Abstract
Although the majority of gastrointestinal stromal tumors (GISTs) possess KIT mutations that induce constitutive signal transduction, the clinical outcomes are variable. The ETS translocation variant 1 (ETV1) gene encodes a transcription factor that is reported to cooperate with KIT in GISTs. However, the clinical role of ETV1 is largely unknown. The aim of this study was to examine ETV1 expression and its associations with clinical features in GISTs. We conducted a cohort study involving 64 patients with GISTs who underwent surgical resection between October 2008 and February 2015. ETV1 mRNA expression was compared with that in non-GISTs and was analyzed among risk classifications or clinical outcomes. The GIST samples exhibited significantly higher ETV1 mRNA expression than the non-GIST samples (P < 0.0001). Sixty-four GISTs were stratified into high or low ETV1 mRNA expression groups based on the median relative abundance of ETV1 mRNA. The multivariate analysis showed that low ETV1 expression, as well as tumor size and mitotic index, was an independent factor of recurrence (hazard ratio: 8.1). Patients with high ETV1 expression achieved significantly longer recurrence-free survival (RFS) times than those with low ETV1 expression (P = 0.025). Our study revealed that low ETV1 expression is an independent factor of recurrence after surgery in patients with GISTs, and thus, low ETV1 expression might be a marker of more aggressive malignant GISTs.
Similar content being viewed by others
Introduction
Gastrointestinal stromal tumors (GISTs) are the most common mesenchymal tumors of the gastrointestinal tract. Most of them specifically express the c-kit protein and carry mutations in the KIT or PDGFRA gene1,2,3. Gene mutations found in GISTs affect the constitutive activation of the c-kit receptor, which is closely related to tumorigenesis, and are classified as “gain-of-function” mutations1,4,5,6,7. In 2001, it was reported that the tyrosine kinase inhibitor (TKI) imatinib markedly suppressed the progression of unresectable GISTs by inhibiting the constitutive c-kit signaling pathway8,9. Therefore, KIT is considered a key molecule in the oncogenesis of GISTs.
In general, GISTs exhibit variable biological behaviors in the clinical setting. Some GISTs grow rapidly or produce liver metastases, which can lead to death10. On the other hand, some GISTs are found incidentally without any clinical symptoms, even though they have the same KIT mutations11,12. The relationships between clinical aggressiveness and gene mutations in GISTs remain unclear. Currently, gene mutations are recognized as an initiating factor of transformation to malignancy, but other factors are also required13,14. Thus, the discovery of new biomarkers that could be used to predict the malignancy and clinical course of GISTs is desired.
In 2010, it was reported that the transcription factor ETS translocation variant 1 (ETV1) is universally expressed in GISTs and contributes to tumorigenesis by cooperating with KIT15. However, other studies concluded that ETV1 is not expressed in all GISTs16,17. Moreover, there was no significant difference in the relapse-free survival rate between patients with GISTs that did and did not express ETV117. Thus, the relationships between ETV1 expression and prognosis in patients with GISTs remain unclear. Thus, we conducted this study to evaluate ETV1 mRNA expression and to clarify the relationships between ETV1 mRNA expression and risk classifications or clinical significance in GISTs.
Materials and methods
Patient cohort
From October 2008 to February 2015, we registered 64 consecutive patients who underwent surgery for GISTs at Nagoya University Hospital. GISTs were categorized as very low-, low-, intermediate-, and high-risk tumors according to the modified National Institutes of Health (NIH) consensus criteria18. In March 2020, medical records were surveyed to determine whether the patients had suffered recurrence and whether they had survived (median interval: 60 months; range 2–96 months). Protocols for the collection of tumor samples and clinical information were approved by the Institutional Review Board at Nagoya University Hospital. Written informed consent was obtained from all patients before surgery. All methods were performed in accordance with the relevant guidelines and regulations.
RNA extraction and cDNA reverse transcription
In each case, a tissue with a side of 5 mm was cored out from the resected fresh tumor in the operating room and immediately immersed in RNAlater stabilization solution (Invitrogen, Carlsbad, CA). The sample was stored at − 80 °C until just before the total RNA extraction procedure using TRIzol reagent (Invitrogen, Carlsbad, CA), chloroform, and isopropanol according to the manufacturer’s instructions. For cDNA synthesis, 300 ng of total RNA was reverse transcribed using PrimeScript Reverse Transcriptase (Takara Bio, Inc., Shiga, Japan) according to the manufacturer’s protocol.
Mutation analysis
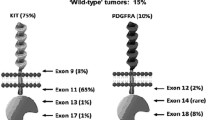
Full-length KIT cDNA was amplified with over 40 cycles of polymerase chain reaction (PCR) using appropriate primers (Table S1). The amplified full-length KIT cDNA was used as a template, and exons 8, 9, 11, 13, and 17 were analyzed using the direct sequencing method on an ABI 310 system (Applied Biosystems, Foster City, CA). In patients who were negative for KIT mutations, exons 12, 14, and 18 of the platelet-derived growth factor receptor A (PDGFRA) gene were further analyzed after PCR amplification of the full-length PDGFRA cDNA. GISTs that did not possess KIT or PDGFRA mutations were defined as wild-type GISTs. If no mutation was detected in either KIT or PDGFRA using cDNA, we also performed mutation analysis by conventional analysis using DNA6,13,19 and confirmed whether wild-type GISTs were present.
Quantitative real-time PCR
cDNA was synthesized starting from 100 ng/µl of total RNA. The expression level of each gene was analyzed with the StepOne real-time PCR system (Applied Biosystems, Foster City, CA). The following primers were used: ETV1 (Hs 00951941_m 1) and GAPDH (Hs 02758991_g 1). GAPDH expression served as an internal control. The sample that showed the lowest ETV1 mRNA expression was set as a reference. The 2-ΔΔCT method was used to calculate relative gene expression levels during direct comparisons between each of the samples. To compare ETV1 mRNA expression with GIST samples, 13 non-GIST samples that were preoperatively suspected as a GIST (one pancreatic ductal cancer, one metastatic lesion of cecal cancer, one gastric cancer, five schwannomas and five leiomyomas) were also analyzed for ETV1 mRNA expression by the same method. All experiments were performed in duplicate.
Western blot analysis
Protein lysates from frozen tumor tissues were prepared in RIPA buffer (Cell Signaling, Danvers, MA). Twenty micrograms of protein in sample buffer was resolved by 8% SDS-PAGE, blotted onto a PVDF membrane, and processed for Western blotting. As primary antibodies, mouse monoclonal ETV1 (clone 2A8, 1:1,000; Abnova, Taiwan) and rabbit monoclonal βactin (4970S, 1:1,000, Cell Signaling Technology, Danvers, MA) were utilized.
Statistical analysis
In the statistical analyses, the comparison of two groups was performed using the Mann–Whitney U test. Univariate and multivariate analyses regarding recurrence or survival after surgery were performed using the Cox proportional hazards model. The Kaplan–Meier method was then used to calculate estimates of the proportions of patients who were alive and free of recurrence. Overall survival (OS) and recurrence-free survival (RFS) rates were compared between the high and low ETV1 mRNA expression groups using the log-rank test. Patients who had not experienced either event at the last follow-up were censored. Statistical significance was defined as P < 0.05. All statistical analyses were performed using the Statistical Package for the Social Sciences (SPSS) version 24 (IBM, Armonk, NY, USA).
Ethical approval
This research involved human participants. Protocols for the collection of tumor samples and clinical information were approved by the Institutional Review Board at Nagoya University Hospital. Written informed consent was obtained from all patients before surgery.
Results
Patient characteristics
The clinical features of the 64 GIST patients are shown in Table 1. Among the 64 patients, 11 suffered from postoperative recurrence. Two of the six patients who underwent adjuvant therapy experience recurrence. We defined a GIST that recurred after surgery as an aggressive malignant GIST. All 11 patients who experienced recurrence were treated with TKIs such as imatinib, sunitinib, and regorafenib. However, five patients died of GISTs. The characteristics of these patients are described in Table S2.
Gene mutations in GISTs
The results of the mutation analysis are shown in Table 2. Fifty-eight GISTs (91%) carried KIT mutations, whereas two GISTs carried PDGFRA mutations. Four GISTs had neither KIT nor PDGFRA mutations and were confirmed by additional DNA analysis as wild-type GISTs. Regarding the mutation status of the 11 aggressive malignant GISTs, KIT exon 11 mutations were detected in 9 patients (deletions were detected in 6 patients, point mutations were detected in 2 patients, and one insertion was detected). A KIT exon 9 insertion was detected in one of these GISTs. The remaining aggressive malignant GISTs had wild-type KIT.
ETV1 mRNA expression in GISTs
GISTs exhibited significantly higher relative ETV1 mRNA expression levels (median: 44.3, range 1.0–192.3) than non-GISTs (median: 0.008, range: 0.0–0.13) (P < 0.0001) (Fig. 1a,b). The minimum ETV1 expression level in the GISTs was higher than the maximum level in the non-GISTs.
(a,b) Comparison of ETV1 mRNA expression in GISTs and non-GISTs. The data are shown on a semilogarithmic graph to base 10 (a) and on a linear graph (b). Patients with GISTs showed significantly higher relative ETV1 mRNA expression levels than those without GISTs (P < 0.0001). The comparison of two groups was performed using the Mann–Whitney U test. Error bars represent the standard deviation. (c) Comparison of ETV1 mRNA expression between high- or intermediate-risk GISTs and low- or very low-risk GISTs. ETV1 mRNA expression was higher in patients with very low- or low-risk GISTs than in those with intermediate- or high-risk GISTs (P = 0.005). The comparison of two groups was performed using the Mann–Whitney U test. Error bars represent the standard deviation. (d) Comparison of ETV1 mRNA expression between aggressive malignant GISTs and other types of GISTs. ETV1 mRNA expression was higher in patients with other types of GISTs than in those with aggressive malignant GISTs (P = 0.001). The comparison of two groups was performed using the Mann–Whitney U test. Error bars represent the standard deviation.
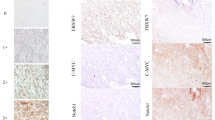
To investigate whether ETV1 is an indicator of an aggressive malignant GIST, we examined the relationships between ETV1 mRNA expression and risk classification. ETV1 mRNA was relatively highly expressed in very low- and low-risk GISTs (n = 47) (median: 46.7, range 1.0–192.3) compared with intermediate- and high-risk GISTs (n = 17) (median: 21.9, range 2.6–94.8) (P = 0.005) (Fig. 1c). In addition, the aggressive malignant GISTs (n = 11) exhibited significantly lower ETV1 mRNA levels (median: 23.2, range 2.6–51.5) than the other types of GISTs (n = 53) (median: 47.7, range 1.0–95.7) (P = 0.001) (Fig. 1d). When the protein expression of ETV1 in the three groups was examined by Western blot, ETV1 expression was detected only in GISTs. One GIST in the low ETV1 group did not display protein expression, and the others expressed less protein than those in the high ETV1 group (Supplementary Fig. S1).
Low ETV1 expression was associated with recurrence after surgery
Regarding postoperative recurrence, ETV1 mRNA expression was a significant factor in the univariate analysis (Table 3). On the other hand, the mutation type, such as the KIT exon 11 deletion, was not identified as a significant factor. The multivariate analysis showed that ETV1 expression (hazard ratio [HR] 8.1; 95% confidence interval [CI] 1.4–48.0, P = 0.022), tumor size (HR 4.8; 95% CI 1.1–21.3, P = 0.039), location (HR 11.7; 95% CI 1.5–88.0, P = 0.017) and mitotic count (HR 13.9; 95% CI 2.8–69.0, P < 0.001) were independent factors for recurrence after surgery. Regarding OS, only a mitotic count over 10/5 mm2 after surgery was selected as a significant factor.
We then examined the relationships between ETV1 mRNA expression and OS or RFS.
Sixty-four patients with GISTs were assigned to the low (n = 32) or high (n = 32) ETV1 expression level groups based on the median ETV1 mRNA expression level (cut-off: 44.3) in all GISTs. The clinical features and mutation status of each group are shown in Tables 1 and 2, respectively. As shown in Fig. 2, the 5-year OS rate was 90.9 ± 8.7% in the high ETV1 group and 86.8 ± 6.2% in the low ETV1 group. There was no significant difference between groups (P = 0.183). On the other hand, patients in the high ETV1 group displayed significantly prolonged RFS compared with patients in the low ETV1 group (P = 0.025) (Fig. 3). The 5-year RFS rate was 92.1 ± 5.4% in the high ETV1 group and 71.8 ± 7.9% in the low ETV1 group. The mean RFS time was 63 months (95% CI 52–74) in the low ETV1 group and 77 months (95% CI 72–81) in the high ETV1 group (HR 4.8; 95% CI 1.1–22.4; P = 0.043). Nine of the 32 (28%) patients in the low ETV1 group developed recurrent disease during the follow-up period, whereas only two patients (6.3%) in the high ETV1 group experienced recurrence.
Kaplan–Meier curve of recurrence-free survival stratified by high or low ETV1 expression. Patients in the high ETV1 expression group displayed significantly prolonged RFS compared with patients in the low ETV1 expression group (P = 0.025). The 5-year RFS rate was 92.1 ± 5.4% in the high ETV1 expression group and 71.8 ± 7.9% in the low ETV1 expression group.
Discussion
Although numerous studies of GISTs have been conducted to discover the genetic and molecular mechanisms responsible for the disease, no critical biomarkers that can be used to predict malignancy have been identified. Here, we clarified the relationships between ETV1 expression and GISTs. First, we found that ETV1 expression was markedly upregulated at the mRNA level in GISTs. Second, ETV1 mRNA expression was significantly attenuated in aggressive malignant GISTs. Third, the multivariate analysis showed that ETV1 expression, in addition to conventional risk factors, such as tumor size, location and mitotic count, was an independent factor of recurrence. Moreover, the patients with GISTs that exhibited low ETV1 expression experienced shorter RFS than patients with GISTs that exhibited high ETV1 expression. Thus, ETV1 mRNA expression is negatively associated with malignancy in GISTs.
After it was first suggested that a relationship might exist between ETV1 expression and GISTs15, several researchers examined ETV1 expression in GISTs using immunohistochemical (IHC) staining and reported that it was present in approximately 50–60% of GISTs16,17, which disagreed with the findings of the initial study15. In the present study, real-time PCR analysis confirmed that ETV1 mRNA was expressed in all GISTs. On the other hand, non-GISTs (adenocarcinoma, schwannoma and leiomyoma) exhibited significantly lower ETV1 mRNA expression than GISTs. A similar result was reported in 2019: ETV1 expression was significantly higher in GISTs than in other sarcomas20. This discrepancy in positive frequency may depend on the detectability of the examination, such as IHC staining or RT-qPCR. Our finding that all GISTs express ETV1 mRNA universally supports that ETV1 is related to GIST formation. Furthermore, we determined that ETV1 mRNA expression was related to clinical outcomes in patients with GISTs, i.e., an inverse relationship was detected between ETV1 mRNA expression and RFS. This result implies that ETV1 expression is a useful molecular marker in GISTs.
After ETV1 was reported to contribute to GIST proliferation in cooperation with KIT in 2010, several studies on ETV1 expression in GISTs were published17,21,22,23. However, there is no consensus about ETV1 expression in GISTs. One of the reasons may be the evaluation method using IHC staining, which depends on the affinity of the antibody. On the other hand, there is only one report regarding mRNA, in which 95% of GISTs and 5% of non-GISTs were positive for ETV1 mRNA in situ hybridization21. However, our assessment of ETV1 mRNA expression using real-time PCR was different and showed that ETV1 was ubiquitously expressed at the mRNA level in GISTs, and low ETV1 expression was associated with poor RFS. However, there was no difference in OS between the low and high ETV1 expression groups. Eleven patients who experienced recurrence were treated with TKIs. Although two patients died within one month, five patients achieved 5-year survival. We presume that the development of molecular targeted therapies such as imatinib, sunitinib, and regorafenib might explain the marked discrepancy between OS and RFS in patients with GISTs.
The role of ETV1 in GISTs implied in the current study may be controversial. However, our finding applies to only ETV1 mRNA, which is completely different from the ETV1 protein. Moreover, protein expression does not always reflect mRNA expression. There are several reports that ETV1 at the protein level is stabilized by activated KIT signaling15,24,25. Therefore, a negative feedback system to regulate mRNA by the ETV1 protein might explain both our and previous results.
There are some limitations to this study. First, the sample size was small, although several data showed a statistically significant difference. Second, the cut-off value used in this study to distinguish between high and low expression was decided by the median. The value was not an absolute value and may have been altered by the study population, especially the rate of high-risk GISTs. Third, most GISTs in this study originated from the stomach. Our results may be applied only to gastric GISTs. Fourth, the protein level of ETV1 expression was not assessed simultaneously by Western blotting in all cases. Fifth, the molecular mechanism of attenuated ETV1 expression was not examined in this study. Further experimental and clinical studies are necessary to more clearly reveal the relationship between ETV1 expression and the aggressiveness of GISTs.
In conclusion, low ETV1 expression was associated with worse RFS and was revealed as an independent factor of recurrence in addition to conventional factors. ETV1 expression could be a predictor of aggressive malignant GISTs. An evaluation system based on ETV1 expression may help choose patients who need adjuvant chemotherapy.
Abbreviations
- GIST:
-
Gastrointestinal stromal tumor
- ETV1:
-
ETS translocation variant 1
- PCR:
-
Polymerase chain reaction
- PDGFRA:
-
Platelet-derived growth factor receptor A
- NIH:
-
National Institutes of Health
- OS:
-
Overall survival
- RFS:
-
Recurrence-free survival
- IHC:
-
Immunohistochemical
References
Hirota, S. et al. Gain-of-function mutations of c-kit in human gastrointestinal stromal tumors. Science 279, 577–580 (1998).
Rubin, B. P., Fletcher, J. A. & Fletcher, C. D. Molecular insights into the histogenesis and pathogenesis of gastrointestinal stromal tumors. Int. J. Surg. Pathol. 8, 5–10. https://doi.org/10.1177/106689690000800105 (2000).
Miettinen, M. & Lasota, J. Gastrointestinal stromal tumors–definition, clinical, histological, immunohistochemical, and molecular genetic features and differential diagnosis. Virchows Arch. 438, 1–12 (2001).
Taniguchi, M. et al. Effect of c-kit mutation on prognosis of gastrointestinal stromal tumors. Cancer Res. 59, 4297–4300 (1999).
Rubin, B. P. et al. KIT activation is a ubiquitous feature of gastrointestinal stromal tumors. Cancer Res. 61, 8118–8121 (2001).
Lasota, J., Jasinski, M., Sarlomo-Rikala, M. & Miettinen, M. Mutations in exon 11 of c-Kit occur preferentially in malignant versus benign gastrointestinal stromal tumors and do not occur in leiomyomas or leiomyosarcomas. Am. J. Pathol. 154, 53–60. https://doi.org/10.1016/s0002-9440(10)65250-9 (1999).
Hirota, S. et al. Gain-of-function mutation at the extracellular domain of KIT in gastrointestinal stromal tumours. J. Pathol. 193, 505–510. https://doi.org/10.1002/1096-9896(2000)9999:9999%3c::aid-path818%3e3.0.co;2-e (2001).
Joensuu, H. et al. Effect of the tyrosine kinase inhibitor STI571 in a patient with a metastatic gastrointestinal stromal tumor. N. Engl. J. Med. 344, 1052–1056. https://doi.org/10.1056/nejm200104053441404 (2001).
Demetri, G. D. et al. Efficacy and safety of imatinib mesylate in advanced gastrointestinal stromal tumors. N. Engl. J. Med. 347, 472–480. https://doi.org/10.1056/NEJMoa020461 (2002).
Tanaka, J. et al. Small gastrointestinal stromal tumor of the stomach showing rapid growth and early metastasis to the liver. Dig. Endosc. 22, 354–356. https://doi.org/10.1111/j.1443-1661.2010.01032.x (2010).
Fernandez, J. A. et al. Clinical and pathological features of “small” GIST (≤2 cm). What is their prognostic value?. Eur. J. Surg. Oncol. 44, 580–586. https://doi.org/10.1016/j.ejso.2018.01.087 (2018).
Rossi, S. et al. Molecular and clinicopathologic characterization of gastrointestinal stromal tumors (GISTs) of small size. Am. J. Surg. Pathol. 34, 1480–1491. https://doi.org/10.1097/PAS.0b013e3181ef7431 (2010).
Corless, C. L., McGreevey, L., Haley, A., Town, A. & Heinrich, M. C. KIT mutations are common in incidental gastrointestinal stromal tumors one centimeter or less in size. Am. J. Pathol. 160, 1567–1572. https://doi.org/10.1016/s0002-9440(10)61103-0 (2002).
Agaimy, A. et al. Minute gastric sclerosing stromal tumors (GIST tumorlets) are common in adults and frequently show c-KIT mutations. Am. J. Surg. Pathol. 31, 113–120. https://doi.org/10.1097/01.pas.0000213307.05811.f0 (2007).
Chi, P. et al. ETV1 is a lineage survival factor that cooperates with KIT in gastrointestinal stromal tumours. Nature 467, 849–853. https://doi.org/10.1038/nature09409 (2010).
Birner, P. et al. MAPKAP kinase 2 overexpression influences prognosis in gastrointestinal stromal tumors and associates with copy number variations on chromosome 1 and expression of p38 MAP kinase and ETV1. Clin. Cancer Res. 18, 1879–1887. https://doi.org/10.1158/1078-0432.ccr-11-2364 (2012).
Kubota, D. et al. Gene expression network analysis of ETV1 reveals KCTD10 as a novel prognostic biomarker in gastrointestinal stromal tumor. PLoS ONE 8, e73896. https://doi.org/10.1371/journal.pone.0073896 (2013).
Joensuu, H. Risk stratification of patients diagnosed with gastrointestinal stromal tumor. Hum. Pathol. 39, 1411–1419. https://doi.org/10.1016/j.humpath.2008.06.025 (2008).
Huss, S. et al. A subset of gastrointestinal stromal tumors previously regarded as wild-type tumors carries somatic activating mutations in KIT exon 8 (p.D419del). Mod. Pathol. 26, 1004–1012. https://doi.org/10.1038/modpathol.2013.47 (2013).
Yen, C. C. et al. Identification of phenothiazine as an ETV1-targeting agent in gastrointestinal stromal tumors using the Connectivity Map. Int. J. Oncol. 55, 536–546. https://doi.org/10.3892/ijo.2019.4829 (2019).
Jang, B. G., Lee, H. E. & Kim, W. H. ETV1 mRNA is specifically expressed in gastrointestinal stromal tumors. Virchows Arch. 467, 393–403. https://doi.org/10.1007/s00428-015-1813-9 (2015).
Wang, H. C. et al. KIT Exon 11 Codons 557–558 deletion mutation promotes liver metastasis through the CXCL12/CXCR4 axis in gastrointestinal stromal tumors. Clin. Cancer Res. 22, 3477–3487. https://doi.org/10.1158/1078-0432.ccr-15-2748 (2016).
Zeng, S. et al. ETV4 collaborates with Wnt/beta-catenin signaling to alter cell cycle activity and promote tumor aggressiveness in gastrointestinal stromal tumor. Oncotarget 8, 114195–114209. https://doi.org/10.18632/oncotarget.23173 (2017).
Hayashi, Y. et al. Platelet-derived growth factor receptor-α regulates proliferation of gastrointestinal stromal tumor cells with mutations in KIT by stabilizing ETV1. Gastroenterology 149, 420–432. https://doi.org/10.1053/j.gastro.2015.04.006 (2015).
Ran, L. et al. Combined inhibition of MAP kinase and KIT signaling synergistically destabilizes ETV1 and suppresses GIST tumor growth. Cancer Discov. 5, 304–315. https://doi.org/10.1158/2159-8290.cd-14-0985 (2015).
Acknowledgements
The authors wish to thank Dr. Takeshi Senga for his valuable help regarding the molecular biology techniques and experiments. This research was founded by the JSPS KAKENHI Grant-in-Aid for Young Scientists (B) (Grant number: JP24790689).
Author information
Authors and Affiliations
Contributions
K.S. and K.F.: concept and design, drafting of the manuscript. K.F. and R.M.: analysis and interpretation of data. T.Y., E.O., M.N., and H.K.: acquisition of the data. Y.H. and M.F.: observers. H.G.: director of this study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sakamaki, K., Funasaka, K., Miyahara, R. et al. Low ETV1 mRNA expression is associated with recurrence in gastrointestinal stromal tumors. Sci Rep 10, 14767 (2020). https://doi.org/10.1038/s41598-020-71719-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71719-y
- Springer Nature Limited