Abstract
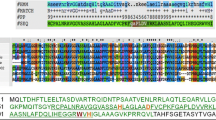
Two families of specific inhibitors of type C lysozyme (Ivy and PliC) secreted from the periplasmic space are known in enterobacteria. Microbial capacity for distant lysozyme inactivation (antilysozyme activity) is most pronounced in the strains and species carrying homologues of the pliC gene. The pliC homologue localized in a ∼200-kb megaplasmid of Klebsiella pneumoniae was shown to differ significantly in the amino acid composition of the coded polypeptide. Similar to the Salmonella enterica pliC homologue, it possesses a detachable signal part and contains identical functionally critical amino acids of the active center of the inhibitor. Antilysozyme activity of the pliC-positive K. pneumoniae strains was observed at the level corresponding to the highest values found in pliC-positive S. enterica. High level of the antilysozyme activity in K. pneumoniae strains containing the plasmid pliC homologue was found in all studied strains, unlike S. enterica strains carrying the known chromosomal pliC homologue.
Similar content being viewed by others
References
Callewaert, L., Aertsen, A., Deckers, D., Vanoirbeek, K.G., Vanderkelen, L., Van Herreweghe, J.M., Masschalck, B., Nakimbugwe, D., Robben, J., and Michiels, C.W., A new family of lysozyme inhibitors contributing to lysozyme tolerance in gram-negative bacteria, PLoS Pathog., 2008, vol. 4, no. 3, e1000019.
Bukharin, O.V., Persistentsiya patogennykh bakterii (Persistence of Pathogenic Bacteria), Moscow: Meditsina, 1999.
Monchois, V., Abergel, C., Sturgis, J., Jeudy, S., and Claverie, J.-M., Escherichia coli ykfE ORFan gene encodes a potent inhibitor of C-type lysozyme, J. Biol. Chem., 2001, vol. 276, pp. 18437–18441.
Abergel, C., Monchois, V., Byrne, D., Chenivesse, S., Lembo, F., Lazzaroni, J.C., and Claverie, J.-M., Structure and evolution of the Ivy protein family, unexpected lysozyme inhibitors in gram-negative bacteria, Proc. Natl. Acad. Sci. U. S. A., 2007, vol. 104, no. 15, pp. 6394–6399.
Chen, Y.T., Chang, H.Y., Lai, Y.C., Pan, C.C., Tsai, S.F., and Peng, H.L., Sequencing and analysis of the large virulence plasmid pLVPK of Klebsiella pneumoniae CG43, Gene, 2004, vol. 337, pp. 189–198.
Wu, K.M., Li, L.H., Yan, J.J., Tsao, N., Liao, T.L., Tsai, H.C., Fung, C.P., Chen, H.J., Liu, Y.M., Wang, J.T., Fang, C.T., Chang, S.C., Shu, H.Y., Liu, T.T., Chen, Y.T., Shiau, Y.R., Lauderdale, T.L., Su, I.J., Kirby, R., and Tsai, S.F., Genome sequencing and comparative analysis of Klebsiella pneumoniae NTUH-K2044, a strain causing liver abscess and meningitis, J. Bacteriol., 2009, vol. 191, no. 14, pp. 4492–4501.
Bondarenko, V.M., Petrovskaya, V.G., and Yablochkov, A.L., Antilysozyme factor of Klebsiella pneumoniae: nature, biological functions, and genetic control, Zh. Mikrobiol. Epidemiol. Immunobiol., 1994, no. 1, pp. 22–28.
Bukharin, O.V., Kirillov, D.A., and Kirillov, V.A., Plasmid screening in Bacillus strains with antilysozyme activity, Zh. Mikrobiol. Epidemiol. Immunobiol., 2006, no. 1, pp. 3–6.
Higgins, D., Thompson, J., and Gibson, T., CLUSTALW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice, Nucleic Acid Res., 1994, vol. 22, no. 22, pp. 4673–4680.
Petersen, T.N., Brunak, S., von Heijne, G., and Nielsen, H., SignalP 4.0: discriminating signal peptides from transmembrane regions, Nature Methods, 2011, no. 8, pp. 785–786.
Maniatis T., Fritsch, E.F., and Sambrook, J., Molecular Cloning-A Laboratory Manual, New York: Cold Spring Harbor Laboratory, 1982.
Rebrikov, D.V., Samatov, G.A., Trofimov, D.Yu., Semenov, P.A., Savilova, A.M., Kofiadi, I.A., and Abramov, D.D., PTsR “v real’nom vremeni” (Real-Time PCR), Moscow: BINOM, 2009.
Cherkasov S.V. Associative symbiosis as the biological basis for the host colonization resistance, Doctoral (Med.) Dissertation, Orenburg: Orenburg. Gos. Med. Acad., 2011.
Hiroshi, N., Molecular basis of bacterial outer membrane permeability revisited, Microbiol. Mol. Biol. Rev., 2003, vol. 67, no. 4, pp. 593–656.
Deckers, D., Masschalck, B., Aertsen, A., Callewaert, L., Van Tiggelen, C.G., Atanassova, M., and Michiels, C.W., Periplasmic lysozyme inhibitor contributes to lysozyme resistance in Escherichia coli, Cell Mol. Life Sci., 2004, vol. 61, no. 10, pp. 1229–1237.
Leysen, S., Van Herreweghe, J.M., Callewaert, L., Heirbaut, M., Buntinx, P., Michiels, C.W., and Strelkov, S.V., Molecular basis of bacterial defense against host lysozymes: X-ray structures of periplasmic lysozyme inhibitors PliI and PliC, J. Mol. Biol., 2011, vol. 405, no. 5, pp. 1233–1245.
Fernie-King, B.A., Seilly, D.J., Binks, M.J., Sriprakash, K.S., and Lachmann, P.J., Streptococcal DRS (distantly related to SIC) and SIC inhibit antimicrobial peptides, components of mucosal innate immunity: a comparison of their activities, Microb. Infect., 2007, vol. 9, no. 3, pp. 300–307.
Callewaert, L., Vanoirbeek, K.G., Lurquin, I., Michiels, C.W., and Aertsen, A., The Rcs two-component system regulates expression of lysozyme inhibitors and is induced by exposure to lysozyme, J. Bacteriol., 2009, vol. 191, no. 6, pp. 1979–1981.
Ellison, R.T. and Giehl, T.J., Killing of gram-negative bacteria by lactoferrin and lysozyme, J. Clin. Investig., 1991, vol. 88, no. 4, pp. 1080–1091.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © S.V. Andryushchenko, N.B. Perunova, O.V. Bukharin, 2014, published in Mikrobiologiya, 2014, Vol. 83, No. 3, pp. 302–311.
Rights and permissions
About this article
Cite this article
Andryushchenko, S.V., Perunova, N.B. & Bukharin, O.V. Periplasmic lysozime inhibitior pliC and its role in antilysozime activity of enterobacteria. Microbiology 83, 211–219 (2014). https://doi.org/10.1134/S0026261714030047
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261714030047