blokhin National Medical Research Center of Oncology (Ministry of Health of the Russian Federation). All patients gave their written informed consent to their tissue samples being used for research purposes.
Additional information
Translated by T. Tkacheva
Rights and permissions
About this article
Cite this article
Abramov, I.S., Emelyanova, M.A., Ryabaya, O.O. et al. Somatic Mutations Associated with Metastasis in Acral Melanoma. Mol Biol 53, 580–585 (2019). https://doi.org/10.1134/S0026893319040022
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893319040022