Abstract
Dairy cattle experience a profound nutrient deficit postpartum that is associated with immune dysfunction characterized by heightened inflammation and reduced pathogen clearance. The activation of the central nutrient-sensing mTOR pathway is comparatively reduced in leukocytes of early postpartum dairy cows during this time of most pronounced nutrient deficit. We assessed the effect of pharmacological mTOR inhibition (Torin-1, rapamycin) on differentiation of monocyte derived classically (M1) and alternatively (M2) activated macrophages (MPh) and dendritic cells (moDC) from 12 adult dairy cows. Treatment with mTOR inhibitors generated M1 MPh with increased oxidative burst and expression of IL12 subunits but decreased phagocytosis and expression of IL1B, IL6, and IL10. In M2 MPh, treatment inhibited expression of regulatory features (CD163, ARG2, IL10) skewing the cells toward an M1-like phenotype. In moDC, mTOR inhibition increased expression of pro-inflammatory cytokines (IL12A, IL12B, IL1B, IL6) and surface CD80. In co-culture with mixed lymphocytes, mTOR-inhibited moDC exhibited a cytokine profile favoring a Th1 response with increased TNF and IFNG production and decreased IL10 concentrations. We conclude that mTOR inhibition in vitro promoted differentiation of inflammatory macrophages with reduced regulatory features and generation of Th1-favoring dendritic cells. These mechanisms could contribute to immune dysregulation in postpartum dairy cows.
Similar content being viewed by others
Introduction
Epidemiological studies have long established the association between alterations in metabolism and innate immune function and increased disease susceptibility in metabolically challenged cattle1,2,3, but the mechanisms behind this relationship are still under investigation. Morbidity and mortality are higher postpartum, a time of transient, but pronounced nutrient deficit, than at any other time in the lactation cycle, and cows show signs of systemic inflammation, greater susceptibility to infection, and reduced ability for efficient pathogen clearance compared with other stages of the lactation cycle4,5. The resulting health disorders in the early postpartum cow lead to increased economic losses and sustained use of antimicrobials in the dairy industry6,7,8. Postpartum cows show distinct changes in their immune cell phenotype and typical response to antigenic stimulation. In vitro-matured monocyte derived dendritic cells (moDC) from postpartum cows exhibited higher co-stimulatory activity and antigen presenting potential along with a reduced production of IL10 compared with moDC from late gestation cows9. Pro-inflammatory M1 macrophages (MPh) were found to infiltrate adipose tissue in diseased postpartum cows10. Although an increase in pro-inflammatory function is considered necessary postpartum to allow the cow to respond appropriately to infections, a dysregulation of this transition potentially induced by severe nutrient deficiencies might lead to an exaggerated immune response with detrimental consequences for the host11.
The mechanistic target of rapamycin (mTOR) is a conserved central intracellular pathway that senses nutrient balance and has emerged as a key orchestrator that integrates metabolic cues and reconciles these with immune activation12,13. At least two complexes exist for mTOR, mTORC1 and mTORC2. While mTORC1 integrates growth factor signaling, energy status (through AMP-activated protein kinase), and oxygen and amino acid availability14, mTORC2 is predominantly activated by growth factors13. Our previous work showed that the mTORC1 pathway is downregulated in muscle and adipose tissue postpartum, and both upstream AKT and mTORC1 kinase activity cannot be activated to the same degree postpartum compared with prepartum in vivo15,16. We have extended our characterization of the mTORC1 pathway to show reduced activation of this pathway in bovine peripheral blood mononuclear cells in the immediate postpartum period17 and that pathway activation can be partially rescued ex vivo by exogenous nutrient supply18. Interestingly, work in other species has shown that mTOR inhibition modulates pro-inflammatory responses of monocytes, MPh, and dendritic cells after lipopolysaccharide (LPS) stimulation in vitro19, and skews MPh to pro-inflammatory M1 cells20,21. Therefore, the observed characteristics of the immune cell phenotype of postpartum cows and the demonstrated reduction in mTOR activation may be linked. However, the possible role of the mTOR pathway in regulating MPh and moDC during the pronounced postpartum nutrient deficit in dairy cows has not been studied to date.
We hypothesized that the inhibition of mTOR during in vitro differentiation of bovine monocyte-derived MPh and moDC leads to alterations of phenotype and function that might be translatable to an exaggerated pro-inflammatory response during the postpartum period in vivo. Our objective was to evaluate the effects of incomplete and complete pharmacological mTOR pathway inhibition (1st and 2nd generation inhibitors, respectively) on MPh and dendritic cells, key regulators of the inflammatory response. Given the different roles that polarized MPh subsets are thought to have in vivo, we studied the effect of mTOR inhibition separately during differentiation of classically (M1) and alternatively (M2) activated MPh from circulating monocytes.
Materials and methods
Animals and blood sampling
All procedures were evaluated and approved by the Cornell University Institutional Animal Care and Use Committee (protocol no. 2019-0030) and all experiments were performed in accordance with relevant guidelines and regulations. We also complied with the ARRIVE guidelines22,23,24,25,26,27,28. We sampled multiparous, lactating animals (n = 12) between January and March of 2020. Average (± SD) parity, days since calving, and days pregnant were 2.7 ± 0.7, 253 ± 28 days, and 178 ± 22 days, respectively. Milk composite sample linear score of 2.1 ± 1.2 and lack of clinical signs indicated absence of mastitis. To select animals in positive energy balance and avoid sampling from immune activated animals, all cows had to lack clinical signs of other detectable diseases at time of sampling, had no record of any disease in the last 30 days and could not have received a vaccine within 30 d prior to the time of sampling. We collected all blood samples before the morning feeding (0700–0800 h) from the coccygeal vessels in an aseptic manner using a 20 g × 2.5 cm needle into sterile vacutainers containing either K3EDTA, 143 USP units of freeze-dried sodium heparin, or into vacutainers without additive for serum separation. Samples were transported immediately to the laboratory.
Blood was allowed to clot for 30 min at room temperature before harvesting serum by centrifugation at 2800 × g for 20 min at 4 °C. Whole EDTA blood was submitted to the Cornell University Animal Health Diagnostic Center Clinical Pathology Lab (Cornell University, Ithaca, NY, USA) for automated complete blood cell counts (Advia 2120, Siemens, Munich, Germany). Average ± SD blood cell counts for WBC, neutrophils, lymphocytes, and monocytes were 9.4 ± 5.0 × 103/µL, 3.2 ± 0.7 × 103/µL, 5.2 ± 4.3 × 103/µL, and 0.5 ± 0.3 × 103/µL, respectively.
Cell separation and sorting
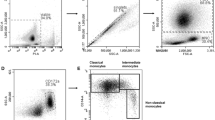
All primary antibodies are listed in Table 1. We obtained peripheral blood mononuclear cells (PBMC) from an average ± SD of 125 ± 15 mL of heparinized whole blood by density gradient separation as previously described23. Subsequently, we enriched CD14+ monocytes by magnetic activated cell sorting (MACS, all reagents Miltenyi Biotech, Bergisch Gladbach, Germany) as previously described9. Briefly, PBMC were labeled with a mouse anti-human monoclonal CD14 antibody (confirmed cross-reactive with bovine CD14 per manufacturer) coupled to paramagnetic beads and passed over a MACS separation column. Positively-selected enriched monocytes were counted, washed, and resuspended in complete medium (phenol-free RPMI 1640 with l-glutamine, 5% v/v FBS, 1× non-essential amino acids, and 1× insulin-transferrin-selenium) at 106 cells/mL for in vitro cell culture to generate MPh and moDC in the presence of mTOR inhibitors as described below. CD14-depleted cells were counted, washed with PBS, and frozen at − 80 °C at 15 × 106 cells per mL in freezing medium (45% v/v FBS, 9% v/v DMSO, 1% v/v HEPES) for later use in monocyte-derived dendritic cell (moDC) co-culture as autologous mixed lymphocytes. We determined purity and viability of CD14+ monocytes and CD14-depleted cells by flow cytometry. An aliquot of PBMC, CD14+ selected, and depleted cells were used to prepare cytospins (5 × 104 cells diluted in 100 µL PBS) for differential cell counts performed by a board-certified clinical pathologist (Animal Health Diagnostic Center, Ithaca, NY). Successful cell separation with our protocol was confirmed with cytospin manual differential cell counts showing that CD14+ enriched cells were 91.0 ± 11.6% monocytes and CD14+ depleted cells were 86.5 ± 7.8% lymphocytes. Cell viability following Ficoll separation was 93.1 ± 7.8% for PBMC and cell viability following MACS separation was 96.2 ± 1.8% for CD14+ and 93.0 ± 2.5% for CD14+ depleted cells. Further flow cytometric characterization of the CD14- cells showed an average (± SD) of 52.7 ± 9.3%, 20.7 ± 6.5%, 14.5 ± 6.4%, and 30.2 ± 10.4% of cells positive for MHCII, CD21, CD4, and CD3, respectively.
mTOR inhibitor treatments
Rapamycin (RAPA, #9904, Cell Signaling Technology, Danvers, MA, USA) and Torin-1 (#14379, Cell Signaling Technology) inhibitors were resuspended in DMSO, diluted in complete RPMI, and for all treatments added to cell suspensions at a final concentration of 100 nM24.
Generation of M1- and M2-polarized monocyte-derived macrophages (MPh)
Our objective was to study the effect of mTOR inhibition in polarized macrophage subsets. Cells differentiated in the presence of M1 polarization stimuli IFNG and CSF2, referred to as classically activated M1 MPh, are characterized by NOS2 expression and a pro-inflammatory cytokine response. Cells differentiated in the presence of M2 polarization stimuli IL4 and CSF1, referred to as alternatively activated M2 MPh, are characterized by CD163 expression and a greater anti-inflammatory IL10 cytokine response. For the generation of M1 MPh, CD14+ monocytes were differentiated in complete medium with bovine recombinant IFNG (10 ng/mL) and CSF2 (20 ng/mL). To generate M2 MPh, bovine recombinant IL4 and CSF1 (both 20 ng/mL) were added to complete medium. Cells were cultured at 2.5 to 5 × 105 cells/well in 48-well plates at 37 ºC and 5% CO2. After 2 days, differentiation medium was changed and RAPA, Torin-1, or DMSO vehicle control were added. Replicate treatments were cultured in parallel to allow for measurement of different outcomes, described below. One day after addition of inhibitor treatments, all cells were stimulated with LPS (100 ng/mL, E. coli O111:B4, Sigma–Aldrich, Burlington, MA, USA) for either 16 h (cell surface marker expression and cytokine production), 4 h (mRNA expression), or left as unstimulated controls. Culture plates were centrifuged (250 × g, 5 min) and supernatants were collected and stored at -80ºC until cytokine analysis. Cells were detached with Accutase® (Thermo-Fisher Scientific, Waltham, MA, USA) for 10 min (37 ºC, 5% CO2), washed with PBS, and immediately analyzed for surface maker expression. For rt-qPCR cells were lysed in 350 µL RLT (Qiagen, Hilden, Germany) plus 0.1% v/v ß-mercaptoethanol and stored at – 80 ºC until RNA isolation.
Monocyte-derived dendritic cells (moDC) and mixed lymphocyte reaction (MLR) co-culture
We were interested in studying the effect of mTOR inhibition on both naïve and mature moDC as well as on the effect of mTOR inhibited mature moDC on T-cell IFNG response. To generate naïve moDC, CD14+ monocytes were cultured in complete medium plus 20 ng/mL recombinant bovine CSF2 and IL4 as previously described25. Cells were cultured at 2.5 to 5 × 105 cells/well in 48-well culture plates at 37 ºC and 5% CO2. On day 3, differentiation medium was changed and RAPA, Torin-1, or DMSO vehicle control were added with fresh medium. Replicate treatments were cultured in parallel to allow for measurement of different outcomes.
For cell surface marker expression, cytokine production, and gene expression profiles, culture medium was changed to inhibitor-free complete medium after 4 days of culture with inhibitor treatments, and moDC were matured for 16 h (surface marker, protein secretion), or 4 h (gene expression profiles) with LPS (100 ng/mL) or left as naïve moDC, and samples harvested as described above.
To study the effect of mTOR inhibited mature moDC on T-cell activation, we used co-cultures with the CD14-depleted cell fraction of PBMC from the same individual (mixed lymphocytes) in an autologous mixed lymphocyte reaction (MLR). Culture medium was changed to inhibitor-free complete medium after 4 days of culture with inhibitor treatments, and moDC were matured for 24 h with UV irradiated E. coli strain ECC-Z at a multiplicity of infection of 10 as previously described25 or left as naïve cells. Autologous MLR were thawed and incubated overnight in complete medium. Mature moDC and MLR were co-cultured at a ratio of 1:526,27,28 in the presence of anti-bovine CD3 and anti-bovine CD28 (Table 1) at 1 µg/mL for 24 h. Then, supernatants were harvested and stored at – 80 ºC until cytokine analysis.
We used parallel moDC-MLR co-cultures to measure the intracellular IFNG production of T-cells by adding brefeldin A (500 ng/mL) in addition to anti-bovine CD3 and anti-bovine CD28 for 6 h. Monocultures of MLR were incubated in parallel and stimulated with 10 ng/mL PMA and 100 ng/mL ionomycin as positive control or left unstimulated as negative control. Cells were stained with fixable LIVE/DEAD® Near-IR Dead Cell Stain Kit (Invitrogen, Thermo Fisher Scientific), fixed with 2% v/v para-formaldehyde in PBS for 10 min, washed, and stored at 4 ºC until antibody labeling and flow cytometry.
Membrane and intracellular immune fluorescence
We performed membrane immune fluorescence for CD14 expression on PBMC, CD14+ selected and depleted cells, CD163 and MHCII on M1 and M2 MPh, CD80 and MHCII on moDC, and for CD4 expression on autologous mixed lymphocytes to describe cell lineage. Cells were washed in FACS buffer (0.05% w/v sodium azide and 0.5% w/v BSA in PBS), pelleted, and incubated with primary antibodies for 30 min at 4 ºC. Then, cells were washed twice and incubated with FITC-conjugated goat pAb: anti-mouse IgG1 (Bio-Rad Laboratories) diluted in FACS buffer at 1:100 for 30 min at 4 °C and washed twice with FACS buffer. Samples were transferred to microfuge vials containing 100 μL propidium iodide (PI; 20 μg/mL), and immediately analyzed by flow cytometry. Intracellular fluorescence was performed with fixed moDC-MLR co-cultures after membrane immune fluorescence labeling was performed as described above. Samples were analyzed using an Attune acoustic focusing flow cytometer (Thermo Fisher Scientific) or, for intracellular fluorescence, a FACS Canto II flow cytometer (BD Biosciences). Doublets were excluded based on forward scatter (FSC) height and FSC area. Dead cells were excluded by gating on PI-negative events (unfixed cells) or on LIVE/DEAD Near-IR negative events (fixed cells). Cell morphology gates were drawn in forward and side scatter and 10,000 events were acquired for each measurement. Mean fluorescence intensity was corrected for background from bioparticles and loading dye, or background fluorescence of unstained or secondary controls, respectively. Data were analyzed using FlowJo V10 software (BD Biosciences, San Jose, CA).
Phagocytosis and oxidative burst assays
Phagocytosis and oxidative burst of M1 and M2 MPh were analyzed in duplicate after 2 days of culture with inhibitor treatments. For measurement of phagocytosis, we first sonicated pHrodo bioparticles (pHrodo Green Zymosan Bioparticles Conjugate, Invitrogen, Carlsbad, CA), diluted these in 1X HEPES buffer (pH, 7.4), and opsonized with 1% v/v autologous serum for 1 h at 37 °C. Cells were incubated with pHrodo bioparticles or left as unstimulated control for 1 h at 37 °C.
For measurement of oxidative burst, we incubated cells with 10 µM H2DCFDA fluorescein loading dye (Invitrogen, Carlsbad, CA) for the detection of reactive oxygen species or left unstained as background control for 15 min at 37 °C. Subsequently, cells were stimulated with PMA at final concentrations of 0, 10, and 25 ng/mL, or left as unstimulated control for 15 min at 37 °C. Following incubation, cells were placed on ice, mixed with 100 μL propidium iodide (20 μg/mL) to assess cell viability, and analyzed by flow cytometry.
Cytokine production and reverse-transcription quantitative PCR
Culture supernatants were used to quantify the concentration of IL10, TNF, and IFNG in a bead-based multiplex assay as described29.
For RNA extraction, cell lysates in RLT buffer were rapidly thawed at 37 °C for 10 min and homogenized using a spin column (QIAshredder, Qiagen, Venlo, Netherlands). RNA was extracted using the RNeasy Plus Mini Kit (Qiagen) and characterized and quantified using a NanoDrop One spectrophotometer (Thermo Fisher Scientific). Samples with an optical density 260:280 ratio ≥ 1.8 but ≤ 2.2 were processed further. A volume containing 0.25 μg of RNA was reverse transcribed (SuperScript IV First-Strand cDNA Synthesis, Life Technologies, Carlsbad, CA, USA) and stored at − 20 °C. A pool of cDNA from all samples served as the calibrator sample. Bovine-specific primer probe sets (TaqMan Gene Expression Assays, Applied BioSystems, Thermo Fisher Scientific) with exon spanning probes were purchased for genes of interest [IL12A (p35): Bt03213922, IL12B (p40): Bt03213923, IL1B: Bt03212741, IL6: Bt03211905, NOS2: Bt03249586, ARG2: Bt03219827] and 2 reference genes (RPLP0: Bt03218087, TBP: Bt03241946). Both reference genes were previously shown to be stably expressed in bovine leukocytes and used in a similar experimental setup21. Real-time reverse-transcription quantitative PCR was performed in triplicate using a 1:3 (moDC) or 1:6 (M1 and M2 Mph) dilution of cDNA at 20% v/v of the final 10-μL reaction volume using TaqMan Gene Expression Master Mix (Applied BioSystems, Thermo Fisher Scientific) in a QuantStudio 12 K Flex Real-Time PCR System (Applied BioSystems, Thermo Fisher Scientific). Run conditions for all genes of interest were 1 cycle at 50 °C for 2 min, 1 cycle at 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Raw data were analyzed using the comparative quantification method (ΔΔCt method, QuantStudio 12 K Flex Software, v1.2.2) and expressed as relative quantity (RQ = 2−ΔΔCt) before export for statistical analysis.
Analytical approach
Sample size was based on the expected difference in IFNG production of co-culturing MLR with rapamycin/Torin-1-conditioned moDC. Our conservative expectation was to find an effect at least as large as 50% of the effect of mTOR inhibition on increased IFNG production as a marker for Th1-cell response performed in human cells described previously26. The sample size of 12 animals is based on an expected difference of 25 ng/mL (90 vs. 115 ng/mL) with a conservative SD of 20 ng/mL of IFNG with probabilities of type I and II error set to 0.05 (alpha = 0.05; power = 0.95), using a statistical software program (GPower v. 3.1.9.230) with assumed correlation between samples within a cow of 0.5.
All data were analyzed in the statistical software package SAS (v. 9.4, SAS Institute Inc., Cary, NC, USA). Data describing the study population and cell subsets were summarized in PROC UNIVARIATE. All other data were analyzed using mixed effects ANOVA in PROC MIXED. Models were corrected for parity and DIM and included a random effect of cow. Outlier diagnostics were performed as defined a priori with the INFLUENCE statement and removed if Cook’s D was > 0.5. Model residuals were visually inspected for normality and homoscedasticity and data log transformed to fulfill model assumptions as indicated. Results are presented as least squares means (LSM) and standard error (SE), or geometric LSM and backtransformed 95% confidence interval (CI) unless otherwise specified. Graphs were produced in GraphPad Prism (v. 9.3.1, La Jolla, CA).
For all models, pairwise comparisons within fixed effects were performed if the F-test for the effect yielded a P ≤ 0.05. Multiple comparisons corrections were performed with Bonferroni or Tukey’s correction as indicated.
Differences between polarized MPh surface markers, gene expression, and cytokine production with and without LPS stimulation were analyzed in a model that included fixed effect of cell type, stimulus, and cell type × stimulus interaction. Pairwise comparisons were applied to LSM to test the effect of stimulus within and between MPh with Bonferroni correction. Difference in phagocytosis was analyzed in a model with fixed effect of MPh type and differences in oxidative burst were analyzed in a model with fixed effect of MPh type, PMA dose, and cell type × PMA interaction; pairwise comparisons were corrected using Tukey’s posthoc procedure. Data for phagocytosis, gene expression, and cytokine production were log transformed before analysis. Differences between naïve and LPS-matured moDC, and between naïve and E. coli-matured moDC in MLR co-cultures, were analyzed in a model with fixed effect of LPS or E. coli treatment, respectively. Gene expression and cytokine production data were log transformed. Treatment differences in polarized MPh and moDC phenotype and function were analyzed separately by stimulation (without or with LPS or PMA in MPh), or maturity (without or with LPS or E. coli in naïve and mature moDC, respectively) before analysis. Surface marker, phagocytosis, oxidative burst, and cytokine production data were all expressed relative to CTRL and log transformed. Gene expression data were analyzed as the fold change (FC) from CTRL and log transformed. All data were analyzed with fixed effect of inhibitor treatment to test the effect of RAPA vs. TORIN-1 and pairwise comparison of RAPA vs. 0 and TORIN-1 vs. 0 in LSMEANS, all with Bonferroni correction. Gene expression data are presented as FC from CTRL, and all other data are presented as the relative change from CTRL.
Results
We investigated the effect of pharmacological mTOR inhibition during differentiation of monocytes to M1 (IFNG, CSF2) and M2 (IL4, CSF1) MPh, as well as moDC (IL4, CSF2) to simulate prolonged cellular nutrient deficit as experienced by the transition dairy cow. Generated M1 and M2 MPh were clearly distinct in phenotype and function, with differences in expression of MHCII, CD163, and NOS2 expression (Supplementary Fig. 1), as well as phagocytosis, oxidative burst, and cytokine mRNA expression (Supplementary Fig. 2).
Inhibition of mTOR generated inflammatory M1 with decreased phagocytosis and IL10 production
We found distinct effects of both mTOR inhibitors on the phenotype of classically activated M1 MPh (with IFNG) that were additionally stimulated with LPS or left unstimulated. Surface MHCII expression was increased (P < 0.0001) in Torin-1-treated, but not (P = 0.30) RAPA-treated unstimulated M1 cells; no effect (P > 0.70) of inhibitor treatment on MHCII expression was found in LPS-stimulated cells (Fig. 1A). Both RAPA and Torin-1-treated cells had decreased (P ≤ 0.06) ARG2 expression in unstimulated and stimulated M1 MPh (Fig. 1B).
Changes in surface maker expression, mRNA abundance, oxidative burst, and phagocytosis in M1 macrophages polarized for 3 days with 10 ng/mL IFNG and 20 ng/mL CSF2 and concurrently treated with vehicle (CTRL), or inhibitors rapamycin (RAPA) or Torin-1 (T1) at 100 nM. Cells were incubated with vehicle (− LPS) or stimulated with 100 nM LPS (+LPS) for 16 h to measure change in MHCII expression (A), or 4 h to measure changes in mRNA abundance of arginase 2 (ARG2, B) and inducible nitric oxide synthase (NOS2, C) as well as their ratio (D). Cells were incubated with 10 µM H2DCFDA fluorescein loading dye for 15 min and stimulated with phorbol myristate acetate (PMA) at 0 (− PMA), or 25 (+ PMA) ng/mL for 15 min to measure oxidative burst (E) or with pHrodo bioparticles for 1 h to measure cellular phagocytosis (F). Data were expressed relative to CTRL and presented as the relative change from CTRL in unstimulated and stimulated cells for MHCII, ARG2:NOS2, phagocytosis, and oxidative burst. mRNA abundance was expressed as the fold change from CTRL in unstimulated and stimulated cells. Data from cell isolation of 12 cows are presented as the geometric mean and backtransformed 95% confidence interval. ANOVA tests between CTRL and inhibitors, and between inhibitors are presented with Bonferroni-adjustment for multiple comparisons: P ≤ 0.10 (#), P ≤ 0.05 (*), P ≤ 0.01 (**), P ≤ 0.0001 (***).
Expression of NOS2 increased (P ≤ 0.04) in RAPA-treated and LPS-simulated cells and the ratio of ARG2:NOS2 was decreased (P < 0.01) in both RAPA and Torin-1 for unstimulated and stimulated cells (Fig. 1C, D). As expected, expression of CD163 was not detected or was minimal in M1 MPh and was therefore not analyzed further.
The inhibition of mTOR during differentiation also affected intracellular killing potential of M1 MPh. Phagocytosis was decreased by both RAPA (P = 0.01) and Torin-1 (P = 0.01) treatment (Fig. 1). Moreover, oxidative burst was increased (P < 0.01) in Torin-1 treated cells, but not in RAPA treated cells (P > 0.99) (Fig. 1E).
The two mTOR inhibitors also affected mRNA expression of the two IL12 subunits. Expression of IL12A was decreased (P ≤ 0.01) in Torin-1-treated unstimulated and stimulated M1 MPh but increased by two-fold (P = 0.0009) in RAPA-treated stimulated cells (Fig. 2A). Expression of IL12B was increased more than two-fold (P ≤ 0.05) in Torin-1-treated unstimulated and more than ten-fold in stimulated M1 MPh (Fig. 2B). Treatment with either inhibitor decreased IL1B and IL6 expression (P ≤ 0.07) in unstimulated cells, but only Torin-1 decreased (P < 0.0001) cytokine expression in LPS-stimulated M1 MPh (Fig. 2C, D). Importantly, the secretion of IL10 in culture medium was decreased for both RAPA and Torin-1-treated naïve and stimulated M1 MPh (P ≤ 0.02). Inhibitor treatment did not change (P ≥ 0.14) the secretion of TNF from cells (Fig. 2E, F). Overall, these changes in cytokine expression profile show a selective effect of mTOR inhibition on the pro-inflammatory cytokine profile of M1 MPh, but a consistent decrease in the anti-inflammatory cytokine IL10.
Changes in cytokine mRNA abundance and production in M1 macrophages polarized for 3 days with 10 ng/mL IFNG and 20 ng/mL CSF2 and concurrently treated with vehicle (CTRL), or inhibitors rapamycin (RAPA) or Torin-1 (TR1) at 100 nM. Cells were incubated with vehicle (− LPS) or stimulated with 100 nM LPS (+ LPS) for 4 h to measure changes in mRNA abundance of IL12A, IL12B, IL1B, and IL6 gene expression (A–D) or for 16 h to measure production of IL10 and TNF protein (E, F). mRNA abundance was expressed as the fold change from CTRL in unstimulated and stimulated cells. Data were expressed relative to CTRL and presented as the relative change from CTRL in unstimulated and stimulated cells for cytokine production. Data from cell isolation of 12 cows are presented as the geometric mean and backtransformed 95% confidence interval. ANOVA tests between CTRL and inhibitors, and between inhibitors are presented with Bonferroni-adjustment for multiple comparisons: P ≤ 0.10 (#), P ≤ 0.05 (*), P ≤ 0.01 (**), P ≤ 0.0001 (***).
Inhibition of mTOR interfered with the differentiation of M2 macrophages
Next, we examined the effect of mTOR inhibitors during polarization on M2 MPh that were left unstimulated or stimulated with LPS. Expression of the M2 marker CD163 was decreased (P ≤ 0.04) by RAPA and Torin-1 in unstimulated M2 MPh. Only Torin-1 significantly decreased (P = 0.003) CD163 expression in LPS-stimulated M2 (Fig. 3A). Expression of ARG2 was decreased (P ≤ 0.01) in both RAPA and Torin-1-treated unstimulated and LPS-stimulated cells, whereas expression of NOS2 was decreased (P ≤ 0.009) in both RAPA and Torin-1-treated stimulated cells (Fig. 3B, C). Changes in expression of ARG2 and NOS2 resulted in decreased (P ≤ 0.05) ratio of ARG2:NOS2 in RAPA- and Torin-1-treated unstimulated M2, but the effect did not differ (P = 0.36) by inhibitor treatment, and inhibitor treatment did not change (P ≥ 0.99) the ratio in stimulated M2 (Fig. 3D).
Changes in surface maker expression, mRNA abundance, oxidative burst, and phagocytosis in M2 macrophages polarized for 3 days with 20 ng/mL IL4 and 20 ng/mL CSF1 and concurrently treated with vehicle (CTRL), or inhibitors rapamycin (RAPA) or Torin-1 (TR1) at 100 nM. Cells were incubated with vehicle (− LPS) or stimulated with 100 nM LPS (+ LPS) for 16 h to measure change in CD163 expression (A), or 4 h to measure changes in mRNA abundance of arginase 2 (ARG2, B) and inducible nitric oxide synthase (NOS2, C) and their ratio (D). Cells were incubated with 10 µM H2DCFDA fluorescein loading dye for 15 min and stimulated with phorbol myristate acetate (PMA) at 0 (− PMA), or 25 (+ PMA) ng/mL for 15 min to measure oxidative burst (E), or for 1 h with pHrodo bioparticles to measure cellular phagocytosis (F). Data were expressed relative to CTRL and presented as the relative change from CTRL in unstimulated and stimulated cells for CD163, ARG2:NOS2, phagocytosis, and oxidative burst. mRNA abundance was expressed as the fold change from CTRL in unstimulated and stimulated cells. Data from cell isolation of 12 cows are presented as the geometric mean and backtransformed 95% confidence interval. ANOVA tests between CTRL and inhibitors, and between inhibitors are presented with Bonferroni-adjustment for multiple comparisons: P ≤ 0.10 (#), P ≤ 0.05 (*), P ≤ 0.01 (**), P ≤ 0.0001 (***).
Different from M1 MPh, we only observed moderate changes in functional assays in response to mTOR inhibition in M2 MPh. Compared with CTRL, RAPA treatment did not affect (P ≥ 0.34) oxidative burst, but Torin-1 decreased (P = 0.08) the oxidative burst response (Fig. 3E). No effect (P ≥ 0.81) of inhibitor treatment on phagocytosis was found (Fig. 3F).
Unlike the opposing effects that mTOR inhibition had on M1, expression of IL12A and IL12B was only affected by Torin-1 in M2 MPh. Expression of IL12A was decreased (P < 0.0001) in Torin-1-treated unstimulated and LPS-stimulated M2 whereas LPS-stimulated Torin-1-treated M2 cells had decreased (P = 0.0009) IL12B (Fig. 4A, B). Expression of IL1B was decreased (P < 0.0001) in both unstimulated and stimulated Torin-1-treated M2 and decreased (P ≤ 0.009) in RAPA and Torin-1-treated M2 MPh (Fig. 4C). Treatment with RAPA during polarization was associated with decreased (P ≤ 0.05) IL6 expression in unstimulated and stimulated cells, but Torin-1 increased IL6 (P = 0.09) in unstimulated M2 cells whereas it decreased IL6 (P = 0.03) in LPS-stimulated cells (Fig. 4D). Concentration of IL10 was decreased (P < 0.001) in cell culture supernatant from unstimulated and stimulated cells that were treated with Torin-1 during polarization, and to a smaller degree in LPS-stimulated cells treated with RAPA (P < 0.0001) (Fig. 4E). Concentration of TNF was decreased (P < 0.0001) in cell culture supernatant from stimulated cells that were treated with Torin-1 during polarization (Fig. 4F). Overall, mTOR inhibition during differentiation of M2 MPh resulted in decreased expression of pro-inflammatory cytokines except for IL12B and IL6 in unstimulated M2, and the same consistent decrease in IL10 production was observed as in M1 MPh under mTOR inhibition.
Changes in cytokine mRNA abundance and production in M2 macrophages polarized for 3 days with 20 ng/mL IL4 and 20 ng/mL CSF1 and concurrently treated with vehicle (CTRL), or inhibitors rapamycin (RAPA) or Torin-1 (TR1) at 100 nM. Cells were incubated with vehicle (− LPS) or stimulated with 100 ng/mL LPS (+ LPS) for 4 h to measure changes in mRNA abundance of IL12A, IL12B, IL1B, and IL6 (A–D), or for 16 h to measure production of IL10 and TNF protein (E, F). mRNA abundance was expressed as the fold change from CTRL in unstimulated and stimulated cells. Data from cell isolation of 12 cows were expressed relative to CTRL and presented as the relative change from CTRL in unstimulated and stimulated cells for cytokine production. Data are presented as the geometric mean and backtransformed 95% confidence interval. ANOVA tests between CTRL and inhibitors, and between inhibitors are presented with Bonferroni-adjustment for multiple comparisons: P ≤ 0.10 (#), P ≤ 0.05 (*), P ≤ 0.01 (**), P ≤ 0.0001 (***).
Inhibition of mTOR during moDC differentiation increased co-stimulatory potential and favored a Th1 cytokine profile
As a third monocyte-derived cell population we investigated how treatment with mTOR inhibitors affected the differentiation of moDC, their response when matured with LPS, and cytokine production when co-cultured with an autologous MLR.
First, we were interested in examining possible changes in naïve moDC to characterize the potential effects of mTOR inhibition on DC prior to antigen encounter, with results indicating no significant changes in phenotype (Fig. 5) with the exception of greater surface MHCII expression (P = 0.08) in RAPA-treated naïve moDC.
Changes in surface maker expression, mRNA abundance, and cytokine production in moDC after differentiation for 7 days with 20 ng/mL CSF2 and IL4 and concurrently treated with vehicle (CTRL), or inhibitors rapamycin (RAPA) or Torin-1 (TR1) at 100 nM during the last 4 days of differentiation. Naïve moDC (Naïve) were matured with 100 ng/mL LPS (Mature) for 16 h to characterize surface markers CD80 (A) and MHCII (B) and measure cytokine production of IL10 (G) and TNF (H) protein, and for 4 h to quantify mRNA abundance of IL12A, IL12B, IL1B, and IL6 (C–F). Data were expressed relative to CTRL and presented as relative change from CTRL in naïve and mature moDC for surface marker expression and cytokine production. mRNA abundance was expressed as the fold change from CTRL in naïve and mature moDC. Data from cell isolation of 12 cows are presented as geometric mean and backtransformed 95% confidence interval. ANOVA tests between CTRL and inhibitors, and between inhibitors are presented with Bonferroni-adjustment for multiple comparisons: P ≤ 0.10 (#), P ≤ 0.05 (*), P ≤ 0.01 (**), P ≤ 0.0001 (***).
Next, we analyzed the effects of the inhibitors on mature moDC. The LPS-matured moDC were characterized by up-regulation of antigen-presenting and co-stimulatory molecules needed to activate T cells and by their cytokine profile (Supplemental Fig. 3). Surface CD80 expression was increased (P = 0.01) in Torin-1-treated mature moDC (Fig. 5A) and MHCII tended to decrease (P = 0.10) in Torin-1-treated mature moDC (Fig. 5B). Treatment with mTOR-inhibitors during moDC differentiation changed the expression profile of selected cytokines in mature moDC (Fig. 5C–F): both Torin-1 (P = 0.0006) and RAPA (P = 0.02) increased expression of IL12A, only Torin-1 increased expression of IL12B, IL1B and IL6, whereas secretion of IL10 and TNF in cell culture supernatants of mature moDC was not altered (P ≥ 0.16) by inhibitor treatment (Fig. 5G, H). In summary, mTOR inhibition with Torin-1 altered moDC surface marker expression, increased expression of pro-inflammatory cytokines, but did not affect the concentration of anti-inflammatory IL10, whereas RAPA showed only minor effects.
Mature mTOR inhibitor-treated moDC were co-cultured with autologous MLR to study changes in Th1-derived IFNG production as an indirect assessment of moDC IL12 cytokine production. Concentration of IFNG was 20-fold greater (P < 0.0001) in co-cultures with mature moDC [3.0 (1.5–6.0) vs. 69 (36–131) ng/mL in naïve and mature moDC, respectively]. Concentration of IFNG was increased (P ≤ 0.07) in co-cultures of naïve and mature moDC that were treated with Torin-1 (Fig. 6C), but no treatment differences were found in intracellular IFNG measurements of CD4+ cells (Fig. 6D, E). As described above, the sample size of 12 individual cows for this study was based on an expected difference of 25 ng/mL and a SD of 20 ng/mL IFNG between treatments. Concentration of IFNG in mature co-cultures averaged (± SD) 89.0 ± 46.8, 134 ± 49, and 147 ± 40 ng/mL for CTRL, RAPA, and Torin-1 respectively, showing that both magnitude of difference between the mTOR inhibitor treatments and CTRL, as well as SD were about twice as high as described above. Regarding the concentration of other cytokines of interest in co-culture, IL10 was decreased (P ≤ 0.04) in supernatants of Torin-1-treated naïve and mature moDC, whereas TNF concentrations increased (P ≤ 0.03) in co-cultures of both naïve and mature moDC that were treated with RAPA or Torin-1 (Fig. 6A, B).
Changes in cytokine concentration of IL10 (A), TNF (B), and IFNG (C) measured in supernatants from moDC that were concurrently treated with vehicle (CTRL), or inhibitors rapamycin (RAPA) or Torin-1 (TR1) at 100 nM during the last 4 days of differentiation before maturation with E. coli (Mature), co-cultured with mixed lymphocyte reaction (MLR), and T-cell activated with anti CD3/CD28. Naïve moDC that were not matured with E. coli were co-cultured and activated in parallel (Naïve). (D) Intracellular IFNG expression in CD4+ MLR was analyzed via flow cytometry of MLR co-cultured for 1 day with in vitro-generated moDC (7-day differentiation protocol with 20 ng/mL CSF2 and IL4, matured for 1 day with E. coli) before activation with anti CD3/CD28 and concurrent incubation with brefeldin A for 6 h. Cells were labeled with anti-bovine CD4 FTIC-conjugated antibodies and after fixation and permeabilization with anti-bovine IFNG A647. Contour plots show fluorescence for CD4-FITC and IFNG A647 of MLR negative controls (w/o moDC), positive controls (stimulated with phorbol myristate acetate (PMA) and ionomycin (Iono), and co-cultures of MLR with moDC treated with vehicle, RAPA or Torin-1. (E) Mean fluorescence intensity (MFI) of IFNG staining in CD4+ MLR relative to CRTL. Data from cell isolation of 12 cows were expressed relative to CTRL and presented as relative change from CTRL in naïve and mature moDC. Data are presented as geometric mean and backtransformed 95% confidence interval. ANOVA tests between CTRL and inhibitors, and between inhibitors are presented with Bonferroni-adjustment for multiple comparisons: P ≤ 0.10 (#), P ≤ 0.05 (*), P ≤ 0.01 (**), P ≤ 0.0001 (***).
Discussion
Our objective in this study was to investigate the role of mTOR pathway inhibition on key regulators of the bovine innate and early adaptive immune response. We simulated nutrient deficit-induced mTOR suppression by using pharmacological mTOR inhibition in an in vitro model. Postpartum dairy cows undergo a time of pronounced nutrient deficit with decreased availability of energy and essential amino acids, as well as reduced circulating concentrations of insulin and IGF131,32,33. At the same time, these animals often display immune dysfunction that is characterized by a heightened inflammatory response and decreased ability to clear pathogens7,34. Several studies in cattle have shown an association of the severity of negative nutrient balance and immune dysfunction35,36, but the causal mechanisms are not well understood to date. In other species, research has shown that the mammalian metabolic status is integrally linked with immune cell function, and that the mTOR pathway is a key integrator of nutrient-sensing and immune activation37. Our previous work indicates a reduced activation of components of the AKT-mTOR pathway in muscle, adipose tissue and peripheral blood mononuclear cells harvested from postpartum dairy cows15,16,17,38.
The observed net effect of mTOR inhibition on M1 MPh in our model suggests a selective regulation of M1 features with potentially decreased pathogen clearance and activation of other phagocytic cells, but concurrent increase in production of potentially damaging free radicals. Activation of M1 MPh typically occurs upon encountering a pathogen, whereas M2 MPh play a role in the resolution of this response39,40. Short-term (48 h) addition of RAPA at the end of a differentiation and polarization protocol shifted M2 polarized human MPh towards an M1-like profile, and increased cell apoptosis20. In our LPS-stimulated bovine M1 MPh model, long-term exposure to mTOR inhibitors during cell differentiation appeared to selectively affect the pro-inflammatory M1 response to stimulation. This was represented by a decrease of the ARG2:NOS2 ratio (RAPA, Torin-1), increase of IL12A (RAPA) or IL12B (Torin-1), as well as increased ROS production and decreased anti-inflammatory IL10 production (RAPA, Torin-1). This is in line with other findings in murine models where mTOR inhibition induced an M1 phenotype and genetic deletion of mTORC1 enhanced pro-inflammatory cytokines41,42. In contrast to those findings, we observed that both inhibitors decreased phagocytosis of bovine M1, and treatment with Torin-1 specifically decreased IL6, and IL1B expression in LPS-stimulated M1.
Furthermore, mTOR inhibition impaired M2 polarization, measured by lower production of IL10, drastically reduced expression of CD163 and ARG2, which are hallmark features of this regulatory cell type43. Polarization towards an M2 phenotype is mTOR-dependent44, and inhibition of this pathway during M2 polarization may reduce survival20. This shift toward an M1 phenotype is consistent with changes observed in human MPh20. The results from our model suggest that mTOR inhibition might prevent the IL4 driven differentiation of M2 which could impair restoration of homeostasis and extend the pro-inflammatory response. In response to LPS stimulation however, we demonstrated that M2 differentiated under mTOR inhibition were clearly different in response to M1 MPh and showed decreased expression of pro-inflammatory cytokines (IL12A, IL12B, IL1β, IL6) as well as TNF concentration in addition to drastically decreasing CD163 surface expression. These results indicate that mTOR inhibition during M2 differentiation did not completely change polarization of these cells to M1 but rather induced an M1-like cell type as we could show that mTOR inhibited M2 did not develop the same pro-inflammatory profile as M1 polarized MPh.
In mature bovine moDC, mTOR inhibition increased expression of co-stimulatory CD80, as previously described in other species13 and in a previous experiment using bovine moDC generated from pregnant, non-lactating cows45. In an in-vitro model comparing bovine moDC from cows in different stages of gestation and lactation, Pomeroy et al.9 showed an increase of CD80 and MHCII in moDC from cows 4–6 days post-calving compared with moDC from mid and late gestation cows. Our results indicate that mTOR inhibition could be a mechanism contributing to this postpartum change in expression of co-stimulatory molecules. Complete mTOR pathway inhibition with Torin-1 in our model also increased expression of pro-inflammatory cytokines (IL12A, IL12B, IL1β, IL6) in E. coli- matured moDC which is consistent with findings in other species19,46.
As hypothesized, our study found effects of mTOR inhibition on immune phenotype, function, and cytokine profile of the cell types under study, albeit results differed between incomplete (RAPA) and complete (Torin-1) mTOR-pathway inhibition. Rapamycin, a naturally occurring first-generation mTOR inhibitor, interacts specifically with mTORC1, thereby inhibiting some of its functions47. However, rapamycin inhibition of mTORC1 shows feedback regulation, whereby mTORC1 4E-BP1 substrate phosphorylation is partially recovered over time47,48. Torin-1 is a second-generation mTOR inhibitor that reduces the kinase activity of both mTORC1 and mTORC2, blocking mTORC1 more effectively47 by evading the feedback loop and rapamycin-resistant functions of mTORC1 that complicates the use of rapamycin49. In general, the two inhibitors behaved similarly in our model with more pronounced changes in Torin-1-treated cells, which is encouraging to enhance our understanding of mTOR inhibition in bovine immune cells. Some effects, however, differed between RAPA and Torin-1. Most remarkably in this context we found opposing effects of the two inhibitors on expression of IL12A in M1 and M2 MPh, where RAPA induced up-regulation but treatment with Torin-1 lead to down-regulation. Additionally, we observed opposing effects of RAPA and Torin-1 on MHCII expression in naïve and mature moDC with RAPA increasing expression in naïve moDC, and Torin-1 decreasing MHCII expression in mature moDC. In a previous study, both rapamycin as well as PP242, a pan-mTOR inhibitor similar in action to Torin-1, increased MHCII expression, albeit not statistically significant for PP24245. Complete pathway inhibition by mTORC1/2 inhibitors such as Torin-1 may elicit different effects on moDC in our model than possibly incomplete mTORC1-specific inhibition by rapamycin as previously discussed by Weichhart et al.12. While mTORC1 regulates a cell´s growth and autophagy in response to nutrients, mTORC2 mediates other cell functions such as cytoskeletal organization50. Alternatively, effects of the type of mTOR inhibition might be cell-type specific, or inhibitor effects might be dose-dependent, and this was not investigated in our current work. In a recent study on mTOR-driven aging, authors showed that at low concentrations mTORC1/2 inhibitors acted like rapamycin, and that was particularly true for Torin-1 and PP242, although effects varied when concentrations increased51. A cytostatic effect in the model of Leontieva et al.51 was found at 30 nM for Torin-1 and 285 nM for PP242. Therefore, differences due to cell-type or dose-dependent responses to each type of inhibitor merit further attention in follow-up studies.
Ideally, the effects of mTOR inhibition could be tested in vivo in dairy cattle to help clarify the whole-body response to reduced mTOR activation. The use of mTOR inhibitors in this food-producing animal species would require authorization and removal of the animals and their products from the food chain. Alternatively, the mTOR pathway could be stimulated by supplying nutrients that are sensed by this pathway during times of deficit, either parenterally or through feed supplementation. Lastly, given that the greatest nutrient needs in the postpartum period are due to the requirements of lactation, cows that produce very little or no milk after calving could be compared with high producing cows regarding their mTOR activity. All aforementioned model systems carry their own limitations but could each contribute significantly to the further understanding of the mTOR pathway on postpartum immune phenotype of the dairy cow.
By using IFNG and LPS for polarization and stimulation of M1, and LPS and irradiated E. coli for maturation of moDC, we biased M1 and moDC towards a pro-inflammatory response52,53. However, gram-negative pathogens, specifically E. coli, are the most common infectious cause of severe inflammation in postpartum cows7,54.
We recognize our bovine model favored a Th1-promoting response a priori by using LPS for moDC maturation52. However, under such conditions we found further Th1-enhancing effects in mTOR-suppressed moDC compared with untreated controls. Decreased IL10 and increased IFNG and TNF concentrations in co-culture supernatants of Torin-1-treated moDC and MLR supports our hypothesis that inhibition of the mTOR-pathway in bovine moDC favors Th1 skewing. These findings are in line with our previous experiment showing increased TNF and decreased IL10 concentrations in rapamycin inhibited moDC, suggesting that mTOR inhibited moDC would favor a Th1 immune response45. When using mTOR-inhibited mature DC as stimulators for murine lymphocytes in co-culture, proliferating CD4+ T-cell populations displayed a Th1 phenotype26,55. We interpret observations in mTOR inhibited moDC in this model such that excessive nutrient deficit induced mTOR suppression in postpartum dairy cows might lead to a more pronounced Th1-dominated inflammatory response with stronger co-stimulatory potential of moDC and increased expression of pro-inflammatory cytokines.
Summarizing the results of our study, we showed in this in-vitro model that reduced activation of the mTOR pathway alters the phenotype and cytokine profile in bovine monocyte-derived innate immune cells. An overview of these changes is given in Fig. 7. Selective reduction of cytokine expression and increased ROS production in LPS-stimulated M1, inhibition of M2 differentiation, and enhanced activation of moDC with a more pronounced Th1 response could contribute to a pro-inflammatory environment, delayed resolution of inflammatory response, and decreased pathogen clearance in vivo.
Summary schematic of hypothesized effects of mTOR inhibition during differentiation of myeloid cell populations in the presence of gram-negative bacterial components (E. coli lipopolysaccharide, LPS). Cows experience a nutrient deficit postpartum at the same time as immune dysfunction. We previously showed that the activation of the mTOR pathway is reduced in postpartum dairy cows. Summarized results are derived from in-vitro pharmacological mTOR inhibition with rapamycin (RAPA) or Torin-1 (TR1) in monocyte derived macrophages (M1, M2), dendritic cells (moDC), and co-culture of moDC with autologous mixed lymphocytes (MLR). We propose that the inflammatory response during nutrient deficit associated mTOR-pathway inhibition may result in the following effects: (1) increased production of reactive oxygen species (ROS) and expression of nitric oxide synthase 2 (NOS2) in M1, (2) decreased pathogen clearance by phagocytosis in M1, (3) selective regulation of pro-inflammatory mediators in M1 (IL12A, IL12B, IL6, IL1β), (4) increased expression of pro-inflammatory, Th1-promoting cytokines in moDC (IL12A, IL12B, IL6, tumor necrosis factor alpha, TNF) and a more potent activation of T cells by increased expression of co-stimulatory molecules (CD80) and cytokine profile changes (decreased IL10, increased TNF and IFNG) in moDC-MLR co-cultures, and (5) decrease in the M2 restorative phenotype hallmark features such as expression of arginase 2 (ARG2), surface CD163, and production of anti-inflammatory IL10. Overall, the observed changes indicate an inflammatory response that is heightened in magnitude and potentially in duration.
The resulting alteration of phenotype and cytokine profile in these key regulators of inflammation indicate that reduced activation of the mTOR pathway could off-set an adequate inflammatory response in postpartum dairy cows. A certain degree of increase in the reactive and pro-inflammatory immune response is considered physiological in the postpartum period56,57 and we previously showed that pro-inflammatory cytokines are upregulated postpartum in bovine whole blood leukocytes17. However, cows overexpressing pro-inflammatory cytokines over a prolonged period systemically and locally are at higher disease risk and show overall decreased efficiency of nutrient metabolism and milk production7. An excessive nutrient deficit might skew this balance towards heightened and prolonged inflammation with a decrease in pathogen clearance and restoration of homeostasis.
In conclusion, our results based on in vitro pharmacological mTOR inhibition indicate that suppression of mTOR-pathway activation caused by the lack of nutrient availability in postpartum dairy cows might contribute to the immune dysfunction observed during the critical postpartum time of dairy cows.
Data availability
The data that support the findings of this study are available, on reasonable request, from the corresponding author.
References
Schukken, Y. H., Erb, H. N. & Scarlett, J. M. A hospital-based study of the relationship between retained placenta and mastitis in dairy cows. Cornell Vet. 79, 319–326 (1989).
Esposito, G., Irons, P. C., Webb, E. C. & Chapwanya, A. Interactions between negative energy balance, metabolic diseases, uterine health and immune response in transition dairy cows. Anim. Reprod. Sci. 144, 60–71. https://doi.org/10.1016/j.anireprosci.2013.11.007 (2014).
Janosi, S. et al. Energy imbalance related predisposition to mastitis in group-fed high-producing postpartum dairy cows. Acta Vet. Hung. 51, 409–424. https://doi.org/10.1556/AVet.51.2003.3.14 (2003).
Alvasen, K. et al. Risk factors associated with on-farm mortality in Swedish dairy cows. Prev. Vet. Med. 117, 110–120. https://doi.org/10.1016/j.prevetmed.2014.08.011 (2014).
Shahid, M. Q., Reneau, J. K., Chester-Jones, H., Chebel, R. C. & Endres, M. I. Cow- and herd-level risk factors for on-farm mortality in Midwest US dairy herds. J. Dairy Sci. 98, 4401–4413. https://doi.org/10.3168/jds.2014-8513 (2015).
Sordillo, L. M. & Raphael, W. Significance of metabolic stress, lipid mobilization, and inflammation on transition cow disorders. Vet. Clin. N. Am. Food Anim. Pract. 29, 267–278. https://doi.org/10.1016/j.cvfa.2013.03.002 (2013).
Bradford, B. J., Yuan, K., Farney, J. K., Mamedova, L. K. & Carpenter, A. J. Invited review: Inflammation during the transition to lactation: New adventures with an old flame. J. Dairy Sci. 98, 6631–6650. https://doi.org/10.3168/jds.2015-9683 (2015).
Yuan, K., Farney, J. K., Mamedova, L. K., Sordillo, L. M. & Bradford, B. J. TNFalpha altered inflammatory responses, impaired health and productivity, but did not affect glucose or lipid metabolism in early-lactation dairy cows. PLoS ONE 8, e80316. https://doi.org/10.1371/journal.pone.0080316 (2013).
Pomeroy, B., Sipka, A., Klaessig, S. & Schukken, Y. Longitudinal characterization of bovine monocyte-derived dendritic cells from mid-gestation into subsequent lactation reveals nadir in phenotypic maturation and macrophage-like cytokine profile in late gestation. J. Reprod. Immunol. 118, 1–8. https://doi.org/10.1016/j.jri.2016.08.003 (2016).
Contreras, G. A., Kabara, E., Brester, J., Neuder, L. & Kiupel, M. Macrophage infiltration in the omental and subcutaneous adipose tissues of dairy cows with displaced abomasum. J. Dairy Sci. 98, 6176–6187. https://doi.org/10.3168/jds.2015-9370 (2015).
Ballou, M. A. Growth and development symposium: Inflammation: Role in the etiology and pathophysiology of clinical mastitis in dairy cows. J. Anim. Sci. 90, 1466–1478. https://doi.org/10.2527/jas.2011-4663 (2012).
Weichhart, T., Hengstschlager, M. & Linke, M. Regulation of innate immune cell function by mTOR. Nat. Rev. Immunol. 15, 599–614. https://doi.org/10.1038/nri3901 (2015).
Braun, C. & Weichhart, T.A.-O. mTOR-dependent immunometabolism as Achilles’ heel of anticancer therapy. Eur. J. Immunol. 51, 3161–3175. https://doi.org/10.1002/eji.202149270 (2021).
Laplante, M. & Sabatini, D. M. mTOR signaling at a glance. J. Cell Sci. 122, 3589. https://doi.org/10.1242/jcs.051011 (2009).
Mann, S. et al. Insulin signaling and skeletal muscle atrophy and autophagy in transition dairy cows either overfed energy or fed a controlled energy diet prepartum. J. Comparat. Physiol. B Biochem. Syst. Environ. Physiol. 186, 513–525. https://doi.org/10.1007/s00360-016-0969-1 (2016).
Mann, S. et al. Insulin signaling, inflammation, and lipolysis in subcutaneous adipose tissue of transition dairy cows either overfed energy during the prepartum period or fed a controlled-energy diet. J. Dairy Sci. 99, 6737–6752. https://doi.org/10.3168/jds.2016-10969 (2016).
Mann, S. et al. Nutrient-sensing kinase signaling in bovine immune cells is altered during the postpartum nutrient deficit: A possible role in transition cow inflammatory response. J. Dairy Sci. 101, 9360–9370. https://doi.org/10.3168/jds.2018-14549 (2018).
Sipka, A. S., Chandler, T. L., Behling-Kelly, E. L., Overton, T. R. & Mann, S. The effect of ex vivo lipopolysaccharide stimulation and nutrient availability on transition cow innate immune cell AKT/mTOR pathway responsiveness. J. Dairy Sci. 103, 1956–1968. https://doi.org/10.3168/jds.2019-17307 (2020).
Weichhart, T. et al. The TSC-mTOR signaling pathway regulates the innate inflammatory response. Immunity 29, 565–577. https://doi.org/10.1016/j.immuni.2008.08.012 (2008).
Mercalli, A. et al. Rapamycin unbalances the polarization of human macrophages to M1. Immunology 140, 179–190. https://doi.org/10.1111/imm.12126 (2013).
Vergadi, E., Ieronymaki, E., Lyroni, K., Vaporidi, K. & Tsatsanis, C. Akt signaling pathway in macrophage activation and M1/M2 polarization. J. Immunol. 198, 1006–1014. https://doi.org/10.4049/jimmunol.1601515 (2017).
Percie-du-Sert, N. et al. The ARRIVE guidelines 2.0: Updated guidelines for reporting animal research. PLOS Biol. 18, 00410. https://doi.org/10.1371/journal.pbio.3000410 (2020).
Sipka, A., Pomeroy, B., Klaessig, S. & Schukken, Y. Bovine natural killer cells are present in Escherichia coli infected mammary gland tissue and show antimicrobial activity in vitro. Comp. Immunol. Microbiol. Infect. Dis. 48, 54–60. https://doi.org/10.1016/j.cimid.2016.08.001 (2016).
Haidinger, M. et al. A versatile role of mammalian target of rapamycin in human dendritic cell function and differentiation. J. Immunol. 185, 3919–3931. https://doi.org/10.4049/jimmunol.1000296 (2010).
Pomeroy, B., Sipka, A., Klaessig, S. & Schukken, Y. Monocyte-derived dendritic cells from late gestation cows have an impaired ability to mature in response to E. coli stimulation in a receptor and cytokine-mediated fashion. Vet. Immunol. Immunopathol. 167, 22–29. https://doi.org/10.1016/j.vetimm.2015.06.016 (2015).
Ohtani, M. et al. Mammalian target of rapamycin and glycogen synthase kinase 3 differentially regulate lipopolysaccharide-induced interleukin-12 production in dendritic cells. Blood 112, 635–643. https://doi.org/10.1182/blood-2008-02-137430 (2008).
Turnquist, H. R. et al. Rapamycin-conditioned dendritic cells are poor stimulators of allogeneic CD4+ T cells, but enrich for antigen-specific Foxp3+ T regulatory cells and promote organ transplant tolerance. J. Immunol. 178, 7018–7031 (2007).
Fan, Y. et al. Miniaturized high-throughput multiparameter flow cytometry assays measuring in vitro human dendritic cell maturation and T-cell activation in mixed lymphocyte reactions. SLAS Discov. 23, 742–750. https://doi.org/10.1177/2472555218775409 (2018).
Sipka, A., Mann, S., Babasyan, S., Freer, H. & Wagner, B. Development of a bead-based multiplex assay to quantify bovine interleukin-10, tumor necrosis factor-α, and interferon-γ concentrations in plasma and cell culture supernatant. JDS Commun. https://doi.org/10.3168/jdsc.2021-0191 (2022).
Faul, F., Erdfelder, E., Lang, A. G. & Buchner, A. G*Power 3: A flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav. Res. Methods 39, 175–191 (2007).
Mann, S. et al. Dry period plane of energy: Effects on glucose tolerance in transition dairy cows. J. Dairy Sci. 99, 701–717 (2016).
Meijer, G. A., Van der Meulen, J., Bakker, J. G., Van der Koelen, C. J. & Van Vuuren, A. M. Free amino acids in plasma and muscle of high yielding dairy cows in early lactation. J. Dairy Sci. 78, 1131–1141. https://doi.org/10.3168/jds.S0022-0302(95)76730-3 (1995).
Beltman, M. E., McNally, J. C., Kelly, E. & Crowe, M. A. Relationship between plasma concentrations of IGF-I and clinical endometritis, and response to progesterone synchrony in dairy cows during early lactation. J. Dairy Sci. 103, 9493–9501. https://doi.org/10.3168/jds.2019-17974 (2020).
Sordillo, L. M., Contreras, G. A. & Aitken, S. L. Metabolic factors affecting the inflammatory response of periparturient dairy cows. Anim. Health Res. Rev. 10, 53–63. https://doi.org/10.1017/s1466252309990016 (2009).
Zerbe, H. et al. Altered functional and immunophenotypical properties of neutrophilic granulocytes in postpartum cows associated with fatty liver. Theriogenology 54, 771–786. https://doi.org/10.1016/S0093-691X(00)00389-7 (2000).
LeBlanc, S. Interactions of metabolism, inflammation, and reproductive tract health in the postpartum period in dairy cattle. Reprod. Domest. Anim. 47, 18–30. https://doi.org/10.1111/j.1439-0531.2012.02109.x (2012).
Linke, M., Fritsch-Stephanie, D., Sukhbaatar, N., Hengstschläger, M. & Weichhart, T. mTORC1 and mTORC2 as regulators of cell metabolism in immunity. FEBS Lett. 591, 3089–3103. https://doi.org/10.1002/1873-3468.12711 (2017).
Mann, S., Sipka, A. S. & Grenier, J. K. The degree of postpartum metabolic challenge in dairy cows is associated with peripheral blood mononuclear cell transcriptome changes of the innate immune system. Dev. Comp. Immunol. 93, 28–36. https://doi.org/10.1016/j.dci.2018.11.021 (2019).
Gordon, S. Alternative activation of macrophages. Nat. Rev. Immunol. 3, 23–35. https://doi.org/10.1038/nri978 (2003).
Gordon, S. & Taylor, P. R. Monocyte and macrophage heterogeneity. Nat. Rev. Immunol. 5, 953–964. https://doi.org/10.1038/nri1733 (2005).
Collins, S. L. et al. mTORC1 signaling regulates proinflammatory macrophage function and metabolism. J. Immunol. 207, 913. https://doi.org/10.4049/jimmunol.2100230 (2021).
Lisi, L., Laudati, E., Navarra, P. & Dello Russo, C. The mTOR kinase inhibitors polarize glioma-activated microglia to express a M1 phenotype. J Neuroinflammation 11, 125. https://doi.org/10.1186/1742-2094-11-125 (2014).
El Chartouni, C. & Rehli, M. Comprehensive analysis of TLR4-induced transcriptional responses in interleukin 4-primed mouse macrophages. Immunobiology 215, 780–787. https://doi.org/10.1016/j.imbio.2010.05.032 (2010).
Lim, J. E., Chung, E. & Son, Y. A neuropeptide, Substance-P, directly induces tissue-repairing M2 like macrophages by activating the PI3K/Akt/mTOR pathway even in the presence of IFNgamma. Sci Rep 7, 9417. https://doi.org/10.1038/s41598-017-09639-7 (2017).
Sipka, A., Weichhart, T. & Mann, S. Pharmacological inhibition of the mTOR pathway alters phenotype and cytokine expression in bovine monocyte-derived dendritic cells. Vet. Immunol. Immunopathol. 249, 110441. https://doi.org/10.1016/j.vetimm.2022.110441 (2022).
Zhang, Y. L. et al. Soybean glycinin impaired immune function and caused inflammation associated with PKC-zeta/NF-kappab and mTORC1 signaling in the intestine of juvenile grass carp (Ctenopharyngodon idella). Fish Shellfish Immunol. 106, 393–403. https://doi.org/10.1016/j.fsi.2020.08.008 (2020).
Guertin, D. A. & Sabatini, D. M. The pharmacology of mTOR inhibition. Sci. Signal. 2, pe24 (2009).
Choo, A. Y., Yoon, S.-O., Kim, S. G., Roux, P. P. & Blenis, J. Rapamycin differentially inhibits S6Ks and 4E-BP1 to mediate cell-type-specific repression of mRNA translation. Proc. Natl. Acad. Sci. 105, 17414–17419. https://doi.org/10.1073/pnas.0809136105 (2008).
Thoreen, C. C. et al. An ATP-competitive mammalian target of rapamycin inhibitor reveals rapamycin-resistant functions of mTORC1. J. Biol. Chem. 284, 8023–8032. https://doi.org/10.1074/jbc.M900301200 (2009).
Benjamin, D., Colombi, M., Moroni, C. & Hall, M. N. Rapamycin passes the torch: A new generation of mTOR inhibitors. Nat. Rev. Drug Discov. 10, 868–880. https://doi.org/10.1038/nrd3531 (2011).
Leontieva, O. V. & Blagosklonny, M. V. Gerosuppression by pan-mTOR inhibitors. Aging 8, 3535–3551. https://doi.org/10.18632/aging.101155 (2016).
Agrawal, S. et al. Cutting edge: Different Toll-like receptor agonists instruct dendritic cells to induce distinct Th responses via differential modulation of extracellular signal-regulated kinase-mitogen-activated protein kinase and c-Fos. J. Immunol. 171, 4984–4989. https://doi.org/10.4049/jimmunol.171.10.4984 (2003).
Murray, P. J. et al. Macrophage activation and polarization: Nomenclature and experimental guidelines. Immunity 41, 14–20. https://doi.org/10.1016/j.immuni.2014.06.008 (2014).
Green, M. J., Green, L. E., Medley, G. F., Schukken, Y. H. & Bradley, A. J. Influence of dry period bacterial intramammary infection on clinical mastitis in dairy cows. J. Dairy Sci. 85, 2589–2599. https://doi.org/10.3168/jds.S0022-0302(02)74343-9 (2002).
Fukao, T. et al. PI3K-mediated negative feedback regulation of IL-12 production in DCs. Nat. Immunol. 3, 875–881. https://doi.org/10.1038/ni825 (2002).
Oliveira, L. J. & Hansen, P. J. Deviations in populations of peripheral blood mononuclear cells and endometrial macrophages in the cow during pregnancy. Reproduction 136, 481–490. https://doi.org/10.1530/REP-08-0218 (2008).
Maeda, Y., Ohtsuka, H., Tomioka, M. & Oikawa, M. Effect of progesterone on Th1/Th2/Th17 and regulatory T cell-related genes in peripheral blood mononuclear cells during pregnancy in cows. Vet. Res. Commun. 37, 43–49. https://doi.org/10.1007/s11259-012-9545-7 (2013).
Acknowledgements
The authors acknowledge the expertise of Tracy Stokol in establishing differential cell counts of individual cell populations and the technical assistance of Suzanne Klaessig (both Cornell University, Ithaca, NY). We thank Bettina Wagner (Cornell University) for her expertise in developing and validating the multiplex cytokine assay used in this study.
Funding
This project was supported by Agriculture & Food Research Initiative Competitive Grant no. 2019-67015-29834 from the USDA National Institute of Food and Agriculture.
Author information
Authors and Affiliations
Contributions
T.L.C. performed experiments, analyzed data, prepared figures, drafted the manuscript. A.S.S. designed and performed experiments, analyzed data, prepared figures, drafted the manuscript. H.J.S. assisted with acquiring grant funding, designed experiments, edited the manuscript. T.W. assisted with acquiring grant funding, designed experiments, edited the manuscript, S.M. supervised the overall project, acquired grant funding, designed experiments, and edited the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sipka, A.S., Chandler, T.L., Weichhart, T. et al. Inhibition of mTOR in bovine monocyte derived macrophages and dendritic cells provides a potential mechanism for postpartum immune dysfunction in dairy cows. Sci Rep 12, 15084 (2022). https://doi.org/10.1038/s41598-022-19295-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-19295-1
- Springer Nature Limited