Abstract
Candida albicans is an opportunistic human fungal pathogen. The ability to switch among multiple cellular forms is key to its pathogenesis. The Dbf4-dependent protein kinase gene CDC7 is conserved due to its role in initiating DNA replication. Because a C. albicans Cdc7 (Cacdc7) homozygous null was not viable, we generated a C. albicans strain with a deleted C. albicans CDC7 (CaCDC7) allele and an expression-repressible allele. Surprisingly, cells of the strain grew as hyphae under the repressed conditions. The in vitro kinase assays confirmed that CaCdc7 (K232) and CaCdc7 (T437) are critical for catalytic and phosphoacceptor of activation activity, respectively. C. albicans cells formed hyphae when expressing either the catalytically inactive CaCdc7 (K232R) or the phosphoacceptor-deficient CaCdc7 (T437A). While CaCdc7 interacted with CaDbf4, cells of the strain in which CaCDC7 was repressed were not rescued by constitutively expressing C. albicans DBF4 or vice versa. We conclude that CaDBF4-dependent CaCDC7 is an essential gene suppressing the hyphal development.
Similar content being viewed by others
Introduction
Candida albicans is an opportunistic human fungal pathogen without a complete sexual cycle. The virulence of C. albicans stems from its ability to alter morphology from the ellipsoid blastospore to various filamentous forms1, although the ability to morphological switch and virulence might be decoupled2. Switching among diverse morphological forms is influenced by many environmental factors3 and is mediated by several signaling pathways. The mechanism by which hyphal growth is controlled has largely been elucidated and was recently reviewed by Sudbery, P.E.4, but the exact roles of many genes known to regulate morphogenesis in polarized growth, cell separation, and the integration of signaling pathways to hyphal growth remain to be determined. Additionally, novel genes involved in morphogenesis remain to be uncovered to understand the overall-control network of the yeast-to-hyphae (YTH) transition. We have identified S. cerevisiae homologs of CDC28, two G1 cyclins in C. albicans5, indicating conservation between C. albicans and S. cerevisiae in the control of the mitotic cell cycle in the G1 phase. However, we and others have shown that C. albicans CDC4 (CaCDC4) suppresses filamentation, in contrast to its S. cerevisiae counterpart, whose function is required for progression from the G1 to the S phase of the mitotic cell cycle6,7. We have previously identified several novel Cdc4-associated proteins8. Consequently, we investigated other genes required for cells to advance from the G1 to the S phase.
We were particularly interested in the catalytic subunit of the serine/threonine protein kinase encoded by CDC7 and its regulatory subunit, encoded by DBF4, known as Dbf4-dependent Cdc7 kinase (DDK), because they play an essential role in the initiation of DNA replication in S. cerevisiae9,10 and are conserved throughout evolution11. In addition to its key role in replication initiation, DDK responds to replication fork stalling12,13,14 and DNA damage14,15 that maintain genome integrity. Moreover, the roles of DDK are extended to many other areas, including checkpoint control, trans-lesion DNA synthesis, meiosis, chromatin reconstruction, and histone, which have been reviewed recently16. Diverse genotoxic insults, including those that block DNA replication, lead to filamentous growth in S. cerevisiae17,18,19 and C. albicans20,21,22,23. The genes involved in DNA replication checkpoints appear to require the induction of filamentous growth23. However, how DNA replication stress leads to filamentous growth remains incompletely understood. Moreover, the importance or requirement of some of the key factors, such as Swe1, for DNA replication stress-induced filamentous development does not seem to be conserved between the two yeasts24,25,26,27. Until a recent global analysis of C. albicans morphology28, no reports pointed to a direct involvement of DDK in filamentation.
To verify the role of DDK in C. albicans, we have characterized CaDBF429 and C. albicans CDC7 (CaCDC7). We generated a C. albicans strain capable of repressing the expression of CaCDC7 and examined the cellular morphology when CaCDC7 is depleted. Strains constitutively expressing the catalytically inactive CaCdc7 or the phosphoacceptor-deficient CaCdc7 were generated to verify the requirement of kinase activity for cellular morphology. Additionally, the functional dependency of CaCdc7 and CaDbf4 was tested by the yeast two-hybrid assay and by constitutively expressing CaCdc7 in a CaDBF-deletion strain and vice versa.
Results and Discussion
C. albicans CDC7 is a structural homolog of S. cerevisiae CDC7
By using Blast to compare the Candida Genome Database with the entire sequence of the S. cerevisiae Cdc7 protein, a single CaCDC7 located on chromosome 2 was identified that belongs to Contig19-10183 [70303..72273] orf19.3561, Assembly 19, which has one reading frame of 1971 bp and potentially encodes a 72 kD protein of 657 amino acid residues. To analyze the structure of the protein encoded by CaCDC7, the protein sequence derived from the 1971 bp ORF was aligned and compared to other Cdc7 protein sequences across the evolutionary spectrum by ClustalW30,31. As shown in Supplementary Fig. S1, all twelve conserved serine/threonine protein kinase subdomains as defined by Hanks, S.K., and Quinn, A.M.32 can clearly be identified in the protein sequence of CaCdc7. Moreover, the invariant amino acid residues in the subdomains, which are found throughout protein serine/threonine kinases, are also conserved in CaCdc7. Based on Supplementary Fig. S1, Fig. 1 was generated as a schematic diagram comparing C. albicans Cdc7 (CaCdc7), S. cerevisiae Cdc7 (ScCdc7), and human Cdc7 (HsCdc7). As shown in Fig. 1, in domain II of the ATP-binding region, CaCdc7 contains a lysine at residue 232 (K232), the same as lysine 76 (K76)33,34 in the budding yeast ScCdc7 and lysine 90 (K90) in human Cdc735, which is the site required for catalytic activity. In subdomain VIII of the phosphoacceptor region, CaCdc7 also contains a threonine at residue 437 (T437), equivalent to threonine 281 (T281) in ScCdc7 and threonine 376 (T376) in humans, which is the phosphorylation site for kinase activation33,34. Additionally, the spacing between subdomains VII and VIII is worth noting, because the same region in ScCdc7 is needed for its mitotic function36 and is unique among all known kinases32. Notably, CaCdc7 possesses Cdc7 characteristic insertions of KI-0, KI-1, KI-2, and KI-3, and distinct regions located at the N- and C-terminus. Importantly, the C-terminal tail of Cdc7 is known to be essential for interacting with Dbf437. Together with KI-2 and KI-3, such an interaction becomes efficient38. Nonetheless, the functional significance of the less-conserved C-terminal tail of CaCdc7 is unclear. Unique features are also visible in CaCdc7. The most striking is an extended stretch of approximately 140 amino acid residues, which are quite hydrophobic, at the amino-terminus of CaCdc7. The region of insertion KI-2 of CaCdc7 between 355 and 393 is rich in threonine and serine (Supplementary Fig. S1). Nevertheless, the functional significance of these differences is unknown. Despite the differences between CaCdc7 and its counterparts, particularly ScCdc7, the overall organization and sequence of CaCdc7 is very similar to Cdc7. We conclude that CaCdc7 encodes a serine/threonine kinase with homology both in sequence and organization to Cdc7 across the evolutionary spectrum.
The indicated conserved subdomains of the serine/threonine kinases are shown, each of which is specified by a roman numeral (from I to XI). The subdomain VI is further separated into the subdomains VIa and VIb. The kinase insert with conserved residues (KI-0) is shown. The kinase inserts of KI-1, KI-2, and KI-3 are indicated. The positions of the essential lysine residue for catalytic activity of the three Cdc7s, CaCdc7 (K232), ScCdc7 (K76), and HsCdc7 (K90), are shown. The positions of the phosphoacceptor threonine residue for kinase activation of the three Cdc7s, CaCdc7 (T437), ScCdc7 (T281), and HsCdc7 (T376), are shown.
Construction of the CDC7 expression-repressible C. albicans strain
To establish the function of CaCDC7 in C. albicans, we sought to construct a Cacdc7 deletion mutant. However, as CaCDC7 encodes a protein with the structural homologue of known Cdc7 proteins whose function is the initiation of DNA replication, we predicted that the cdc7 homozygous null mutant is lethal. We addressed this issue by generating a strain with one CaCDC7 allele deleted and the other under the control of MET3 promoter (MET3p). To delete one CaCDC7 allele, we used the mini-Ura-blaster approach39. We PCR-generated a cassette using the plasmid pDDB5739 with the dpl200 flanked by URA3 as a template, together with primers having sequences homologous to URA3-dpl200 and the up- and down-stream sequences of CaCDC7. We then introduced the cassette into the C. albicans auxotrophic strain BWP17 (ura3 arg4 his1) to obtain strain CaCDC7+/U3−. Then, CaCDC7+/U3− cells were treated with 5-FOA to obtain strain CaCDC7+/− whose URA3 was removed. To obtain a strain capable of conditionally expressing CaCDC7 under MET3p control, we PCR-amplified the partial CaCDC7 and cloned it into plasmid vector pFA-HIS1-MET3p to generate plasmid pFA-HIS1-MET3p-pCaCDC7. We then linearized the plasmid at a unique restriction site within the partial CaCDC7. By introducing the linearized plasmid into strain CaCDC7+/U3−, we obtained strain CaCDC7 M3/U3−. To remove the URA3 cassette, we treated cells of strain CaCDC7 M3/U3− with 5-FOA to obtain strain CaCDC7 M3/−. The strains CaCDC7 M3/U3−, CaCDC7 M3/−, CaCDC7+/U3−, and BWP17 were subjected to Southern blotting analysis.
The size of the XbaI fragment containing CaCDC7 shifted from 8392 bp to 1924 bp and from 8392 bp to 6698 bp, respectively, demonstrating that the CaCDC7 alleles were either integrated with the mini-Ura-blaster cassette or MET3p-driven CaCDC7 (Fig. 2A). The size of the HindIII fragment containing CaCDC7 shifted from 7578 bp to 6223 bp, demonstrating that the CaCDC7 alleles integrated with the mini-Ura-blaster cassette had lost URA3 (Fig. 2B). These results indicate that the strain CaCDC7 M3/− has the expected genome organization of C. albicans.
Cells of BWP17 (CaCDC7+/+) (#1) were consecutively introduced a cassette of mini-Ura-blaster to obtain CaCDC7+/U3− (#2) and MET3p-driven CaCDC7 to obtain CaCDC7 M3/U3− (#3), after which the CaCDC7 M3/U3− was treated with 5-FOA to obtain CaCDC7 M3/− (#4). The genomic DNA from cells of each strain was extracted and subjected to either XbaI (A) or Hind III (B) digestion before electrophoresis and Southern blotting analysis (the bottom panel). Strain CaCDC7+/−(*) was made by introducing a cassette of mini-Ura-blaster to BWP17 (CaCDC7+/+) to obtain CaCDC7+/U3−, which was then treated with 5-FOA. The CaCDC7+/−(*) is used as a control to highlight the relative positions of allele CaCDC7 and Cacdc7::dpl200. The relative positions of the probes are shown.
CDC7 is an essential gene and depletion of CDC7 leads to hyphal growth in C. albicans
To test the repressibility of CaCDC7 in the strain CaCDC7 M3/−, we grew strains CaCDC7 M3/−, BWP17 (CaCDC7+/+), and CaCDC7+/− cells in SD medium with or without 2.5 mM methionine/cysteine (Met/Cys) and extracted RNA for RT-PCR analysis. The expression of CaCDC7 in the strain CaCDC7 M3/− was significantly reduced under the repressed conditions compared to the de-repressed conditions (Fig. 3A), suggesting that the expression of CaCDC7 in the strain CaCDC7 M3/− was solely controlled by MET3p and the other allele was deleted.
(A) Cells of strains CaCDC7+/+ (BWP17), CaCDC7+/−, and CaCDC7 M3/− were grown in the SD medium with required supplements in the absence (−Met/Cys) or presence (+Met/Cys) of each of 2.5 mM methionine and cysteine at 16 h prior to collection for RT-PCR to verify the repression of CaCDC7. The numbers shown are relative fold change in the expression of those strains to BWP17 under the de-repressed condition (−Met/Cys), normalized to ACTIN expression. (B) The same cultures were grown for the indicated times prior to the assessment of morphological alterations under the microscope. Bars represent 10 μm.
We next established the requirement for CaCDC7 in C. albicans by growing strains on selective media in the presence or absence of Met/Cys. Strain CaCDC7 M3/− formed colonies in the presence of Met/Cys with wrinkled surfaces (unpublished data). The results suggested that CaCDC7 suppresses filamentous development and may not be essential. This result is consistent with our finding that repression of CaDBF4 expression leads to filamentous growth in C. albicans29. To confirm the necessity of CaCDC7 in C. albicans, we established a Cacdc7 homozygous null mutant. However, whereas the Cacdc7 heterozygous null mutants were generated with ease, no Cacdc7 homozygous null mutants were obtained. Our result is in agreement with a recent report where the functional copy of the essential CaCDC7 gene in the heterozygous strain was governed by a tetracycline-repressible promoter to allow functional study28. This result is also consistent with the CaDBF4 data, in which Cadbf4 homozygous null mutants were unable to survive29. These results suggest that C. albicans CDC7 and its regulator encoded CaDBF4 gene, like their counterparts across evolutionary spectrum, possesses a conserved role in the initiation of DNA replication. In the MET3 promoter-driven system, the difference in expression can reach 85-fold between repressed and de-repressed conditions40. We reasoned that under the repressed condition, CaCDC7 was depleted while a limited amount of CaCdc7 was able to function, although to a lesser extent. While Cdc7 depletion substantially inhibits proliferation in cancer cells41,42, the depletion of CaCDC7 appeared to reduce proliferation, as the growth rate of CaCDC7 M3/− was lower than that of its parental BWP17 when cells were grown in medium with Met/Cys (Supplementary Fig. S2). Taken together, these results suggest that the function of CaCDC7 is tightly associated to that of CaDBF4 in C. albicans and that the CaDbf4-dependent CaCdc7 kinase is essential, as is its counterpart in S. cerevisiae9,10.
To determine the phenotypic consequences, in particular the cellular morphology, of CaCDC7 M3/− under the repressed condition, we grew CaCDC7 M3/− cells in medium with or without 2.5 mM Met/Cys and examined the phenotypic consequences microscopically. The cells formed germ tubes after 4 h of repression and continued to grow as hyphae from 8 h to 24 h under the repressed condition (Fig. 3B). These observations were comparable to those of cells repressing CaDBF4 expression29, suggesting that CaCDC7 and CaDBF4 function as a DDK for the YTH transition and that CaCDC7 may have an additional role in morphogenesis. To definitely determine that CaCDC7 is the gene for suppression of the yeast-to-hypha transition, we performed a rescue assay where a constitutive ACT1 promoter (ACT1p)-driven CaCDC7 was introduced into the CaCDC7 M3/− strain. These cells grew in the yeast form even in the presence of Met/Cys (Supplementary Fig. S3), confirming that CaCDC7 is responsible for the inhibition of hyphal growth in C. albicans. Together with the fact that repression of CaDBF4 expression led to filamentous growth in C. albicans29, these data suggest that the CaCDC7 and CaDBF4 encoded proteins likely act together to perform their function. The hyphal growth during the CaCDC7 or CaDBF4 depleted condition may be the consequence of constrained function that exerts as a stress condition. In S. cerevisiae, genotoxic stress conditions that reduce DNA synthesis can induce filamentous differentiation through Mec1-Rad53-Swe1-Cdc28-Clb217. However, in C. albicans, DNA damage-induced cell cycle delay leading to polarized growth is only partially dependent on Swe124, suggesting that genotoxic stress-induced filamentous growth is not or is only partially mediated by Swe1 in C. albicans. However, no evidence indicates that CDC7 interacts with the above components in C. albicans. Hence, whether genotoxic stress-induced filamentous growth is mediated by CaCDC7 in C. albicans remains to be verified.
The conserved residues are essential for the catalytic activity of CaCdc7 and the phosphoacceptor of activation
To verify if the CaCDC7-encoded protein product has kinase activity and is capable of being phosphorylated, we created plasmids p6HF-ACT1p-CaCDC7, p6HF-ACT1p-CaCDC7 (K232R) and p6HF-ACT1p-CaCDC7 (T437A) capable of constitutively expressing either the wild-type CaCDC7, the catalytically inactive CaCdc7 (K232R), or the phosphoacceptor-deficient CaCdc7 (T437A) (Fig. 4A), each of which was introduced into cells of the strain CaCDC7 M3/−. We grew cells of these strains exponentially in the medium with Met/Cys. After harvesting and lysing, each of the wild-type and mutant CaCdc7s was purified by Ni2+-NTA agarose and subjected to an in vitro kinase assay. While the wild-type CaCdc7 was able to phosphorylate histone H1, this phosphorylation was significantly inhibited by FSBA, which blocks the catalytic activity of a kinase (Fig. 4B). This result was consistent with the inability of catalytically inactive CaCdc7 (K232R) to phosphorylate histone H1 (Fig. 4B). We noted that the FSBA appeared to block autophosphorylation of CaCdc7, which was indicated by the decrease in the upper band of the CaCdc7 doublet (Fig. 4B). These results confirmed that CaCdc7 possesses kinase activity and that the structurally predicted K232 (Fig. 1)33,34 is indeed the site essential for catalytic activity. Additionally, the phosphoacceptor-deficient CaCdc7 (T437A) almost lost the ability to phosphorylate histone H1 (Fig. 4B), indicating that the site is the conserved phosphorylation site (Fig. 1)33,34 that is required for activating kinase activity. The reduction of the upper band of the CaCdc7 doublet from the purified wild-type CaCdc7 in a time-dependent manner when treated with phosphatase confirmed that the upper band of the CaCdc7 doublet is the phosphorylated form of CaCdc7 (Fig. 4C). A reduction in the phosphorylated form of CaCdc7 was observed when cells were treated with HU for 1 h before western blot analysis (Fig. 4C). This result is in agreement with the downregulation of S. cerevisiae DDK activity mediated by the phosphorylation of Dbf4 through Rad53 in S. cerevisiae12,15,43,44 and in Xenopus egg extracts45 under DNA stress, which likely resulted from inhibiting the phosphorylation site of kinase activation.
(A) The alignment of subdomains II and VII among the homologues of Cdc7, revealing lysine 232 as a conserved residue for catalytic activity and threonine 437 as a conserved phosphoacceptor residue. These were used to generate the catalytically inactive CaCdc7 (K232R) or the phosphoacceptor-deficient CaCdc7 (T437A). (B) An in vitro kinase assay was used to verify that CaCdc7 (K232) and CaCdc7 (T437) are required for the catalytic activity and activation by phosphorylation. Cells of strain CaCDC7 M3/− were transformed with p6HF-ACT1p-CaCDC7, which is capable of constitutively expressing wild-type CaCdc7 (CaCDC7), catalytically inactive CaCdc7 (K232R) or the phosphoacceptor-deficient CaCdc7 (T437A). Cells of each strain were grown in SD medium with the required supplements in the presence (+Met/Cys) of each of 2.5 mM methionine and cysteine for 12 h prior to purification of CaCdc7 for the in vitro kinase assay, followed by western blot analysis with specific antibodies to FLAG (CaCdc7-6xHisFLAG tagged), histone H1, and phosphorylated histone H1. The purified CaCdc7 was also subjected to FSBA treatment as described in the Materials and Methods. (C) Phosphorylation of CaCdc7 is activated by hydroxyurea (HU). The cells carrying p6HF-ACT1p-CaCDC7 were grown in same the conditions and media as in (B) treated with or without 0.1 mM HU for 1 h. The purified CaCdc7 from non-HU treated cells was exposed to CIP phosphatase for the indicated times as described in the Materials and Methods.
The catalytic activity of CaCdc7 and the phosphorylated activation of CaCdc7 are required for suppression of hyphal growth in C. albicans
To investigate whether the ability of cells to suppress the YTH transition requires kinase activity and phosphorylation of DDK, CaCDC7 M3/− cells containing plasmids p6HF-ACT1p- CaCDC7, p6HF-ACT1p-CaCDC7 (K232R), p6HF-ACT1p- CaCDC7 (T437A) or the empty plasmid p6HF-ACT1p were grown in medium with or without Met/Cys. Regardless of the expression of endogenous CaCDC7 under the control of MET3p being repressed or de-repressed, the expression levels of different versions of the CaCdc7 protein driven under ACT1p control were similar (Fig. 5A). It was apparent that cells with repressed MET3p-controlled endogenous CaCDC7 expression but constitutive expression of the ACT1p-driven mutants CaCdc7 (K232R) or CaCdc7 (T437A) grew as the hyphal form (Fig. 5B), which is in contrast to those expressing ACT1p-driven wild type CaCdc7, which remained as yeast cells. These results suggest that the catalytic activity of CaCdc7 and the phosphorylation of CaCdc7 are required for the function of CaCdc7 on the suppression of the YTH transition.
(A) Cells of strain CaCDC7 M3/− were transformed with either the empty p6HF-ACT1p or p6HF-ACT1p-CaCDC7, capable of constitutively expressing wild-type CaCdc7 (CaCDC7), catalytically inactive CaCdc7 (K232R) or the phosphoacceptor-deficient CaCdc7 (T437A). Cells of each strain, together with the BWP17 from which CaCDC7 M3/− was derived, were grown in the SD medium with required supplements in the presence (+Met/Cys) or absence (−Met/Cys) of 2.5 mM methionine and cysteine for 12 h at 30 °C prior to the assessment of ACT1p-driven protein expression by western blot analysis. (B) The same cultures were collected at 12 h for the assessment of morphological alteration under a microscope. Bars represent 10 μm.
By examining cells of the strains repressing the expression of CaCDC7 in a longer time, we confirmed that the mutant strains were able to proliferate, though in a reduced rate as verified by the growth curve (Supplementary Fig. S2). The constitutive expression of CaCDC7 but none of the two CaCDC7 mutants allows recovery of this growth defect (Supplementary Fig. S2). These data indicate that cells depleted with CaCDC7 or expressing the defective CaCdc7 mutants delay the cell cycle progression. We note that cdc7 conditional mutants of S. cerevisiae show a dumbbell shape46, unlike polarized growth of C. albicans expressing the CDC7 mutations. It is known that pseudohyphae and true hyphae emerge in response to cell-cycle arrest in C. albicans47. Hence, the C. albicans expressing the mutant CDC7 may delay the cell cycle at the S phase. However, the cells were still able to grow, and the extension of hyphal development was more prominent at a later time point of 32 h (Supplementary Fig. S4). Moreover, because the cell cycle stage of cells repressing with CaCDC7 was different, the budded daughter cells, like the mother cells, switch to the hyphal mode of growth can grow as extended hyphal form (Fig. 5). These results suggest that cells depleted with CaCDC7 or expressing the defective CaCDC7 mutants delay in the cell cycle, which accompanies the reduced rate of hyphal extension. To assess further if CaCDC7 is required for YTH, cells of the strains, together with BWP17 were subjected to grow in 37 °C, a condition known to induce hyphal growth. As shown in the Supplementary Fig. S5A, however, the hyphal formation appeared in the CaCDC7-repressede but not the CaCDC7-derepressed condition at 37 °C, suggesting that the initiation and continuation of filamentation are dependent on CaCdc7 in C. albicans. Importantly, unlike the wild-type strain SC5314, BWP17was unable to induce hyphal development but pseudohyphae-like type when cultured in the SD minimum medium at 37 °C (Supplementary Fig. S5B).
We are not aware of any Cdc7 homologs across the evolutionary spectrum that are involved in morphogenesis until the latest report. Based on the GRACE strain collection48,49, the report identified over a hundred strains as negative regulators of filamentation. The expression of CaCDC7 of the GRACE strain was controlled by the tet-operator; hence, cells formed filaments under the repressed condition28. However, the kinase activity of CaCdc7 and its activation by phosphorylation has not been determined.
CDC7 and DBF4 are interdependent for the function of morphological control in C. albicans
The protein products encoded by CDC7 and DBF4 function together as a DDK for the initiation of DNA replication10. Although our study suggests that DDK in C. albicans plays a significant role in morphogenesis, we wondered whether the presence of C. albicans DDK functions in hypha-suppression in C. albicans. The definitive determination of how CaCdc7 and CaDbf4 constitute DDK required verification of their direct interaction. To do so, we adopted the yeast two-hybrid assay. The diploid S. cerevisiae cells were able to grow on media lacking leucine and tryptophan, indicating the presence of both plasmids based on the Gal4 DNA binding domain with the TRP1 selection marker and the Gal4 activation domain with the LEU2 selection marker (Fig. 6A). However, only those cells expressing CaCdc7 fused with the Gal4 DNA binding domain and CaDbf4 fused with the Gal4 activation domain concurrently were able to grow on the plate without histidine (Fig. 6B). These results suggest that CaCdc7 and CaDbf4 can associate physically. Moreover, diploid S. cerevisiae cells capable of simultaneously expressing CaCdc7 fused with the Gal4 DNA binding domain and CaDbf4 fused with the Gal4 activation domain were able to grow on the plate without histidine, even in the presence of 20 mM 3-amino-1,2,4-triazole (3-AT), the inhibitor of imidazole glycerol-phosphate dehydratase encoded by the reporter gene HIS3. These results indicate that the interaction between CaCdc7 and CaDbf4 is relatively stable and thus likely to be genuine (Fig. 6C).
(A) Verification of diploid S. cerevisiae cells carrying the plasmids required for the test. The plasmids pACT2 and pACT2-CaDBF4 (capable of expressing Gal4 activation domain fused with CaDbf4) were transformed into AH109, from which AD and AD CaDBF4 were obtained, respectively. The plasmids pGBKT7 and pGBKT7-CaCDC7 (capable of expressing Gal4 DNA biding domain fused with CaCdc7) were transformed into Y187, from which BD and BD CaCDC7 were obtained, respectively. AH109 and Y187 derivatives were mated to obtain diploid S. cerevisiae cells of AD/BD, AD CaDBF4/BD, AD/BD CaCDC7, and AD CaDBF4/BD CaCDC7. Ten-fold serially diluted S. cerevisiae diploid cells (104∼101) were grown on semi-solid agar plates with selective minimum media lacking leucine and tryptophan (SD−LW). (B) Verification of the interaction between CaCdc7 and CaDbf4. Ten-fold serially diluted S. cerevisiae diploid cells (104∼101), the same as in (A) were grown on semi-solid agar plates with selective minimum media lacking leucine, tryptophan, and histidine, supplemented with 2.5 mM 3-AT, the antagonist of the HIS3 gene product (SD−LWH+2.5 mM 3-AT). (C) Strong interaction occurs between CaCdc7 and CaDbf4. Serially diluted S. cerevisiae diploid cells of AD CaDBF4/BD CaCDC7 were grown on semi-solid agar plates with selective media lacking leucine, tryptophan, and histidine but with the indicated concentration of 3-amino-triazole (3-AT). The cells were able to proliferate normally at the concentration of 20 mM due to a relatively strong interaction between CaCdc7 and CaDbf4.
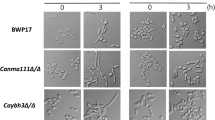
To assess the functional dependency between CaCDC7 and CaDBF4, we generated plasmids p6HF-ACT1p-CaCDC7 and p6HF-ACT1p-CaDBF4 that are capable of constitutively expressing either CaCDC7 or CaDBF4, and introduced them into CaDBF4 M3/−/−29 and CaCDC7 M3/−, respectively. The ability of a DDK gene that can be constitutively expressed to suppress the loss of the other DDK gene that is repressed in the presence of Met/Cys was assessed. Constitutive expression of CaCDC7 (Fig. 7A) did not suppress the loss of CaDBF4 (Fig. 7B), suggesting that the function of CaDBF4 requires the presence of CaCDC7. Similarly, constitutive expression of CaDBF4 (Fig. 7C) was unable to suppress the loss of CaCDC7 (Fig. 7D). The epistasis analyses and yeast two-hybrid assay results confirm that CaCDC7 and CaDBF4 encode proteins that form functional DDK and are functionally interdependent. Additionally, we introduced a Tet-on expression system50 into BWP17 where CaCDC7 or CaDBF4 were massively overproduced in the presence of doxycycline under serum-induced hyphal growth conditions. Cells under such conditions grew as hyphae (Supplemmentary Fig. S6), suggesting that either DDK control of the yeast-to-hypha transition is not via the serum-induced signaling pathway, but instead through other pathways, such as DNA replication stress, or that the blockage of serum-induced filamentous growth could be bypassed by other factors. It is equally possible that DDK control of the yeast-to-hypha transition constitutes a novel pathway.
Cells of strain CaCDC7 M3/− and those with either the empty p6HF-ACT1p or p6HF-ACT1p-CaDBF4 and p6HF-ACT1p-CaCDC7, which are capable of constitutively expressing CaDbf4 and CaCdc7, respectively, were grown in SD medium with the required supplements in the presence (+Met/Cys) or absence (−Met/Cys) of 2.5 mM methionine and cysteine for 12 h prior to the assessment of protein expression by western blotting analysis (A) and assessment of the morphological consequences microscopically (B). Cells of strain CaDBF4 M3/−/− (ref) and those with either the empty p6HF-ACT1p or p6HF-ACT1p-CaCDC7 and p6HF-ACT1p-CaDBF4, which are capable of constitutively expressing wild-type CaCdc7 and CaDbf4, respectively, were grown in the same conditions as above prior to the assessment of protein expression (C) and microscopic assessment of the morphological consequences (D). β-Actin was used as a loading control in the analyses of protein expression by western blotting. Bars represent 10 μm in the microscopic observations.
Conclusion
In this paper, we describe the characterization of the CaCDC7 gene. To our surprise, we found that C. albicans CDC7 plays a role as a negative regulator of the YTH transition. Additionally, kinase activity is required for the function of CaCdc7 and the presence of other kinases for the function of CaCdc7. Moreover, the function of CaCdc7 is dependent on CaDbf4 for suppression of the hyphal mode of growth. To the best of our knowledge, no known role for Dbf4-dependent Cdc7 homologs across the evolutionary spectrum relevant to morphogenesis has been identified. The uncovering of CaCdc7 function in morphogenesis may be of evolutionary significance in that essential elements in the cell cycle have evolved in the direction of morphogenesis for adapting the host-pathogen interaction. It would be crucial to elucidate the regulation of DDK in the network controlling morphogenesis. Elucidating the regulation of DDK in the system that controls morphogenesis should lend insight into understanding why DDK is involved in morphogenesis.
Methods
General manipulation, media and growth conditions
The E. coli strain DH5α was used for routine manipulation of the plasmids, and all C. albicans strains (Table 1) were derived from the auxotrophic strain BWP17 (arg4/arg4 his1/his1 ura3/ura3)51. The wild-type SC5314 strain52 was also used. The media usage and routine growth conditions of the strains of E. coli and C. albicans were as previously described53. E. coli strain DH5α was transformed with plasmid DNA by CaCl2 as described54 or by electroporation55. Candida albicans strains were transformed by the LiAc/PEG/ssDNA method56 or electroporation57.
Strain construction
The nucleotide sequence of C. albicans was obtained from the Candida Genome Database (http://www.candidagenome.org/). The CaCDC7 gene including regions flanking the protein coding sequence was PCR-generated with the primers CaCDC7-xba-F and CaCDC7-xba-R (Table 2) and genomic DNA extracted from C. albicans strain BWP17. The CaCDC7 gene was cloned into the plasmid vector pBluescript II (+) to obtain the plasmid pBII-CaCDC7. CaCDC7 was deleted in the C. albicans auxotrophic strain BWP17 with the mini-Ura-blaster cassette dpl200-URA3-dpl200 derived from pDDB5739. Briefly, the mini-Ura-blaster cassette flanked by 60 bp of upstream and downstream homology to the CaCDC7 open reading frame was amplified by PCR with the template pBII-CaCDC7 and a pair of primers, CaCDC7-Spe-F and CaCDC7-Spe-R (Table 2). The mini-Ura-blaster cassette flanked by the short homology regions of CaCDC7 was then transformed into BWP17 cells and the uridine prototrophic strains (Ura+) CaCDC7+/U3− (Table 1) containing the heterozygous deletion of CaCDC7 were selected for. To allow CaCDC7 expression under the control of MET3p, a partial CaCDC7 ORF (1–884 bp) flanked by an SpeI site was PCR generated with the template pBII-CaCDC7 and a pair of primers, CaCDC7-spe-F and CaCDC7-spe-R (Table 2). The PCR product was cloned into pFA-HIS1-MET3p40 at the SpeI site and the plasmid pFA-HIS1-MET3p-CaCDC7 was obtained. Plasmid pFA-HIS1-MET3p-CaCDC7 was linearized by digesting with a unique EcoRI on partial CaCDC7 and was transformed into CaCDC7+/U3− to generate the His+ prototrophic strain CaCDC7 M3/U3−. To remove the URA3 marker, the cells of CaCDC7 M3/U3− were then treated with 1 mg/ml 5-FOA to generate CaCDC7 M3/− (Table 1). To repress the CaCDC7 expression that is controlled by MET3p, strains were grown in SD medium or on a plate with 2.5 mM Met/Cys, which turns off the expression of MET3p-driven downstream genes40. To allow constitutive expression of CaCDC7 in C. albicans cells, the protein coding sequence of CaCDC7 was cloned from the plasmid pBII-CaCDC7 with a pair of primers, CaCDC7-Sph-F and CaCDC7-Sph-R (Table 2), and introduced into plasmid p6HF-ACT1p58 to generate p6HF-ACT1p-CaCDC7. The p6HF-ACT1p-CaCDC7 and the empty plasmid p6HF-ACT1p were linearized with NcoI and introduced into CaCDC7 M3/−. These cells were selected for Ura+ transformants that targeted and integrated at the RP10 locus to generate CaCDC7M3/−|CaCDC7 and CaCDC7M3/−|p6HF-ACT1p (Table 2), respectively. The linearized plasmids were also introduced into CaDBF4 M3/−/−29 to obtain CaDBF4 M3/−/−|CaCDC7 and CaDBF4 M3/−/−|p6HF-ACT1p (Table 2), respectively. In addition, the linearized plasmid p6HF-ACT1p-CaDBF4 (manuscript submitted) was introduced into CaCDC7 M3/− to generate CaCDC7M3/−|CaDBF4.
Site-directed mutagenesis
The QuickChangeTM Site-Directed Mutagenesis Kit (Stratagene, San Diego, CA) was used as instructed by the manufacturer to introduce mutations in CaCDC7 using the plasmid p6HF-ACT1p-CaCDC7. Following site-directed mutagenesis with two pairs of primers, CaCDC7-K232R-F/CaCDC7-K232R-R and CaCDC7-T436A-F/ CaCDC7-T436A-R (Table 2), the plasmids p6HF–ACT1p-CaCDC7 (K232R) and p6HF-ACT1p- CaCDC7 (T437A) were generated, respectively. Each of the plasmids (p6HF–ACT1p-CaCDC7 (K232R), p6HF-ACT1p–CaCDC7 (T437A) was NcoI-linearized and introduced into C. albicans strain CaCDC7 M3/− and targeted and integrated at the RP10 locus to generate CaCDC7M3/−|K232R and CaCDC7M3/−|T437A, respectively.
Yeast two-hybrid assay
The yeast two-hybrid assay was used. Plasmid constructs capable of expressing the Gal4 activation domain fused N-terminally to CaDbf4 (pACT2-CaDBF4) or the Gal4 DNA binding domain fused N-terminally to CaCdc7 (pGBKT7-CaCDC7) were made using genomic DNA extracted from BWP17 with two pairs of oligonucleotides, CaDBF4-XmaI-F and CaDBF4-XhoI-R (Table 2) and CaCDC7-NcoI(hybrid)-F and CaCDC7-PstI(hybrid)-R (Table 2), respectively. The plasmids pGBKT7-CaCDC7 and pACT2-CaDBF4 were introduced into the haploid S. cerevisiae strain Y18759 and AH109, a derivative of strain PJ69-2A60 resulting from the introduction of the lacZ reporter gene into PJ69-2A (Table 1), respectively, with opposite mating types and with the reporter systems HIS3, ADE2, and LacZ. These strains were used to determine the activation of the system and the interaction between CaCdc7 and CaDbf4 upon fusion to become diploid.
Isolation of genomic DNA and Southern blotting
Genomic DNA from the C. albicans strains was isolated by the MasterPureTM Yeast DNA Purification Kit (EPICENTRE). Southern blotting was performed according to standard protocols with the aid of the Rapid Downward Transfer System (TURBOBLOTTERTM) using 10 μg of the restriction enzyme-digested genomic DNA. The probe was generated by the PCR DIG probe synthesis kit (Roche) with a genomic DNA template extracted from BWP17 and a pair of primers, CaCDC7-Probe-F and CaCDC7-Probe-R (Table 2). The DNA on the blot was hybridized with the probe using DIG Easy Hyb (Roche). To reveal the gene deletion, the DIG Luminescent Detection Kit (Roche) was used after hybridization, and the blot was exposed to X-ray film for up to 24 h as appropriate.
RT-PCR analysis
Cells were grown to mid-log phase and total RNA was extracted using the MasterPureTM Yeast RNA Purification Kit (EPICENTRE) following the manufacturer’s instructions. Then, 5 μg of total RNA was used to generate cDNA by using the SuperScript III Reverse Transcriptase Kit (Invitrogen) following the manufacturer’s instructions. The cDNA was then subjected to PCR with a pair of CaCDC7-specific primers, CaCDC7-internal-F-2 and CaCDC7-BspEI-R (Table 2), targeted downstream of the coding sequence. A 366 bp product was generated.
Immunoblot analysis
The total protein was extracted from cultured cells as previously described61. The protein was partially purified from cells bearing the p6HF-ACT1p plasmid with the open reading frame of the gene integrated at RP10 capable of generating a tagged (6 × His and FLAG) protein using Ni2+-NTA-agarose beads (Qiagen Inc.) essentially as previously described61. Precipitated proteins were resolved by 10% SDS-PAGE and transferred electrophoretically to PVDF membranes (PerkinElmer, Boston, USA) and probed with a polyclonal antibody to FLAG (Sigma) in a 1:2000 dilution and visualized using the SuperSignal West Pico Chemiluminescent Substrate Kit (PIERCE). The detected proteins were recorded with the Luminescent Image Analyzer (FUJIFILM LAS-1000) and analyzed by ImageGauge 3.46 and L Process v 1.96 (FUJIFILM).
In vitro kinase assay
The in vitro kinase assays using CaCdc7-6xHisFlAG and purified bovine histone H1 (Upstate) as substrates were performed using a non-radioactive assay essentially as previously described62. Briefly, the Ni2+-NTA-agarose bead (Qiagen Inc.)-purified CaCdc7 was washed three times with kinase reaction buffer [25 mM Tris pH 7.5, 5 mM β-glycerophosphate, 2 mM DTT, 0.1 mM Na3VO4, 10 mM MgCl2 and a serine/threonine phosphatase inhibitor cocktail (Calbiochem)] and used in the kinase reaction with 5 μg histone H1 as substrate in the presence of 200 μM ATP in a 20 μl kinase assay buffer. The kinase reaction was incubated at 30 °C for 45 minutes, and the end products were resolved by 12% SDS-PAGE and detected by immunoblotting using anti-Histone H1 (H11-4, Sigma; 1:500) and anti-phospho-Histone H1 (12D11, Millipore; 1:1000 monoclonal antibodies) according to the manufacturer’s instructions.
Chemical treatment
Phosphatase treatment was performed with calf intestinal alkaline phosphatase (CIP) (NEB). Briefly, the Ni2+-NTA-agarose bead (Qiagen Inc.)-purified CaCdc7 was washed three times with CIP buffer (1XNE Buffer 3) and resuspended in 20 μl CIP buffer before the addition of 20 units of CIP and incubation at 37 °C for the required period of time. The CIP was inactivated by heating to 75 °C for 10 minutes in the presence of 5 mM EDTA. Irreversible protein kinase inhibitor treatment was performed with 5′-fluorosulfonylbenzoyl-5′-adenosine (FSBA), essentially as previously described63. Briefly, the Ni2+-NTA- agarose bead-purified CaCdc7 was subjected to three successive treatments by 1 mM FSBA (Sigma) at 30 °C for 15 min in kinase buffer without DTT. The product was then washed three times with kinase buffer to eliminate the FSBA before performing the in vitro kinase assay. Hydroxyurea (HU) was used to block DNA replication, which is known to attenuate Cdc7 kinase activity15. Cells in culture were treated with 0.1 M HU before the Ni2+-NTA- agarose bead purification of CaCdc7.
Cellular image observation and recording
Cells in liquid culture were visualized and recorded with a Nikon 50i microscope at 400x magnification. Colonies were photographed with a MEIJI stereoscopic microscope EMZ5 at 40x magnification.
Additional Information
How to cite this article: Lai, W.-C. et al. Candida albicans Dbf4-dependent Cdc7 kinase plays a novel role in the inhibition of hyphal development. Sci. Rep. 6, 33716; doi: 10.1038/srep33716 (2016).
References
Sudbery, P., Gow, N. & Berman, J. The distinct morphogenic states of Candida albicans. Trends Microbiol 12, 317–324, doi: 10.1016/j.tim.2004.05.008 (2004).
Noble, S. M., French, S., Kohn, L. A., Chen, V. & Johnson, A. D. Systematic screens of a Candida albicans homozygous deletion library decouple morphogenetic switching and pathogenicity. Nat Genet 42, 590–598, doi: 10.1038/ng.605 (2010).
Schinabeck, M. K. et al. Rabbit model of Candida albicans biofilm infection: liposomal amphotericin B antifungal lock therapy. Antimicrob Agents Chemother 48, 1727–1732 (2004).
Sudbery, P. E. Growth of Candida albicans hyphae. Nat Rev Microbiol 9, 737–748, doi: 10.1038/nrmicro2636 (2011).
Sherlock, G. et al. Molecular cloning and analysis of CDC28 and cyclin homologues from the human fungal pathogen Candida albicans. Mol Gen Genet 245, 716–723 (1994).
Atir-Lande, A., Gildor, T. & Kornitzer, D. Role for the SCFCDC4 Ubiquitin Ligase in Candida albicans Morphogenesis. Mol Biol Cell 16, 2772–2785 (2005).
Shieh, J. C., White, A., Cheng, Y. C. & Rosamond, J. Identification and functional characterization of Candida albicans CDC4. J Biomed Sci 12, 913–924, doi: 10.1007/s11373-005-9027-9 (2005).
Tseng, T. L. et al. Affinity purification of Candida albicans CaCdc4-associated proteins reveals the presence of novel proteins involved in morphogenesis. Biochem Biophys Res Commun 395, 152–157, doi: 10.1016/j.bbrc.2010.03.162 (2010).
Hartwell, L. H. Three additional genes required for deoxyribonucleic acid synthesis in Saccharomyces cerevisiae. J Bacteriol 115, 966–974 (1973).
Jackson, A. L., Pahl, P. M., Harrison, K., Rosamond, J. & Sclafani, R. A. Cell cycle regulation of the yeast Cdc7 protein kinase by association with the Dbf4 protein. Mol Cell Biol 13, 2899–2908 (1993).
Masai, H. & Arai, K. Cdc7 kinase complex: a key regulator in the initiation of DNA replication. J Cell Physiol 190, 287–296 (2002).
Zegerman, P. & Diffley, J. F. Checkpoint-dependent inhibition of DNA replication initiation by Sld3 and Dbf4 phosphorylation. Nature 467, 474–478, doi: 10.1038/nature09373 (2010).
Bousset, K. & Diffley, J. F. The Cdc7 protein kinase is required for origin firing during S phase. Genes Dev 12, 480–490 (1998).
Yamada, M. et al. ATR-Chk1-APC/CCdh1-dependent stabilization of Cdc7-ASK (Dbf4) kinase is required for DNA lesion bypass under replication stress. Genes Dev 27, 2459–2472, doi: 10.1101/gad.224568.113 (2013).
Weinreich, M. & Stillman, B. Cdc7p-Dbf4p kinase binds to chromatin during S phase and is regulated by both the APC and the RAD53 checkpoint pathway. EMBO J 18, 5334–5346, doi: 10.1093/emboj/18.19.5334 (1999).
Matsumoto, S. & Masai, H. Regulation of chromosome dynamics by Hsk1/Cdc7 kinase. Biochem Soc Trans 41, 1712–1719, doi: 10.1042/BST20130217 (2013).
Jiang, Y. W. & Kang, C. M. Induction of S. cerevisiae filamentous differentiation by slowed DNA synthesis involves Mec1, Rad53 and Swe1 checkpoint proteins. Mol Biol Cell 14, 5116–5124, doi: 10.1091/mbc.E03-06-0375 (2003).
McIntosh, E. M., Kunz, B. A. & Haynes, R. H. Inhibition of DNA replication in Saccharomyces cerevisiae by araCMP. Curr Genet 10, 579–585 (1986).
Tercero, J. A. & Diffley, J. F. Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 412, 553–557, doi: 10.1038/35087607 (2001).
Bachewich, C. & Whiteway, M. Cyclin Cln3p links G1 progression to hyphal and pseudohyphal development in Candida albicans. Eukaryot Cell 4, 95–102 (2005).
Loll-Krippleber, R. et al. A study of the DNA damage checkpoint in Candida albicans: uncoupling of the functions of Rad53 in DNA repair, cell cycle regulation and genotoxic stress-induced polarized growth. Mol Microbiol 91, 452–471, doi: 10.1111/mmi.12471 (2014).
da Silva Dantas, A. et al. Thioredoxin regulates multiple hydrogen peroxide-induced signaling pathways in Candida albicans. Mol Cell Biol 30, 4550–4563, doi: 10.1128/MCB.00313-10 (2010).
Shi, Q. M., Wang, Y. M., Zheng, X. D., Lee, R. T. & Wang, Y. Critical role of DNA checkpoints in mediating genotoxic-stress-induced filamentous growth in Candida albicans. Mol Biol Cell 18, 815–826, doi: 10.1091/mbc.E06-05-0442 (2007).
Andaluz, E., Ciudad, T., Gomez-Raja, J., Calderone, R. & Larriba, G. Rad52 depletion in Candida albicans triggers both the DNA-damage checkpoint and filamentation accompanied by but independent of expression of hypha-specific genes. Mol Microbiol 59, 1452–1472, doi: 10.1111/j.1365-2958.2005.05038.x (2006).
Hazan, I., Sepulveda-Becerra, M. & Liu, H. Hyphal elongation is regulated independently of cell cycle in Candida albicans. Mol Biol Cell 13, 134–145, doi: 10.1091/mbc.01-03-0116 (2002).
Umeyama, T. et al. Candida albicans protein kinase CaHsl1p regulates cell elongation and virulence. Mol Microbiol 55, 381–395, doi: 10.1111/j.1365-2958.2004.04405.x (2005).
Wightman, R., Bates, S., Amornrrattanapan, P. & Sudbery, P. In Candida albicans, the Nim1 kinases Gin4 and Hsl1 negatively regulate pseudohypha formation and Gin4 also controls septin organization. J Cell Biol 164, 581–591, doi: 10.1083/jcb.200307176 (2004).
O’Meara, T. R. et al. Global analysis of fungal morphology exposes mechanisms of host cell escape. Nat Commun 6, 6741, doi: 10.1038/ncomms7741 (2015).
Chien, T. et al. Candida albicans DBF4 gene inducibly duplicated by the mini-Ura-blaster is involved in hypha-suppression. Mutat Res 779, 78–85, doi: 10.1016/j.mrfmmm.2015.06.013 (2015).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948, doi: 10.1093/bioinformatics/btm404 (2007).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22, 4673–4680 (1994).
Hanks, S. K. & Quinn, A. M. Protein kinase catalytic domain sequence database: identification of conserved features of primary structure and classification of family members. Methods Enzymol 200, 38–62 (1991).
Buck, V., White, A. & Rosamond, J. CDC7 protein kinase activity is required for mitosis and meiosis in Saccharomyces cerevisiae. Mol Gen Genet 227, 452–457 (1991).
Sclafani, R. A. & Jackson, A. L. Cdc7 protein kinase for DNA metabolism comes of age. Mol Microbiol 11, 805–810 (1994).
Masai, H. et al. Human Cdc7-related kinase complex. In vitro phosphorylation of MCM by concerted actions of Cdks and Cdc7 and that of a criticial threonine residue of Cdc7 bY Cdks. J Biol Chem 275, 29042–29052, doi: 10.1074/jbc.M002713200 (2000).
Bahman, M., Buck, V., White, A. & Rosamond, J. Characterisation of the CDC7 gene product of Saccharomyces cerevisiae as a protein kinase needed for the initiation of mitotic DNA synthesis. Biochim Biophys Acta 951, 335–343 (1988).
Patterson, M., Sclafani, R. A., Fangman, W. L. & Rosamond, J. Molecular characterization of cell cycle gene CDC7 from Saccharomyces cerevisiae. Mol Cell Biol 6, 1590–1598 (1986).
Sato, N., Arai, K. & Masai, H. Human and Xenopus cDNAs encoding budding yeast Cdc7-related kinases: in vitro phosphorylation of MCM subunits by a putative human homologue of Cdc7. Embo J 16, 4340–4351 (1997).
Wilson, R. B., Davis, D., Enloe, B. M. & Mitchell, A. P. A recyclable Candida albicans URA3 cassette for PCR product-directed gene disruptions. Yeast 16, 65–70 (2000).
Care, R. S., Trevethick, J., Binley, K. M. & Sudbery, P. E. The MET3 promoter: a new tool for Candida albicans molecular genetics. Mol Microbiol 34, 792–798 (1999).
Cao, J. X. et al. MiR-630 inhibits proliferation by targeting CDC7 kinase, but maintains the apoptotic balance by targeting multiple modulators in human lung cancer A549 cells. Cell Death Dis 5, e1426, doi: 10.1038/cddis.2014.386 (2014).
Rodriguez-Acebes, S. et al. Targeting DNA replication before it starts: Cdc7 as a therapeutic target in p53-mutant breast cancers. Am J Pathol 177, 2034–2045, doi: 10.2353/ajpath.2010.100421 (2010).
Kihara, M. et al. Characterization of the yeast Cdc7p/Dbf4p complex purified from insect cells. Its protein kinase activity is regulated by Rad53p. J Biol Chem 275, 35051–35062, doi: 10.1074/jbc.M003491200 (2000).
Lopez-Mosqueda, J. et al. Damage-induced phosphorylation of Sld3 is important to block late origin firing. Nature 467, 479–483, doi: 10.1038/nature09377 (2010).
Costanzo, V. et al. An ATR- and Cdc7-dependent DNA damage checkpoint that inhibits initiation of DNA replication. Mol Cell 11, 203–213 (2003).
Hartwell, L. H., Mortimer, R. K., Culotti, J. & Culotti, M. Genetic Control of the Cell Division Cycle in Yeast: V. Genetic Analysis of cdc Mutants. Genetics 74, 267–286 (1973).
Berman, J. Morphogenesis and cell cycle progression in Candida albicans. Curr Opin Microbiol 9, 595–601, doi: 10.1016/j.mib.2006.10.007 (2006).
Roemer, T. et al. Large-scale essential gene identification in Candida albicans and applications to antifungal drug discovery. Mol Microbiol 50, 167–181 (2003).
Ryan, O. et al. Global gene deletion analysis exploring yeast filamentous growth. Science 337, 1353–1356, doi: 10.1126/science.1224339 (2012).
Lai, W. C. et al. Construction of Candida albicans Tet-on tagging vectors with a Ura-blaster cassette. Yeast 28, 253–263, doi: 10.1002/yea.1833 (2011).
Wilson, R. B., Davis, D. & Mitchell, A. P. Rapid hypothesis testing with Candida albicans through gene disruption with short homology regions. J Bacteriol 181, 1868–1874 (1999).
Gillum, A. M., Tsay, E. Y. & Kirsch, D. R. Isolation of the Candida albicans gene for orotidine-5′-phosphate decarboxylase by complementation of S. cerevisiae ura3 and E. coli pyrF mutations. Mol Gen Genet 198, 179–182 (1984).
Chin, C., Lai, W. C., Lee, T. L., Tseng, T. L. & Shieh, J. C. Dissection of the Candida albicans Cdc4 protein reveals the involvement of domains in morphogenesis and cell flocculation. J Biomed Sci 20, 97, doi: 10.1186/1423-0127-20-97 (2013).
Warren, G. & Sherratt, D. Incompatibility and transforming efficiency of ColE1 and related plasmids. Mol Gen Genet 161, 39–47 (1978).
Dower, W. J., Miller, J. F. & Ragsdale, C. W. High efficiency transformation of E. coli by high voltage electroporation. Nucleic Acids Res 16, 6127–6145 (1988).
Gietz, R. D. & Woods, R. A. Yeast transformation by the LiAc/SS Carrier DNA/PEG method. Methods Mol Biol 313, 107–120 (2006).
Becker, D. M. & Lundblad, V. Introduction of DNA into yeast cells. Curr Protoc Mol Biol Chapter 13, Unit13 17, doi: 10.1002/0471142727.mb1307s27 (2001).
Kaneko, A. et al. Tandem affinity purification of the Candida albicans septin protein complex. Yeast 21, 1025–1033 (2004).
Harper, J. W., Adami, G. R., Wei, N., Keyomarsi, K. & Elledge, S. J. The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 75, 805–816, doi: 0092-8674(93)90499-G (1993).
James, P., Halladay, J. & Craig, E. A. Genomic libraries and a host strain designed for highly efficient two-hybrid selection in yeast. Genetics 144, 1425–1436 (1996).
Shieh, J. C. et al. Tailor-made zinc-finger transcription factors activate FLO11 gene expression with phenotypic consequences in the yeast Saccharomyces cerevisiae. PLoS ONE 2, e746, doi: 10.1371/journal.pone.0000746 (2007).
Maddika, S. et al. Akt-mediated phosphorylation of CDK2 regulates its dual role in cell cycle progression and apoptosis. J Cell Sci 121, 979–988, doi: 10.1242/jcs.009530 (2008).
Lopez-Girona, A., Mondesert, O., Leatherwood, J. & Russell, P. Negative regulation of Cdc18 DNA replication protein by Cdc2. Mol Biol Cell 9, 63–73 (1998).
Acknowledgements
The authors thank Dr. A. Mitchell (Columbia University, USA) for C. albicans strain BWP17 and plasmid pDDB57, Dr. A.J.P. Brown (University of Aberdeen, United Kingdom) for C. albicans strain SC5314, Dr. J. Wendland (Friedrich-Schiller- University, Jena, Germany) for pFA-HIS1-MET3p, and Dr. Dr. M. Niimi (National Institute of Infectious Diseases, Tokyo, Japan) for p6HF-ACT1p. The DIG-based Southern blotting assay was established with the help of Dr. JH Lo (the National Health Research Institutes, Taiwan). We thank Hsiao-Chi Hsu for technical assistance in generating the growth curves. We gratefully acknowledge the support for this work provided by grants from the National Science Council of Taiwan, Republic of China to J.C.S. (NSC 97-2320-B-040-014-MY3 & NSC 101-2629-B-040-001-MY3), C.H.W. (NSC 96-2815-C-040-026-B), and S.Y.Y (NSC 98-2815-C-040 -045-B), from the Ministry of Science and Technology of Taiwan, Republic of China to J.C.S. (MOST 105-2320-B-040-027-MY3), and from the National Health Research Institutes of Taiwan, Republic of China to J.C.S. (NHRI-EX99-9808SI & NHRI-EX100-9808SI).
Author information
Authors and Affiliations
Contributions
J.-C.S. conceived of and designed the study and supervised the project. Wei-Chung Lai (W.-C.L.) made critical analyses of the western blotting and the Southern blotting as well as morphological observation and growth evaluation. T.-w.C., C.-H.W. and Y.-C.C. established the initial strains and verified the phenotypes. S.-Y.Y. and T.C. constructed the strains and performed the Southern blotting analysis and microscopic observations. S.-Y.Y performed site-directed mutagenesis and the yeast two-hybrid analysis manuscript. Wan Chen Li (W.C.L.), S.-Y.Y. performed the western blotting analysis. T.-L.L. provided critical reagents and advice. All authors analyzed the data, discussed the results, and commented on the manuscript. J.-C.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lai, WC., Chang, Tw., Wu, C. et al. Candida albicans Dbf4-dependent Cdc7 kinase plays a novel role in the inhibition of hyphal development. Sci Rep 6, 33716 (2016). https://doi.org/10.1038/srep33716
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33716
- Springer Nature Limited