Abstract
Long-term monocultures have severely inhibited the cultivation of Chinese peanut (Arachis hypogaea L.). In this study, the effects of continuous cropping on soil chemical properties and microbial communities were investigated in peanut fields that had been in crop rotation for 10 years and in monoculture for 10 years. The results found that long-term monoculture increased the activities of available potassium, available phosphorus, available nitrogen, soil organic matter, urease, acid phosphatase and catalase; while decreasing the activity of catalase. The diversity and abundance of soil bacteria and fungi is higher under continuous peanut cultivation. At the genus level, the relative abundance of potentially beneficial microflora genera was higher in the rhizosphere soil of rotational cropping than in continuous cropping, while the opposite was true for the relative abundance of potentially pathogenic fungal genera. Principal coordinates and cluster analysis indicated that continuous cropping altered the structure of the microbial community. The results of the functional predictions showed significant differences in the functioning of the rhizosphere microbial community between continuous and rotational cropping. In conclusion, long-term continuous cropping changed the chemical properties of the soil, altered the structure and function of the soil bacterial and fungal communities in peanut rhizosphere, which to some extent reduced the relative abundance of potentially beneficial microbial genera and increased the relative abundance of potentially pathogenic fungal genera, thus increasing the potential risk of soil-borne diseases and reducing the yield and quality of peanut. Therefore, in the actual production process, attention should be paid not only to the application of chemical fertilizers, but also to crop rotation and the application of microbial fertilizers.
Similar content being viewed by others
Introduction
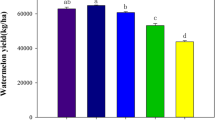
The economic crop peanut (Arachis hypogaea L.) is a major source of oil; it originated in South America and has been cultivated since 350 BCE1. China is a major producer of peanuts, with approximately 5 million hectares planted annually, accounting for approximately 60–85% of the nation’s dry farming area2,3. The production scale of peanuts has continued to expand, and peanut production areas in China are now relatively concentrated. However, long-term consecutive monoculture problems are among the key factors currently affecting the productivity and quality of peanuts in China4. Monoculture is common in agricultural planting, but it causes problems such as reduced crop yield, nutritional disorder, soil microbiological deterioration, autotoxin accumulation, and the aggravation of pests and diseases5,6,7,8,9. After investigation, it was found that the yield of the sample plots selected for this study was about 3450 kg ha−1 for continuous cropping and 3900 kg ha−1 for rotational cropping. The yield of peanut under long-term monoculture was reduced by about 11%.
Previous research into continuous cropping has mainly focused on nutritional disorders and autotoxicity caused by allelochemicals in root exudates10,11,12. However, recent research has found that the rhizosphere is rich in microorganisms, which have been referred to as the second genome of plants; these microorganisms are crucial to plant health and crop yield13. Wu et al. found that during the growth and continuous cropping of Rehmannia glutinosa, the number of bacterial, and fungal populations in rhizosphere soil changed significantly, which destroyed the balance between beneficial and harmful microorganisms14. Galazka et al. found that soil enzyme activity and the total number of bacteria and actinomycetes increased significantly in soil following continuous corn cropping15. Wu et al. found that replanting diseases regarding Pistacia chinensis were closely related to changes in the structure and potential functions of the rhizosphere bacterial community16. Although reports have suggested that plant-microbial community interactions play a vital role in crop health, however, few reports have simultaneously researched the effects of continuous cropping on the structure and potential function of the bacterial and fungal communities of peanut rhizosphere soil. In addition, multiple studies have shown that changes in residential soil microflora are related to changes in soil physical and chemical properties, soil enzyme activities, plant species, and the type of soil environment17,18,19. Therefore, herein, it was hypothesized that long-term consecutive monoculture in peanuts may directly influence the soil microbial community and its chemical properties and may further negatively affect plant growth.
Rhizosphere soils from continuous cropping and rotation peanut fields were sampled in this study. The physical and chemical characteristics of the total soil organic matter (SOM), available nitrogen (AN), available phosphorus (AP), available potassium (AK), soil bacteria structures, and fungal community structures of both types of fields were analyzed and compared. The bacterial and fungal communities and physical and chemical characteristics of soil under the two different peanut planting modes were studied. The relationship between the rhizosphere microbial community structure and the obstacles to continuous peanut cropping was then discussed, providing a theoretical basis for its control.
Results
Bacterial and fungal community composition
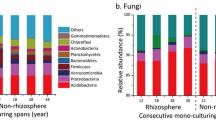
After filtering reads according to basic quality control and the removal of a single OTU, 758,780 sequences were obtained from 12 samples, including 3,611 bacterial OTUs. The lengths of high-quality sequences per sample ranged from 202 to 516 bp. The bacterial OTUs were assigned to 29 phyla, 85 classes, and 685 genera. The dominant phyla (average relative abundance > 1%) across all samples were Proteobacteria (38.13%), Actinobacteria (17.56%), Firmicutes (10.46%), Chloroflexi (8.12%), Acidobacteria (6.52%), Bacteroidetes (4.43%), Patescibacteria (2.52%), WPS-2 (2.51%), Myxococcota (2.30%), Cyanobacteria (1.93%), and Planctomycetota (1.65%; Fig. 1a).
After filtering the reads as indicated above, 868,491 fungal sequences were obtained from 12 samples, and sequences were clustered into 760 OTUs. The lengths of these high-quality sequences per sample ranged from 141 to 511 bp. The fungal OTUs were assigned to 12 different phyla, 36 classes, and 271 genera. Ascomycota accounted for 76.60% of all fungal OTUs. The other two dominant phyla were Basidiomycota (17.44%) and Mortierellomycota (3.45%) (Fig. 1b).
The relative abundances of different phyla in monoculture (LIZ) and rotation (LUZ) soils were compared (Fig. 1). As shown in Fig. 1a, LIZ resulted in a significantly higher relative abundance of Firmicutes and a significantly lower relative abundance of WPS-2 in the dominant phyla (P < 0.05), compared to LUZ (Fig. S1). Although there were differences between the other dominant bacteria phyla, these differences were not significant. The relative abundance of Ascomycetes in the dominant fungal phyla (top three) was significantly lower in LIZ, while the relative abundance of Mortierellomycota was significantly higher (P < 0.05), compared to LUZ (Figs. 1b, S2).
At the genus level, the dominant bacterial genus was Burkholderia–Caballeronia–Paraburkholderia. The bacterial genera that showed significant differences were Acidibacter, Clostridium_Sensu_Stricto_1, Turicibacter, Puia, Romboutsia, Streptomyces, Bryobacter, Paeniclostridium and Ralstonia (Figs. 2, S3). Regarding fungi, the dominant genus was Talaromyces. The genera with significant differences between monocillium and rotated rhizosphere soils were as follows: Talaromyces, Chaetomium, Mortoerella, Neocosmospora, Solicoccozyma, and Papulaspora(P < 0.05) (Figs. 2, S4).
Bacterial and fungal α-diversity
As shown in Table 1, the coverage rate was above 98% for all samples, which indicates that the depth of sequencing met the needs of the experiment. The Chao1 and ACE indices of bacterial and fungal communities were significantly higher for LIZ than for LUZ (P < 0.05). The Shannon index of bacteria showed no significant difference between the two tillage practices, whereas the Shannon index of fungi was significantly higher in LIZ. These results showed that long-term continuous cropping resulted in relatively high abundance and diversity of fungi and high abundance of bacteria without significant changes in diversity in peanut fields.
Bacterial and fungal β-diversity
In the cluster analysis of bacteria (Fig. 3a), all samples were grouped into two, among which samples from the same farming method were grouped. Tillage methods appeared to have some influence on the bacterial community structure of the soil in the rhizosphere of peanuts. Similar results were also observed for fungi (Fig. 3b).
PCoA of bacteria and fungi in the rhizosphere soils of LUZ and LIZ revealed that the first two axes explained 44.88% of the variation in the bacterial data and 44.92% of the variation in the fungal data (Fig. 4). These results indicated that there were significant differences in the composition and structure of the bacterial and fungal communities between LUZ and LIZ. PCoA and cluster analysis revealed that the microbial community structure of peanut rhizosphere soil was affected by the tillage method used.
Soil chemical properties and enzyme activity on bacterial or fungal taxa
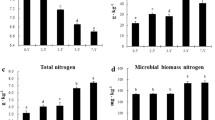
The chemical properties of rhizosphere soil under the different tillage methods are shown in Table 2. The activities of AN, AK, AP, SOM, urease, phosphatase, and sucrase in soil were significantly higher in LIZ than in LUZ, while catalase activity in rotation soil was slightly higher than that in monocropping.
Distance-based redundancy analysis (db-RDA) based on the relative abundances of bacterial and fungal genus and environmental variables. Db-RDA showed that the first and second CAP components accounted for 34.62 and 36.09% of the total bacterial and fungal variations, respectively (Fig. 5). For bacteria and fungi, the LIZ treatment was separated from the LUZ treatment by the first component (CAP1). Soil AK was the most important factor in shifting the bacterial and fungal community structures.
Prediction of microbial community function
In order to study the bacterial function of the different samples, functional prediction analysis was carried out using FAPROTAX software. The predicted results showed that the main functions of the bacterial community were chemoheterotrophy, aerobic_chemoheterotrophy, nitrogen_fixation, animal_parasites_or_symbionts, fermentation and ureolysis (Fig. 6). Nitrogen_fixation, ureolysis, chloroplasts, intracellular_parasites, aromatic_compound_degradation, aromatic_hydrocarbon_degradation and hydrocarbon_degradation functions were significantly higher in the percentage of rotational cropped fields than in the percentage of continuous cropped fields, while the opposite was true for chemoheterotrophic, fermentation and predatory_or_exoparasitic. The figure indicated that the relative abundance of functional groups associated with nitrogen and carbon cycling was mostly higher in rotational cropped fields than in continuous cropped fields.
Wilcoxon rank-sum test for functional prediction of the top 20 bacterial genera. The Y-axis indicate the function name and the X-axis indicate the percentage value of the abundance of a particular function for that sample, with different colours indicating different groupings. P values are on the far right, *0.01 < P ≤ 0.05, **0.001 < P ≤ 0.01.
FUNGuild was used to predict the trophic and functional groups of fungal communities for the different treatments (Fig. 7). FUNGuild detected all three main trophic modes and 60 guilds. There were nine trophic patterns in the rhizosphere fungi, of which the most abundant was Undefined Saprotroph (41%), followed by Fungal Parasite-Undefined Saprotroph(11.5%), Fungal Parasite (3.8%), Endophyte-Litter Saprotroph-Soil Saprotroph-Undefined Saprotroph (3.4%), Plant Pathogen (2.8%), Animal Pathogen-Endophyte-Lichen Parasite-Plant Pathogen-Soil Saprotroph-Wood Saprotroph (1.8%), Fungal Parasite-Wood Saprotroph (1.4%), Lichenized (1.4%) and Animal Pathogen-Dung Saprotroph-Endophyte-Epiphyte-Plant Saprotroph-Wood Saprotroph (1.3%). The results showed that the relative abundance of Saprotroph, Fungal Parasite, Plant Pathogen and Animal Pathogen was higher in the continuous crop sample plots than in the rotation sample plots.
Discussion
Long-term continuous monoculture can lead to a severe decrease in peanut yield, as shown in previous studies4,20. Here, PCoA and hierarchical clustering analysis showed that there were significant differences in bacterial and fungal community structures between LUZ and LIZ. There were also differences in microbial richness and diversity in the rhizosphere soil between LUZ and LIZ. The soil bacterial richness and diversity index showed an increasing trend after continuous cropping, which was consistent with the findings of previous studies into continuous cropping for Andrographis paniculata, Vanilla planifolia, and Rehmannia glutinosa, contrary to findings regarding coffee, American ginseng, and Panax notoginseng21,22,23,24,25,26. Here, the fungal richness and diversity index were found to be higher under continuous cropping than under rotation, consistent with previously reported results for sweet potato (Ipomoea batatas), Pseudostellaria heterophylla, and alfalfa (Medicago sativa) under continuous cropping conditions, contrary to results regarding Atractylodes macrocephala27,28,29,30. The above results indicated that continuous cropping can had certain effects on the structure and composition of soil microbial communities and that differences in microbial diversity and richness in different studies may be related to not only long-term monoculture but also environmental conditions, plant types, fertilization, and other factors.
Here, changes in bacterial and fungal communities in the rhizosphere soil of peanuts grown under continuous cropping were identified through taxonomic analysis. For bacteria, the relative abundances of Proteobacteria, Chloroflexi, Acidobacteria, and WPS-2 were lower for LIZ than for LUZ, while the relative abundances of Actinobacteria and Firmicutes were higher. Proteobacteria was shown to be dominating the bacterial community and soil types in different geographic regions31. This phylum is the most dynamic taxon associated with rhizoctonia disease suppression, and play a key role in phylogenetic, ecological and pathogenic values and participates in energy metabolism, such as the oxidation of organic and inorganic compounds and obtaining energy from light32,33. In this study, Proteobacteria is the most abundant phylum in the rhizosphere soil of peanut monoculture. This result was consistent with the research result that proteobacteria were the most dominant in the continuous cropping soil of pineapple34, strawberry35 and coffee24. The relative abundance of Proteobacteria increased after monoculture of potato36 and black pepper37, while decreased in sugar beet5 and Panax notoginseng38. Many factors affect the composition of rhizosphere bacteria, among which root exudates and plant species play key roles. This results in the presence of different plant genotype-specific community structures even in the same soil21. Acidobacteria are the most abundant bacterial phylum in soils with meager resource utilization39. Long-term continuous cropping can lead to the decrease of soil nutrients, which may be the cause of the increase in the relative abundance of Acidobacteria. Regarding fungi, Ascomycota and Basidiomycota were the dominant phyla of soil fungi in the rhizosphere soil after peanut continuous cropping, which was consistent with previous studies27,40. After continuous cropping, the relative abundance of Ascomycota decreased, while the relative abundance of Basidiomycota and Mortierellomycota increased. Ascomycetes play an important role in the degradation of organic matter in rhizosphere soil, and the decrease of this fungi phylum content may affect soil fertility41.
At the genus level, the relative abundances of beneficial bacterial genera, such as Burkholderia, Bradyrhizobium, Rhodanobacter, Dyella, and Bacillus, were lower for LIZ than for LUZ. Some Burkholderia bacteria promote plant growth and antagonize nematodes42. Peanuts mainly form a symbiotic relationship with different Bradyrhizobium species, which fix atmospheric nitrogen into binding nitrogen (ammonia), which can then be used by host plants42,43,44. Dyella species have been reported to have a variety of plant growth-promoting properties, while Bacillus and Rhodanobacter have biological control activities45,46,47,48. Here, LIZ rhizosphere soil also exhibited significant differences in its fungal community at the genus level, compared with that of LUZ. The relative abundance of Talaromyces was significantly higher in LIZ than LUZ. Talaromyces species have antagonistic fungal functions on species such as Cylindrocarpon destructans, Fusarium oxysporum, Rhizoctonia solani, Sclerotinia nivalis, Botrytis cinerea, and Phytophthora capsica49. The relative abundance of pathogens such as Fusarium, Penicillium, Gibberella and Colletotrichum in LUZ rhizosphere soils was lower than in LIZ, and Athelia were not detected in the LUZ rhizosphere soil, while the relative abundance of Athelia in the LIZ field was 0.0916% (Table S1). Penicillium is a toxin-producing genus that can cause fruit, vegetable, and meat rots, as well as citrus penicillium50. Fusarium, which can cause plant rots, stem rot, flower rot and spike rot51. Gibberella is often used as the sexual stage of Fusarium fungi, many of which can cause devastating plant diseases and produce specific toxins or active metabolites that are toxic to humans and animals52. The prediction of fungal function revealed that the relative abundance of pathotroph fungi in the rhizosphere soil was higher in LIZ than in LUZ. The long-term continuous cropping of peanuts resulted in fewer beneficial fungi and more pathogenic fungi, which might be one of the important factors for peanut production, which is consistent with the results of existing studies53.
Table 2 shows that the contents of SOM, AN, AP, and AK were all higher in LIZ than in LUZ, which may have arisen from the overuse of inorganic fertilizers36,37 or the insufficient utilization of nutrients in the soil by peanuts after a long period of monoculture54. The study on cucumber continuous cropping showed that the soil nutrient contents of OM, AN, AK and AP were the highest in the seventh crop, while the biomass of cucumber was the lowest in the ninth crop. However, the soil nutrient content decreased and the biomass of cucumber increased55. Shao et al. also found that as the continuous cropping of peanuts progressed, the contents of AN, AP, AK, and organic carbon increased significantly56. Differences in nutrient content may occur due to significant differences in nutrient absorption and utilization during the continuous cropping of peanuts24.
The activities of urease, phosphatase, and invertase were significantly higher in LIZ than in LUZ. The plant type, continuous planting time, and fertilization regime can have different effects on the activities of various enzymes57,58,59,60. The volume of plant residue deep in the root system can also affect enzyme activity61. Galazka et al. suggested that enzyme activity is dependent not only on the type of plant but also on the volume of plant residue deep in the root system15. Therefore, the accumulation of plant residue during long-term peanut monoculture may have increased enzyme activity. Agomoh et al. found that increasing crop diversity through wheat rotation increased yields but reduced soil health62. These results indicate that the effects of the observed changes in enzyme activity might have been less influential than the effects of the change in microbial community structure regarding obstacles to continuous cropping.
The PCoA results showed that the bacterial and fungal communities in the soil in LIZ and LUZ were different. This phenomenon has also been observed in plants such as Andrographis paniculata, Pineapple (Ananas comosus), and soybean (Glycine max)21,34,63. Combined with the RDA results, the significant changes observed in the bacterial and fungal community structures might have been caused by changes in the soil's chemical properties. Investigating the correlation between microbial community diversity and soil environmental factors helps us to better understand the mechanisms of succession barriers. In our study, RDA results suggest that many soil properties may influence microbial community structure. We found that the structure of bacterial and fungal communities was mainly driven by AK and AP. Previous studies have shown that environmental variables affects the structure of microbial communities. For example, the Basidiomycota in the rhizosphere soil of coffee with 57 years of continuous crop was closely related to the content of AP24. In a research on long-term continuous cropping in strawberries, environmental variables were found to be positively correlated with non-continuously cropped strawberry rhizosphere soil and negatively correlated with continuously cropped strawberry rhizosphere soil35. Notably, changes in microbial community structure during long-term peanut continuous cropping may also be caused by the long-term effects of peanut residues or root exudates24.
Soil microbial biomass is an important indicator of soil quality, which reflects the processes of nutrient transfer and energy cycling64. The nitrogen cycle consists of complex interaction pathways including assimilative nitrate reduction, isochemical nitrate reduction, denitrification, nitrogen fixation, nitrification and anaerobic ammonia oxidation. We hypothesized, based on the predicted results of microbial functions, that reduced nitrogen fixation (significant difference) due to continuous cropping leads to less nitrogen being taken up by plants from the atmosphere and that reduced nitrate reduction (no significant difference) leads to more ammonia being produced intracellularly, and that the accumulated nitrite and ammonia may then become toxins, leading to a reduction in biomass (Fig. 6). Another cause of succession disorders may be related to the increase in plant pathogenic fungi (Fig. 7).
Conclusion
These results provide new insights into changes in peanut rhizosphere microbial community structure based on high-throughput analysis of rhizosphere soils in continuous cropping and crop rotation. In summary, the main causes of growth inhibition and yield decline in peanut under long-term continuous cropping may be related to changes in soil microbial structure and potential functions, and the relative abundance of pathogenic microorganisms such as Fusarium and Athelia was higher in continuous rhizosphere soils than in rotational cropping sample plot (Table S1). Future research is needed to further elucidate the triangular relationship between soil characteristics, potential beneficial microbiota and peanut growth. This research provides a theoretical basis for developing sustainable agricultural practices, improving soil microbial activity and promoting peanut production in China.
Materials and methods
Ethics
The samples in the study were collected on private land where the owner allowed the study. The experimental materials did not involve any humans or animals. All methods comply with local and national guidelines.
Sampling site
Soil samples were collected from Yishui County, Linyi City, Shandong Province, China (35°40′ 05′ N, 118°42 ′45′ E). The altitude of the study area is 160 m, the annual average temperature is 12.3 °C, and the annual rainfall is 750–850 mm. Under continuous cropping conditions, peanuts were planted continuously for at least 10 years (LIZ). For the rotation fields, vegetables, maize, wheat and peanuts had been rotated for more than 10 years (LUZ), crop rotation sequence of rape (or turnip), peanut, winter wheat, with summer maize in alternate years. The amount of fertilizer applied was 600 kg ha−1 of compound fertilizer (N:P2O5:K2O = 16:13:16), 120 kg ha−1 of micronutrient fertilizer and 600 kg ha−1 of organic fertilizer. Six sample sites were randomly selected from each sample site as replicates. First, litter was removed from the ground, following which the peanut roots were carefully removed. These roots were shaken to remove loosely attached soil. Any soil that was still firmly attached to the roots was then collected to sample the rhizosphere soil. The samples were collected in September 2020. Twelve soil samples were placed in sterile plastic bags and shipped to the laboratory in ice boxes. A portion of each sample was stored at − 80 °C for deoxyribose nucleic acid (DNA) extraction, while the remainder were air-dried for analysis of soil properties.
Soil DNA extraction and polymerase chain reaction amplification
Soil total DNA was extracted from 12 soil samples using a TIANamp Soil DNA Kit, according to the manufacturer’s instructions. Genomic DNA concentration and purity were measured using a NanoDropND-2000 spectrophotometer. DNA integrity was determined via agarose gel electrophoresis. Specific primers of the bacterial 16S ribosomal ribonucleic acid (rRNA) genes and partial internal transcribed spacer regions of fungi were used for amplification. The primer set 338 F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806 R (5′-GGACTACHVGGGTWTCTAAT-3′) was used to amplify the V3-V4 hypervariable region of the bacterial 16S rRNA gene. ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and ITS2 (5′-GCTGCGTTCTTCATCGATGC-3′) were selected to target the fungal ITS1 region. Polymerase chain reactions (PCR) were performed in 20 ul reactions using the following temperature program: 95 °C for 3 min, 27 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 45 s. A final step of 72 °C for 10 min is followed by a holding step at 4 °C. The DNA was then sequenced using the Illumina Miseq PE300 platform (Illumina, San Diego, USA) at Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China).
Bioinformatics and data analysis
After removing the adaptors and primer sequences, raw sequences were assembled for each sample according to the unique barcode using quantitative insights into microbial ecology (QIIME)65. Flash v. 1.2.7 was used to merge the split sequences of each sample66. The valid labels of all samples were clustered using Uparse software67. Valuable sequences were divided into operational taxonomic units (OTUs) using USEARCH software, with a 97% identity threshold as the cut-off point. For species annotation, representative bacterial sequences were matched against the SILVA database (version 132)68, and the fungal representative sequences were classified using the UNITE database (version 8.0)69. The Mothur software was used to analyze the alpha diversity. The R language was used for distance-based redundancy analysis (db-RDA), principal coordinate analysis (PCoA), and unweighted paired group algorithm clustering. The unweighted-unifrac index was used as the distance measure. The Functional Annotation of Prokaryotic Taxa database (FAPROTAX) was used to predict bacterial functional groups. FUNGuild (http://www.funguild.org/) database was used to annotate fungal functions.
Sequence accession numbers
Sequences data were deposited in the NCBI Sequence Read Archive (SRA) database under the BioProject: PRJNA746121 (the accession number SRR15112779-SRR15112790 for bacteria and SRR15115453–SRR15115464 for fungi).
Analysis of soil chemical properties
The chemical properties were determined according to the method described by Bao70. Briefly, AK was determined photometrically using a flame spectrophotometer. AP was detected through colorimetry using a spectrophotometer. SOM content was assayed using acidified potassium dichromate (K2Cr2O7–H2SO4). AN was determined using the alkaline hydrolysis diffusion method.
Analysis of soil enzymes activity
Soil urease activity was determined using the phenyl disodium phosphate colorimetric method71. Catalase activity was determined using K2Cr2O7/acetic acid reagent colorimetry72. The 3,5-dinitrosalicylic acid colorimetric method was used to measure soil invertase activity73. Phosphatase activity was determined using p-nitrophenol colorimetry74.
Statistical analysis
All experimental data were analyzed using Microsoft Excel 2019 and IBM SPSS 25.0. Statistical significance was set at P < 0.05.
References
Jaiswal, S. K., Msimbira, L. A. & Dakora, F. D. Phylogenetically diverse group of native bacterial symbionts isolated from root nodules of groundnut (Arachis hypogaea L.) in South Africa. Syst. Appl. Microbiol. 40, 215–226. https://doi.org/10.1016/j.syapm.2017.02.002 (2017).
Tahir, M., Lv, Y., Gao, L., Hallett, P. D. & Peng, X. Soil water dynamics and availability for citrus and peanut along a hillslope at the Sunjia Red Soil Critical Zone Observatory (CZO). Soil Tillage Res. 163, 110–118. https://doi.org/10.1016/j.still.2016.05.017 (2016).
Xiaogang, L. The composition of root exudates from two different resistant peanut cultivars and their effects on the growth of soil-borne pathogen. Int. J. Biol. Sci. https://doi.org/10.7150/ijbs.5579 (2013).
Chen, M. et al. Dynamic succession of soil bacterial community during continuous cropping of peanut (Arachis hypogaea L.). PLoS ONE https://doi.org/10.1371/journal.pone.0101355 (2014).
Huang, W. et al. Effects of continuous sugar beet cropping on rhizospheric microbial communities. Genes https://doi.org/10.3390/genes11010013 (2019).
Wang, Y. et al. Effect of continuous cropping on the rhizosphere soil and growth of common buckwheat. Plant. Prod. Sci. 23, 81–90. https://doi.org/10.1080/1343943X.2019.1685895 (2020).
Meng, L. B. et al. Changes in soil microbial diversity and control of Fusarium oxysporum in continuous cropping cucumber greenhouses following biofumigation. Emir. J. Food Agric. 30, 644–653. https://doi.org/10.9755/ejfa.2018.v30.i8.1752 (2018).
Li, X., Ding, C., Zhang, T. & Wang, X. Fungal pathogen accumulation at the expense of plant-beneficial fungi as a consequence of consecutive peanut monoculturing. Soil Biol. Biochem. 72, 11–18. https://doi.org/10.1016/j.soilbio.2014.01.019 (2014).
Wang, H. W. et al. Fungal endophyte Phomopsis liquidambari biodegrades soil resveratrol: A potential allelochemical in peanut monocropping systems. J. Sci. Food Agric. 99, 5899–5909. https://doi.org/10.1002/jsfa.9865 (2019).
Huang, L. et al. Plant-soil feedbacks and soil sickness: From mechanisms to application in agriculture. J. Chem. Ecol. 39, 232–242. https://doi.org/10.1007/s10886-013-0244-9 (2013).
Deng, J. J. et al. Autotoxicity of phthalate esters in tobacco root exudates: Effects on seed germination and seedling growth. Pedosphere 27, 1073–1082. https://doi.org/10.1016/s1002-0160(17)60374-6 (2017).
Chen, S. L., Zhou, B. L., Lin, S. S., Li, X. & Ye, X. L. Accumulation of cinnamic acid and vanillin in eggplant root exudates and the relationship with continuous cropping obstacle. Afr. J. Biotechnol. 10, 2659–2665. https://doi.org/10.5897/AJB10.1338 (2011).
Berendsen, R. L., Pieterse, C. M. J. & Bakker, P. A. H. M. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486. https://doi.org/10.1016/j.tplants.2012.04.001 (2012).
Wu, L. K. et al. Comparative metagenomic analysis of rhizosphere microbial community composition and functional potentials under Rehmannia glutinosa consecutive monoculture. Int. J. Mol. Sci. https://doi.org/10.3390/ijms19082394 (2018).
Galazka, A., Gawryjolek, K., Perzynski, A., Galazka, R. & Ksiezak, J. Changes in enzymatic activities and microbial communities in soil under long-term maize monoculture and crop rotation. Pol. J. Environ. Stud. 26, 39–46. https://doi.org/10.15244/pjoes/64745 (2017).
Wu, L. K. et al. Modification of rhizosphere bacterial community structure and functional potentials to control Pseudostellaria heterophylla replant disease. Plant Dis. 104, 25–34. https://doi.org/10.1094/pdis-04-19-0833-re (2020).
Becker, J., Rodibaugh, K., Hahn, D. & Nowlin, W. Bacterial community composition and carbon metabolism in a subtropical riverscape. Hydrobiologia 792, 209–226. https://doi.org/10.1007/s10750-016-3058-2 (2017).
Zheng, Q. et al. Soil multifunctionality is affected by the soil environment and by microbial community composition and diversity. Soil Biol. Biochem. 136, 107521. https://doi.org/10.1016/j.soilbio.2019.107521 (2019).
Berg, G. & Smalla, K. Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol. Ecol. 68, 1–13. https://doi.org/10.1111/j.1574-6941.2009.00654.x (2009).
Yang, D., Liu, Y., Wang, Y., Gao, F. & Li, X. Effects of soil tillage, management practices, and mulching film application on soil health and peanut yield in a continuous cropping system. Front. Microbiol. 11, 570924. https://doi.org/10.3389/fmicb.2020.570924 (2020).
Li, J. et al. Variations of rhizospheric soil microbial communities in response to continuous Andrographis paniculata cropping practices. Bot. Stud. https://doi.org/10.1186/s40529-020-00295-1 (2020).
Xiong, W. et al. Distinct roles for soil fungal and bacterial communities associated with the suppression of vanilla Fusarium wilt disease. Soil Biol. Biochem. 107, 198–207. https://doi.org/10.1016/j.soilbio.2017.01.010 (2017).
Wu, L. et al. Barcoded pyrosequencing reveals a shift in the bacterial community in the rhizosphere and rhizoplane of Rehmannia glutinosa under consecutive monoculture. Int. J. Mol. Sci. 19, 850. https://doi.org/10.3390/ijms19030850 (2018).
Zhao, Q. et al. Long-term coffee monoculture alters soil chemical properties and microbial communities. Sci. Rep. https://doi.org/10.1038/s41598-018-24537-2 (2018).
Dong, L. et al. High-throughput sequencing technology reveals that continuous cropping of American ginseng results in changes in the microbial community in arable soil. Chin. Med. https://doi.org/10.1186/s13020-017-0139-8 (2017).
Dong, L., Xu, J., Feng, G., Li, X. & Chen, S. Soil bacterial and fungal community dynamics in relation to Panax notoginseng death rate in a continuous cropping system. Sci. Rep. https://doi.org/10.1038/srep31802 (2016).
Gao, Z. et al. Effects of continuous cropping of sweet potato on the fungal community structure in rhizospheric soil. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.02269 (2019).
Wu, L. et al. Effects of consecutive monoculture of Pseudostellaria heterophylla on soil fungal community as determined by pyrosequencing. Sci. Rep. 6, 26601. https://doi.org/10.1038/srep26601 (2016).
Yao, Q. et al. Dynamics of soil properties and fungal community structure in continuous-cropped alfalfa fields in Northeast China. PeerJ 7, 7125. https://doi.org/10.7717/peerj.7127 (2019).
Zhu, B., Wu, J., Ji, Q., Wu, W. & Qin, L. Diversity of rhizosphere and endophytic fungi in Atractylodes macrocephala during continuous cropping. PeerJ 8, e8905. https://doi.org/10.7717/peerj.8905 (2020).
Janssen, P. H. Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl. Environ. Microb. 72, 1719–1728 (2006).
Mendes, R. et al. Deciphering the rhizosphere microbiome for disease-sppressive bacteria. Science https://doi.org/10.1126/science.1203980 (2011).
Zhou, H. et al. Changes in the soil microbial communities of alpine steppe at Qinghai-Tibetan Plateau under different degradation levels. Sci. Total Environ. 651, 2281–2291. https://doi.org/10.1016/j.scitotenv.2018.09.336 (2019).
Chen, J., Gong, J. L. & Xu, M. G. Implications of continuous and rotational cropping practices on soil bacterial communities in pineapple cultivation. Eur. J. Soil Biol. 97, 103172. https://doi.org/10.1016/j.ejsobi.2020.103172 (2020).
Li, W., Liu, Q. & Chen, P. Effect of long-term continuous cropping of strawberry on soil bacterial community structure and diversity. J. Integr. Agr. 17, 206–218. https://doi.org/10.1016/S2095-3119(18)61944-6 (2018).
Liu, X. et al. Microbial community diversities and taxa abundances in soils along a seven-year gradient of potato monoculture using high throughput pyrosequencing approach. PLoS ONE 9, e86610–e86610. https://doi.org/10.1371/journal.pone.0086610 (2014).
Xiong, W. et al. The effect of long-term continuous cropping of black pepper on soil bacterial communities as determined by 454 pyrosequencing. PLoS ONE 10, e0136946. https://doi.org/10.1371/journal.pone.0136946 (2015).
Tan, Y. et al. Diversity and composition of rhizospheric soil and root endogenous bacteria in Panax notoginseng during continuous cropping practices. J. Basic Microb. 57, 337. https://doi.org/10.1002/jobm.201600464 (2017).
Fierer, N., Bradford, M. A. & Jackson, R. B. Toward an ecological classification of soil bacteria. Ecology 88, 1354–1364. https://doi.org/10.1890/05-1839 (2007).
Yang, Y. et al. Effects of microbiological fertilizer on rhizosphere soil fungus communities under long-term continuous cropping of protected Hami melon. Chin. J. App. Environ. Biol. https://doi.org/10.19675/j.cnki.1006-687x.2017.03014 (2018).
Schoch, C. L. et al. The Ascomycota tree of life: A phylum-wide phylogeny clarifies the origin and evolution of fundamental reproductive and ecological traits. Syst. Biol. 58, 224–239. https://doi.org/10.1093/sysbio/syp020 (2009).
Hayat, R., Ali, S., Amara, U., Khalid, R. & Ahmed, I. Soil beneficial bacteria and their role in plant growth promotion: A review. Ann. Microbiol. 60, 579–598. https://doi.org/10.1007/s13213-010-0117-1 (2010).
Jann Lasse, G., Hurek, T., Wiebke, B. & Reinhold-Hurek, B. Bradyrhizobium vignae sp. nov., a nitrogen-fixing symbiont isolated from effective nodules of Vigna and Arachis. Int. J. Syst. Evol. Microbiol. 66, 62. https://doi.org/10.1099/ijsem.0.000674 (2015).
Ormeo-Orrillo, E. & Esperanza, M.-R. A genomotaxonomy view of the bradyrhizobium genus. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.01334 (2019).
Palaniappan, P., Chauhan, P. S., Saravanan, V. S., Anandham, R. & Sa, T. Isolation and characterization of plant growth promoting endophytic bacterial isolates from root nodule of Lespedeza sp. Biol. Fertil. Soils 46, 807–816. https://doi.org/10.1007/s00374-010-0485-5 (2010).
Wang, H. et al. Impact of soybean nodulation phenotypes and nitrogen fertilizer levels on the rhizosphere bacterial community. Front. Microbiol. https://doi.org/10.3389/fmicb.2020.00750 (2020).
Wang, M. X. et al. Streptomyces lydicusM01 regulates soil microbial community and alleviates foliar disease caused by Alternaria alternataon cucumbers. Front. Microbiol. https://doi.org/10.3389/fmicb.2020.00942 (2020).
Li, Y. S. et al. Biological fertilizer containing Bacillus subtilis BY-2 for control of Sclerotinia sclerotiorum on oilseed rape. Crop Prot. https://doi.org/10.1016/j.cropro.2020.105340 (2020).
Kim, M. J. et al. Enhancement of seed dehiscence by seed treatment with talaromyces flavus GG01 and GG04 in ginseng (Panax ginseng). Plant Pathol. J. 33, 1–8. https://doi.org/10.5423/ppj.Oa.06.2016.0146 (2017).
Chen, W. et al. Occurrence and characterization of fungi and mycotoxins in contaminated medicinal herbs. Toxins 12, 30. https://doi.org/10.3390/toxins12010030 (2020).
Naeem, M. et al. Characterization and pathogenicity of fusarium species associated with soybean pods in maize/soybean strip intercropping. Pathogens 8, 117. https://doi.org/10.3390/pathogens8040245 (2019).
Desjardins, A. Gibberella from A (Venaceae) to Z (eae). Ann. Rev. Phytopathol. 41, 177–198. https://doi.org/10.1146/annurev.phyto.41.011703.115501 (2003).
Mingna, C. et al. Soil eukaryotic microorganism succession as affected by continuous cropping of peanut: Pathogenic and beneficial fungi were selected. PLoS ONE 7, e40659. https://doi.org/10.1371/journal.pone.0040659 (2012).
Arafat, Y. et al. Long-term monoculture negatively regulates fungal community composition and abundance of tea orchards. Agronomy https://doi.org/10.3390/agronomy9080466 (2019).
Zhou, X. G. & Wu, F. Z. Changes in soil chemical characters and enzyme activities during continuous monocropping of cucumber (Cucumis sativus). Pak. J. Bot. 47, 691–697 (2015).
Shao, S., Chen, M., Liu, W., Hu, X. & Li, Y. Long-term monoculture reduces the symbiotic rhizobial biodiversity of peanut. Syst. Appl. Microbiol. 43, 126101. https://doi.org/10.1016/j.syapm.2020.126101 (2020).
Zhang, Y., Zheng, Y. J., Xia, P. G., Xun, L. L. & Liang, Z. S. Impact of continuous Panax notoginseng plantation on soil microbial and biochemical properties. Sci. Rep. https://doi.org/10.1038/s41598-019-49625-9 (2019).
Zhang, L. C. et al. Comparison of soil enzyme activity and microbial community structure between rapeseed-rice and rice-rice plantings. Int. J. Agric. Biol. 20, 1801–1808. https://doi.org/10.17957/ijab/15.0692 (2018).
Hansen, J. C., Schillinger, W. F., Sullivan, T. S. & Paulitz, T. C. Soil microbial biomass and fungi reduced with canola introduced into long-term monoculture wheat rotations. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.01488 (2019).
Guo, Z. B. et al. Fertilization regime has a greater effect on soil microbial community structure than crop rotation and growth stage in an agroecosystem. Appl. Soil. Ecol. https://doi.org/10.1016/j.apsoil.2020.103510 (2020).
Zhao, H. L. et al. Effect of different straw return modes on soil bacterial community, enzyme activities and organic carbon fractions. Soil Sci. Soc. Am. J. 83, 638–648. https://doi.org/10.2136/sssaj2018.03.0101 (2019).
Agomoh, I. V., Drury, C. F., Phillips, L. A., Reynolds, W. D. & Yang, X. Increasing crop diversity in wheat rotations increases yields but decreases soil health. Soil Sci. Soc. Am. J. https://doi.org/10.1002/saj2.20000 (2020).
Liu, Z. X. et al. Long-term continuous cropping of soybean is comparable to crop rotation in mediating microbial abundance, diversity and community composition. Soil Tillage Res. https://doi.org/10.1016/j.still.2019.104503 (2020).
Powlson, D. S., Prookes, P. C. & Christensen, B. T. Measurement of soil microbial biomass provides an early indication of changes in total soil organic matter due to straw incorporation. Soil Biol. Biochem. 19, 159–164. https://doi.org/10.1016/0038-0717(87)90076-9 (1987).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. https://doi.org/10.1038/nmeth.f.303 (2010).
Magoc, T. & Salzberg, S. L. FLASH: Fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963. https://doi.org/10.1093/bioinformatics/btr507 (2011).
Edgar, R. C. UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998. https://doi.org/10.1038/nmeth.2604 (2013).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596. https://doi.org/10.1093/nar/gks1219 (2013).
Kõljalg, U. et al. Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 22, 5271–5277. https://doi.org/10.1111/mec.12481 (2013).
Bao, S. Soil and Agricultural Chemistry Analysis (Agriculture Press Publisher, 2013).
Guan, S. Y., Zhang, D. & Zhang, Z. Soil Enzyme and its Research Methods (Springer, 1986).
Sinha, A. K. Colorimetric assay of catalase. Anal. Biochem. 47, 389–394. https://doi.org/10.1016/0003-2697(72)90132-7 (1972).
Schinner, F. & Mersi, W. V. Xylanase-, CM-cellulase- and invertase activity in soil: An improved method. Soil Biol. Biochem. 22, 511–515. https://doi.org/10.1016/0038-0717(90)90187-5 (1990).
Tabatabai, M. A. & Bremner, J. M. Use of p-nitrophenyl phosphate for assay of soil phosphatase activity. Soil Biol. Biochem. 1, 301–307. https://doi.org/10.1016/0038-0717(69)90012-1 (1969).
Acknowledgements
This research was supported by the Shandong Agricultural Science and Technology Fund (Forestry, Science, and Technology Innovation) (2019LY003-5), the West Coast Science and Technology Foundation of Qingdao (2019-23), and the Major Science and Technology Innovation project of the Shandong province (2019JZZY020614).
Author information
Authors and Affiliations
Contributions
H.L., C.L., and X.L. conceived and designed the study. H.L., X.S., Y.L., Q.G., R.Z., J.L., and P.Z. performed the experiments. H.L. analyzed the data and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, H., Li, C., Song, X. et al. Impacts of continuous and rotational cropping practices on soil chemical properties and microbial communities during peanut cultivation. Sci Rep 12, 2758 (2022). https://doi.org/10.1038/s41598-022-06789-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-06789-1
- Springer Nature Limited
This article is cited by
-
Impacts of continuous cropping on the rhizospheric and endospheric microbial communities and root exudates of Astragalus mongholicus
BMC Plant Biology (2024)
-
Response of rhizosphere soil physicochemical properties and microbial community structure to continuous cultivation of tobacco
Annals of Microbiology (2024)
-
Combined metagenomics and metabolomic analysis of microbial community structure and metabolic function in continuous soybean cropping soils of Songnen Plain, China
Chemical and Biological Technologies in Agriculture (2024)
-
Distinct prokaryotic and eukaryotic communities and networks in two agricultural fields of central Japan with different histories of maize–cabbage rotation
Scientific Reports (2023)
-
Eliciting the Response of Rhizospheric Soil Microbial Community Structure to Zinc Amendment: A Case Study of Sugar Beet Cultivation in Black Soil
Sugar Tech (2023)