Abstract
Bacillus thuringiensis (Bt) isolates native to Maranhão (BtMA) that are highly toxic to Aedes aegypti larvae and seven standard subspecies of Bt were analyzed for genetic diversity using the rep-PRC technique with BOX, ERIC, REP, MB1, and GTG5 markers. The rep-PCR technique is considered an extremely reliable, reproducible, fast and highly discriminatory technique that may be used even among populations of the same species. These five markers revealed a total of 38 polymorphic DNA fragments for 30 BtMA isolates. Eight groups were obtained with the dendrogram generated through Pearson's correlation analysis, with four groups formed only with BtMA isolates and four comprised of isolates of BtMA and the standard subspecies toxic to dipterans and lepidopterans. Despite the high genetic diversity of BtMA, a low correlation between the collection site, gene content and mortality against A. aegypti larvae was evidenced. The clustering of the standard subspecies of Bt that were toxic against dipterans with BtMA isolates confirm the mosquitocidal action of the native isolates from Maranhão, and they can be used as an alternative for A. aegypti control and other insects of medical importance and for the control of agricultural pests.
Similar content being viewed by others
Introduction
Bacillus thuringiensis (Bt) is a gram-positive, aerobic, rod-shaped and spore-forming entomopathogenic bacterium that is highly effective in controlling immature forms of mosquitoes, since several strains produce protein crystals containing δ-endotoxin (Cry and Cyt) with mosquitocidal action1,2,3,4,5.
Several subspecies of Bt have been reported to be highly toxic against dipteran insects, such as Bacillus thuringiensis israelensis (Bti), Bacillus thuringiensis entomocidus (Bte), Bacillus thuringiensis sotto (Bts), Bacillus thuringiensis jegathesan (Btj), Bacillus thuringiensis darmstadiensis (Btd), Bacillus thuringiensis medellin (Btm), Bacillus thuringiensis fukuokaensis (Btf), and Bacillus thuringiensis higo (Bth)6,7,8,9,10,11,12,13. Bacillus thuringiensis israelensis is considered the most toxic to dipteran larvae1,3,14. The mosquitocidal activity of Bti is due to pesticidal proteins encoded by the cry4Aa, cry4Ba, cry11Aa, cyt1Aa, cry10Aa and cyt2Ba genes1,5,15.
Molecular typing has been used to characterize and discriminate species belonging to Bacillus, in addition to monitoring the evolution and phylogeny of the group16,17,18,19. Thus, the rep-PCR technique is widely used to investigate specific genetic relationships between Bt isolates20,21,22,23,24,25.
The rep-PCR technique uses primers specific to the Bacillus cereus group and is considered an extremely reliable, reproducible, fast and highly discriminatory technique that may be used even among populations of the same species26,27,28,29,30 or even among microorganisms20,23,27,30.
The objective of the current study was to estimate the genetic diversity of Bt isolates from the Entomopathogenic Bacteria Collection of Maranhão (CBENMA) and the Entomopathogenic Bacilli Bank of Maranhão (BBENMA) that are toxic to A. aegypti larvae31,32,33 using the repetitive sequences BOX, ERIC, REP, MB1, and GTG5, to evaluate the association between Bt isolate clusters and standard subspecies of this bacterium, geographic regions of collection, mortality against A. aegypti larvae, and amplification of dipteran-specific genes (cry and cyt genes).
Results
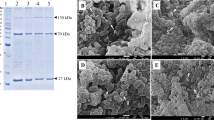
The diversity of Bacillus thuringiensis isolates native to Maranhão (BtMA) was determined by rep-PCR, and a complex fingerprint pattern was obtained for the 30 BtMA isolates with the BOX, ERIC, REP, MB1 and GTG5 primers. The results revealed a total of 38 polymorphic DNA fragments (Fig. 1).
Rep-PCR fingerprint showing the amplification of the BOX (a), ERIC (b), REP (c), MB1 (d) and GTG5 (e) molecular markers in 30 isolates of Bacillus thuringiensis and seven standard subspecies. MM: Molecular marker; Bta: B. thuringiensis aizawai; Bte: B. thuringiensis entomocidus; Btf: B. thuringiensis fukuokaensis; Bti: B. thuringiensis israelensis; Btk: B. thuringiensis kurstaki; Bts: B. thuringiensis sotto; Bty: B. thuringiensis yunnanensis; BtMA: Bacillus thuringiensis from Maranhão; NC: negative control.
The number of fragments per isolate for the BOX primer ranged from one to seven, with the smallest fragment of 100 bp and the largest exceeding 10.000 bp, with a total of 24 electrophoretic patterns (Fig. 1a). The ERIC and REP primers provided fragments ranging from one to six per isolate. For the ERIC primer, the base pairs ranged from 250 to 1250 bp, and for REP, they ranged from 250 to 2000 bp, with a total of 13 and 12 electrophoretic patterns, respectively (Fig. 1b,c). The MB1 primer provided between one to eight fragments and a total of 31 electrophoretic patterns with fragment sizes between 250 and 6000 bp (Fig. 1d). Regarding the GTG5 primer, the BtMA isolates provided fragments of repetitive sequences that ranged from 2 to 11 bands, with the majority being similar, from 200 to 3000 bp, with a total of 11 electrophoretic patterns (Fig. 1e). Although some isolates shared the same bands, most of them presented different fragment profiles.
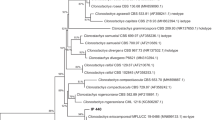
A dendrogram was generated through Pearson's correlation for genetic diversity analyses using the sequences BOX, ERIC, REP, MB1, and GTG5, with an average distance of 45% (cut-off point for the formation of groups with less genetic variation), in which eight groups (designated I, II, III, IV, V, VI, VII and VIII) were obtained, which comprised 67.5% of the isolates (Fig. 2). The other Bt isolates showed an average distance greater than the cut-off point.
Groups I, V, VI and VIII were formed only with BtMA isolates (Fig. 2). Group I (BtMA-212, BtMA-527 and BtMA-459) and group V (BtMA-687, BtMA-694 and BtMA-703) included isolates obtained from different biomes. The BtMA-212 isolate obtained from Amazonia grouped with the BtMA-527 and BtMA-459 isolates from the Cerrado. These three isolates, in addition to presenting 100% mortality to A. aegypti larvae in the second evaluation (48 h), amplified the cry and cyt genes.
Groups VI and VIII were formed by isolates from the same biome, with isolates of group VI (BtMA-679, BtMA-684, BtMA-685, BtMA-688 and BtMA-689) all from the Cerrado and those from group VIII (BtMA-229 and BtMA-237) from the Amazon. All isolates in group VI caused 100% mortality within a 24-h period and amplified all analyzed dipteran-specific genes, with the exception of the BTMA-689 isolate, which did not present the cry11Ba gene. For the genetic profile of isolates from group VIII, it was found that they did not share any of the cry and cyt genes analyzed. An A. aegypti larval mortality of 100% was caused by isolates of BtMA in this group at 48 h.
Groups II, III, IV and VII comprised isolates of BtMA and the standard subspecies. The BtMA-179 isolate that grouped with the subspecies Bacillus thuringiensis aizawai (Bta) (group II) was isolated from the soil of the Amazon biome, caused 100% mortality in 48 h and amplified only the dipteran-specific cry4Ba gene, and the BtMA-233 and BtMA-241 isolates grouped with subspecies Bacillus thuringiensis yunnanensis (Bty) (IV) were isolated from the soil of the Amazon and Cerrado biomes, respectively. These three isolates were toxic to A. aegypti larvae over a period of 48 h, and isolates BtMA-233 and BtMA-241 presented the cyt2Aa gene in common. The subspecies Bta and Bty are toxic to insects of the order Lepidoptera34,35,36, which suggests that isolates from groups II and IV may also be active against insects of that order.
The BtMA-215 isolate, obtained from Amazonian soil, did not amplify any of the analyzed genes, caused 100% mortality in A. aegypti larvae in 48 h, and grouped with two subspecies, Bacillus thuringiensis. israelensis (Bti) and Bacillus thuringiensis sotto (Bts) (group III). These subspecies possibly confirm the mosquitocidal action of the isolate BtMA-215. However, more research should be carried out with the isolate BtMA-215 to predict it as a strain with potential for the control of dipterans and lepidopterans.
The isolates BtMA-682, BtMA-690 and BtMA-691 were found in group VII, all from the Cerrado soil, with different mortality rates and genetic profiles, and grouped with the subspecies Bacillus thuringiensis entomocidus (Bte), which is at the same time effective against insects of the orders Diptera and Lepidoptera8,37. It is worth mentioning that the BtMA-690 isolate amplified all the cry and cyt genes analyzed and provided 100% mortality of A. aegypti larvae in 24 h.
Discussion
The repetitive element palindromic-polymerase chain reaction (rep-PCR) technique has been used to investigate the interspecific and intraspecific genetic diversity of Bt isolates, allowing us to correlate them with the geographic regions of collection, the origin of the substrate, genetic characteristics, and rates of mortality and toxicity against insect22,23,29,38,39.
In this research, the genetic diversity of BtMA isolates, collected in different biomes, with a high mortality against larvae of A. aegypti and that amplified for cry and cyt dipteran-specific genes31,32,33, was studied using group analysis with data obtained with the BOX, ERIC, REP, MB1 and GTG5 primers.
The BOX and MB1-PCRs were the most informative, producing the most complex fragment patterns20,25,40. Robust and, commonly, highly complex fingerprints are obtained using the BOX primer41,42. The MB1 primer generated extremely complex fingerprints, showing that almost all 30 BtMA isolates had their own unique MB1-PCR standard, suggesting high genetic variability among the isolates21,40.
Bacillus thuringiensis is a genetically diverse species, so its large number of strains can form several different profiles according to their genetic profile43. The high genetic variability of Bt may be related to both the influence of geographic and ecological factors22,23,24 because this bacterium is isolated from several substrates and has a wide range of target hosts.
Several studies have reported that ERIC markers are more informative than REP markers in discriminating Bt strains20,23,39,44, as was also verified in this research, suggesting that ERIC-like sequences may be more widely distributed than REP-like sequences in Bt44.
Among all the markers, GTG5 was not as effective in discriminating between isolates of BtMA, indicating low genetic diversity. A similar result was verified when this marker was used to determine the diversity among the different Bacillus species25,45.
It was possible to verify that groups VI, VII and VIII included isolates of BtMA from the same biome. The similarity of the BtMA isolates from the Cerrado (groups VI and VII) and the Amazon (VIII) biomes may be related to the geographic location. However, groups I, IV and V included isolates of BtMA from the Cerrado with other biomes. The tendency to form groups or subgroups in relation to the similarity of Bt according to geographic origin, using the Rep-PCR technique, is quite controversial.
Huerta et al.38, using REP primers, evaluated the genetic diversity of 53 Bt isolates collected in soil from different geographical regions of Peru and correlated the similarity of these isolates according to geographic origin. Similarly, Da Silva and Valicente20 pointed out that the similarity of Bt isolates may be related to geographic location when they used the ERIC, REP and BOX primers simultaneously. Katara et al.44 observed that the REP-PCR and ERIC-PCR patterns, generated with 113 Bt native isolates from diverse habitats of India, formed four main groups with isolates of diverse origin.
All isolates of BtMA from the VI group (BtMA-684, BtMA-685, BtMA-688, BtMA-689, and BtMA-679) presented the same mortality rate (100% in 24 h), and all isolates of BtMA from the I, II and VIII groups caused 100% mortality within 48 h. Groups V and VII included isolates with different mortality rates (24 and 48 h). However, further studies are needed to confirm the hypothesis that similarity is related with the pathogenicity.
When correlating the mortality of the BtMA isolates and the site of origin, only group VI, constituted by the isolates BtMA-684, BtMA-685, BtMA-688, BtMA-689 and BtMA-679, showed a genetic relationship. This low correlation was also verified by Huerta et al.38 when verifying subgroups of Bt formed according to the geographical origin and potential use against A. aegypti.
Likewise, Da Silva and Valicente20 and Machado et al.23, comparing the genetic relationship of Bt isolates toxic to Spodoptera spp. and the collection sites of these isolates reported a low correlation. In addition, in agreement with Reyes-Ramirez and Ibarra43 and Da Silva and Valicente20, the genomic relationship between Bt strains is not defined only by their specific toxicity but by several characteristics, such as the content of the cry gene, crystal morphology and plasmid pattern.
The BtMA isolates showed a great diversity of the diptera-specific genes cry4, cry10, cry11, cyt1 and cyt231,32,33. Groups V and VI included BtMA isolates with all the investigated genes, with the exception of the BtMA-694 isolate from group V, which did not amplify the cyt2Ba gene, and the BtMA-689 isolate from group VI, which did not amplify the cry11Ba gene. The great diversity of genes encoding mosquito-specific toxins in BtMA isolates, presenting the same genetic content as Bti, represents a great opportunity to control vector insects and enables strategies to manage the evolution of insect resistance to Bt pesticidal proteins.
One factor that may or may not explain the correlation with the Bt collection sites is that the diversity may also be associated with the region's climate, soil conditions where the samples were collected, and the influence of other microbial communities present in the region24.
Interestingly, the isolates of group V (BtMA-687 BtMA-694 BtMA-703) did not show any relation to the geographic origin and mortality of the A. aegypti larvae, and the isolates of VI (BtMA-684, BtMA-685, BtMA-688, BtMA-689 and BtMA-679) were all from the same biome and caused 100% mortality to A. aegypti larvae in the first evaluation (24 h). Based on these results, the correlation between the genetic relationship of BtMA isolates and their investigated gene content is very low. Lima et al.39 highlighted that repetitive sequences showed no relation to the types of pesticidal protein produced by Bt, which demonstrates that the majority of clusters that occur between isolates that do not carry the same combinations of dipteran-specific genes.
For the groups formed with the standard subspecies and the BtMA isolates (groups II, III, IV and VII), it was possible, in addition to confirming the toxic activity of these isolates against A. aegypti larvae, to suggest the potential of the BtMA isolates for the control of insects of the order Lepidoptera. However, it will be necessary to carry out bioassays with larvae of pest lepidopterans and to characterize these isolates with lepidopteran-specific genes of Bt.
An interesting fact was the grouping of the isolate BtMA-215 with the subspecies Bti (group III). Unlike the subspecies Bti, this isolate caused mortality to A. aegypti larvae within 48 h and did not amplify any of the diptera-specific cry and cyt genes selected for this study. However, Soares-da-Silva et al.32 reported that the BtMA-215 isolate amplified the cry32 gene. There are 29 Cry32 proteins46, and according to Van Frankenhuyzen47 and Rajchanuwong et al. 48, these proteins are toxic to insects of the order Diptera, especially mosquito larvae.
In this work, we provided evidence of high genetic variability of Bt isolates native to Maranhão (BtMA) and toxic to A. aegypti, providing important information about their phylogenetic relationships and similarity with the standard subspecies (Bti, Bta, Bte, Bty, and Bts).
In addition, the clusters with the standard subspecies of Bt toxic to dipterans with BtMA isolates confirm the mosquitocidal action of the native isolates from Maranhão and that they can be used as an alternative for control of A. aegypti and other insects of medical importance. This fact also demonstrated that isolates that grouped with subspecies toxic to lepidopterans, such as Bta and Bty, could constitute new tools in integrated pest management, providing different Cry pesticidal proteins than those expressed in ordinary Bt formulations49.
We must note that detailed information on the Bt cry gene profile is highly important not only for approaches to insect resistance management and host spectrum47,50,51 but also for providing important insights into Bt formulation because different pesticidal proteins can require specific fermentation conditions, such as dissolved oxygen and temperature52,53.
Methods
Bacillus thuringiensis isolates
Thirty Bacillus thuringiensis isolates native to Maranhão (BtMA) obtained from soil samples, with high toxicity to A. aegypti larvae and that amplified for cry (cry4Aa, cry4Ba, cry10Aa, cry11Aa and cry11Ba) and cyt (cyt1Aa, cyt1Ab, cyt2Aa and cyt2Ba) genes31,32,33, and seven standard subspecies of Bt were used in this survey (Tables 1 and 2). All these isolates are stored in CBENMA located in the Laboratory of Entomopathogenic Bacteria and Molecular Markers (BEMMOL) at the Centro de Estudos Superiores de Caxias of Universidade Estadual do Maranhão (CESC/UEMA).
DNA extraction and PCR conditions
Genomic DNA from BtMA isolates and standard subspecies of Bt was extracted using the Instagene matrix (Bio-Rad) according to the manufacturer's recommendations. PCR was performed in final volume of 20 μL. Each reaction mixture contained 50 ng of genomic DNA of Bt isolates, 1 × buffer, 2 mM MgCl2, each dNTP at a final concentration of 200 µM, 1 U of Taq DNA Polymerase and 1 μM each primer (BOX, ERIC, REP, MB1, and GTG5) (Table 3). Amplification was accomplished with the Gencycler-G96G thermocycler (Biosystems), following the standard program at 94 °C for 5 min; 36 cycles at 94 °C for 1 min; annealing (with temperatures varying for each primer) for 1 min; extension at 72 °C for 2 min; and final extension at 72 °C for 7 min. The PCR products were visualized in a 1,5% agarose gel using a photo documentation system (L-PIX EX Loccus).
Data analysis
The Rep-PCR profiles (banding patterns) obtained with the 30 BtMA isolates and the seven standard subspecies of Bt generated binary matrices that were used as input data into Bionumerics software (Applied Maths, Belgium) after Pearson’s correlation analysis. The Dice similarity coefficient was used to calculate the similarity matrix from binary data. Clustering analysis was performed using this coefficient and UPGMA (unweighted pair-group method with arithmetic mean) with 1000 bootstrapping replicates to evaluate the consistency of the group, and Bionumerics was used to produce both the similarity matrix and dendrogram (Supplementary Fig. S1).
References
Ben-Dov, E. Bacillus thuringiensis subsp. israelensis and its dipteran-specific toxins. Toxins 6, 1222–1243. https://doi.org/10.3390/toxins6041222 (2014).
De Bortoli, C. P. & Jurat-Fuentes, J. L. Mechanisms of resistance to commercially relevant entomopathogenic bacteria. Curr. Opin. Insect Sci. 33, 56–62. https://doi.org/10.1016/j.cois.2019.03.007 (2019).
Federici, B. A., Park, H.-W. & Bideshi, D. K. Overview of the basic biology of Bacillus thuringiensis with emphasis on genetic engineering of bacterial larvicides for mosquito control. Open Toxinol. J. 3, 154–171. https://doi.org/10.2174/1875414701003010083 (2010).
Jurat-Fuentes, J. L., Heckel, D. G. & Ferre, J. Mechanisms of resistance to insecticidal proteins from Bacillus thuringiensis. Ann. Rev. Entomol. 66, 121–140. https://doi.org/10.1146/annurev-ento-052620-073348 (2021).
Valtierra-de-Luis, D., Villanueva, M., Lai, L., Williams, T. & Caballero, P. Potential of Cry10Aa and Cyt2Ba, two minority δ-endotoxins produced by Bacillus thuringiensis ser. israelensis, for the control of Aedes aegypti larvae. Toxins. 12, 355. https://doi.org/10.3390/toxins12060355 (2020).
Bourgouin, C., Delécluse, A., Ribier, J., Klier, A. & Rapoport, G. A Bacillus thuringiensis subsp. israelensis gene encoding a 125-kilodalton larvicidal polypeptide is associated with inverted repeat sequences. J. Bacteriol. 170, 3575–3583. https://doi.org/10.1128/jb.170.8.3575-3583.1988 (1988).
Drobniewski, F. A. & Ellar, D. J. Purification and properties of a 28-kilodalton hemolytic and mosquitocidal protein toxin of Bacillus thuringiensis subsp. darmstadiensis 73–E10–2. J. Bacteriol. 171, 3060–3067. https://doi.org/10.1128/jb.171.6.3060-3067.1989 (1989).
Ito, T., Ikeya, T., Sahara, K., Bando, H. & Asano, S. Cloning and expression of two crystal protein genes, cry30Ba1 and cry44Aa1, obtained from a highly mosquitocidal strain, Bacillus thuringiensis subsp. entomocidus INA288. Appl. Environ. Microbiol. 72, 5673–5676. https://doi.org/10.1128/AEM.01894-05x (2008).
Lee, H.-K. & Gill, S. S. Molecular cloning and characterization of a novel mosquitocidal protein gene from Bacillus thuringiensis subsp. fukuokaensis. Appl. Environ. Microbiol. 63, 4664–4670. https://doi.org/10.1128/aem.63.12.4664-4670.1997 (1997).
Ohba, M., Saitoh, H., Miyamoto, K., Higuchi, K. & Mizuki, E. Bacillus thuringiensis serovar higo (flagellar serotype 44), a new serogroup with a larvicidal activity preferential for the anopheline mosquito. Lett. Appl. Microbiol. 21(316–3), 18. https://doi.org/10.1111/j.1472-765x.1995.tb01068.x (1995).
Ohgushi, A., Saitoh, H., Wasano, N., Uemori, A. & Ohba, M. Cloning and characterization of two novel genes, cry24B and s1orf2, from mosquitocidal strain of Bacillus thuringiensis serovar sotto. Curr. Microbiol. 51, 131–136. https://doi.org/10.1007/s00284-005-7529-3 (2005).
Ruiz, L. M., Segura, C., Trujillo, J. & Orduz, S. In vivo binding of the Cry11Bb toxin of Bacillus thuringiensis subsp. medellin to the midgut of mosquito larvae (Diptera: Culicidae). Mem. Inst. Oswaldo Cruz 99, 73–79. https://doi.org/10.1590/S0074-02762004000100013 (2004).
Wirth, M. C., Delécluse, A., Federici, B. A. & Walton, W. E. Variable cross-resistance to Cry11B from Bacillus thuringiensis subsp. jegathesan in Culex quinquefasciatus (Diptera: Culicidae) resistant to single or multiple toxins of Bacillus thuringienisis subsp. israelensis. Appl. Environ. Microbiol. 64, 4174–4179. https://doi.org/10.1128/AEM.64.11.4174-4179.1998 (1998).
Diaz-Mendoza, M., Bideshi, D. K. & Federici, B. A. A 54-kilodalton protein encoded by pBtoxis is required for parasporal body structural integrity in Bacillus thuringiensis subsp. israelensis. J. Bacteriol. 194, 1562–1571. https://doi.org/10.1128/JB.06095-11 (2011).
Berry, C. et al. Complete sequence and organization of pBtoxis, the toxin-coding plasmid of Bacillus thuringiensis subsp. israelensis. Appl. Environ. Microbiol. 68, 5082–5095. https://doi.org/10.1128/AEM.68.10.5082-5095.2002 (2002).
Ehling-Schulz, M. & Messelhäusser, U. Bacillus, “next generation” diagnostics: Moving from detection toward subtyping and risk-related strain profiling. Front. Microbiol. 4, 1–8. https://doi.org/10.3389/fmicb.2013.00032 (2013).
Kumar, A., Kumar, A. & Pratush, A. Molecular diversity and functional variability of environmental isolates of Bacillus species. Springerplus 3, 312. https://doi.org/10.1186/2193-1801-3-312 (2014).
Lee, J. et al. A RAPD-PCR method for the rapid detection of Bacillus cereus. J. Microbiol. Biotechnol. 21, 274–276. https://doi.org/10.4014/jmb.1008.08031 (2011).
Mohkam, M., Nezafat, N., Berenjian, A., Mobasher, M. A. & Ghasemi, Y. Identification of Bacillus probiotics isolated from soil rhizosphere using 16S rRNA, recA, rpoB gene sequencing and RAPD-PCR. Probiotics Antimicrob. Proteins 8, 8–18. https://doi.org/10.1007/s12602-016-9208-z (2016).
Da Silva, R. B. & Valicente, F. H. Molecular characterization of Bacillus thuringiensis using rep-PCR. Springerplus 2, 641. https://doi.org/10.1186/2193-1801-2-641 (2013).
Galvis, F. & Moreno, L. Y. Caracterización molecular mediante rep-PCR de aislados nativos de Bacillus thuringiensis, obtenidos de muestras de suelo. Agron. Costarric. 38, 223–229 (2014).
García, K. et al. Variability of Bacillus thuringiensis strains by ERIC-PCR and biofilm formation. Curr. Microbiol. 70, 10–18. https://doi.org/10.1007/s00284-014-0675-8 (2015).
Machado, D. H. B. et al. Molecular characterization of Bacillus thuringiensis strains to control Spodoptera eridania (Cramer) (Lepidoptera: Noctuidae) population. Rev. Bras. Entomol. 64, e201947. https://doi.org/10.1590/1806-9665-RBENT-2019-47 (2020).
Mishra, P. K. et al. Genetic diversity and functional characterization of endophytic Bacillus thuringiensis isolates from the North Western Indian Himalayas. Ann. Microbiol. 67, 143–155. https://doi.org/10.1007/s13213-016-1244-0 (2017).
Subbanna, A. R. N. S. et al. Interspecies diversity of Bacillus thuringiensis isolates native from North Western Indian Himalayas. J. Environ. Biol. 39, 306–313. https://doi.org/10.22438/jeb/39/3/MRN-574 (2018).
Abriouel, H. et al. Differentiation and characterization by molecular techniques of Bacillus cereus group isolates from poto poto and dégué, two traditional cereal-based fermented foods of Burkina Faso and Republic of Congo. J. Food Prot. 70, 1165–1173. https://doi.org/10.4315/0362-028x-70.5.1165 (2007).
Ishii, S. & Sadowsky, M. J. Applications of the rep-PCR DNA fingerprinting technique to study microbial diversity, ecology and evolution. Environ. Microbiol. 11, 733–740. https://doi.org/10.1111/j.1462-2920.2008.01856.x (2009).
Rademaker, J. L. W. et al. Comparison of AFLP and rep-PCR genomic fingerprinting with DNA–DNA homology studies: Xanthomonas as a model system. Int. J. Syst. Evol. Microbiol. 50, 665–677. https://doi.org/10.1099/00207713-50-2-665 (2000).
Sauka, D. H., Basile, J. I. & Benintende, G. Evidence of Bacillus thuringiensis intra-serovar diversity revealed by Bacillus cereus group-specific repetitive extragenic palindromic sequence-based PCR genomic fingerprinting. J. Mol. Microbiol. Biotechnol. 21, 184–190. https://doi.org/10.1159/000335532 (2011).
Versalovic, J., Schneider, M., de Bruijn, F. J. & Lupski, J. R. Genomic fingerprinting of bacteria using repetitive sequence-based polymerase chain reaction. Methods Mol. Cell Biol. 5, 25–40 (1994).
Lobo, K. S. et al. Isolation and molecular characterization of Bacillus thuringiensis found in soils of the Cerrado region of Brazil, and their toxicity to Aedes aegypti larvae. Rev. Bras. Entomol. 62, 5–12. https://doi.org/10.1016/j.rbe.2017.11.004 (2018).
Soares-da-Silva, J. et al. Molecular characterization of the gene profile of Bacillus thuringiensis Berliner isolated from brazilian ecosystems and showing pathogenic activity against mosquito larvae of medical importance. Acta Trop. 176, 197–205. https://doi.org/10.1016/j.actatropica.2017.08.006 (2017).
Vieira-Neta, M. R. A. et al. Estirpe de Bacillus thuringiensis da restinga, tóxico ao Aedes (Stegomyia) aegypti (Linnaeus) (Diptera, Culicidae). Braz. J. Biol. 81, 872–880. https://doi.org/10.1590/1519-6984.228790 (2021).
Balasubramanian, P. et al. Cloning and characterization of the crystal protein-encoding gene of Bacillus thuringiensis subsp. yunnanensis. Appl. Environ. Microbiol. 68, 408–411. https://doi.org/10.1128/AEM.68.1.408-411.2002 (2002).
Dequech, S. T. B., Fiuza, L. M., Silva, R. F. P. & Zumba, R. C. Histopatologia de lagartas de Spodoptera frugiperda (Lep., Noctuidae) infectadas por Bacillus thuringiensis aizawai e com ovos de Campoletis flavicincta (Hym., Ichneumonidae). Cienc. Rural. 37, 273–276. https://doi.org/10.1590/S0103-84782007000100045 (2007).
Lima, M. P. L., Oliveira, J. V., Gondim Junior, M. G. C., Marques, E. J. & Correia, A. A. Bioatividade de formulações de NIM (Azadirachta indica A. Juss, 1797) e de Bacillus thuringiensis subsp. aizawai em lagartas de Spodoptera frugiperda (J.E. Smith) (Lepidoptera: Noctuidae). Ciênc. Agrotec. 34, 1381–1389. https://doi.org/10.1590/S1413-70542010000600004 (2010).
Shin, B.-S. et al. Distribution of cryV-type insecticidal protein genes in Bacillus thuringiensis and cloning of cryV-Type genes from Bacillus thuringiensis subsp. kurstaki and Bacillus thuringiensis subsp. entomocidus. Appl. Environ. Microbiol. 61, 2402–2407. https://doi.org/10.1128/aem.61.6.2402-2407 (1995).
Huerta, D., Acosta, O. & Chang, M. Diversidad genético molecular de cepas de Bacillus thuringiensis con potencial tóxico contra Aedes aegypti. An. Fac. Med. 73, 205–210 (2012).
Lima, A. S. G., Guidelli, A. M., Abreu, I. L. & Lemos, M. V. F. Identification of new isolates of Bacillus thuringiensis using rep-PCR products and δ-endotoxin electron microscopy. Genet. Mol. Biol. 25, 225–229. https://doi.org/10.1590/S1415-47572002000200017 (2002).
Brumlik, M. J. et al. Genetic diversity among Bacillus anthracis, Bacillus cereus and Bacillus thuringiensis strains using repetitive element polymorphism-PCR. Pol. J. Microbiol. 53, 215–225 (2004).
Rademaker, J. L. W., Aaris, H. J. M. & Vinuesa, P. Molecular typing of environmental isolates (eds. Osborn, A. C. & Smith, C. J.) 97–134 (Taylor & Francis Group, 2005).
Rademaker, J. L. W., Louws, F. J., Versalovic, J. & de Bruijn, F. J. Characterization of the diversity of ecological important microbes by rep-PCR genomic fingerprinting. (eds. Kowalchuk, G. A., de Bruijn, F. J., Head, I. M., Akkermans, A. D. & van Elsas, J. D.) 611–644 (Kluwer Academic Publishers 2004).
Reyes-Ramirez, A. & Ibarra, J. E. Fingerprinting of Bacillus thuringiensis type strains and isolates by using Bacillus cereus group-specific repetitive extragenic palindromic sequence-based PCR analysis. Appl. Environ. Microbiol. 71, 1346–1355. https://doi.org/10.1128/AEM.71.3.1346-1355.2005 (2005).
Katara, J., Deshmukh, R., Singh, N. K. & Kaur, S. Molecular typing of native Bacillus thuringiensis isolates from diverse habitats in India using REP-PCR and ERIC-PCR analysis. J. Gen. Appl. Microbiol. 58, 83–94. https://doi.org/10.2323/jgam.58.83 (2012).
Freitas, D. B. et al. Genotypic and phenotypic diversity of Bacillus spp. isolated from steel plant waste. BMC Res. Notes 1, 92. https://doi.org/10.1186/1756-0500-1-92 (2008).
Crickmore, N. et al. Bacterial Pesticidal Protein Resource Center. Accessed 15 July 2021. https://www.bpprc.org/. (2020).
Van Frankenhuyzen, K. Insecticidal activity of Bacillus thuringiensis crystal proteins. J. Inverteb. Pathol. 101, 1–16. https://doi.org/10.1016/j.jip.2009.02.009 (2009).
Rajchanuwong, P., Chanpaisaeng, J. & Kaewsompong, S. Characterization and toxicity of Bacillus thuringiensis serovar chanpaisis (H46): A serovar from Thailand. Songklanakarin J. Sci. Technol. 41, 804–812. https://doi.org/10.14456/sjst-psu.2019.103 (2019).
Polanczyk, R. A., Van Frankenhuyzen, K & Pauli, G. The American Bacillus thuringiensis based biopesticides market (eds. Fiuza, L. M., Polanczyk, R. A. & Crickmore, N.) 173–184 (Springer International Publishing, 2017).
Liang, H. et al. Characterization of cry2-type genes of Bacillus thuringiensis strains from soil-isolated of Sichuan basin, China. Braz. J. Microbiol. 42, 140–146. https://doi.org/10.1590/S1517-83822011000100018 (2011).
Zorzetti, J. et al. Isolation, morphological and molecular characterization of Bacillus thuringiensis strains against Hypothenemus hampei Ferrari (Coleoptera: Curculionidae: Scolytinae). Rev. Bras. Entomol. 62, 198–204. https://doi.org/10.1016/j.rbe.2018.07.002 (2018).
Ghribi, D., Zouari, N., Trabelsi, H. & Jaoua, S. Improvement of Bacillus thuringiensis delta-endotoxin production by overcome of carbon catabolite repression through adequate control of aeration. Enzyme Microb. Tech. 40, 614–622. https://doi.org/10.1016/j.enzmictec.2006.05.015 (2007).
Özkan, M., Dilek, F. B., Yetis, Ü. & Özcengiz, G. Nutritional and cultural parameters influencing antidipteran delta-endotoxin production. Res. Microbiol. 154, 49–53. https://doi.org/10.1016/s0923-2508(02)00006-2 (2003).
Thorne, L. et al. Structural similarity between the Lepidoptera- and Diptera-specific insecticidal endotoxin genes of Bacillus thuringiensis subsp. “kurstaki” and “israelensis”. J. Bacteriol. 166, 801–811. https://doi.org/10.1128/jb.166.3.801-811.1986 (1986).
Bourgouin, C., Klier, A. & Rapoport, G. Characterization of the genes encoding the haemolytic toxin and the mosquitocidal delta-endotoxin of Bacillus thuringiensis israelensis. Mol. Gen. Genet. 205, 390–397. https://doi.org/10.1007/BF00338072 (1986).
Dankocsik, C., Donovan, W. P. & Jany, C. S. Activation of a cryptic crystal protein gene of Bacillus thuringiensis subspecies kurstaki by gene fusion and determination of the crystal protein insecticidal specificity. Mol. Microbiol. 4, 2087–2094. https://doi.org/10.1111/j.1365-2958.1990.tb00569.x (1990).
Moar, W. J., Masson, L., Brousseau, R. & Trumble, J. T. Toxicity to Spodoptera exigua and Trichoplusia ni of individual P1 protoxins and sporulated cultures of Bacillus thuringiensis subsp. kurstaki HD-1 and NRD-12. Appl. Environ. Microbiol. 56, 2480–2483. https://doi.org/10.1128/aem.56.8.2480-2483.1990 (1990).
Ishii, T. & Ohba, M. Diversity of Bacillus thuringiensis environmental isolates showing larvicidal activity specific for mosquitoes. J. Gen. Microbiol. 139, 2849–2854. https://doi.org/10.1099/00221287-139-11-2849 (1993).
Magda, S. & Moharam, M. E. Comparing the effect of seven isolated Bacillus thuringiensis against Tuta absoluta infesting in laboratory and field condition. Int. J. Sci. Res. 4, 458–462. https://doi.org/10.21275/v4i11.01111502 (2015).
Koeuth, T., Versalovic, J. & Lupski, J. R. Differential subsequence conservation of interspersed repetitive Streptococcus pneumoniae BOX elements in diverse bacteria. Genome Res. 5, 408–418. https://doi.org/10.1101/gr.5.4.408 (1995).
Versalovic, J., Koeuth, T. & Lupski, J. R. Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucleic Acids Res. 19, 6823–6831. https://doi.org/10.1093/nar/19.24.6823 (1991).
Acknowledgements
We wish to thank FAPEMA (Maranhão Research and Development Support Foundation) for financially supporting this research and UEMA (State University of Maranhão) for providing the studentships to the students.
Author information
Authors and Affiliations
Contributions
G.C.F.; D.K.P.C.; N.S.O.; E.C.P.S., these authors performed all stages of the experiments, from the maintenance of Bacillus thuringiensis isolates and DNA extraction to the rep-PCR technique. D.H.B.M., prepared figure 2 (dendrogram), R.A.P. and H.Á.A.S. and M.C.S. wrote the main manuscript text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
da Costa Fernandes, G., de Prado Costa, D.K., de Oliveira, N.S. et al. Genetic diversity of Brazilian Bacillus thuringiensis isolates with toxicity against Aedes aegypti (Diptera: Culicidae). Sci Rep 12, 14408 (2022). https://doi.org/10.1038/s41598-022-18559-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-18559-0
- Springer Nature Limited
This article is cited by
-
New native Bacillus thuringiensis strains induce high insecticidal action against Culex pipiens pallens larvae and adults
BMC Microbiology (2023)
-
Genotyping-driven diversity assessment of biocontrol potent Bacillus spp. strain collection as a potential method for the development of strain-specific biomarkers
Archives of Microbiology (2023)