Abstract
Tight and accurate regulation of immunity and thiamine biosynthesis is critical for proper defence mechanisms and several primary metabolic cycles in plants. Although thiamine is known to enhance plant defence by priming, the mechanism by which thiamine biosynthesis responds to immune signals remains poorly understood. Here we identified a novel rice (Oryza sativa L.) NB-LRR gene via an insertion mutation, this mutant confesses a low seed setting phenotype and the corresponding genetic locus was named OsLSR (Low seed setting related). Comparing with wildtype plant, both overexpression and suppression of OsLSR lead to the autoactivation of the rice immune system and accumulation of thiamine, which result in a great fitness cost and yield penalty. Moreover, when fused with eGFP at their C terminus, two fragments, OsLSR1-178 and OsLSR464-546, localized to chloroplasts where thiamine is produced. Our result suggests that OsLSR differs from traditional NB-LRR genes. Its expression is closely related to the immune status and thiamine level in plant cells and should be maintained within a narrow range for rice growth.
Similar content being viewed by others
Introduction
Plants continually face attempted pathogen invasions and, in response, they use multiple mechanisms to fight against pathogen attacks. The resistance (R) gene-mediated defence response is one of these strategies. Most plant R genes belong to the nucleotide-binding leucine-rich repeat (NB-LRR) gene family.
A typical NB-LRR protein contains a C-terminal LRR domain, a central NB domain and a variable N-terminal domain, which usually exhibits either a Toll/Interleukin-1 receptor (TIR) domain or a coiled-coil (CC) domain1. Most NB-LRR proteins function as intracellular immune receptors and are expressed at a constant basal level in plant cells. They detect pathogen-derived effectors either directly or indirectly and then activate immune responses, including the accumulation of the defence hormones salicylic acid (SA) or jasmonic acid (JA), the production of reactive oxygen species (ROS) and the secretion of numerous PR (pathogenesis-related) proteins. The activation of these various responses eventually leads to localized killing of infected cells and restricted pathogen spread2,3.
NB-LRR proteins can activate the defence response in multiple subcellular compartments. Arabidopsis RPM1 is located at the plasma membrane, where it activates and triggers defence responses4,5. Some NB-LRR proteins need to be relocated to the nucleus to trigger full resistance during activation, for example, barley MLA10 and tobacco N6,7. The chloroplast plays an important role in defence signalling because it is involved in the generation of defence signalling molecules, such as ROS, JA and SA8. While some NB-LRR proteins were predicted to be chloroplastic9, to date, the chloroplast localization of a NB-LRR protein lacks experimental evidence.
The chloroplast is also where thiamine (vitamin B1) is created10. Thiamine and its phosphorylated forms play a fundamental role as an enzymatic cofactor in plant metabolism, including in the Calvin Benson cycle (C3), the pentose phosphate pathway (PPP) and the tricarboxylic acid cycle (TCA). Furthermore, thiamine is well known to trigger the immune response in plants11. Thiamine enhances plant disease resistance by priming, leading to a rapid counter attack against pathogen invasion and perturbation of disease progression12. Several studies have further shown that thiamine’s ability to enhance or trigger disease resistance is SA-dependent, as this induction or priming fails in mutants that do not accumulate SA12,13,14. Thiamine is de novo synthesized in plants and bacteria. In plants, thiamine is biosynthesized through the separate formation of HMP-PP (4-amino-2-methyl-5-hydroxymethylpyrimidine diphosphate) and HET-P (4-methyl-5-b-hydroxyethylthiazole phosphate); these two molecules then couple to form thiamine monophosphate (TMP) that is subsequently phosphorylated to form thiamine triphosphate (TPP), which is the active form of vitamin B110. Plants encode three enzymes to catalyse these committed steps: HET-P synthase (THI4), HMP-P synthase (THIC) and a bifunctional enzyme HMPPK (HMP-P kinase). HET-P biosynthesis is similar to that in yeast, in which HET-P synthase (THI4p) catalyses the formation of the thiazole moiety from NAD+, glycine and a sulphur from a backbone cysteine within itself15. Homologs of plant THI4 (THI1) have been cloned from maize16, Alnusglutinosa17, Arabidopsis thaliana18 and Oryza sativa19. The HMP-P synthase THIC converts aminoimidazole ribonucleotide (AIR) to hydroxymethylpyrimidine phosphate (HMP-P). THICs have been characterized in Arabidopsis20 and tomato21. HMP-P is further phosphorylated to HMP-PP by HMPPK and the latter catalyses the condensation of HET-P and HMP-PP to form TMP. All these processes occur in the chloroplast22. HMPPKs have been characterized in Arabidopsis as TH123, Brassica napus (BTH1)24 and Z.mays (THI3)25. TMP is dephosphorylated to thiamine, which is then phosphorylated by thiamine phosphorylase (TPKs) to become the active form TPP. Arabidopsis encodes two thiamine phosphorylases, TPK1 and TPK226.
Thiamine biosynthesis is tightly and negatively regulated by TPP through a riboswitch in the THIC mRNA27. A high level of TPP in the cell leads to an increased level of unstable THIC mRNA in which the second intron in the 3′UTR is spliced out. When the TPP level is low, this intron is retained and results in enhanced stability of the mRNA, which increases the translation of THIC protein and consequently increases thiamine biosynthesis. A recent study showed that overexpression of the enzyme transketolase (TK) involved in the C3 cycle provides precursors for producing thiamine, leading to thiamine auxotrophy in tobacco28. This result suggests that the thiamine level in a plant cell can also be affected by a basic metabolic cycle. Thiamine biosynthesis is activated during a plant’s response to abiotic and biotic stresses, including oxidative, salt and osmotic stress29,30 and also by pathogen inoculation19. A recent study found that thiamine biosynthesis in response to abiotic stress is mediated by ABA at an early stage31. However, the mechanism by which thiamine biosynthesis responds to immune signals remains to be elucidated.
In our current study, we characterized the NB-LRR gene OsLSR (Gene Bank Accession No. Os10g0183000), both of its up- and down- regulation triggering immune response of rice plants. These plants constitutively express PR genes, accumulate thiamine and showed a weakness. RT-qPCR analysis of thiamine biosynthesis related genes revealed that OsLSR overexpression lines accumulate thiamine by enhanced expression of OsDR8 (Defence-response protein 8), while RNA silence of OsLSR leads to a boost in OsTHIC transcripts. Our results broaden our understanding of the function of the NB-LRR protein and suggest a possible link between the NB-LRR-mediated immune response and thiamine accumulation, which could be used in rice improvement.
Results
OsLSR encodes a CC-NB-LRR protein
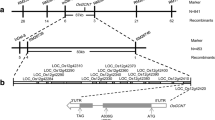
OsLSR (Low Seed setting Related) was initially cloned as genomic sequences flanking the transgene insertion site in the Bt T51-1 line, which has an MH63 background32,33. The exogenous Bt gene was inserted within its promoter region at a distance of 797 bp upstream of the start codon ATG. The T51-1 line showed a yellowish colour with a necrotic tip at the seeding stage (Fig. 1a,b). T51-1 also showed weakness in seed setting and was 10–20% lower than the wild type controls (Supplementary Table S1).
Up regulation of OsLSR and its effects on rice morphology and activation of the immune response.
(a) Yellowish leaf phenotype of T51-1 compared with MH63. (b) Leaf tip necrosis in T51-1. (c) Transcript levels of OsLSR and disease-resistant marker genes in MH63 and T51-1. For the expression analysis, OsUBQ1 was used as the internal control. Scale bars indicate the means ± SE (n = 3), *P < 0.05(ANOVA) comparing with T51-1 and MH63.
A protein sequence analysis revealed that OsLSR encodes a putative disease resistance-like protein (R protein) that has an N-terminal CC domain (1–128aa), a C-terminal LRR domain (458–894aa) and a central NB-ARC domain (165–457aa) (a nucleotide-binding domain shared by APAF-1, certain R gene products and CED-4) (http://www.ebi.ac.uk/interpro/).
Up-regulation of OsLSR triggers the rice immune response
To determine the effect of the insertion, the OsLSR transcripts were analysed by RT-qPCR in T51-1 leaf tissue. OsUBQ1 was used as an internal control and the analysis showed a slight increase of OsLSR in T51-1 compared to MH63 (Fig. 1c). Thus, OsLSR is up-regulated in T51-1.
The up-regulation of NB-LRR genes often triggers immune responses34,35. An activated immune system is often accompanied by high expression of pathogenesis-related (PR) and defence-related genes35,36. Thus, we examined the transcripts of several pathogenesis-related (PR) genes in T51-1 and MH63 at the three-leaf stage. Although these seedlings were free from pathogens, the results showed that the expression of OsPR1a, OsPR4a and OsPAD4 in T51-1 were up-regulated compared to MH63 (Fig. 1c); OsPR2 was unchanged; and OsPR5, OsPR8 and OsPR10a decreased. These data indicate that OsLSR up-regulation activates the expression of some disease response genes and the immune system in T51-1 is activated at a low level. The metabolic investment of a plant for a constitutively activated immune system consequently results in a high fitness cost and yield penalty37,38. Thus, the weakness of T51-1 can be attributed to its activated immune system.
Transient expression of OsLSR triggers hypersensitive response liked cell death in tobacco leaf
Transient overexpression of plant NB-LRR commonly leads to a hypersensitive response, which results in programmed cell death of infiltrated cells in tobacco leaf. Thus, we transiently expressed OsLSR::eGFP under control of the 35 S promoter in tobacco Nicotiana benthamiana. The construct triggered localized cell death in tobacco leaf (Fig. 2a–b), which was accompanied by H2O2 production and NtPR1 expression in the infiltration area (Supplementary Fig. S1). This result demonstrated that transient overexpression of OsLSR can trigger the plant immune response and programmed cell death.
OsLSR induces HR-like cell death in tobacco leaf and activation of thiamine biosynthesis in rice.
(a,b) OsLSR transient expression assays. HR-like cell death appears in the infiltration area of OsLSR::eGFP (a), eGFP (b) was used as a control. (c) Expression of rice genes homologous to plant thiamine biosynthesis OsDR8, OsTHIC in T51-1 and MH63. (d) Thiamine level in MH63 and T51-1 plants. (e) Thiamine level in brown rice of T51-1 and MH63. For the expression analysis, OsUBQ1 was used as the internal control. Values are means ± SE (n = 3), *P < 0.05(ANOVA) comparing with T51-1 and MH63.
Up-regulation of thiamine biosynthesis genes in T51-1
Plant NB-LRR genes activate the plant immune system through various pathways, including SA (salicylic acid), JA (jasmonic acid), etc. However, a RT-qPCR analysis showed that the SA and JA pathway-related genes were down regulated in T51-1 compared with MH63 (Supplementary Fig. S1). This result suggests that the upregulation of OsLSR did not activate JA or SA pathway. This was consistent with the results showing that some JA- and SA-responsive PR genes were not increased in T51-1(Fig. 1c).
A previous study showed that the basal expression of OsPR1a and OsPR4a is related to OsDR8 because RNAi of OsDR8 leads to the down regulation of OsPR1a and OsPR4a19. OsDR8 encodes an enzyme involved in the biosynthesis of the thiazole precursor of thiamine (VB1). Thus, the transcript levels of OsDR8 and another rice gene OsTHIC (Os03g0679700) homologous to the Arabidopsis AtTHIC, which is critical in thiamine biosynthesis and regulation, were analysed in T51-1 and MH63. The results showed that their transcript levels were all increased in T51-1 (Fig. 2c). Then, we examined the thiamine content in the leaf tissue of both T51-1 and MH63. Correlating with the expression analysis, T51-1 showed a 0.5-fold increase in the thiamine level compared with that of MH63 (Fig. 2d). However, both T51-1 and MH63 accumulated a similar level of thiamine in their brown rice (P = 0.578) (Fig. 2e). Thus, our results showed that the slight upregulation of OsLSR expression activated the de novo synthesis of VB1 in rice leaf.
Overexpression of OsLSR::eGFP results in autoactivation of immune response and accumulation of thiamine through up regulation of OsDR8
To further understand the functional characteristics of OsLSR, we created OsLSR overexpressing (OsLSROE) lines in Nipponbare under the control of the ACTIN1 promoter. These lines showed high expression of OsLSR (Supplementary Fig. S2). A phenotypic evaluation showed that overexpressing OsLSR in rice results in an extremely low seed setting rate (Supplementary Table S1). Moreover, the OsLSROE lines showed a severe weakness compared with the control Nipponbare (Supplementary Fig. S2). Seeds carrying the OsLSROE construct failed to germinate and form seedlings (Supplementary Fig. S2).
Thus we further overexpressed an eGFP fusion OsLSR::eGFP in Nipponbare under the control of the 35 S promoter. Transgenic rice overexpressing OsLSR::eGFP showed minor weakness compared with the wildtype at three leaf stage (Fig. 3a). Relative quantitative PCR showed that the two OsLSR::eGFPOE-11,-16 lines of T3 generation highly expressed the foreign fusion and resistance marker genes showed a similar expression pattern as that in T51-1, OsPR1a, OsPR4a and OsPAD4 were up regulated while the rest were down-regulated (Fig. 3b), indicating the function of OsLSR was unlikely affected by the fused eGFP. Further transcript screening of thiamine synthesis-related genes revealed that the expressions of OsTHIC and OsDR8 were increased particularly for OsDR8 in these lines compared with wildtype Nipponbare (Fig. 3c). These changes correlated with the expression level of OsLSR::eGFP (Fig. 3b). In parallel, the thiamine contents in these two OsLSR::eGFPOE lines increased by 0.5~1-fold compared with the wild type Nipponbare (Fig. 3d). These results further verified that the up regulation of OsLSR results in autoactivation of immune response and accumulation of thiamine. It was well documented that maintaining an activated immune system is resource-consuming and a high level of thiamine results in increased activity of TPP-requiring enzymes to result in enhanced carbohydrate oxidation37,39. Thus, these plants suffered a great fitness cost and showed weakness, especially in growth and seed setting (Supplementary Table S1).
Molecular characterization of OsLSR::eGFPOE plants.
(a) Growth retardation of OsLSR::eGFPOE-11,-16 (T3) plants. (b) Expression level of OsLSR and the immune response related genes in OsLSR::eGFPOE lines. (c) Transcript level of thiamine biosynthesis genes in OsLSR::eGFPOE lines. (d) Thiamine level examined in the leaf tissue of Nipponbare (wild type) and OsLSR::eGFPOE plants. For the expression analysis, OsUBQ1 was used as the internal control. Values are means ± SE (n = 3). *P < 0.05 (ANOVA) comparing WT plants and two OsLSR::eGFPOE lines.
Overexpression of OsLSR1-437 activates PR genes expression in rice
Several studies have shown that ectopic expression of certain parts of the NB-LRR protein can trigger an immune response40. Based on this consideration, we overexpressed the N-terminal 437aa of OsLSR under the ACTIN1 promoter in Nipponbare (OsLSR1-437OE) and eventually obtained one transgenic line. T3 seedlings positive for OsLSR1-437 at the three-leaf stage were used for RT-qPCR analysis. The results showed that the fragment was efficiently expressed, whereas the immune response genes showed a different pattern from OsLSR::eGEPOE lines. All the detected PR genes were up regulated compared with wild type (Fig. 4a) and at a level much higher than that in OsLSR::eGFPOE and T51-1, but OsPAD4 was decreased. These results suggest that the N-terminal portion of OsLSR (the CC plus NB domain) can successfully trigger transcriptional reprogramming when overexpressed in rice. However, when OsLSR1-437::eGFP under the control of the 35 S promoter was transiently expressed in tobacco leaf, the construct failed to trigger HR-like cell death comparing with 35 S::OsLSR (Fig. 4b–d). Furthermore, an expression analysis of thiamine biosynthesis genes showed that only OsTHIC increased slightly, while OsDR8 was down regulated (Fig. 4e). These results suggest that the N-terminal portion of OsLSR (the CC plus NB domain) partly lost OsLSR’s signalling property when overexpressed in rice or transient expressed in tobacco leaf. LRR domain was reported also critical in signalling function for some NB-LRR proteins41. These results indicate that remove of the LRR domain disables OsLSR in cell death and thiamine biosynthesis signalling, while the N-terminus could conducts transcriptional reprogramming when overexpressed. These results also imply that OsLSR-triggered transcriptional reprogramming and thiamine biosynthesis are two independent processes.
Molecular analyses and transient expression assay of OsLSR1-437OE plants.
(a) Expression analysis of immune response genes in OsLSR1-437OE plants. (b–d) Transient expression assay of OsLSR1-437::eGFP in tobacco leaf. Free eGFP was used as a negative control (b), OsLSR1-437::eGFP (c), (d) 35 S::OsLSR was used as a positive control. (e) Expression analysis of two thiamine biosynthesis enzyme-encoding genes in OsLSR1-437OE plants. For the expression analysis, OsUBQ1 was used as the internal control. Values are means ± SE (n = 3). *P < 0.05 (ANOVA) comparing WT plants and OsLSR1-437OE plants.
Down regulation of OsLSR by RNA interference also leads to autoactivation of immune response and accumulation of thiamine by promoting OsTHIC
To investigate the effect of knocking out OsLSR, the expression of OsLSR in Nipponbare was blocked by RNA interference (OsLSRRI). OsLSRRI lines showed not only delayed growth compared to wild type Nipponbare, but even more severe growth retardant than OsLSR::eGFPOE plants (Fig. 5a) and they also displayed a low seed setting phenotype (Supplementary Table S1). The expression levels of OsLSR in two representative lines, OsLSRRI-7 and OsLSRRI-8, of the T3 generation at seedling stage were analysed. The results showed that the lines had 95% and 96% inhibition, respectively (Fig. 5b). However, an expression profile analysis revealed a different pattern of thiamine biosynthesis genes from wild type. The down regulation of OsLSR resulted in increased OsDR8, OsTHIC; in particular, OsTHIC was considerably increased (Fig. 5c). Like OsLSR::eGFPOE plants the two OsLSRRI lines showed increased thiamine content compared to wild type plants (Fig. 5d) and the level was even higher than that in OsLSR::eGFPOE plants. Transcript analyse of PR genes showed they were all increased by a certain amount, indicating silence of OsLSR also results in autoactivation of immune response (Fig. 5e). Interestingly, OsPAD4 was down regulated in the OsLSRRI lines (Fig. 5e), indicating that OsLSR plays a signalling role upstream of OsPAD4 and positively regulates its expression.
RNA interference of OsLSR and its impacts on rice growth and thiamine biosynthesis.
(a) Weakness of OsLSRRI plants (right and middle) and wild type Nipponbare (left). (b) Expression of OsLSR in Nipponbare and two typical OsLSRRI lines. (c) Transcripts of thiamine biosynthesis genes in OsLSRRI plants compared with wild type Nipponbare. (d) Thiamine level in Nipponbare and two OsLSRRI independent lines. (e) Expression of immune response genes in OsLSRRI lines compared with Nipponbare. For the expression analysis, OsUBQ1 was used as the internal control. Values are means ± SE (n = 3). *P < 0.05 (ANOVA) comparing Nipponbare plants and two RNAi lines.
Gene expression changes in OsLSR transgenic plants
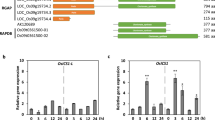
The biosynthesis of thiamine is tightly and negatively regulated by TPP through a riboswitch on the THIC mRNA27. A high level of TPP in cells leads to an increased level of the unstable mRNA form of THIC, in which the second intron in the 3′UTR is spliced out; when the TPP level is low, this intron is retained, resulting in enhanced stability of the mRNA and increased translation of the THIC protein. An expression analysis of these types of OsTHIC mRNA in OsLSR::eGFPOE and OsLSRRI plants showed an increase in the intron-retained type (Fig. 6a,b). In the OsLSRRI lines, all three types of OsTHIC mRNA were increased. Taken together, these results suggest that the expression pattern of OsTHIC alternative transcripts in OsLSR transgenic plants is consistent with a high thiamine level.
Gene expression analysis of OsLSR transgenic plants.
Expression level of OsTHIC alternative transcripts in OsLSR::eGFPOE (a) and OsLSRRI (b) plants. Gene expressions of rice transketolase (TK) and deoyxylulose-5-phosphate synthase (DXS) in T51-1 and MH63 (c), OsLSR::eGFPOE (d) and OsLSRRI (e) lines. For the expression analysis, OsUBQ1 was used as the internal control. Values are means ± SE (n = 3). *P < 0.05 (ANOVA) comparing Nipponbare plants and two RNAi lines.
The plant Calvin Benson (C3) cycle provides two major precursors for thiamine biosynthesis, G3P (Glyceraldehyde 3-phosphate) and R5-P (Ribose 5-phosphate). Two C3 cycle enzymes, transketolase (TK) and deoyxylulose-5-phosphate synthase (DXS) are critical for their metabolism22,42. We analysed the expression of these enzymes in the transgenic plants. Results showed OsTK (Os01g0931400) and OsDSX (Os05g0408900) were increased in both OsLSR::eGFPOE and OsLSRRI lines, however, the two genes were not significant up regulated in T51-1 (Fig. 6c–e). This result indicates that OsLSR might affect thiamine biosynthesis at an upstream step and the accurate regulation of OsLSR is important for plant growth and development because a disturbance in its expression results in metabolic chaos.
Two fragments, OsLSR1–178 and OsLSR464–546, are targeted to the chloroplast
Plant thiamine is biosynthesized in the chloroplast, whereas most cloned NB-LRR proteins are distributed in the nucleus or cytoplasm. To better understand the function of OsLSR and its relationship with thiamine biosynthesis, we constructed a C-terminal fusion with eGFP to study the distribution of OsLSR in the plant cell. However, we were unable to detect an eGFP signal after transient expression in tobacco leaves. In addition, we were still unable to detect the eGFP signal after steady overexpression in rice plants. One possibility is that the fusion protein triggers cell death, tobacco cells were breakdown before the fusion accumulated to a visible level, transgenic rice cells with high fusion protein level were aborted during early stage of Agrobacterium mediated transformation and only these compromised transgenic lines were survived.
Several studies on the subcellular distribution of NB-LRRs have shown that its individual domains play a role and contribute to the distribution of the intact protein43,44. Thus, we examined the distribution of individual domains of OsLSR to speculate about its subcellular location. We constructed a series of eGFP fusions that each contained a different truncation of OsLSR (Supplementary Fig. S3). All of the fusions could be visualized. The fusions containing OsLSR1–31 and OsLSR1–89 exhibited a nucleo-cytoplasmic distribution similar to control 35 S::eGFP (Supplementary Fig. S3). The fusion containing OsLSR1–178, which includes the CC domain plus 50aa, showed a distinct distribution pattern. At 24HPI (hour post infiltration), the eGFP signal was widely distributed in chloroplasts, the nucleus and the cytoplasm. As time progressed to 48HPI, the fusion was observable only in chloroplasts (Fig. 7a).
Subcellular localization of constructs carrying truncated OsLSR domains.
(a) Subcellular distribution pattern of OsLSR1-178::eGFP showing a nuclear, chloroplast and cytoplasmic distribution pattern at 24 h after infiltration (upper panel). At 48 h, only chloroplast localization was observed (bottom panel). (b) Subcellular localization of eGFP (CK). (c) Subcellular localization pattern of OsLSR32–178::eGFP and DFOsLSR32–178::eGFP (d). Both constructs showed a nuclear-preferred distribution. (e) Subcellular distribution of OsLSR161-437::eGFP. Subcellular localization pattern of OsLSR464–894::eGFP (f) and OsLSR717–894::eGFP (g). Both constructs showed cytoplasmic distribution with non-uniform, punctate staining. (h) Subcellular localization of a truncated OsLSR LRR fragment, OsLSR464–731. Two sets of images are shown: a large-scale view (top panel) and a detailed view of the area indicated by the square (bottom panel). The fusion construct showed cytoplasmic distribution in a cell with strong signal (shown in upper panel), while localization to the chloroplast was observed in a cell with a weak signal (bottom panel). Scale bar, 50 μm. For OsLSR464–731::eGFP, the scale bar is 50 μm in the upper panel and 10 μm in the bottom panels.
A fusion containing OsLSR32–178 was constructed to test whether a chloroplast signal was still produced. This fusion did not produce a signal in chloroplasts and tended to be located in the nucleus with very little signal in the cytoplasm (Fig. 7c). To determine whether the nuclear accumulation of OsLSR32–178::eGFP resulted from active nuclear import via a nuclear targeting signal or passive import through diffusion, we added another eGFP to its N-terminus and named the construct DFLSR32–178::eGFP. This construct still displayed nuclear localization, although the additional eGFP increased its retention in the cytoplasm in tobacco leaf cells (Fig. 7d). These results indicated that OsLSR has a positive NLS in the OsLSR32–178 region and full-length OsLSR1–178 is necessary to target the protein to the chloroplast.
The construct containing OsLSR161–437, which corresponds to the ATPase domain, displayed nuclear localization with very little fluorescence in the cytoplasm (Fig. 7e). The fusion containing OsLSR464–894, which includes the putative LRR domain or its truncated fragment OsLSR717–894, localized in the cytoplasm (Fig. 7f,g).
Unexpectedly, constructs carrying truncated sequences from the N-terminus of OsLSR464–894, such as OsLSR464–546, OsLSR464–731 and OsLSR464–867, showed a distribution pattern that was very different from either OsLSR464–894 or OsLSR717–894. In cells with strong signals, the fluorescence pattern clearly outlined the tobacco epidermis cells, indicating that the fusions were located in the cytoplasm (Fig. 7h, Supplementary Fig. S3).By contrast, in cells with weak signals, the eGFP fluorescence overlapped with chloroplasts, indicating that the fusions located to the chloroplasts (Fig. 7h, Supplementary Fig. S3). Thus, the original target of these constructs was the chloroplast; however, the constructs localized to the cytoplasm when they were transiently overexpressed. This pattern of subcellular distribution is similar to that of OsLSR1–178::eGFP. The difference between the patterns is that the former three fusions no longer remained in the chloroplast during transient overexpression. These results revealed a subtle relationship between OsLSR truncations and the chloroplast.
Discussion
Our results demonstrate that up- or down- regulation of OsLSR results in activated immune system of transgenic rice. These plants showed a weakness and yellowish phenotype, they constitutively express PR genes compared with wild type. Moreover these transgenic plants had a higher thiamine level in cells.
OsLSR can trigger immune response. This characteristic of OsLSR was evidenced by several experiments. First, in OsLSR up regulation lines, including T51-1 and OsLSR::eGFPOE, increases in defence-related and PR genes were observed. An increase in a set of PR and defence-related gene transcripts is a reliable molecular marker to indicate whether the plant immune system is activated35,36. Second, transiently overexpressed OsLSR or its eGFP fusion in tobacco leaf resulted in HR-like cell death. Timely programmed cell death of infected cells in the R gene-mediated defence response is an effective strategy for plants to restrict pathogens from spreading to other healthy cells45. Third, strong expression of PR genes was observed in rice ectopically expressing the CC plus NB domains of OsLSR, suggesting that overexpressed OsLSR1–437 triggers rice transcripts reprogram which is common in an immune response. The N-terminal portion of NB-LRR was thought to be involved in downstream signal transduction. Previous reports indicated that overexpression of the N-terminal portion of some NB-LRR proteins results in auto-activation of the plant immune system40. Thus, OsLSR clearly acts as an activator of the plant immune system and can activate the plant defence response after activation or up-regulation; its N-terminal domain is involved in downstream defence signalling.
Different subsets of defence signalling cascades can be activated by R genes mediated resistance46,47. Our results showed that the expression of OsPAD4 was associated with the transcriptional level of OsLSR. In OsLSR up regulation lines, OsPAD4 expression was increased, while in OsLSRRI lines, expression was decreased. These results indicate that OsLSR functions upstream of OsPAD4 and positively regulates its expression. Furthermore, OsLSR may function as a signalling factor downstream of immune receptors because it is required for proper expression of OsPAD4. Interestingly, strong expression of PR genes but with the down regulation of OsPAD4 was observed in rice plants ectopically expressing OsLSR1–437, suggest OsLSR1–437 lost the native specificity of OsLSR in downstream immune signalling when overexpressed, which was also observed in several cases exploring NB-LRR protein function40. OsPAD4 encodes a putative triacylglycerol lipase and positively regulates rice defence responses to bacterial Xanthomonas oryzae pv. Oryzae (Xoo) in JA pathway48. In Arabidopsis, pathogen-induced PR1a expression is dependent on AtPAD449, this observation aligns with the increased expression of OsPR1a in OsLSR up regulated lines. However, different from OsPAD4-mediated bacterial blight resistance is JA dependent, in T51-1 SA and JA synthetic genes were inhibited, indicating OsLSR activated immune signalling does not activate JA and SA. This might because plants possess multi mechanisms to activate immune system besides SA and JA and some time one activated immune signal interact antagonism to another. Antagonism interaction was found between SA and JA when plants facing biotrophic or necrotrophic pathogen invasion in Arabidopsis. Suppression of JA and SA synthesis related genes with enhanced resistance to Xoo infection was also reported in rice overexpressing an IAA-amino acid synthetase gene GH3–850,51. More efforts are needed to elucidate details of the immune signalling pathway in OsLSR up- and down- regulated lines and it is of interesting to systematically evaluate the disease resistance property of OsLSR transgenic rice plants in future study.
Another interesting thing in the present study is we found that up-regulation or silencing of OsLSR result in accumulation of thiamine in rice. In T51-1, 0.8 fold increase of OsLSR results in 0.5 fold higher of thiamine level compared with MH63 in its leaf tissue. However, the thiamine level in its brown rice was increased not significantly. Similar results were also observed in the Arabidopsis plants overexpressing AtTHIC, which had three-fold increase of the thiamine level in their leaf tissue but only 20% increase in its seeds compared with wildtype plants39,52. These results indicated that simply up-regulation of thiamine synthetic genes in plants is insufficiently to cause considerable increase of the thiamine level in its storage organ. The interesting thing is that in OsLSR::eGFPOE-11, while the expression of OsLSR, OsDR8 and OsTHIC was much higher than that in T51-1, the two lines had similar thiamine level. This may be because the two lines are of different genetic background, as the thiamine level in OsLSR::eGFPOE lines with the same background is positively related with the expression of OsLSR, OsTHIC and OsDR8 (Fig. 3).
The higher thiamine level in OsLSRRI lines could be explained by the fact that in OsLSR::eGFPOE lines, OsDR8 increased more than OsTHIC; however, in OsLSRRI plants, a strong increase in the OsTHIC intron-retained variant was observed compared to a minor increase in OsDR8. The THIC intron-retained variant is more effective than TH1 in the induction of thiamine biosynthesis when it is overexpressed23,39. Further, OsTK and OsDXS which provides precursors for thiamine biosynthesis is more enhanced in OsLSRRI lines than OsLSR::eGFPOE lines. These results imply that the signal triggered for thiamine biosynthesis in these two lines is different. OsLSR::eGFPOE plants tend to activate thiamine biosynthesis through thiazole moieties pathway, while OsLSRRI plants mainly by activating the other pyrimidine moiety pathway. In addition, the above results also indicate that OsLSR might have a role in balancing the provision of both moieties of HMPP and HET-P used for thiamine biosynthesis. This balancing was found to be critical in Chlamydomonas reinhardtii, which guarantees that sufficient but not excess thiamine is available34,53.
Thiamine is synthesized in the chloroplast22,42. In the experiment, the data from the assay of eGFP-tagged OsLSR fragments showed that the chloroplast is the primary target of OsLSR1–178 and OsLSR428–546 after transient expression in tobacco leaf cells. And further experiments showed the two fragments had an interaction in yeast. Taking together, these results indicate that these two fragments may help OsLSR target to chloroplast and affects thiamine synthesis there.
In the present study, we have shown that both overexpression and RNA silencing of OsLSR results in weakness of the transgenic rice plants, including growth retardation, chlorosis and deceased seed setting. One explanation for the phenotype is their constitutively activated immune system, as these defects were common in plant mutants with an auto-activated immune system54,55,56. Maintaining an activated immune system is costly and fitness-limiting resources and energy that are allocated to resistance are unavailable for fitness-relevant processes, such as growth and cause penalties in reproduction37,38. These results indicate that NB-LRR genes should be tightly controlled to avoid unnecessary activation, which is harmful to plants per se.
An alternative explanation for the weakness of OsLSR transgenic plants is the elevated thiamine level in the plants. Thiamine and its active form TPP are an essential cofactor for enzymes involved in a number of critical metabolic processes, such as the production of acetyl-CoA, the tricarboxylic acid cycle (TCA), the pentose phosphate pathway (PPP), the C3 cycle, etc.57. Excess available TPP in a cell increased the activity of TPP-requiring enzymes, resulting in an increase in carbohydrate oxidation through TCA, PPP and the accumulation of amino acids. This augmented flux through central metabolism that perturbs metabolic homeostasis results in chlorosis, growth retardation, etc.39,52. In this study, abnormal expression of OsTK and OsDXS which evolve in C3 cycle is observed indicating a metabolic chaos in these transgenic rice plants.
Our results presented here showed that both up-and down-regulation of OsLSR leads to autoactivation of rice immune system and accumulation of thiamine results in a weakness phenotype. Thus, there exists an unknown mechanism regulates OsLSR within a narrow range for the normal growth and development of rice plants. More efforts are needed to explore the function of OsLSR as its expression is closely related to thiamine level in cells and immunity which could be used for rice improvement.
Materials and Methods
Plant materials
Oryza sativa L. subsp. Indica line MH63 (wild type controls), T51-1 (MH63 background), the japonica line Nipponbare and transgenic plants or lines were grown in the field or greenhouse at Zijingang campus, Zhejiang University. The growth conditions in the greenhouse were set to a photoperiod of 12 h light at 28 °C/12 h dark at 25 °C, with a relative humidity of approximately 60%. Nicotiana benthamiana was grown in a growth chamber under a photoperiod of 12 h light at 28 °C/12 h dark at 22 °C, with a relative humidity of approximately 50%.
Vector construction
Total RNA was isolated from the leaf tissue of Nipponbare plants using TRIzol reagent (Invitrogen), according to the manufacturer’s instruction. First-strand cDNA synthesis was conducted using the M-MLV first-strand synthesis system (Promega). To construct the overexpression vector, the complete coding sequence (CDS) of OsLSR was amplified using the gene-specific primers LSROE-F and LSROE-R (Supplementary Table S2) with XbaI and KpnI restriction sites, respectively. The amplified cDNA products were cloned into p1300-actin:NOS58 to produce p1300-actin::OsLSR::NOS, which was named pOELSR.
The RNAi vector was constructed using a 350 bp fragment corresponding to the 2806 bp to 3155 bp region of Os10g0183000; the fragment was amplified using the primers LSRRi-F and LSRRi-R. The product was digested with two pairs of restriction enzymes (BamHI/KpnI and SpeI/SacI) and the products were cloned into pTCK30359 to produce the RNAi repression construct pRiLSR. The pOELSR and pRiLSR vectors were introduced into Agrobacterium tumefaciens strain EHA105, which was used to infect rice embryonic calli from Nipponbare to generate OsLSR overexpressing plants (OsLSROE) and RNA interfering (OsLSRRI) plants, respectively.
To construct the subcellular localization vectors, the complete CDS or truncations of OsLSR were amplified using the primers listed in Supplementary Table S2; all forward and reverse primers contained XbaI and KpnI restriction sites, respectively. The PCR products were then inserted into p1300-35S::eGFP::NOS to generate the corresponding vectors.
Quantification of thiamine content
The thiamine level in the leaf tissue of rice plants was determined using a fluorescent method19,21. In brief, ~2–4 g of rice leaf tissue was ground into a powder in liquid nitrogen. Thiamine was extracted in 0.1 M HCl and thiamine was oxidized into thiochrome in a solution containing 0.1% kalium ferricyanide and 15% sodium hydroxide. The fluorescence emitted by the thiochrome was detected at 365 nm excitation and 435 nm emission wavelengths using an ELISA (Biotech Synergy H1).
Transient expression assay in Nicotiana benthamiana
The vectors were transformed into A.tumefaciens EH105 for transient expression. A. tumefaciens cells were grown in 5 ml LB with the appropriate antibiotics overnight. One hundred microliters of the overnight culture was used to inoculate 25 ml of fresh LB (with the same antibiotics as the overnight culture, plus 20 μM acetosyringone added after autoclaving and immediately before use) and grown to an OD600 of 1.0. The cells were then collected by centrifugation and resuspended in the induction medium (10 mM MES, pH 5.7, 10 mM MgCl2, 200 μM acetosyringone) to an OD600 of 0.4. After incubation at room temperature for 2–3 h, the A. tumefaciens cells were infiltrated into the abaxial surface of tobacco leaves using 1 ml needle syringes. eGFP signals were observed using a laser scanning confocal microscope (Leica TCS 710).
Real-time quantitative RT-PCR
For real-time quantitative RT-PCR, leaf samples were harvested at the three-leaf stage; three biological repeats for each sample were used. Total RNA from the leaves was isolated using an RNA extraction kit (TRIzol reagent, Invitrogen). Approximately 1 μg of total RNA was reverse-transcribed using a PrimeScriptTM RT reagent kit with gDNA Eraser (TAKARA) in a final volume of 20 μl and then diluted to 100 μl. Quantitative RT-PCR was performed in a total volume of 20 μl, using 2 μl of the reverse-transcribed product above, 0.2 μM primers and 10 μl of Fast Start Essential DNA Green Master (Roche) on a Roche LC96. The amounts of transcript were estimated using the relative quantification method60 with UBQ1 as internal standards for normalization. The primers used in this experiment are listed in Supplementary Table S3.
Phenotype evaluation of transgenic rice
Seed setting rate of MH63, T51-1 was the mean value of ten independent plants. For each plant only productive tillerings are taken in consider. Seed setting for transgenic material: certain independent lines indicated in tables were evaluated. For each independent line, ten plants were evaluated and the mean value of each transgenic line is further statistically analyzed to get standard error for each transgenic material.
Additional Information
Accession codes: OsLSR: Os10g0183000, OsDXS: Os05g0408900, OsTHIC: Os03g0679700, OsTK: Os06g0133800.
How to cite this article: Wang, L. et al. Both overexpression and suppression of an Oryza sativa NB-LRR-like gene OsLSR result in autoactivation of immune response and thiamine accumulation. Sci. Rep. 6, 24079; doi: 10.1038/srep24079 (2016).
References
Pan, Q., Wendel, J. & Fluhr, R. Divergent evolution of plant NBS-LRR resistance gene homologues in dicot and cereal genomes. J Mol Evo 50, 203–213 (2000).
Hammond-Kosack, K. E. & Jones, J. D. Resistance gene-dependent plant defense responses. Plant Cell 8, 1773–1791 (1996).
Fu, Z. Q. & Dong, X. Systemic acquired resistance: turning local infection into global defense. ANNU REV PLANT BIOL 64, 839–863 (2013).
Zhiyong, G., Zhiyoug, G., Eui-Hwan, C., Eitas, T. K. & Dangl, J. L. Plant intracellular innate immune receptor Resistance to Pseudomonas syringae pv. maculicola 1 (RPM1) is activated at and functions on, the plasma membrane. Proc Natl Acad Sci USA 108, 7619–7624 (2011).
Chung, E. H. et al. Specific threonine phosphorylation of a host target by two unrelated type III effectors activates a host innate immune receptor in plants. Cell Host Microbe 9, 125–136 (2011).
Qian-Hua, S. et al. Nuclear Activity of MLA Immune Receptors Links Isolate-Specific and Basal Disease-Resistance Responses. Science 315, 1098–1103 (2007).
Burch-Smith, T. M. et al. A novel role for the TIR domain in association with pathogen-derived elicitors. PLoS Biol 5, e68 (2007).
Padmanabhan, M. S. & Dinesh-Kumar, S. P. All hands on deck—the role of chloroplasts, endoplasmic reticulum and the nucleus in driving plant innate immunity. Mol Plant Microbe Interact 23, 1368–1380 (2010).
Meyers, B. C., Alexander, K., Alyssa, G., Hanhui, K. & Michelmore, R. W. Genome-Wide Analysis of NBS-LRR-Encoding Genes in Arabidopsis. Plant Cell Online 15, 809–834 (2003).
Goyer, A. Thiamine in plants: Aspects of its metabolism and functions. Phytochemistry 71, 1615–1624 (2010).
Asselin, A. et al. Light-influenced extracellular accumulation of b (pathogenesis-related) proteins in Nicotiana green tissue induced by various chemicals or prolonged floating on water. Canadian Journal of Botany 63, 1276–1283 (1985).
Il-Pyung, A., Soonok, K. & Yong-Hwan, L. Vitamin B1 functions as an activator of plant disease resistance. Plant Physiology 138, 1505–1515 (2005).
Klessig, D. F. Dissection of the Salicylic Acid Signaling Pathway in Tobacco. Mol Plant Microbe Interact 9, 474–482 (1996).
Il-Pyung, A., Soonok, K., Yong-Hwan, L. & Seok-Cheol, S. Vitamin B1-induced priming is dependent on hydrogen peroxide and the NPR1 gene in Arabidopsis. Plant Physiol 143, 838–848 (2007).
Abhishek, C. et al. Saccharomyces cerevisiae THI4p is a suicide thiamine thiazole synthase. Nature 478, 542–546 (2011).
Belanger, F. C., Leustek, T., Chu, B. & Kriz, A. L. Evidence for the thiamine biosynthetic pathway in higher-plant plastids and its developmental regulation. Plant Mol Biol 29, 809–821 (1995).
Ribeiro, A. et al. Identification of agthi1, whose product is involved in biosynthesis of the thiamine precursor thiazole, in actinorhizal nodules of Alnus glutinosa. Plant J 10, 361–368 (1996).
Machado, C. R. et al. Thi1, a thiamine biosynthetic gene inArabidopsis thaliana, complements bacterial defects in DNA repair. Plant Mol Biol 31, 585–593 (1996).
Wang, G. et al. Dual function of rice OsDR8 gene in disease resistance and thiamine accumulation. Plant Mol Biol 60, 437–449 (2006).
Danyu et al. AtTHIC, a gene involved in thiamine biosynthesis in Arabidopsis thaliana. Cell Res 18, 566–576 (2008).
Weina, Z. et al. Tomato LeTHIC is an Fe-requiring HMP-P synthase involved in thiamine synthesis and regulated by multiple factors. Plant Cell Physiol 52, 967–982 (2011).
Julliard, J.-H. Biosynthesis of the pyridoxal ring (vitamin B6) in higher plant chloroplasts and its relationship with the biosynthesis of the thiazol ring (vitamin B1). Comptes Rendus De Lacadémie Des Sciences Série Sciences De La Vie 314, 285–290 (1992).
Ajjawi, I., Tsegaye, Y. & Shintani, D. Determination of the genetic, molecular and biochemical basis of the Arabidopsis thaliana thiamin auxotroph th1. Arch Biochem Biophys 459, 107–114 (2007).
Yang, S. K. et al. A Brassica cDNA clone encoding a bifunctional hydroxymethylpyrimidine kinase/thiamin-phosphate pyrophosphorylase involved in thiamin biosynthesis. Plant Mol Biol 37, 955–966 (1998).
Rapala-Kozik, M., Olczak, M., Ostrowska, K., Starosta, A. & Kozik, A. Molecular characterization of the thi3 gene involved in thiamine biosynthesis in Zea mays: cDNA sequence and enzymatic and structural properties of the recombinant bifunctional protein with 4-amino-5-hydroxymethyl-2- methylpyrimidine (phosphate) kinase and thiamine monophosphate synthase activities. Biochem. J 408, 149–159 (2007).
Ajjawi, I., Milla, M. A. R., Cushman, J. & Shintani, D. K. Thiamin pyrophosphokinase is required for thiamin cofactor activation in Arabidopsis. Plant Mol Biol 65, 151–162 (2007).
Wachter, A. et al. Riboswitch control of gene expression in plants by splicing and alternative 3′ end processing of mRNAs. The Plant Cell 19, 3437–3450 (2007).
Khozaei, M. et al. Overexpression of Plastid Transketolase in Tobacco Results in a Thiamine Auxotrophic Phenotype. The Plant Cell Online 27, 432–447 (2015).
Rapala-Kozik, M., Kowalska, E. & Ostrowska, K. Modulation of thiamine metabolism in Zea mays seedlings under conditions of abiotic stress. J Exp Bot 59, 4133–4143 (2008).
Tunc-Ozdemir, M. et al. Thiamin confers enhanced tolerance to oxidative stress in Arabidopsis. Plant Physiol 151, 421–432 (2009).
Rapala-Kozik, M., Wolak, N., Kujda, M. & Banas, A. K. The upregulation of thiamine (vitamin B1) biosynthesis in Arabidopsis thaliana seedlings under salt and osmotic stress conditions is mediated by abscisic acid at the early stages of this stress response. BMC Plant Biology 12, 2 (2012).
Tu, J. et al. Field performance of transgenic elite commercial hybrid rice expressing Bacillus thuringiensis δ-endotoxin. Nature Biotechnol 18, 1101–1104 (2000).
LAI, Y.-s. et al. Development of Insect-Resistant Hybrid Rice by Introgressing the Bt Gene from Bt Rice Huahui 1 into II-32A/B, a Widely Used Cytogenic Male Sterile System. J Integr Agr 13, 2081–2090 (2014).
Lukasik, E. & Takken, F. L. STANDing strong, resistance proteins instigators of plant defence. Curr Opin Plant Biol 12, 427–436 (2009).
Friedrich, L. et al. A benzothiadiazole derivative induces systemic acquired resistance in tobacco. Plant J 10, 61–70 (1996).
Guo, A., Durner, J. & Klessig, D. F. Characterization of a tobacco epoxide hydrolase gene induced during the resistance response to TMV. Plant J 15, 647–656 (1998).
Tian, D., Traw, M., Chen, J., Kreitman, M. & Bergelson, J. Fitness costs of R-gene-mediated resistance in Arabidopsis thaliana. Nature 423, 74–77 (2003).
van Hulten, M., Pelser, M., Van Loon, L., Pieterse, C. M. & Ton, J. Costs and benefits of priming for defense in Arabidopsis. Proc Natl Acad Sci USA 103, 5602–5607 (2006).
Bocobza, S. E. et al. Orchestration of Thiamin Biosynthesis and Central Metabolism by Combined Action of the Thiamin Pyrophosphate Riboswitch and the Circadian Clock in Arabidopsis. Plant Cell 25, 288–307 (2013).
Bernoux, M., Ellis, J. G. & Dodds, P. N. New insights in plant immunity signaling activation. Curr Opin Plant Biol 14, 512–518 (2011).
Hwang, C.-F., Bhakta, A. V., Truesdell, G. M., Pudlo, W. M. & Williamson, V. M. Evidence for a role of the N terminus and leucine-rich repeat region of the Mi gene product in regulation of localized cell death. The Plant Cell 12, 1319–1329 (2000).
Julliard, J.-H. & DoUCE, R. Biosynthesis of the thiazole moiety of thiamin (vitamin B1) in higher plant chloroplasts. Proc Natl Acad Sci USA 88, 2042–2045 (1991).
Botella, M. A. et al. Three genes of the Arabidopsis RPP1 complex resistance locus recognize distinct Peronospora parasitica avirulence determinants. The Plant Cell 10, 1847–1860 (1998).
Takemoto, D. et al. N-terminal motifs in some plant disease resistance proteins function in membrane attachment and contribute to disease resistance. Mol Plant Microbe Interact 25, 379–392 (2012).
Steller, H. Mechanisms and genes of cellular suicide. Science 267, 1445–1449 (1995).
Durner, J., Shah, J. & Klessig, D. F. Salicylic acid and disease resistance in plants. Trends Plant Sci 2, 266–274 (1997).
Kachroo, A. et al. Induction of H2O2 in transgenic rice leads to cell death and enhanced resistance to both bacterial and fungal pathogens. Transgenic Res 12, 577–586 (2003).
Ke, Y., Liu, H., Li, X., Xiao, J. & Wang, S. Rice OsPAD4 functions differently from Arabidopsis AtPAD4 in host-pathogen interactions. Plant J 78, 619–631 (2014).
Zhou, N., Tootle, T. L., Tsui, F., Klessig, D. F. & Glazebrook, J. PAD4 functions upstream from salicylic acid to control defense responses in Arabidopsis. The Plant Cell 10, 1021–1030 (1998).
Ding, X. et al. Activation of the indole-3-acetic acid–amido synthetase GH3-8 suppresses expansin expression and promotes salicylate-and jasmonate-independent basal immunity in rice. The Plant Cell 20, 228–240 (2008).
Fu, J. et al. Manipulating broad-spectrum disease resistance by suppressing pathogen-induced auxin accumulation in rice. Plant Physiol 155, 589–602 (2011).
Bocobza, S. E. et al. Orchestration of thiamin biosynthesis and central metabolism by combined action of the thiamin pyrophosphate riboswitch and the circadian clock in Arabidopsis. The Plant Cell 25, 288–307 (2013).
Moulin, M., Nguyen, G. T., Scaife, M. A., Smith, A. G. & Fitzpatrick, T. B. Analysis of Chlamydomonas thiamin metabolism in vivo reveals riboswitch plasticity. Proc Natl Acad Sci USA 110, 14622–14627 (2013).
Yang, S. & Hua, J. A haplotype-specific Resistance gene regulated by BONZAI1 mediates temperature-dependent growth control in Arabidopsis. The Plant Cell 16, 1060–1071 (2004).
Shirano, Y., Kachroo, P., Shah, J. & Klessig, D. F. A gain-of-function mutation in an Arabidopsis Toll Interleukin1 Receptor–Nucleotide Binding Site–Leucine-Rich Repeat type R gene triggers defense responses and results in enhanced disease resistance. The Plant Cell 14, 3149–3162 (2002).
Stokes, T. L., Kunkel, B. N. & Richards, E. J. Epigenetic variation in Arabidopsis disease resistance. Gene Dev 16, 171–182 (2002).
Hohmann, S. & Meacock, P. A. Thiamin metabolism and thiamin diphosphate-dependent enzymes in the yeast Saccharomyces cerevisiae: genetic regulation. Biochimica et Biophysica Acta (BBA)-Protein Structure and Molecular Enzymology 1385, 201–219 (1998).
Wang, Y. et al. Inhibition of a basal transcription factor 3-like gene Osj10gBTF3 in rice results in significant plant miniaturization and typical pollen abortion. Plant Cell Physiol 53, 2073–2089 (2012).
Wang, Z. et al. A practical vector for efficient knockdown of gene expression in rice (Oryza sativa L.). Plant Mol Biol Rep 22, 409–417 (2004).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
This work was financially supported by the National ‘863’ High Technology Program (grant No. 2006AA10Z159 and 2006 AA10A102). We gratefully thank Zhang Zhenzhong for his field assistance and Zhang Jiwen for his scientific discussions and suggestions. The photographs of tobacco leaf cell were taken by Li Yuqin.
Author information
Authors and Affiliations
Contributions
L.W. designed and carried out most experiments in this study and wrote the manuscript. X.Y. conducted rice transformation and seed setting analysis. H.L. helped in gene transcripts analysis. X.L. helped in thiamine determination. C.W. helped in tobacco transient expression assay. Y.L. helped in manuscript writing, J.T. took charge in overall experimental design, organization, management and manuscript final review. All authors reviewed this manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wang, L., Ye, Liu, H. et al. Both overexpression and suppression of an Oryza sativa NB-LRR-like gene OsLSR result in autoactivation of immune response and thiamine accumulation. Sci Rep 6, 24079 (2016). https://doi.org/10.1038/srep24079
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep24079
- Springer Nature Limited
This article is cited by
-
Mapping of dwarfing QTL of Ari1327, a semi-dwarf mutant of upland cotton
BMC Plant Biology (2022)
-
Time-series expression profiling of sugarcane leaves infected with Puccinia kuehnii reveals an ineffective defense system leading to susceptibility
Plant Cell Reports (2020)
-
Genome-wide association mapping of gene loci affecting disease resistance in the rice-Fusarium fujikuroi pathosystem
Rice (2019)